Abstract
Extramammary Paget disease (EMPD) is a rare skin cancer that primarily affects older individuals predominantly in areas with apocrine sweat glands. Although most early EMPD lesions are indolent, patients with metastatic EMPD have a poor prognosis due to the lack of effective systemic treatment. In this study, we investigated the role of forkhead box M1 (FOXM1), a potent transcription factor, in EMPD and assessed the potential of FOXM1 as a therapeutic target. Immunohistochemistry of 112 primary and 17 metastatic EMPD samples revealed that FOXM1 expression increased with tumor progression. Patients in whom FOXM1 was expressed in more than 10% of tumor cells had significantly shorter disease-specific survival than the other patients (p = 0.0397). In in vitro studies using our newly established EMPD cell line, KS-EMPD-1, we found high expression of FOXM1. Knockdown of FOXM1 impaired tumor cell viability, migration, and invasion. Inhibition of FOXM1 using thiostrepton also reduced tumor cell viability in a dose-dependent manner. These findings suggest that FOXM1 is a promising therapeutic target for patients with EMPD.
Similar content being viewed by others
Introduction
Extramammary Paget disease (EMPD) is a rare skin cancer that primarily occurs in older individuals predominantly in areas with apocrine sweat glands 1,2,3,4,5,6. The pathogenesis and risk factors of EMPD are not well understood, but it typically affects Caucasian women and Asian men over 60 years old 2,7,8,9. The crude prevalence of EMPD in mainland China was reported to be 0.4 per million people in 2016 and the age-standardized incidence in Europe is 0.6 per million person-years 10,11. Most EMPD tumors are slow-growing and remain indolent as in situ lesions for long periods, resulting in a favorable prognosis for patients 12,13,14. It is worth noting that EMPD lesions can sometimes be difficult to diagnose clinically, as they can bear a striking resemblance to other types of inflammatory skin diseases 2. Furthermore, it is important to be aware that dermal invasion occurs in a significant proportion of cases (between 15 and 40%), which can increase the likelihood of metastasis 2,3,15. Given these potential risks, it is strongly recommended that individuals seek out early surgical removal of EMPD lesions in order to minimize their potential impact on overall health and wellbeing 6. It is also important to distinguish EMPD from mammary Paget disease, which is an epidermal extension from breast cancer, and secondary EMPD, which is an extension from visceral cancers 1,2. A panel of immunohistochemical staining such as CK7, CK20, GCDFP15, CDX2, and TRPS1 can be helpful in making this distinction 1,2,15,16. EMPD lesions often have an unclear tumor border and may have hidden skip lesions 17. To completely remove the tumors, various methods have been used, such as wide local excision with mapping biopsies, photodynamic diagnosis-guided surgery, and Mohs micrographic surgery 18,19,20,21. However, complete removal of the tumors is sometimes challenging due to anatomical constraints 22. Conventional chemotherapies with anticancer drugs such as TS-1, docetaxel (DTX), paclitaxel (PTX), 5-fluorouracil (5-FU), or cisplatin (CDDP) are used for metastatic EMPD, but the prognosis for such cases is generally poor 23,24,25; there is thus an urgent need for alternative treatments.
In the pursuit of novel therapeutic targets for EMPD, researchers have primarily concentrated on human epidermal growth factor receptor 2 (HER2), which is encoded by the ERBB2 gene 26,27. Several sporadic cases have demonstrated promising results with HER2 antibody and chemotherapy combinations 28,29,30. Currently, other potential targets are being extensively studied to broaden our understanding of EMPD treatment options, including androgen receptor 31,32,33,34, programmed death 1/programmed death-ligand 1 35,36,37, cyclin-dependent kinase 4 38, C-X-C chemokine receptor type 4 (CXCR4), CXCR7 39, phosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha (PIK3CA) 40, dynamin-related protein 1 (DRP1) 41, nectin cell adhesion molecule 4 (NECTIN4) 42,43, trophoblast cell surface antigen 2 (TROP2) 44, and forkhead box A1 (FOXA1) 45. However, the results were mainly based on analyzing formalin-fixed paraffin-embedded (FFPE) or fresh/frozen clinical tissue samples using immunohistochemical or nucleic acid-based approaches. To reveal the detailed pathogenetic mechanisms of EMPD, cellular methods including in vivo or in vitro disease models are required.
Forkhead box M1 (FOXM1) is a transcription factor belonging to the FOX family, which is expressed in embryonic tissues 46,47,48,49. Its regulated expression and activity play a crucial role in ensuring proper growth and maturation. FOXM1 is also involved in regulating significant processes in tumor development and progression such as proliferation, migration, angiogenesis, and chemoresistance 46,47,48,49. Previous studies have reported that FOXM1 is overexpressed in various cancers and sarcomas including breast cancer, melanoma, angiosarcoma, lung cancer, pancreatic cancer, gastric cancer, prostate cancer, leukemia, leiomyosarcoma, and synovial sarcoma 48,49,50,51,52,53,54,55,56,57,58,59. The overexpression of FOXM1 is associated with poor patient survival in many of these tumors, in which advanced tumors tend to more strongly express FOXM1. However, the role of FOXM1 expression in EMPD tumorigenesis and tumor progression remains unclear, as little information is available on its expression in EMPD. However, the difficulty in establishing in vivo or in vitro models of EMPD had hampered basic research to explore the pathobiology of EMPD. We recently established a novel EMPD cell line, named KS-EMPD-1 60. In this study, we investigated FOXM1 expression and its role in tumor proliferation and development in EMPD patients’ clinical samples and in this cell line. FOXM1 expression increased with tumor progression in patients’ clinical samples, and high FOXM1 expression in primary tumors was significantly associated with shorter disease-specific survival. In in vitro study using KS-EMPD-1, FOXM1 was associated with tumor cell viability, migration, and invasiveness, and its inhibition strongly impaired tumor cell viability.
Results
FOXM1 expression in EMPD patient tissue
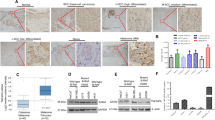
To investigate the expression of FOXM1 in EMPD lesions, 112 samples of primary EMPD and 17 samples of metastatic disease were immunohistochemically stained for FOXM1 and observed. Demographic and clinical data of the patients are listed in Table 1. As shown in Fig. 1A, FOXM1 was expressed in most of the EMPD samples, at least in part. FOXM1-positive cells increased in number with tumor progression; early lesions had fewer positive cells, while metastatic lesions had more of them (*p < 0.05 and ***p < 0.001, Fig. 1B). To further investigate the impact of FOXM1 expression level on patient survival, the patients were categorized into two groups (FOXM1 ≤ 10% and FOXM1 > 10%), and the relationship between FOXM1 expression and survival was analyzed. Patients with FOXM1 > 10% had significantly shorter survival than those with FOXM1 ≤ 10% (p = 0.0397) (Fig. 1C).
