A channel in our cancer community includes background on the PCAWG project and behind-the -paper insights.
A guide to access all the datasets and tools generated by the ICGC/PCAWG consortium
Landing Page July 25, 2019 |
PCAWG Landing Page This is the recommended starting point for users wishing to access the PCAWG data used in the February 2020 publications in Nature and affiliated journals. It provides browsing, download and usage information for frozen PCAWG data files as of July 25, 2019. |
|
Data Portal Open & controlled access |
ICGC Data Portal This is the main entry point for accessing PCAWG datasets using a single uniform web interface and a high performance data download client. Most of the data is open access and can be accessed without prior approval. Researchers wishing to access data that is potentially identifiable, such as germline variants, must obtain approval from the ICGC and TCGA data access committees. |
|
Cloud Portal Open & controlled access |
Cancer Genome Collaboratory https://cancercollaboratory.org/ Academic compute cloud-based access to the PCAWG data set, excepting the TCGA-originated portion of the controlled data tier. |
|
Cloud Portal Controlled access |
The Bionimbus Protected Data Cloud https://bionimbus-pdc.opensciencedatacloud.org Academic compute cloud-based access to the TCGA-originated portion of the controlled data tier. |
|
Data Portal Open access |
UCSC Xena UCSC Xena's unique visualisations and analyses allow users to integrate the many diverse types of omics data generated by the PCAWG consortium, including copy number, gene expression, gene fusion, promoter usage, simple somatic mutations, large somatic structural variation, mutational signatures and phenotypic data. |
|
Data Portal Open access |
Expression Atlas https://www.ebi.ac.uk/gxa/experiments?experimentSet=Pan-Cancer Expression Atlas is an open science resource that gives users a powerful way to find information about gene and protein expression. It enables queries across different tissues, cell types, developmental stages and experimental conditions, across thousands of publicly available RNA-Seq, microarray and proteomics data sets. |
|
Data Portal Open access |
PCAWG-Scout PCAWG-Scout provides a framework for ‘omics workflow and website templating to make on-demand, in-depth analyses over the open access PCAWG data. |
|
Data Portal Open access |
Chromothripsis Explorer http://compbio.med.harvard.edu/chromothripsis/ The Chromothripsis Explorer portal enables exploration of patterns of chromothripsis in the PCAWG dataset. Users can explore properties such as purity and ploidy, and interact with circos plots for all tumors. |
|
Data Portal Open access |
Cancer LncRNA Census The Cancer LncRNA Census is an ongoing effort to identify and catalogue lncRNA genes which have been causally implicated in cancer. |
|
Software Tool Open source |
PCAWG Core Pipelines https://dockstore.org/organizations/PCAWG/collections/PCAWG All core alignment, QC and variation-calling pipelines used by PCAWG have been packaged as portable binaries using Docker, and described using workflow description languages. The binaries, source code, and documentation can be found at the Dockstore site using the URL above. |
|
Software Tool Open source |
Overture Suite Overture comprises a set of open source tools for efficiently managing large genomic data sets and transferring them efficiently and reliably across the Internet. |
|
Software Tool Open source |
Butler https://github.com/llevar/butlerl Butler is a workflow frameworkthat facilitates large-scale genomic analyses on public and academic clouds while offering comprehensive error detection and self-healing capabilities. |
|
Software Tool Open source |
SVClone https://github.com/mcmero/SVclone SVclone is a computational method for inferring the cancer cell fraction of structural variant (SV) breakpoints from whole-genome sequencing data. |
|
Software Tool Open source |
DriverPower https://github.com/smshuai/DriverPower DriverPower is a tool used to discover potential coding and non-coding cancer driver elements from tumor whole-genome or whole-exome somatic mutation sets. |
|
Software Tool Open source |
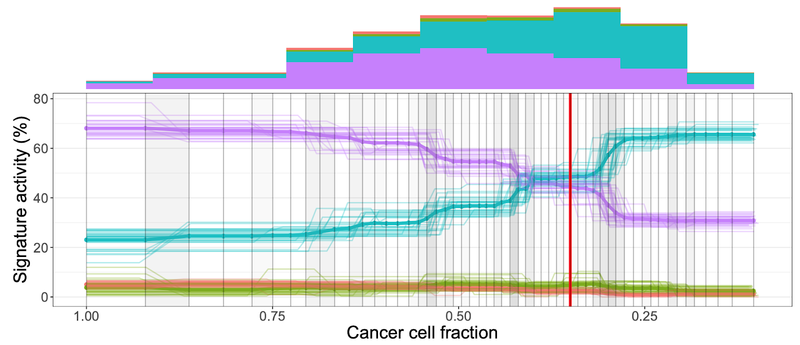
TrackSig https://github.com/morrislab/TrackSig TrackSig is a computational framework to infer changes in somatic mutational signatures over time |
|