Abstract
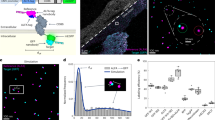
F-actin is a helical assembly of actin, which is a component of muscle fibres essential for contraction and has a crucial role in numerous cellular processes, such as the formation of lamellipodia and filopodia1,2, as the most abundant component and regulator of cytoskeletons by dynamic assembly and disassembly (from G-actin to F-actin and vice versa). Actin is a ubiquitous protein and is involved in important biological functions, but the definitive high-resolution structure of F-actin remains unknown. Although a recent atomic model well reproduced X-ray fibre diffraction intensity data from a highly oriented liquid-crystalline sol specimen3, its refinement without experimental phase information has certain limitations. Direct visualization of the structure by electron cryomicroscopy, however, has been difficult because it is relatively thin and flexible. Here we report the F-actin structure at 6.6 Å resolution, made obtainable by recent advances in electron cryomicroscopy. The density map clearly resolves all the secondary structures of G-actin, such as α-helices, β-structures and loops, and makes unambiguous modelling and refinement possible. Complex domain motions that open the nucleotide-binding pocket on F-actin formation, specific D-loop and terminal conformations, and relatively tight axial but markedly loose interprotofilament interactions hydrophilic in nature are revealed in the F-actin model, and all seem to be important for dynamic functions of actin.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Pollard, T. D. & Borisy, G. G. Cellular motility driven by assembly and disassembly of actin filaments. Cell 112, 453–465 (2003)
Carlier, M. F. & Pantaloni, D. Control of actin assembly dynamics in cell motility. J. Biol. Chem. 282, 23005–23009 (2007)
Oda, T., Iwasa, M., Aihara, T., Maeda, Y. & Narita, A. The nature of the globular- to fibrous-actin transition. Nature 457, 441–445 (2009)
Holmes, K. C., Popp, D., Gebhard, W. & Kabsch, W. Atomic model of the actin filament. Nature 347, 44–49 (1990)
Holmes, K. C. et al. Electron cryo-microscopy shows how strong binding of myosin to actin releases nucleotide. Nature 425, 423–427 (2003)
Namba, K. & Stubbs, G. Structure of tobacco mosaic virus at 3.6 A resolution: implications for assembly. Science 231, 1401–1406 (1986)
Yonekura, K., Maki-Yonekura, S. & Namba, K. Complete atomic model of the bacterial flagellar filament by electron cryomicroscopy. Nature 424, 643–650 (2003)
Miyazawa, A., Fujiyoshi, Y. & Unwin, N. Structure and gating mechanism of the acetylcholine receptor pore. Nature 423, 949–955 (2003)
Sachse, C. et al. High-resolution electron microscopy of helical specimens: a fresh look at tobacco mosaic virus. J. Mol. Biol. 371, 812–835 (2007)
Galkin, V. E., Orlova, A., Cherepanova, O., Lebart, M. C. & Egelman, E. H. High-resolution cryo-EM structure of the F-actin-fimbrin/plastin ABD2 complex. Proc. Natl Acad. Sci. USA 105, 1494–1498 (2008)
Fujii, T., Kato, T. & Namba, K. Specific arrangement of α-helical coiled coils in the core domain of the bacterial flagellar hook for the universal joint function. Structure 17, 1485–1493 (2009)
Sander, B., Golas, M. M. & Stark, H. Advantages of CCD detectors for de novo three-dimensional structure determination in single-particle electron microscopy. J. Struct. Biol. 151, 92–105 (2005)
Egelman, E. H. A robust algorithm for the reconstruction of helical filaments using single-particle methods. Ultramicroscopy 85, 225–234 (2000)
Rosenthal, P. B. & Henderson, R. Optimal determination of particle orientation, absolute hand, and contrast loss in single-particle electron cryomicroscopy. J. Mol. Biol. 333, 721–745 (2003)
Topf, M. et al. Protein structure fitting and refinement guided by cryo-EM density. Structure 16, 295–307 (2008)
Otterbein, L. R., Graceffa, P. & Dominguez, R. The crystal structure of uncomplexed actin in the ADP state. Science 293, 708–711 (2001)
Kabsch, W., Mannherz, H. G., Suck, D., Pai, E. F. & Holmes, K. C. Atomic model of the actin:DNase I complex. Nature 347, 37–44 (1990)
Pollard, T. D. Regulation of actin filament assembly by Arp2/3 complex and formins. Annu. Rev. Biophys. Biomol. Struct. 36, 451–477 (2007)
Iwasa, M. et al. Dual roles of Q137 of actin revealed by recombinant human cardiac muscle alpha-actin mutants. J. Biol. Chem. 283, 21045–21053 (2008)
Béla, N. & Jencks, W. P. Depolymerization of F-actin by concentrated solutions of salts and denaturing agents. J. Am. Chem. Soc. 87, 2480–2488 (1965)
Fujiwara, I., Vavylonis, D. & Pollard, T. D. Polymerization kinetics of ADP- and ADP-Pi-actin determined by fluorescence microscopy. Proc. Natl Acad. Sci. USA 104, 8827–8832 (2007)
Sutoh, K. Identification of myosin-binding sites on the actin sequence. Biochemistry 21, 3654–3661 (1982)