FOXM1 expression in patients’ EMPD tissues. (A) Immunohistochemical images of FOXM1 in patients’ EMPD tissues. Representative images of early lesions [tumor thickness (TT) ≤ 3 mm, n = 102], locally advanced lesions (TT > 3 mm, n = 10), and metastatic lesions (n = 17) are shown. Scale bars = 100 μm. (B) Comparison of FOXM1-positive cells (%) among early, locally advanced, and metastatic lesions of patients’ tissues. *p < 0.05 and ***p < 0.001. (C) Survival curves of patients with ≤ 10% (n = 77) or > 10% (n = 35) FOXM1 staining positivity in immunohistochemistry. Patients with FOXM1 > 10% had significantly shorter disease-specific survival than those with FOXM1 ≤ 10% (p = 0.0397).
FOXM1 expression in an EMPD cell line, KS-EMPD-1
Next, we investigated FOXM1 expression in vitro using an EMPD cell line, KS-EMPD-1. Normal epidermal keratinocytes (NHEKs) were also analyzed to determine FOXM1 expression in non-malignant skin cells for comparison. FOXM1 mRNA was significantly highly expressed in KS-EMPD-1 compared with that in NHEKs (Fig. 2A). In accordance with the findings on mRNA expression, FOXM1 protein was significantly highly expressed in KS-EMPD-1 cells compared with that in NHEKs (Fig. 2B, Supplementary Fig. S1). To further confirm the protein expression and localization, FOXM1 was immunocytochemically stained and observed. As shown in Fig. 2C, FOXM1 was expressed in the nucleus of KS-EMPD-1 cells, but not expressed in NHEKs.
FOXM1 is highly expressed in KS-EMPD-1 cells. (A) FOXM1 mRNA expression in KS-EMPD-1 cells and NHEKs. Mean ± SD of FOXM1 obtained from three independent experiments is shown. ***p < 0.001. (B) FOXM1 protein expression in KS-EMPD-1 cells and NHEKs. Representative blot images (left) and mean ± SD of FOXM1 protein (right) are shown. Experiments were independently repeated three times. Original, uncropped images are shown in Supplementary Fig. S1. ***p < 0.001. (C) Immunocytochemical images of FOXM1 in KS-EMPD-1 cells and NHEKs. FOXM1 (green), DAPI (blue, nuclear), and merged images of FOXM1 and DAPI are shown. Scale bars = 100 μm.
Thiostrepton, a FOXM1 inhibitor, inhibits proliferation of KS-EMPD-1 cells
Thiostrepton is an antibiotic that is also known to potently inhibit FOXM1 by directly interacting with it. When the KS-EMPD-1 cells were treated with thiostrepton, it significantly reduced their viability compared with that under the vehicle (dimethyl sulfoxide: DMSO)-treated conditions. The IC50 of thiostrepton was calculated as 0.606 ± 0.0883 μM in KS-EMPD-1 cells (Fig. 3A). The effects of thiostrepton on NHEKs (non-malignant skin cells) were also assessed. Thiostrepton treatment significantly decreased the number of NHEKs with IC50 of 0.981 ± 0.798 μM (Fig. 3B).
KS-EMPD-1 is sensitive to a FOXM1 inhibitor, thiostrepton. KS-EMPD-1 cells and NHEKs were incubated with DMSO (0.1%) or various concentrations of thiostrepton (0.25–10.0 μM) for 72 h and evaluated for the number of viable cells using a formazan-based method. Mean ± SD of fold change (viable cells) calculated from three independent experiments is shown. The IC50 of thiostrepton is shown in the boxes below the graphs. *p < 0.05, **p < 0.01, and ***p < 0.001.
Effects of FOXM1 knockdown on proliferation of KS-EMPD-1 cells
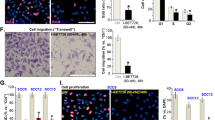
The effect of FOXM1 inhibition on cell proliferation was further tested by knocking down FOXM1 using specific small interfering RNA (siRNA). Knockdown efficiency was over 80% at both mRNA (Fig. 4A) and protein levels (Fig. 4B, Supplementary Fig. S2A). Cell proliferation was significantly inhibited in FOXM1 siRNA-transfected cells compared with that of control siRNA-transfected cells (Fig. 4C). The expression of cyclin B1, one of the downstream molecules of FOXM1 regulating the cell cycle, was further assessed in siRNA-transfected cells. The gene expression of cyclin B1 (CCNB1) was significantly downregulated in FOXM1 siRNA-transfected cells compared with that in control siRNA-transfected cells (Fig. 4D). Western blotting confirmed that cyclin B1 protein expression was also significantly downregulated in FOXM1-inhibited cells (Fig. 4E, Supplementary Fig. S2B). In addition, knockdown of FOXM1 significantly impaired tumor cell migration (Fig. 4F, G) and invasion (*p < 0.05, **p < 0.01, and ***p < 0.001, Fig. 4H, I).
FOXM1 knockdown inhibits proliferation, migration, and invasion of KS-EMPD-1 cells. Cells were transfected with FOXM1 siRNA and evaluated for their proliferation, migration, and invasion. (A) Knockdown efficiency of FOXM1 mRNA. Mean ± SD of FOXM1 calculated from three independent experiments is shown. ***p < 0.001. (B) Knockdown efficiency of FOXM1 protein. Representative blot images (upper) and mean ± SD of FOXM1 protein (lower) are shown. Experiments were independently repeated three times. Original, uncropped images are shown in Supplementary Fig. S2A. *p < 0.05, **p < 0.01, and ***p < 0.001. (C) Viable cells in control and FOXM1 siRNA-transfected conditions were quantified using a formazan-based method. Experiments were independently repeated three times. ***p < 0.001. (D) CCNB1 mRNA expression in control or FOXM1 siRNA-transfected cells at 24 h post-transfection. Mean ± SD of FOXM1 expression calculated from three independent experiments is shown. ***p < 0.001. (E) Cyclin B1 protein expression in control or FOXM1 siRNA-transfected cells at 48 h post-transfection. Experiments were independently repeated three times and representative blot images (upper) and mean ± SD of cyclin B1 protein expression (lower) are shown. Original, uncropped images are shown in Supplementary Fig. S2B. *p < 0.05. Cells were transfected with siRNA for 48 h. The cell monolayers were then scratched and the wound area were captured for 24 h. Representative images (F) and wound area relative to that at 0 h (G) calculated from three independent experiments are shown. *p < 0.05 and **p < 0.01. Cells were transfected with siRNA for 48 h and evaluated for cell invasion. Representative images (H) and mean ± SD of absorbance of cell staining dye (I) are shown. Experiments were independently repeated three times. Scale bars = 1.0 mm. ***p < 0.001.
