Featured
-
-
Letter |
 The structural basis for membrane binding and pore formation by lymphocyte perforin
The structural basis for membrane binding and pore formation by lymphocyte perforin
Natural killer cells and cytotoxic T cells kill virus-infected and malignant cells, releasing the pore-forming protein perforin in the process. Perforin is required for the delivery of pro-apoptotic granzymes to the target cell. These authors present the crystal structure of a perforin monomer together with a cryo-electron microscopy reconstruction of the oligomeric pore. Perforin monomers within the pore are arranged with an inside-out orientation relative to the structurally homologous monomers of cholesterol-dependent cytolysins.
- Ruby H. P. Law
- , Natalya Lukoyanova
- & James C. Whisstock
-
Article |
 Homologue structure of the SLAC1 anion channel for closing stomata in leaves
Homologue structure of the SLAC1 anion channel for closing stomata in leaves
SLAC1 is a plant ion channel that controls turgor pressure in the guard cells of plant stomata, thereby regulating the exchange of water vapour and photosynthetic gases in response to environmental signals. Here, the X-ray crystal structure of a bacterial homologue of SLAC1 has been solved, and structure-inspired mutagenesis has been used to analyse the conductance properties of the channel. The findings indicate that selectivity among different anions is largely a function of the energetic cost of ion dehydration.
- Yu-hang Chen
- , Lei Hu
- & Wayne A. Hendrickson
-
Letter |
 Structure and mechanism of the S component of a bacterial ECF transporter
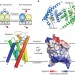
Structure and mechanism of the S component of a bacterial ECF transporter
The energy-coupling factor transporters are responsible for vitamin uptake in prokaryotes. Here, the X-ray crystal structure of the membrane-embedded, substrate-binding domain of a riboflavin transporter from Staphylococcus aureus is reported. The transporter adopts a previously unreported fold and contains a riboflavin molecule bound to a loop and the periplasmic portion of several transmembrane segments.
- Peng Zhang
- , Jiawei Wang
- & Yigong Shi
-
-
News & Views |
 Last of the multidrug transporters
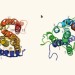
Last of the multidrug transporters
Proteins that pump a wide range of toxic compounds out of cells are ubiquitous in nature, but crystal structures for one family of these transporters have remained elusive. Until now. See Letter p.991
- Hendrik W. van Veen
-
-
News & Views |
 Egocentric pre–T–cell receptors
Egocentric pre–T–cell receptors
The T-cell receptor on the surface of T cells requires antigen recognition to function. Structural studies reveal that its predecessor, the pre-T-cell receptor, is much more independent. See Letter p.844
- Bernard Malissen
- & Hervé Luche
-
Letter |
 The proteasome antechamber maintains substrates in an unfolded state
The proteasome antechamber maintains substrates in an unfolded state
The proteasome is a multi-protein complex that enzymatically degrades proteins. Proteolysis occurs in a barrel-shaped 20S core particle comprising three interconnected cavities, including a pair of antechambers in which substrates are held before degradation. These authors demonstrate that substrates interact actively with the antechamber walls and that the environment in this compartment is optimized to maintain the substrates in unfolded states so as to be accessible for hydrolysis.
- Amy M. Ruschak
- , Tomasz L. Religa
- & Lewis E. Kay
-
Letter |
 The structural basis for autonomous dimerization of the pre-T-cell antigen receptor
The structural basis for autonomous dimerization of the pre-T-cell antigen receptor
The pre-T-cell antigen receptor mediates early T-cell development and differentiation. These authors report its structure and explain how the head-to-tail dimeric arrangement allows the interaction of the pre-Tα domain with any variable β domain, and provides the basis for ligand-independent signalling.
- Siew Siew Pang
- , Richard Berry
- & Jamie Rossjohn
-
Article |
 The Ndc80 kinetochore complex forms oligomeric arrays along microtubules
The Ndc80 kinetochore complex forms oligomeric arrays along microtubules
The Ndc80 complex is a key component of kinetochore that mediates direct interaction with spindle microtubules. These authors present a cryo-electron microscopy reconstruction of Ndc80 bound to microtubules. They find that Ndc80 uses a novel microtubule recognition mode coupling tubulin binding to self-oligomerization of the complex, and present a mechanistic model for the formation of high-affinity kinetochore–microtubule attachments during cell division.
