Abstract
Organelles use specialized molecules to regulate their essential cellular processes. However, systematically elucidating the subcellular distribution and function of molecules such as long non-coding RNAs (lncRNAs) in cellular homeostasis and diseases has not been fully achieved. Here, we reveal the diverse and abundant subcellular distribution of organelle-associated lncRNAs from mitochondria, lysosomes and endoplasmic reticulum. Among them, we identify the mitochondrially localized lncRNA growth-arrest-specific 5 (GAS5) as a tumour suppressor in maintaining cellular energy homeostasis. Mechanistically, energy-stress-induced GAS5 modulates mitochondrial tricarboxylic acid flux by disrupting metabolic enzyme tandem association of fumarate hydratase, malate dehydrogenase and citrate synthase, the canonical members of the tricarboxylic acid cycle. GAS5 negatively correlates with levels of its associated mitochondrial metabolic enzymes in tumours and benefits overall survival in individuals with breast cancer. Together, our detailed annotation of subcellular lncRNA distribution identifies a functional role for lncRNAs in regulating cellular metabolic homeostasis, highlighting organelle-associated lncRNAs as potential clinical targets to manipulate cellular metabolism and diseases.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All sequencing data used in this study have been deposited in the NCBI under accession number ‘BioProject: PRJNA594757’. The source data for gels, blots and column charts are provided in the source data. The original images for IHC and fluorescence staining are uploaded at https://doi.org/10.6084/m9.figshare.13176239.v1. Source data are provided with this paper. Further information and requests for resources and reagents should be directed to and will be fulfilled by the corresponding author.
References
Birsoy, K. et al. An essential role of the mitochondrial electron transport chain in cell proliferation is to enable aspartate synthesis. Cell 162, 540–551 (2015).
Ballabio, A. & Bonifacino, J. S. Lysosomes as dynamic regulators of cell and organismal homeostasis. Nat. Rev. Mol. Cell Biol. 21, 101–118 (2020).
Perera, R. M. & Zoncu, R. The lysosome as a regulatory hub. Annu Rev. Cell Dev. Biol. 32, 223–253 (2016).
Boyman, L., Karbowski, M. & Lederer, W. J. Regulation of mitochondrial ATP production: Ca2+ signaling and quality control. Trends Mol. Med. 26, 21–39 (2019).
Stevenson, J., Huang, E. Y. & Olzmann, J. A. Endoplasmic reticulum-associated degradation and lipid homeostasis. Annu. Rev. Nutr. 36, 511–542 (2016).
Zong, Y. et al. Hierarchical activation of compartmentalized pools of AMPK depends on severity of nutrient or energy stress. Cell Res. 29, 460–473 (2019).
Mottis, A., Herzig, S. & Auwerx, J. Mitocellular communication: shaping health and disease. Science 366, 827–832 (2019).
Efeyan, A., Zoncu, R. & Sabatini, D. M. Amino acids and mTORC1: from lysosomes to disease. Trends Mol. Med. 18, 524–533 (2012).
Thul, P. J. et al. A subcellular map of the human proteome. Science 356, eaal3321 (2017).
Zhang, C. S. et al. Fructose-1,6-bisphosphate and aldolase mediate glucose sensing by AMPK. Nature 548, 112–11 (2017).
Wyant, G. A. et al. mTORC1 activator SLC38A9 is required to efflux essential amino acids from lysosomes and use protein as a nutrient. Cell 171, 642–654 (2017).
Wang, S. et al. Metabolism. Lysosomal amino acid transporter SLC38A9 signals arginine sufficiency to mTORC1. Science 347, 188–194 (2015).
Wagner, G. R. et al. A class of reactive Acyl-CoA species reveals the non-enzymatic origins of protein acylation. Cell Metab. 25, 823–837 (2017).
Lin, A. et al. The LINK-A lncRNA interacts with PI(3,4,5)P3 to hyperactivate AKT and confer resistance to AKT inhibitors. Nat. Cell Biol. 19, 238–251 (2017).
Lin, A. et al. The LINK-A lncRNA activates normoxic HIF1α signalling in triple-negative breast cancer. Nat. Cell Biol. 18, 213–224 (2016).
Sang, L. J. et al. LncRNA CamK-A regulates Ca2+-signaling-mediated tumor microenvironment remodeling. Mol. Cell 72, 71–83 (2018).
Ma, Y., Zhang, J., Wen, L. & Lin, A. Membrane-lipid associated lncRNA: a new regulator in cancer signaling. Cancer Lett. 419, 27–29 (2018).
Chen, X. et al. Visualizing RNA dynamics in live cells with bright and stable fluorescent RNAs. Nat. Biotechnol. 37, 1287–1293 (2019).