Schmid, M. F., Sherman, M. B., Matsudaira, P. & Chiu, W. Structure of the acrosomal bundle. Nature 431, 104–107 (2004)
McGough, A., Pope, B., Chiu, W. & Weeds, A. Cofilin changes the twist of F-actin: implications for actin filament dynamics and cellular function. J. Cell Biol. 138, 771–781 (1997)
Yu, X., Jin, L. & Zhou, Z. H. 3.88 Å structure of cytoplasmic polyhedrosis virus by cryo-electron microscopy. Nature 453, 415–419 (2008)
Zhang, X. et al. Near-atomic resolution using electron cryomicroscopy and single-particle reconstruction. Proc. Natl Acad. Sci. USA 105, 1867–1872 (2008)
Zhang, X., Jin, L., Fang, Q., Hui, W. H. & Zhou, Z. H. 3.3 A cryo-EM structure of a nonenveloped virus reveals a priming mechanism for cell entry. Cell 141, 472–482 (2010)
Cong, Y. et al. 4.0-Å resolution cryo-EM structure of the mammalian chaperonin TRiC/CCT reveals its unique subunit arrangement. Proc. Natl Acad. Sci. USA 107, 4967–4972 (2010)
Zhang, J. et al. Mechanism of folding chamber closure in a group II chaperonin. Nature 463, 379–383 (2010)
Hayward, S. & Berendsen, H. J. Systematic analysis of domain motions in proteins from conformational change: new results on citrate synthase and T4 lysozyme. Proteins 30, 144–154 (1998)
Ludtke, S. J., Baldwin, P. R. & Chiu, W. EMAN: semiautomated software for high-resolution single-particle reconstruction. J. Struct. Biol. 128, 82–97 (1999)
Frank, J. et al. SPIDER and WEB: processing and visualization of images in 3D electron microscopy and related fields. J. Struct. Biol. 116, 190–199 (1996)
Mindell, J. A. & Grigorieff, N. Accurate determination of local defocus and specimen tilt in electron microscopy. J. Struct. Biol. 142, 334–347 (2003)
Yonekura, K., Braunfeld, M. B., Maki-Yonekura, S. & Agard, D. A. Electron energy filtering significantly improves amplitude contrast of frozen-hydrated protein at 300kV. J. Struct. Biol. 156, 524–536 (2006)
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004)
Acknowledgements
We thank T. Kato for setting up and managing our cryoEM and computing facilities for high-throughput, high-resolution image analysis, and I. Tokita, M. Urabe and JEOL for maintaining the electron cryomicroscope in the best conditions. We also thank E. Egelman for his help in the use of the iterative helical real-space refinement method in the early stages of this study, and F. Oosawa, S. Asakura, H. Hotani and D. L. D. Caspar for their discussions, continuous support and encouragement. T.F. was a research fellow of the Japan Society for the Promotion of Science. This work was supported in part by Grants-in-Aid for Scientific Research to K.N. (16087207 and 21227006) from the Ministry of Education, Culture, Sports, Science and Technology of Japan.
Author information
Authors and Affiliations
Contributions
T.F. made various improvements to the cryoEM method and performed all the cryoEM experiments, the image analysis and the model-building of F-actin. A.H.I. prepared G-actin. T.Y. and K.N. planned the project, and K.N. supervised the project. T.F. and K.N. wrote the paper on the basis of discussions with A.H.I. and T.Y.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-7 with legends, Supplementary Tables 1-2, legends for Supplementary Movies 1-2 and additional references. (PDF 15001 kb)
Supplementary Movie 1
This movie shows a side view of the F-actin density map at 6.6 Å resolution obtained by cryoEM image analysis - see Supplementary Information file page 11 for full legend. (MOV 8871 kb)
Supplementary Movie 2
In this move we see the initial model fitting to the cryoEM density map and its refinement process to the final F-actin model - see Supplementary Information file page 11 for full legend. (MOV 4220 kb)
Rights and permissions
About this article
Cite this article
Fujii, T., Iwane, A., Yanagida, T. et al. Direct visualization of secondary structures of F-actin by electron cryomicroscopy. Nature 467, 724–728 (2010). https://doi.org/10.1038/nature09372
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature09372
This article is cited by
-
From amino-acid to disease: the effects of oxidation on actin-myosin interactions in muscle
Journal of Muscle Research and Cell Motility (2023)
-
50 Years of the steric-blocking mechanism in vertebrate skeletal muscle: a retrospective
Journal of Muscle Research and Cell Motility (2023)
-
Cofilactin filaments regulate filopodial structure and dynamics in neuronal growth cones
Nature Communications (2022)
-
Mechanical Behavior of Axonal Actin, Spectrin, and Their Periodic Structure: A Brief Review
Multiscale Science and Engineering (2021)
-
Comprehensive analysis of cytoskeleton regulatory genes identifies ezrin as a prognostic marker and molecular target in acute myeloid leukemia
Cellular Oncology (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.