- Gregory M. Alushin
- , Vincent H. Ramey
- & Eva Nogales
-
Letter |
 A redox switch in angiotensinogen modulates angiotensin release
A redox switch in angiotensinogen modulates angiotensin release
Angiotensins have a crucial role in blood pressure regulation and are generated by cleavage of a larger protein, angiotensinogen, by the enzyme renin. Structures of angiotensinogen alone and in complex with renin show that a large conformational change is required to expose the renin-cleavage site. The authors also show that this transition is regulated by oxidation and that women with pre-eclampsia have higher levels of the more active, oxidized, form.
- Aiwu Zhou
- , Robin W. Carrell
- & Randy J. Read
-
Article |
 Jasmonate perception by inositol-phosphate-potentiated COI1–JAZ co-receptor
Jasmonate perception by inositol-phosphate-potentiated COI1–JAZ co-receptor
The F-box protein CORONATINE INSENSITIVE 1 (COI1) mediates jasmonate signalling by promoting hormone-dependent ubiquitylation and degradation of the JASMONATE ZIM DOMAIN (JAZ) family of transcriptional repressors. These authors elucidate the mechanism of jasmonate perception. They present structural and pharmacological data to show that the true jasmonate receptor is a complex of both COI1 and JAZ. In addition, inositol pentakisphosphate functions as a critical component of the hormone receptor complex.
- Laura B. Sheard
- , Xu Tan
- & Ning Zheng
-
Article |
 An unprecedented nucleic acid capture mechanism for excision of DNA damage
An unprecedented nucleic acid capture mechanism for excision of DNA damage
DNA bases that are alkylated or deaminated are removed by DNA glycosylase repair enzymes. In structures of several other DNA glycosylases, the modified base inserts into the active site. These authors solve the structure of glycosylase AlkD, find that the modified base is extruded in an extrahelical position and propose a model for how this solvent-exposed position allows cleavage of N3- and N7-alkylated bases specifically.
- Emily H. Rubinson
- , A. S. Prakasha Gowda
- & Brandt F. Eichman
-
News |
 Consortium solves its 1,000th protein structure
Consortium solves its 1,000th protein structure
The international Structural Genomics Consortium celebrates its focus on expensive but exciting targets.
- Heidi Ledford
-
Letter |
 Two enzymes bound to one transfer RNA assume alternative conformations for consecutive reactions
Two enzymes bound to one transfer RNA assume alternative conformations for consecutive reactions
In most bacteria and all archaea, glutamyl-tRNA synthetase (GluRS) glutamylates both tRNAGlu and tRNAGln; Glu-tRNAGln is then converted to Gln-tRNAGln by an amidotransferase. Here the structure is reported of a bacterial complex containing tRNAGln, GluRS and the amidotransferase GatCAB. The structure provides an explanation for how the enzymes work consecutively: only one can assume a productive state at any time. There also seems to be an intermediary state in which neither enzyme is productive.
- Takuhiro Ito
- & Shigeyuki Yokoyama
-
Comment |
 The lost correspondence of Francis Crick
The lost correspondence of Francis Crick
Alexander Gann and Jan Witkowski unveil newly found letters between key players in the DNA story. Strained relationships and vivid personalities leap off the pages.
- Alexander Gann
- & Jan Witkowski
-
Letter |
 Structural basis for semaphorin signalling through the plexin receptor
Structural basis for semaphorin signalling through the plexin receptor
Semaphorin proteins mediate signal transduction by interacting with plexin receptors; they have key roles in neuronal development and many other biological processes. Here, crystal structures are presented of the semaphorin 6A receptor-binding fragment and the plexin A2 ligand-binding fragment in their pre-signalling and signalling states. On the basis of these structures, a signalling mechanism is proposed.