Effect of FOXM1 knockdown on chemosensitivity of KS-EMPD-1 cells to anticancer drugs
To investigate the effects of FOXM1 knockdown on chemosensitivity to anticancer drugs, KS-EMPD-1 cells were transfected with FOXM1 siRNA and further treated with anticancer drugs (5-FU, CDDP, DTX, and PTX) used for treating EMPD. Knockdown of FOXM1 did not affect the chemosensitivity of KS-EMPD-1 cells to 5-FU (Fig. 5A). Knockdown of FOXM1 and further treatment with CDDP significantly decreased the proportion of viable cells compared with the control siRNA-transfected and CDDP -treated condition, but the change was only small (Fig. 5B). Similar to the findings for 5-FU, knockdown of FOXM1 did not affect the chemosensitivity of KS-EMPD-1 cells to DTX and PTX (Fig. 5C, D).
Knockdown of FOXM1 increases the chemosensitivity of KS-EMPD-1 cells. KS-EMPD-1 cells were transfected with control or FOXM1 siRNA for 24 h and further treated with vehicle control or anticancer drugs for 72 h. The viable cells were then quantitated using a formazan-based method. 5-Fluorouracil (5-FU, final concentration 100 nM), docetaxel (DTX, final concentration 5 μM), and paclitaxel (PTX, final concentration 1 μM) were dissolved in DMSO and cisplatin (CDDP, final concentration 10 μM) was dissolved in saline and further added to the culture medium. Concentration of anticancer drugs was determined based on the Cmax of each drug. Mean ± SD of fold change of viable cells calculated from three independent experiments is shown. *p < 0.05, **p < 0.01, and ***p < 0.001.
Discussion
Forkhead Box proteins, also known as Forkhead Homeobox proteins, are a group of transcription factors that belong to the winged helix transcription factors. These transcription factors originated from helix-turn-helix motifs of bacteria and are evolutionarily conserved 49,61,62,63. They were first discovered in the eukaryotic Forkhead Homeotic gene and have been found to have various roles in cell development, maintenance of cardiac equilibrium, angiogenesis, and tumorigenesis of different types of cancer 49,61,62,63. The human FOX group proteins have 19 subclasses (FOX-A to FOX-S) based on sequence conservation, and each subclass has different members 49,61,62,63. Among all FOX family members, FOXM1 is a central tissue factor and plays a key role in embryonic development. Its aberrant expression is associated with the development and progression of various types of cancer 49. FOXM1 is responsible for regulating cell proliferation and increasing the expression of genes related to DNA replication, G1/S and G2/M transitions, cyclin A2, Cdc25A phosphatase, ATF2, and JNK1 49,64,65,66. Overexpression and gene amplification of FOXM1 are linked to several types of cancer and this association has been found to result in a poor prognosis 51,52,56,57,58,59. Additional research has shown that FOXM1 plays a role in invasion, migration, angiogenesis, stemness, and therapy resistance, indicating its insidious nature 62,67,68,69,70,71.
In this study, we immunostained a relatively large number of EMPD clinical samples and conducted several in vitro analyses using our newly established EMPD cell line, KS-EMPD-1 60, in order to investigate the role of FOXM1 in the tumorigenesis of this disease and examine FOXM1 as a potential therapeutic target of EMPD. We found that, even in the early stages of EMPD, most cases showed strong and abundant positive expression of FOXM1, and its positivity increased with tumor progression. Considering the non-aggressive nature of EMPD, in that it arises in the epithelium of the skin and mucosa and remains indolent there for a long period as an in situ lesion 2, the overexpression of FOXM1 in EMPD is interesting since such overexpression is commonly related to aggressiveness in many other cancers 57,58,59. The KS-EMPD-1 cell line strongly expressed FOXM1 compared with NHEKs and knockdown of FOXM1 significantly reduced viable EMPD cells, suggesting its pivotal role in EMPD cell survival. Knockdown of FOXM1 also resulted in the downregulation of Cyclin B1, a molecule crucial for cell cycling. Knockdown of FOXM1 significantly impaired tumor cell migration and invasion. In addition, a potent FOXM1 inhibitor, thiostrepton, strongly impaired tumor cell viability in a dose-dependent manner. A topical ointment containing thiostrepton is currently available on the market for use on animals 72, but it is not approved for humans. EMPD is a skin condition that affects the epithelium, which is the outermost layer of the body. In most cases, EMPD cells remain in the epithelium. Therefore, the topical application of thiostrepton could be an effective way of delivering this FOXM1 inhibitor to the tumor cells and alleviating the disease. However, unlike in other cancers in which FOXM1 contributes to chemoresistance 51,52,58,59, sufficient improvement of chemosensitivity by FOXM1 inhibition was not observed in the KS-EMPD-1 cells because every cytotoxic agent used in this experiment (5-FU, CDDP, DTX, and PTX) massively reduced viable cell counts.
A major limitation of this study is the limited number of cell lines used. However, until we established the KS-EMPD-1 cell line 60, there was no cell line available for EMPD. Currently, KE-EMPD-1 is the only cell line available for EMPD.
In conclusion, our findings indicate a significant increase in the expression of FOXM1 in EMPD. Strong and abundant FOXM1 expression was observed in both early and advanced stages of the disease. Further research is needed to explore the potential of FOXM1 as a therapeutic target for treating both early and advanced EMPD.
Methods
Reagents
DMSO (07–4860-5; Sigma-Aldrich, St. Louis, MO) was used as a solvent of chemical compounds. Docetaxel (DTX, 047-31281), paclitaxel (PTX, 163-28163), 5-fluorouracil (5-FU, 068-01401), and cisplatin (CDDP, 033-20091; all purchased from Fujifilm Wako Pure Chemicals, Osaka, Japan) were dissolved in DMSO or saline before being used for the experiments. A potent inhibitor of FOXM1, thiostrepton (T8902; Sigma-Aldrich), was dissolved in DMSO before its use.