Hocine, S., Raymond, P., Zenklusen, D., Chao, J. A. & Singer, R. H. Single-molecule analysis of gene expression using two-color RNA labeling in live yeast. Nat. Methods 10, 119–121 (2013).
Chen, K. H., Boettiger, A. N., Moffitt, J. R., Wang, S. & Zhuang, X. RNA imaging. Spatially resolved, highly multiplexed RNA profiling in single cells. Science 348, aaa6090 (2015).
Mercer, T. R. et al. The human mitochondrial transcriptome. Cell 146, 645–658 (2011).
Kino, T., Hurt, D. E., Ichijo, T., Nader, N. & Chrousos, G. P. Non-coding RNA gas5 is a growth arrest- and starvation-associated repressor of the glucocorticoid receptor. Sci. Signal 3, ra8 (2010).
Patel, N., Murr, M., Lui, A., Shi, Y. & Cai, J. OR01-6 lncRNA GAS5 directed therapeutic increases insulin receptor expression in adipocytes. J. Endocr. Soc. 3 (Suppl. 1), OR01-6 (2019).
Zhao, H. et al. Lowly-expressed lncRNA GAS5 facilitates progression of ovarian cancer through targeting miR-196-5p and thereby regulating HOXA5. Gynecol. Oncol. 151, 345–355 (2018).
Yoshida, H. ER stress and diseases. FEBS J. 274, 630–658 (2007).
Bethune, J., Jansen, R. P., Feldbrugge, M. & Zarnack, K. Membrane-associated RNA-binding proteins orchestrate organelle-coupled translation. Trends Cell Biol. 29, 178–188 (2019).
Fazal, F. M. et al. Atlas of subcellular RNA localization revealed by APEX-seq. Cell 178, 473–490 e426 (2019).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Weinberg, S. E. & Chandel, N. S. Targeting mitochondria metabolism for cancer therapy. Nat. Chem. Biol. 11, 9–15 (2015).
Xing, Z. et al. LncRNA directs cooperative epigenetic regulation downstream of chemokine signals. Cell 159, 1110–1125 (2014).
Nelson, D. L., Cox, M. M. & Lehninger, A. L. Lehninger Principles of Biochemistry 7th edn (Freeman and Company, 2017).
Lv, L. et al. Acetylation targets the M2 isoform of pyruvate kinase for degradation through chaperone-mediated autophagy and promotes tumor growth. Mol. Cell 42, 719–730 (2011).
Someya, S. et al. Sirt3 mediates reduction of oxidative damage and prevention of age-related hearing loss under caloric restriction. Cell 143, 802–812 (2010).
Zhao, S. et al. Regulation of cellular metabolism by protein lysine acetylation. Science 327, 1000–1004 (2010).
Vander Heiden, M. G., Cantley, L. C. & Thompson, C. B. Understanding the Warburg effect: the metabolic requirements of cell proliferation. Science 324, 1029–1033 (2009).
Mourtada-Maarabouni, M., Pickard, M. R., Hedge, V. L., Farzaneh, F. & Williams, G. T. GAS5, a non-protein-coding RNA, controls apoptosis and is downregulated in breast cancer. Oncogene 28, 195–208 (2009).
Graham, J. M. Biological Centrifugation (Bios, 2001).
Work, T. S. & Work, E. Laboratory Techniques in Biochemistry and Molecular Biology (North-Holland Pub. Co., 1969).
Kaewsapsak, P., Shechner, D. M., Mallard, W., Rinn, J. L. & Ting, A. Y. Live-cell mapping of organelle-associated RNAs via proximity biotinylation combined with protein-RNA crosslinking. eLife 6, e29224 (2017).
Chen, W. W., Freinkman, E., Wang, T., Birsoy, K. & Sabatini, D. M. Absolute quantification of matrix metabolites reveals the dynamics of mitochondrial metabolism. Cell 166, 1324–1337 (2016).
Ryan, D. G. et al. Coupling Krebs cycle metabolites to signalling in immunity and cancer. Nat. Metab. 1, 16–33 (2019).
Schneider, C., King, R. M. & Philipson, L. Genes specifically expressed at growth arrest of mammalian cells. Cell 54, 787–793 (1988).
Yu, X. & Li, Z. Long non-coding RNA growth arrest-specific transcript 5 in tumor biology. Oncol. Lett. 10, 1953–1958 (2015).
Wang, T. et al. O-GlcNAcylation of fumarase maintains tumour growth under glucose deficiency. Nat. Cell Biol. 19, 833–843 (2017).
Wang, Y. et al. KAT2A coupled with the alpha-KGDH complex acts as a histone H3 succinyltransferase. Nature 552, 273–277 (2017).