- Terukazu Nogi
- , Norihisa Yasui
- & Junichi Takagi
-
Letter |
 Structure of a fucose transporter in an outward-open conformation
Structure of a fucose transporter in an outward-open conformation
In Escherichia coli, the uptake of L-fucose, an important source of carbon for microorganisms, is mediated by a proton symporter from the major facilitator superfamily (MFS). These authors report the first X-ray crystal structure of the outward-open conformation of an MFS proton transporter, FucP. Building on previous work, they develop a working model for how the substrate is recognized by the transporter and how the protein mediates L-fucose/proton symport.
- Shangyu Dang
- , Linfeng Sun
- & Nieng Yan
-
Letter |
 Structural basis of semaphorin–plexin signalling
Structural basis of semaphorin–plexin signalling
Semaphorin proteins mediate signal transduction by interacting with plexin receptors; they have key roles in neuronal development and many other biological processes. Here, crystal structures are presented of the semaphorin-binding regions of plexin B1 and plexin A2 with their cognate semaphorin ectodomains. On the basis of these structures, a signalling mechanism is proposed.
- Bert J. C. Janssen
- , Ross A. Robinson
- & E. Yvonne Jones
-
Article |
 Selective inhibition of BET bromodomains
Selective inhibition of BET bromodomains
A new approach is used to target BET family bromodomains which are found in transcriptional regulators where they mediate the recognition of acetyl-lysine chromatin marks. Structural data reveal how the compound JQ1 binds to the bromodomain of BRD4. BRD4 has been implicated in a subtype of human squamous carcinomas, and JQ1 is found to inhibit the growth of BRD4 dependent tumours in mouse models.
- Panagis Filippakopoulos
- , Jun Qi
- & James E. Bradner
-
Letter |
 Crystal structures of the CusA efflux pump suggest methionine-mediated metal transport
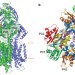
Crystal structures of the CusA efflux pump suggest methionine-mediated metal transport
Gram-negative bacteria, such as Escherichia coli, use tripartite efflux complexes in the resistance-nodulation-cell division family to expel toxic compounds from the cell. The CusCBA system is responsible for removing biocidal Cu(I) and Ag(I) ions. Here, the X-ray crystal structure is reported of CusA in the absence and presence of bound Cu(I) or Ag(I). The structures reveal that the metal-binding sites are located within the cleft region of the periplasmic domain. A potential pathway for ion export is proposed.
- Feng Long
- , Chih-Chia Su
- & Edward W. Yu
-
-
Letter |
 Structure of a cation-bound multidrug and toxic compound extrusion transporter
Structure of a cation-bound multidrug and toxic compound extrusion transporter
Transporter proteins from the MATE (multidrug and toxic compound extrusion) family are involved in metabolite transport in plants, and in multiple-drug resistance in bacteria and mammals. Here, the X-ray crystal structure of a MATE transporter from Vibrio cholerae is reported. The structure is in an outward-facing conformation, and reveals a cation-binding site near to residues previously deemed essential for transport.
- Xiao He
- , Paul Szewczyk
- & Geoffrey Chang
-
Letter |
 Crystal structure of the human symplekin–Ssu72–CTD phosphopeptide complex
Crystal structure of the human symplekin–Ssu72–CTD phosphopeptide complex
The scaffolding protein symplekin affects the initiation and termination of transcription and is involved in cleavage and polyadenylation at the 3′ ends of precursor messenger RNAs. These authors have solved the structure of a ternary complex of symplekin, a short peptide mimicking the phosphorylated carboxy-terminal tail of RNA polymerase II, and Ssu72, which dephosphorylates this residue. The structure suggests explains how Ssu72 binding can facilitate polyadenylation activity when 3′-end processing is coupled to transcription.
- Kehui Xiang
- , Takashi Nagaike
- & Liang Tong
-
News |
 Electron microscopy gets twisted
Electron microscopy gets twisted
Spiralling electron beams have the potential to measure and manipulate the properties of single atoms.
- Zeeya Merali
-
Letter |
 Direct visualization of secondary structures of F-actin by electron cryomicroscopy
Direct visualization of secondary structures of F-actin by electron cryomicroscopy
The formation of filamentous F-actin, through polymerization of globular G-actin, is essential for processes such as cell motility and muscle contraction. These authors report the structure of F-actin as visualized by electron cryomicroscopy, and build a complete atomic model of F-actin. This new structure will improve our understanding of the mechanism of actin assembly and disassembly.