Ethical approval
The current study was carried out following the tenets of the Declaration of Helsinki. Immunohistochemical analysis of patients’ samples was approved by the Ethics Committee of Kyushu University Hospital (approval number 30-363, approved on Nov. 27, 2018). Written informed consent was obtained from each participant before their inclusion in the study.
Immunohistochemistry (IHC)
FFPE EMPD tissues were obtained from the archives of Kyushu University Hospital. Prior to this study, secondary EMPD had been carefully excluded by clinical examination and a panel of IHC (CK7, CK20, GCDFP15 and CDX2, etc.). FFPE samples were sliced into 4-μm-thick sections and stained for FOXM1 via slightly modified versions of previously reported methods 73,74,75. These sections were incubated with primary antibody (rabbit anti-human FOXM1, 1:1,000, sc-502; Santa Cruz Biotechnology Inc., Dallas, TX) for 90 min at room temperature and further incubated with secondary antibody (N-Histofine Simple Stain AP MULTI, 414261; Nichirei Biosciences, Tokyo, Japan) for 30 min at room temperature. Next, the sections were treated with FastRed II (415261; Nichirei Biosciences) and counterstained with hematoxylin (30002; Muto Pure Chemicals, Tokyo, Japan). For CK7, antigen was retrieved by protease (715231; Nichirei Biosciences) incubation. Then, we used mouse anti-human CK7 (prediluted by supplier, 713481; Nichirei Biosciences) as the primary antibody followed by N-Histofine Simple Stain MAX-PO MULTI (724152; Nichirei Biosciences) as the secondary antibody. The 3,3′-diaminobenzidine tetrahydrochloride (725191; Nichirei Biosciences) was used as a chromogenic substrate. Stained sections were observed and images were captured using a Nikon ECLIPSE 80i microscope (Nikon, Tokyo, Japan).
Evaluation of FOXM1 IHC staining
The stained samples were independently observed by two dermatologists (T.I. and Y.K.-I.), who were blinded to the patients’ information. Immunoreactivity of FOXM1 was defined as cells showing nuclear staining with or without cytoplasmic staining patterns. Tumors with strong staining intensity in > 10% of the tumor cells were recorded as having positive immunoreactivity, based on a previously reported method 51,52,56,57.
Cell culture
KS-EMPD-1 cells (originally established by Ito et al. 60) were cultured in Endothelial Cell Growth Medium 2 Kit (C-22111; Takara Bio Inc., Tokyo, Japan) supplemented with 10 ng/mL Heregulin β1 (100-03; PeproTech, Cranbury, NJ). NHEKs (00192907; Lonza, Basel, Switzerland) were cultured in KGM-Gold Basal Medium with SingleQuots supplements (00192060; Lonza). Medium was refreshed every 2–3 days and cells were passaged at 80% confluence using 0.05 w/v% trypsin/EDTA (202-16931; Fujifilm Wako Pure Chemicals). Contamination of mycoplasma was tested using CycleavePCR Mycoplasma Detection Kit (CY232; Takara Bio Inc.). Cells were confirmed to be mycoplasma free.
Immunocytochemistry
Immunocytochemistry was performed as described in a previous report 60. The antibodies used were rabbit anti-human FOXM1 antibody (1:100, 5436; Cell Signaling Technology, Danvers, MA) as primary antibody and AlexaFluor®488-conjugated goat anti-rabbit IgG (1:400, A11008; Thermo Fisher Scientific, Waltham, MA) as secondary antibody. Stained samples were then covered with Vectashield mounting medium with DAPI (H-1200; Vector Laboratories Inc., Newark, CA). Images were observed and captured using an EVOS FL imaging system (Thermo Fisher Scientific).
siRNA transfection
siRNA transfection was performed using Lipofectamine RNAiMAX (13778150; Thermo Fisher Scientific), in accordance with the manufacturer’s protocol. Cells were seeded into 6-well plates (2 × 105 cells/well), 12-well plates (1 × 105 cells/well), or 96-well plates (5000 cells/well) and incubated for 24 h at 37 °C in 5% CO2. Diluted negative control siRNA (AM4635; Invitrogen, Waltham, MA) or FOXM1 siRNA (s5250; Invitrogen) was mixed with Opti-MEM I reduced serum medium (31985-070; Thermo Fisher Scientific) and RNAiMAX, and incubated for 20 min at room temperature. The final siRNA concentration was set to 10 nM. The resulting siRNA-Lipofectamine complex was added to the cells and further incubated at 37 °C in 5% CO2.
Cell proliferation assay
Cells were seeded into 96-well plates (5,000 cells/well) and transfected with control or FOXM1 siRNAs as described above and were cultured for 1–5 days. Then, the viable cells were quantitated with Cell Counting Kit-8 (CCK-8, 343-07623; Dojindo, Kumamoto, Japan). On each day, 10 μL of CCK-8 solution was added to each well and the culture plate was incubated for 2–4 h at 37 °C. After the incubation, absorbance at 450 nm was measured using an iMark microplate reader (Bio-Rad Laboratories, Inc., Hercules, CA).
RNA extraction and quantitative reverse-transcription polymerase chain reaction (qRT-PCR)
RNA extraction, cDNA synthesis, and following qRT-PCR were performed as described in a previous report 60. β-Actin (ACTB) was used as a housekeeping gene. The sequences of the primers were as follows: FOXM1, forward 5′-TCTGCCAATGGCAAGGTCTCCT-3′, reverse 5′-CTGGATTCGGTCGTTTCTGCTG-3′; cyclin B1 (CCNB1), forward 5′-GACCTGTGTCAGGCTTTCTCTG-3′, reverse 5′-GGTATTTTGGTCTGACTGCTTGC-3′; and ACTB, forward 5′-ATTGCCGACAGGATGCAGA-3′, reverse 5′-GAGTACTTGCGCTCAGGAGGA-3′.
Protein extraction and western blotting
Western blotting was performed as described in previous publications 76,77,78. Antibodies used were rabbit anti-human FOXM1 (1:1000, 5436; Cell Signaling Technology), rabbit anti-human cyclin B1 (1:1000, 12231, Cell Signaling Technology), and rabbit anti-human β-actin (1:2000, 4970; Cell Signaling Technology). Resultant immunological bands were observed using the ChemiDoc XRS Plus System (Bio-Rad Laboratories). The intensity of the bands was quantified using ImageJ software (National Institutes of Health, Bethesda, MD). Uncropped, full-length original gel images are shown in Supplementary Figs. S1 and S2.