- Takashi Fujii
- , Atsuko H. Iwane
- & Keiichi Namba
-
Letter |
 Structural basis of Na+-independent and cooperative substrate/product antiport in CaiT
Structural basis of Na+-independent and cooperative substrate/product antiport in CaiT
Transport of solutes across biological membranes is carried out by specialized secondary transport proteins in the lipid bilayer. These authors report structures of the sodium-independent carnitine/butyrobetaine antiporter CaiT from two microorganisms. The three-dimensional architecture of CaiT resembles that of the Na+-dependent transporters LeuT and BetP, but in CaiT a methionine sulphur takes the place of the Na+ ion to coordinate the substrate in the central transport site, enabling Na+-independent transport to occur.
- Sabrina Schulze
- , Stefan Köster
- & Werner Kühlbrandt
-
Letter |
 Genome-wide measurement of RNA secondary structure in yeast
Genome-wide measurement of RNA secondary structure in yeast
Experimental determination of the secondary structure of RNA molecules has usually been carried out on a case-by-case basis. Now, however, a deep-sequencing approach has been used to profile the secondary structure of 3,000 distinct messenger RNA transcripts from Saccharomyces cerevisiae. The results provide interesting hints about the role of secondary structure in protein translation, and set the stage for the examination of how such structures can change in response to environmental conditions.
- Michael Kertesz
- , Yue Wan
- & Eran Segal
-
News & Views |
 Conservation in vesicle coats
Conservation in vesicle coats
Coat proteins of vesicles involved in intracellular membrane trafficking have closely related molecular architectures. The structure of COPI extends known similarities, and strengthens the case for a common evolutionary origin.
- Stephen C. Harrison
- & Tomas Kirchhausen
-
Article |
 Structure of RCC1 chromatin factor bound to the nucleosome core particle
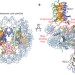
Structure of RCC1 chromatin factor bound to the nucleosome core particle
The small GTPase Ran regulates nuclear transport and cell division by creating a gradient of RanGTP around chromosomes. The RCC1 protein recruits Ran to nucleosomes and activates Ran's nucleotide exchange activity. Here, the crystal structure of the complex between RCC1 and the nucleosome core particle is revealed. It provides an atomic view of how a chromatin protein interacts with the histone and DNA components of the nucleosome.
- Ravindra D. Makde
- , Joseph R. England
- & Song Tan
-
Letter |
 The structure of (CENP-A–H4)2 reveals physical features that mark centromeres
The structure of (CENP-A–H4)2 reveals physical features that mark centromeres
The centromeres of chromosomes are specified epigenetically, and the histone H3 variant CENP-A is assembled into the chromatin of all active centromeres. Here, the crystal structure of CENP-A in a tetrameric complex with histone H4 reveals the physical features of centromeric chromatin. CENP-A seems to mark the centromere by altering nucleosome structure from within its folded histone core.
- Nikolina Sekulic
- , Emily A. Bassett
- & Ben E. Black
-
-
Brief Communications Arising |
Veselka et al. reply
- Nina Veselka
- , David D. McErlain
- & M. Brock Fenton
-
Letter |
 Crystal structure of the α6β6 holoenzyme of propionyl-coenzyme A carboxylase
Crystal structure of the α6β6 holoenzyme of propionyl-coenzyme A carboxylase
Propionyl-coenzyme A carboxylase (PCC) is a biotin-dependent enzyme that is essential for the catabolism of several amino acids, cholesterol and some fatty acids. Here, the crystal structure of a bacterial PCC is presented, along with a cryo-electron microscopy reconstruction showing a similar structure for human PCC. The structural information establishes a molecular basis for understanding the known disease-causing mutations in PCC, and is relevant to the holoenzymes of other biotin-dependent carboxylases.