Invasion assay
The invasion of control or FOXM1 siRNA-transfected cells was evaluated with a CytoSelect 24-well cell invasion assay (CBA-110; Cell Biolabs Inc., San Diego, CA). Cells were transfected with siRNA as explained above and at 48 h post-transfection, cells were harvested and used for the invasion assay. Invasion assay was performed evaluated as reported previously 60.
Migration assay
Cell migration assay was performed as described in a previous report with slight modifications 78. Cells were seeded into a 96-well ImageLock tissue culture microplate (1.5 × 104 cells/well; Essen Bioscience, Ann Arbor, MI) pre-coated with type I collagen (Nitta Gelatin Inc., Osaka, Japan) and incubated at 37 °C in 5% CO2. Twenty-four hours after the incubation, cells were transfected with siRNAs as described above. At 48 h, cell monolayers were scratched with a wound-maker (Essen Bioscience) and images of the scratched area were automatically captured every 2 h for a total of 24 h in the IncuCyte Cell Imaging System (Essen Bioscience). The wound area relative to that at 0 h was calculated by IncuCyte software (Essen Bioscience).
Thiostrepton treatment and IC 50 measurement
Cells were seeded into 96-well plates (5,000 cells/well) and incubated at 37 °C in 5% CO2. At 24 h, the medium was replaced with medium containing vehicle (DMSO, 0.1%) or various concentrations of thiostrepton (0.25–10.0 μM) and the cells were further incubated for 72 h at 37 °C in 5% CO2. The range of thiostrepton concentrations was determined based on previous publications with slight modifications 51,52. After the incubation, viable cells were quantified using CCK-8 solution as described above. The IC50 of thiostrepton was calculated using GraphPad Prism 7 software (GraphPad Software, San Diego, CA).
Chemosensitivity assay
The chemosensitivity of FOXM1-knockdown cells to 5-FU, CDDP, DTX, and PTX was tested. Cells were transfected with negative control siRNA or FOXM1 siRNA as described above. Twenty-four hours post-transfection, cells were incubated with the anticancer drugs at 37 °C in 5% CO2. DMSO was used as a solvent of 5-FU, DTX, and PTX, while saline (0.9% sodium chloride solution) was used as a solvent of CDDP. The concentrations of the drugs were determined considering the Cmax of each drug (100 nM, 10 μM, 5 μM, and 1 μM for 5-FU, CDDP, DTX, and PTX, respectively) 60,79. At 72 h of incubation, viable cells were quantified using CCK-8 solution as described above.
Statistical analysis
All experiments were repeated independently at least three times and the quantitative results are shown as mean ± standard deviation (SD). Statistical analysis was performed using GraphPad Prism 7 software. The normality of the obtained data was investigated using the Shapiro–Wilk test and the significance of differences between two groups was tested by Student’s unpaired two-tailed t-test. If the data were not normally distributed, the Mann–Whitney U test was used. To analyze the significance of differences among three or more groups, one-way ANOVA and subsequent multiple comparison test were performed. The Kaplan–Meier method and the log-rank test were used to evaluate disease-specific survival. Disease-specific survival was calculated from the date of the diagnosis to the date of death due to EMPD. Data on patients without death were censored on the date of the last visit. Data on patients who had died of other causes were censored at the time of death. In the survival analysis, patients were stratified by FOXM1 expression of primary tumor. p < 0.05 was defined as statistically significant.
Data availability
All data obtained or analyzed during this study are included in the main text and figures.
References
Kanitakis, J. Mammary and extramammary Paget’s disease. J. Eur. Acad. Dermatol. Venereol. 21(5), 581–590. https://doi.org/10.1111/j.1468-3083.2007.02154.x (2007).
Ito, T., Kaku-Ito, Y. & Furue, M. The diagnosis and management of extramammary Paget’s disease. Expert Rev. Anticancer Ther. 18(6), 543–553. https://doi.org/10.1080/14737140.2018.1457955 (2018) (Epub 2018 Mar 27).
Shepherd, V., Davidson, E. J. & Davies-Humphreys, J. Extramammary Paget’s disease. BJOG 112(3), 273–279. https://doi.org/10.1111/j.1471-0528.2004.00438.x (2005).
Hashimoto, H. & Ito, T. Current management and treatment of extramammary Paget’s disease. Curr. Treat. Options Oncol. 23(6), 818–830. https://doi.org/10.1007/s11864-021-00923-3 (2022) (Epub 2022 Apr 4).
Ishizuki, S. & Nakamura, Y. Extramammary Paget’s disease: Diagnosis, pathogenesis, and treatment with focus on recent developments. Curr. Oncol. 28(4), 2969–2986. https://doi.org/10.3390/curroncol28040260 (2021).
Kibbi, N. et al. Evidence-based clinical practice guidelines for extramammary Paget disease. JAMA Oncol. 8(4), 618–628. https://doi.org/10.1001/jamaoncol.2021.7148 (2022).
Simonds, R. M., Segal, R. J. & Sharma, A. Extramammary Paget’s disease: A review of the literature. Int. J. Dermatol. 58(8), 871–879. https://doi.org/10.1111/ijd.14328 (2019) (Epub 2018 Dec 19).
Nasioudis, D., Bhadra, M. & Ko, E. M. Extramammary Paget disease of the vulva: Management and prognosis. Gynecol. Oncol. 157(1), 146–150. https://doi.org/10.1016/j.ygyno.2019.11.009 (2020) (Epub 2019 Nov 25).
Funaro, D. et al. Extramammary Paget disease: Epidemiology and association to cancer in a Quebec-based population. J. Low. Genit. Tract Dis. 17(2), 167–174. https://doi.org/10.1097/LGT.0b013e31825f4b4f (2013).
Yin, S. et al. Prevalence of extramammary Paget’s disease in urban China: A population-based study. Orphanet. J. Rare Dis. 16(1), 134. https://doi.org/10.1186/s13023-021-01715-6 (2021).
van der Zwan, J. M., Siesling, S., Blokx, W. A., Pierie, J. P. & Capocaccia, R. Invasive extramammary Paget’s disease and the risk for secondary tumours in Europe. Eur. J. Surg. Oncol. 38(3), 214–221. https://doi.org/10.1016/j.ejso.2011.12.008 (2012) (Epub 2012 Jan 14).
Ito, T. et al. Tumor thickness as a prognostic factor in extramammary Paget’s disease. J. Dermatol. 42(3), 269–275. https://doi.org/10.1111/1346-8138.12764 (2015) (Epub 2014 Dec 30).