- Christine S. Huang
- , Kianoush Sadre-Bazzaz
- & Liang Tong
-
Letter |
 Neurological disease mutations compromise a C-terminal ion pathway in the Na+/K+-ATPase
Neurological disease mutations compromise a C-terminal ion pathway in the Na+/K+-ATPase
The Na+/K+-ATPase pumps three sodium ions out of and two potassium ions into the cell while splitting a single molecule of ATP. Here it is found that the carboxy terminus of the ATPase's α-subunit is also a key regulator of a previously unrecognized ion pathway. The data indicate that, in the ATPase's potassium-bound state, a cytoplasmic proton can enter and stabilize site III when empty. When potassium is released, the proton returns to the cytoplasm, thus permitting an overall asymmetric stoichiometry of the transported ions.
- Hanne Poulsen
- , Himanshu Khandelia
- & Poul Nissen
-
Letter |
 Structure of the LexA–DNA complex and implications for SOS box measurement
Structure of the LexA–DNA complex and implications for SOS box measurement
Normally, expression of bacterial DNA damage repair genes is repressed by the binding of LexA protein to SOS ‘boxes’ in their operators. DNA damage activates the RecA protein, which promotes autocleavage of LexA such that its repression is relieved and repair proteins are expressed. These authors solve several structures of LexA dimer bound to SOS box DNA, and find that the orientation of the DNA-binding wings can account for the strict intersite spacing.
- Adrianna P. P. Zhang
- , Ying Z. Pigli
- & Phoebe A. Rice
-
Letter |
 Predicting protein structures with a multiplayer online game
Predicting protein structures with a multiplayer online game
Predicting the structure of a folded protein from first principles for any given amino-acid sequence remains a formidable computational challenge. To recruit human abilities to the task, these authors turned their Rosetta structure prediction algorithm into an online multiplayer game in which thousands of non-scientists competed and collaborated to produce new algorithms and search strategies for protein structure refinement. This shows that computationally complex problems can be effectively 'crowd-sourced' through interactive multiplayer games.
- Seth Cooper
- , Firas Khatib
- & Foldit players
-
News |
 Feeling the shapes of molecules
Feeling the shapes of molecules
The atomic structure of a small organic molecule can be revealed by atomic force microscopy.
- Philip Ball
-
Letter |
 Structure of the torque ring of the flagellar motor and the molecular basis for rotational switching
Structure of the torque ring of the flagellar motor and the molecular basis for rotational switching
The bacterial flagellar motor drives the rotation of flagellar filaments, propelling bacteria through viscous media. The rotation can switch from an anticlockwise to a clockwise direction, determining a smooth or tumbling motion. A protein called FliG forms a ring in the motor's rotor, and has been proposed to adopt distinct conformations that induce switching. Here, the full-length structure of FliG is presented, and conformational changes are identified that are involved in switching between clockwise and anticlockwise rotations.
- Lawrence K. Lee
- , Michael A. Ginsburg
- & Daniela Stock
-
News |
 Protein mapping gains a human focus
Protein mapping gains a human focus
Next phase of the US Protein Structure Initiative enlists biologists to help crack tough human receptors.
- Heidi Ledford
-
Article |
 Ribosome dynamics and tRNA movement by time-resolved electron cryomicroscopy
Ribosome dynamics and tRNA movement by time-resolved electron cryomicroscopy
During protein synthesis within the ribosome, transfer RNAs (tRNAs) move sequentially through different sites as their attached amino acids are transferred onto the growing protein chain. Large conformational movements accompany this process. Here, a staggering 1.9 million electron cryomicroscopy images of the ribosome have been processed to visualize these changes. The results reveal that the ribosome functions as a Brownian machine that couples spontaneous changes driven by thermal energy to directed movement.
- Niels Fischer
- , Andrey L. Konevega
- & Holger Stark
-
News & Views |
 Translocation in slow motion
Translocation in slow motion
Time-resolved electron microscopy can capture structural changes in active macromolecular complexes, but detailed imaging is essential. The dynamics of one step in protein synthesis has been deduced from two million images.
- Måns Ehrenberg
-
Letter |
 Microtubule nucleating γ-TuSC assembles structures with 13-fold microtubule-like symmetry
Microtubule nucleating γ-TuSC assembles structures with 13-fold microtubule-like symmetry
XXXMicrotubules are nucleated in vivo by γ-tubulin complexes and comprise 13 protofilaments. How this precise geometry is controlled remains unclear. These authors report the cryo-electron microscopic structure of the universally conserved, core microtubule nucleating complex, γ-tubulin small complex. The structure provides insight into how this complex establishes thirteen-fold tubulin symmetry.