Ohara, K. et al. A proposal for a TNM staging system for extramammary Paget disease: Retrospective analysis of 301 patients with invasive primary tumors. J. Dermatol. Sci. 83(3), 234–239. https://doi.org/10.1016/j.jdermsci.2016.06.004 (2016) (Epub 2016 Jun 3).
Hatta, N., Yamada, M., Hirano, T., Fujimoto, A. & Morita, R. Extramammary Paget’s disease: Treatment, prognostic factors and outcome in 76 patients. Br. J. Dermatol. 158(2), 313–318. https://doi.org/10.1111/j.1365-2133.2007.08314.x (2008) (Epub 2007 Nov 19).
Cho, W. C. et al. Immunohistochemical expression of TRPS1 in mammary Paget disease, extramammary Paget disease, and their close histopathologic mimics. J. Cutan. Pathol. 50(5), 434–440. https://doi.org/10.1111/cup.14414 (2023) (Epub 2023 Mar 1).
Liu, Y. A. et al. TRPS1 expression in primary and secondary extramammary Paget diseases: An immunohistochemical analysis of 93 cases. Hum. Pathol. 143, 5–9. https://doi.org/10.1016/j.humpath.2023.11.004 (2024) (Epub 2023 Nov 23).
Murata, T. et al. Three-dimensional evaluation of subclinical extension of extramammary Paget disease: Visualization of the histological border and its comparison to the clinical border. Br. J. Dermatol. 177(1), 229–237. https://doi.org/10.1111/bjd.15282 (2017) (Epub 2017 May 4).
Matsuo, K. et al. Surgical margin status and recurrence pattern in invasive vulvar Paget’s disease: A Japanese Gynecologic Oncology Group study. Gynecol. Oncol. 160(3), 748–754. https://doi.org/10.1016/j.ygyno.2020.12.023 (2021) (Epub 2020 Dec 29).
Kaku-Ito, Y. et al. Evaluation of mapping biopsies for extramammary Paget disease: A retrospective study. J. Am. Acad. Dermatol. 78(6), 1171-1177.e4. https://doi.org/10.1016/j.jaad.2017.12.040 (2018) (Epub 2017 Dec 23).
Li, X. et al. PDD-guided tumor excision combined with photodynamic therapy in patients with extramammary Paget’s disease. Photodiagn. Photodyn. Ther. 38, 102841. https://doi.org/10.1016/j.pdpdt.2022.102841 (2022) (Epub 2022 Mar 31).
Lukowiak, T. M. et al. Mohs micrographic surgery for male genital tumors: Local recurrence rates and patient-reported outcomes. J. Am. Acad. Dermatol. 84(4), 1030–1036. https://doi.org/10.1016/j.jaad.2020.11.060 (2021) (Epub 2020 Dec 3).
Hashimoto, H., Kaku-Ito, Y., Furue, M. & Ito, T. Mucosal invasion, but not incomplete excision, has negative impact on long-term survival in patients with extramammary Paget’s disease. Front. Oncol. 11, 642919. https://doi.org/10.3389/fonc.2021.642919 (2021).
Kato, J. et al. Successful TS-1 monotherapy as the second-line treatment for advanced extramammary Paget’s disease: A report of two cases. J. Dermatol. 45(1), 80–82. https://doi.org/10.1111/1346-8138.14017 (2018) (Epub 2017 Sep 11).
Hashimoto, H., Kaku-Ito, Y., Furue, M. & Ito, T. The outcome of chemotherapy for metastatic extramammary Paget’s disease. J. Clin. Med. 10(4), 739. https://doi.org/10.3390/jcm10040739 (2021).
Matsushita, S. et al. Efficacy of S-1 plus docetaxel in the treatment of metastatic extramammary Paget’s disease: A multicentre retrospective study. Br. J. Dermatol. 185(2), 458–460. https://doi.org/10.1111/bjd.20135 (2021) (Epub 2021 May 25).
Richter, C. E. et al. HER-2/NEU overexpression in vulvar Paget disease: The Yale experience. J. Clin. Pathol. 63(6), 544–547. https://doi.org/10.1136/jcp.2010.077446 (2010) (Epub 2010 Apr 23).
Bartoletti, M. et al. Human epidermal growth factor receptor-2 (HER2) is a potential therapeutic target in extramammary Paget’s disease of the vulva. Int. J. Gynecol. Cancer 30(11), 1672–1677. https://doi.org/10.1136/ijgc-2020-001771 (2020) (Epub 2020 Sep 30).
Kimura, T., Akamatsu, Y., Kajihara, I., Fukushima, S. & Ihn, H. Case of metastatic extramammary Paget’s disease treated with trastuzumab-biosimilar monotherapy after S-1 and docetaxel combination chemotherapy. J. Dermatol. 47(1), e1–e2. https://doi.org/10.1111/1346-8138.15096 (2020) (Epub 2019 Sep 30).
Sekiguchi, N. et al. Experiences of trastuzumab plus paclitaxel combination therapy in metastatic human epidermal growth factor receptor 2-positive extramammary Paget’s disease: Four cases and a review. J. Dermatol. 47(11), 1276–1279. https://doi.org/10.1111/1346-8138.15515 (2020) (Epub 2020 Jul 24).
Zattarin, E. et al. Case report: Prolonged clinical benefit with sequential trastuzumab-containing treatments in a patient with advanced extramammary Paget disease of the groin. Front. Oncol. 12, 925551. https://doi.org/10.3389/fonc.2022.925551 (2022).
Liegl, B., Horn, L. C. & Moinfar, F. Androgen receptors are frequently expressed in mammary and extramammary Paget’s disease. Mod. Pathol. 18(10), 1283–1288. https://doi.org/10.1038/modpathol.3800437 (2005).
Diaz-de-Leon, E. et al. Extramammary Paget disease is characterized by the consistent lack of estrogen and progesterone receptors but frequently expresses androgen receptor. Am. J. Clin. Pathol. 113(4), 572–575. https://doi.org/10.1309/P756-XXCB-TV71-U4XV (2000).
Azmahani, A. et al. Androgen receptor, androgen-producing enzymes and their transcription factors in extramammary Paget disease. Hum. Pathol. 46(11), 1662–1669. https://doi.org/10.1016/j.humpath.2015.07.007 (2015) (Epub 2015 Jul 21).
Yamada-Kanazawa, S. et al. Upregulated androgen receptor variant-7 mRNA and protein in extramammary Paget’s disease. J. Eur. Acad. Dermatol. Venereol. 36(9), e724–e726. https://doi.org/10.1111/jdv.18229 (2022) (Epub 2022 May 28).