- Justin M. Kollman
- , Jessica K. Polka
- & David A. Agard
-
Letter |
 Mechanism and regulation of acetylated histone binding by the tandem PHD finger of DPF3b
Mechanism and regulation of acetylated histone binding by the tandem PHD finger of DPF3b
The lysine residues of histone proteins can be acetylated or methylated, with important effects on gene expression. Until recently the protein modules that bind acetyl-lysine have been limited to bromodomains. However, the tandem plant homeodomain (PHD) finger of human DPF3b — which is involved in gene activation — has also been reported to bind to acetylated histones. Here, three-dimensional solution structures of DPF3b offer mechanistic insight into how this protein recognizes acetylation marks.
- Lei Zeng
- , Qiang Zhang
- & Ming-Ming Zhou
-
Article |
 Structural mechanism of C-type inactivation in K+ channels
Structural mechanism of C-type inactivation in K+ channels
K+ channels can convert between conductive and non-conductive forms through mechanisms that range from flicker transitions (which occur in microseconds) to C-type inactivation (which occurs on millisecond to second timescales). Here, the crystal structures are presented of the potassium channel KcsA in an open-inactivated conformation and 'trapped' in several partially open conformations. The structures indicate a molecular basis for C-type inactivation in K+ channels.
- Luis G. Cuello
- , Vishwanath Jogini
- & Eduardo Perozo
-
Letter |
 Structural basis for the coupling between activation and inactivation gates in K+ channels
Structural basis for the coupling between activation and inactivation gates in K+ channels
K+ channels can convert between conductive and non-conductive forms through mechanisms that range from flicker transitions (which occur in microseconds) to C-type inactivation (which occurs on millisecond to second timescales). Here, the mechanisms are revealed through which movements of the inner gate of the K+ channel KcsA trigger conformational changes at the selectivity filter, leading to the non-conductive C-type inactivated state.
- Luis G. Cuello
- , Vishwanath Jogini
- & Eduardo Perozo
-
Letter |
 Subnanometre single-molecule localization, registration and distance measurements
Subnanometre single-molecule localization, registration and distance measurements
These authors have developed a method that enables them to observe single-molecule fluorescent probes with subnanometre precision and accuracy using conventional far-field fluorescence imaging. The improved resolution will enable researchers to characterize single 'molecules' of large, multisubunit biological complexes in biologically relevant environments.
- Alexandros Pertsinidis
- , Yunxiang Zhang
- & Steven Chu
-
Article |
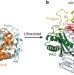 Structural basis for the suppression of skin cancers by DNA polymerase η
Structural basis for the suppression of skin cancers by DNA polymerase η
Ultraviolet radiation causes damage to DNA in skin cells, blocking DNA replication and causing mutations that can lead to cancer. One way in which the cell deals with such damage involves specialized DNA polymerases, such as Polη, that can bypass lesions. Here the crystal structure is presented of Pol? in complex with a thymine–thymine dimer and with undamaged DNA. The bulky thymine dimer is accommodated in an atypically large active site, and stabilized by interactions not found in other polymerases.
- Timothy D. Silverstein
- , Robert E. Johnson
- & Aneel K. Aggarwal
-
Article |
 Structure and mechanism of human DNA polymerase η
Structure and mechanism of human DNA polymerase η
Ultraviolet radiation causes damage to DNA in skin cells, blocking DNA replication and causing mutations that can lead to cancer. One way in which the cell deals with such damage involves specialized DNA polymerases, such as Polη, that can bypass lesions. Here the crystal structure of Polη is presented at four consecutive steps during DNA synthesis through thymine dimers. Polη acts like a 'molecular splint' to stabilize damaged DNA, and accommodates the thymine dimer in an atypically large active site.
- Christian Biertümpfel
- , Ye Zhao
- & Wei Yang

 The amino-terminal disease hotspot of ryanodine receptors forms a cytoplasmic vestibule
The amino-terminal disease hotspot of ryanodine receptors forms a cytoplasmic vestibule Concrete evidence of confusion
Concrete evidence of confusion