Goto, H., Sugita, K. & Yamamoto, O. Expression of programmed death-ligand 1 and programmed death-1 in patients with extramammary Paget’s disease. Indian J. Dermatol. 66(2), 169–173. https://doi.org/10.4103/ijd.IJD_341_18 (2021).
Kato, J. et al. Expression of programmed cell death ligand 1 (PD-L1) at in situ and invasive extramammary Paget’s disease and literature review. Austral. J. Dermatol. 62(3), 412–414. https://doi.org/10.1111/ajd.13607 (2021) (Epub 2021 May 12).
Mauzo, S. H. et al. Expression of PD-1 and PD-L1 in extramammary Paget disease: Implications for immune-targeted therapy. Cancers (Basel) 11(6), 754. https://doi.org/10.3390/cancers11060754 (2019).
Hashimoto, H., Kaku-Ito, Y., Oda, Y. & Ito, T. CDK4: A novel therapeutic target for extramammary Paget’s disease. Front. Oncol. 11, 710378. https://doi.org/10.3389/fonc.2021.710378 (2021).
Chang, K. et al. Chemokine receptors CXCR4 and CXCR7 are associated with tumor aggressiveness and prognosis in extramammary Paget disease. J. Cancer 8(13), 2471–2477. https://doi.org/10.7150/jca.19127 (2017).
Kusaba, Y. et al. Intertumor and intratumor heterogeneity of PIK3CA mutations in extramammary Paget’s disease. J. Dermatol. 49(5), 508–514. https://doi.org/10.1111/1346-8138.16343 (2022) (Epub 2022 Mar 6).
Kitamura, S., Yanagi, T., Maeda, T. & Shimizu, H. Drp1 expression levels correlate with clinical stage in extramammary Paget’s disease. J. Eur. Acad. Dermatol. Venereol. 34(9), e510–e513. https://doi.org/10.1111/jdv.16422 (2020) (Epub 2020 Jun 8).
Murata, M., Ito, T., Tanaka, Y., Kaku-Ito, Y. & Furue, M. NECTIN4 expression in extramammary Paget’s disease: Implication of a new therapeutic target. Int. J. Mol. Sci. 21(16), 5891. https://doi.org/10.3390/ijms21165891 (2020).
Hashimoto, H., Tanaka, Y., Murata, M. & Ito, T. Nectin-4: A novel therapeutic target for skin cancers. Curr. Treat. Options Oncol. 23(4), 578–593. https://doi.org/10.1007/s11864-022-00940-w (2022) (Epub 2022 Mar 21).
Ito, T. et al. Trop2 expression in extramammary Paget’s disease and normal skin. Int. J. Mol. Sci. 22(14), 7706. https://doi.org/10.3390/ijms22147706 (2021).
Takeichi, T. et al. Frequent FOXA1-activating mutations in extramammary Paget’s disease. Cancers (Basel) 12(4), 820. https://doi.org/10.3390/cancers12040820 (2020).
Bella, L., Zona, S., Nestal-de-Moraes, G. & Lam, E. W. FOXM1: A key oncofoetal transcription factor in health and disease. Semin. Cancer Biol. 29, 32–39 (2014).
Halasi, M. & Gartel, A. L. Targeting FOXM1 in cancer. Biochem. Pharmacol. 85(5), 644–652 (2013).
Katoh, M., Igarashi, M., Fukuda, H., Nakagama, H. & Katoh, M. Cancer genetics and genomics of human FOX family genes. Cancer Lett. 328(2), 198–206. https://doi.org/10.1016/j.canlet.2012.09.017 (2013) (Epub 2012 Sep 27).
Khan, M. A., Khan, P., Ahmad, A., Fatima, M. & Nasser, M. W. FOXM1: A small fox that makes more tracks for cancer progression and metastasis. Semin. Cancer Biol. 92, 1–15. https://doi.org/10.1016/j.semcancer.2023.03.007 (2023) (Epub 2023 Mar 22).
Bektas, N. et al. Tight correlation between expression of the Forkhead transcription factor FOXM1 and HER2 in human breast cancer. BMC Cancer 8, 42. https://doi.org/10.1186/1471-2407-8-42 (2008).
Ito, T. et al. Prognostic significance of forkhead box M1 (FoxM1) expression and antitumour effect of FoxM1 inhibition in melanoma. Histopathology 69(1), 63–71. https://doi.org/10.1111/his.12909 (2016) (Epub 2016 Jan 11).
Ito, T. et al. Prognostic significance of forkhead box M1 (FOXM1) expression and antitumor effect of FOXM1 inhibition in angiosarcoma. J. Cancer 7(7), 823–830. https://doi.org/10.7150/jca.14461 (2016).
Ha, S. Y. et al. Differential expression of forkhead box M1 and its downstream cyclin-dependent kinase inhibitors p27(kip1) and p21(waf1/cip1) in the diagnosis of pulmonary neuroendocrine tumours. Histopathology 60(5), 731–739. https://doi.org/10.1111/j.1365-2559.2011.04137.x (2012) (Epub 2012 Feb 1).
Yang, D. K. et al. Forkhead box M1 expression in pulmonary squamous cell carcinoma: Correlation with clinicopathologic features and its prognostic significance. Hum. Pathol. 40(4), 464–470. https://doi.org/10.1016/j.humpath.2008.10.001 (2009) (Epub 2009 Jan 3).
Pilarsky, C., Wenzig, M., Specht, T., Saeger, H. D. & Grützmann, R. Identification and validation of commonly overexpressed genes in solid tumors by comparison of microarray data. Neoplasia 6(6), 744–750. https://doi.org/10.1593/neo.04277 (2004).
Nakamura, S. et al. The FOXM1 transcriptional factor promotes the proliferation of leukemia cells through modulation of cell cycle progression in acute myeloid leukemia. Carcinogenesis 31(11), 2012–2021. https://doi.org/10.1093/carcin/bgq185 (2010) (Epub 2010 Sep 7).
Kuda, M. et al. FOXM1 expression in rhabdomyosarcoma: A novel prognostic factor and therapeutic target. Tumour Biol. 37(4), 5213–5223. https://doi.org/10.1007/s13277-015-4351-9 (2016) (Epub 2015 Nov 9).
Maekawa, A. et al. Expression of Forkhead box M1 in soft tissue leiomyosarcoma: Clinicopathologic and in vitro study using a newly established cell line. Cancer Sci. 107(1), 95–102. https://doi.org/10.1111/cas.12846 (2016) (Epub 2016 Jan 12).
Maekawa, A. et al. Prognostic significance of FOXM1 expression and antitumor effect of FOXM1 inhibition in synovial sarcomas. BMC Cancer 16, 511. https://doi.org/10.1186/s12885-016-2542-4 (2016).
Ito, T., Tanaka, Y., Ichiki, T., Kaku-Ito, Y. & Nakahara, T. KS-EMPD-1: A novel cell line of primary extramammary Paget’s disease. Hum. Cell 36(5), 1813–1829. https://doi.org/10.1007/s13577-023-00951-1 (2023) (Epub 2023 Jul 11).
Liang, S. K. et al. FOXM1 is required for small cell lung cancer tumorigenesis and associated with poor clinical prognosis. Oncogene 40(30), 4847–4858. https://doi.org/10.1038/s41388-021-01895-2 (2021) (Epub 2021 Jun 21).
Sher, G. et al. Dysregulated FOXM1 signaling in the regulation of cancer stem cells. Semin. Cancer Biol. 86(Pt 3), 107–121. https://doi.org/10.1016/j.semcancer.2022.07.009 (2022) (Epub 2022 Aug 2).
Gartel, A. L. FOXM1 in cancer: Interactions and vulnerabilities. Cancer Res. 77(12), 3135–3139. https://doi.org/10.1158/0008-5472.CAN-16-3566 (2017) (Epub 2017 Jun 5).
Yang, E. J. et al. Co-inhibition of ATM and ROCK synergistically improves cell proliferation in replicative senescence by activating FOXM1 and E2F1. Commun. Biol. 5(1), 702. https://doi.org/10.1038/s42003-022-03658-5 (2022).
Ilaslan, E. et al. Distinct roles of NANOS1 and NANOS3 in the cell cycle and NANOS3-PUM1-FOXM1 axis to control G2/M phase in a human primordial germ cell model. Int. J. Mol. Sci. 23(12), 6592. https://doi.org/10.3390/ijms23126592 (2022).
Chen, X., Chen, J., Yu, X., Lin, G. & Chen, T. FOXM1 promotes malignant proliferation of esophageal squamous cell carcinoma through transcriptional activating CDC6. DNA Cell Biol. 41(7), 671–682. https://doi.org/10.1089/dna.2022.0169 (2022) (Epub 2022 May 30).
Gu, C. et al. Upregulation of FOXM1 leads to diminished drug sensitivity in myeloma. BMC Cancer 18(1), 1152. https://doi.org/10.1186/s12885-018-5015-0 (2018).
Luo, W., Gao, F., Li, S. & Liu, L. FoxM1 promotes cell proliferation, invasion, and stem cell properties in nasopharyngeal carcinoma. Front. Oncol. 8, 483. https://doi.org/10.3389/fonc.2018.00483 (2018).
Luo, X. et al. FOXM1 promotes invasion and migration of colorectal cancer cells partially dependent on HSPA5 transactivation. Oncotarget 7(18), 26480–26495. https://doi.org/10.18632/oncotarget.8419 (2016).
Zanin, R. et al. HMGA1 promotes breast cancer angiogenesis supporting the stability, nuclear localization and transcriptional activity of FOXM1. J. Exp. Clin. Cancer Res. 38(1), 313. https://doi.org/10.1186/s13046-019-1307-8 (2019).
Huang, C. et al. FoxM1 induced paclitaxel resistance via activation of the FoxM1/PHB1/RAF-MEK-ERK pathway and enhancement of the ABCA2 transporter. Mol. Ther. Oncolytics 14, 196–212. https://doi.org/10.1016/j.omto.2019.05.005 (2019).
https://www.dechra-us.com/our-products/us/companion-animal/cat/prescription/animax-ointment-nystatin-neomycin-sulfate-thiostrepton-triamcinolone-acetonide. (2023, accessed 27 Aug 2023).
Tanaka, Y. et al. Human epidermal growth factor receptor 3 serves as a novel therapeutic target for acral melanoma. Cell Death Discov. 9(1), 54. https://doi.org/10.1038/s41420-023-01358-5 (2023).
Tanaka, Y., Murata, M., Shen, C. H., Furue, M. & Ito, T. NECTIN4: A novel therapeutic target for melanoma. Int. J. Mol. Sci. 22(2), 976. https://doi.org/10.3390/ijms22020976 (2021).
Murata, M. et al. OVOL2-mediated ZEB1 downregulation may prevent promotion of actinic keratosis to cutaneous squamous cell carcinoma. J. Clin. Med. 9(3), 618. https://doi.org/10.3390/jcm9030618 (2020).
Tanaka, Y., Uchi, H. & Furue, M. Antioxidant cinnamaldehyde attenuates UVB-induced photoaging. J. Dermatol. Sci. 96(3), 151–158. https://doi.org/10.1016/j.jdermsci.2019.11.001 (2019) (Epub 2019 Nov 6).
Tanaka, Y., Ito, T., Tsuji, G. & Furue, M. Baicalein inhibits benzo[a]pyrene-induced toxic response by downregulating Src phosphorylation and by upregulating NRF2-HMOX1 system. Antioxid. (Basel) 9(6), 507. https://doi.org/10.3390/antiox9060507 (2020).
Tanaka, Y., Uchi, H., Ito, T. & Furue, M. Indirubin-pregnane X receptor-JNK axis accelerates skin wound healing. Sci. Rep. 9(1), 18174. https://doi.org/10.1038/s41598-019-54754-2 (2019).
Liston, D. R. & Davis, M. Clinically relevant concentrations of anticancer drugs: A guide for nonclinical studies. Clin. Cancer Res. 23(14), 3489–3498. https://doi.org/10.1158/1078-0432.CCR-16-3083 (2017).
Acknowledgements
We would like to show our appreciation to all of the patients who participated in this research. We also thank Ms. Mieko Ogawa for technical support with the immunostaining of patient samples.
Funding
This work was supported by JSPS KAKENHI, Grant number JP 22K15543 (T.I.).
Author information
Authors and Affiliations
Contributions
Research supervision, research design, data collection, data analysis, manuscript writing and revision: T.I. Data collection, data analysis, manuscript revision: Y.T. Research design, data collection, data analysis, manuscript writing and revision: Y.K.-I. Preparation of patient samples, manuscript revision: Y.O. Research supervision, manuscript revision: T.N.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ito, T., Tanaka, Y., Kaku-Ito, Y. et al. FOXM1: a new therapeutic target of extramammary Paget disease. Sci Rep 14, 4048 (2024). https://doi.org/10.1038/s41598-024-54773-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-54773-8
Keywords
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.