Zhang, X. C. et al. YY1/LncRNA GAS5 complex aggravates cerebral ischemia/reperfusion injury through enhancing neuronal glycolysis. Neuropharmacology 158, 107682 (2019).
Xiao, Z., Dai, Z. & Locasale, J. W. Metabolic landscape of the tumor microenvironment at single-cell resolution. Nat. Commun. 10, 3763 (2019).
Etchegaray, J. P. & Mostoslavsky, R. Interplay between metabolism and epigenetics: a nuclear adaptation to environmental changes. Mol. Cell 62, 695–711 (2016).
Pertea, M., Kim, D., Pertea, G. M., Leek, J. T. & Salzberg, S. L. Transcript-level expression analysis of RNA-seq experiments with HISAT, StringTie and Ballgown. Nat. Protoc. 11, 1650–1667 (2016).
Sun, L. et al. Utilizing sequence intrinsic composition to classify protein-coding and long non-coding transcripts. Nucleic Acids Res. 41, e166 (2013).
Kong, L. et al. CPC: assess the protein-coding potential of transcripts using sequence features and support vector machine. Nucleic Acids Res. 35, W345–W349 (2007).
The Gene Ontology, C. The gene ontology resource: 20 years and still GOing strong. Nucleic Acids Res. 47, D330–D338 (2019).
Kanehisa, M. et al. From genomics to chemical genomics: new developments in KEGG. Nucleic Acids Res. 34, D354–D357 (2006).
Li, C. et al. A ROR1–HER3–lncRNA signalling axis modulates the Hippo–YAP pathway to regulate bone metastasis. Nat. Cell Biol. 19, 106–119 (2017).
Acknowledgements
We thank T.-H. Zhou (Zhejiang University), J. Huang (Zhejiang University) and P.-L. Xu (Zhejiang University) for their support and suggestions during this study. We thank J.-H. Han (Xiamen University) for gifting CS and FH template vectors. We thank X. Li (Westlake University) for gifting SIRT vectors and assistance with protein MS analysis. We thank H.-L. Piao (Chinese Academy of Science) for metabolite analysis. We also thank Cipher Gene for support in generating and processing RNA-sequencing data and for help with data interpretation. This work was supported in part by the National Natural Science Foundation of China (81672791 and 81872300), and Zhejiang Provincial Natural Science Fund for Distinguished Young Scholars of China (LR18C060002; to A.L.)
Author information
Authors and Affiliations
Contributions
A.L. and L.S. conceived and designed the research. L.S., H.-Q.J. and Z.Y. performed most of the biochemical, molecular experiments and bioinformatics analysis, with assistance from Q.G., Z.Z., F.L., L.Y., H.G., C.S., L.Q., Hui Chen, Hao Chen, M.W., R.L. and Q.Z. H.-Q.J. performed ascertainment and processing of clinical specimens. Z.Y., L.Q. and W.Y. conducted the bioinformatics analysis. L.S. and F.L. performed xenograft experiments and IHC analyses. T.Z., W.W., J.L., J.S., L.W., Q.Y., H-Q.J. and H.P. contributed to discussion and data interpretation. T.Z., W.W. and J.L. edited the manuscript. A.L. initiated and supervised the project. A.L. and L.S. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Primary Handling Editor: George Caputa. Nature Metabolism thanks Jan-Wilhelm Kornfeld and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 The landscape of subcellular lncRNAs is established and qualified.
(a) Centrifugation isolated light mitochondria fraction (LMF) from HEK293T cells was smeared and fixed on the glass slide followed by immunofluorescence detection using mitochondria marker Tom20 and lysosome marker Lamp2. Mitochondria was stained in red (anti rabbit, Alex 569) and lysosome in green (anti mouse, Alex 488). Scale bar, 20 µm. (b-c) Mitochondria and lysosome were isolated from HEK293T cells by Tom20 or Lamp1 antibody captured protein A/G magnetic beads. Mitochondria or lysosome-captured beads were incubated with mitotracker or lysotracker and smeared on the glass slide to image the colocalization of mitotracker (b) or lysotracker (c) red to beads. Scale bar, 10 μm. (d) Technical replicates showed by scatter plots. The correlation between biological replicates of each group was analyzed (R > 0.95, P < 0.05). The expression level was calculated by log2(fpkm+1). Pearson’s correlation coefficient followed by Student’s t-distribution. (e) PCA analysis for RNAs. LMF, mitochondrial, lysosome and endoplasmic reticulum relatively cluster together, separately from total group. (f) Violin map for each organelle. The expression value (Y axis) was calculated by log10(fpkm+1). The sequenced genes’ pattern of all samples was shown. (g) Circos plot showing the co-localization of mRNAs to multiple locations. (h) Venn map for overlapping and specific mRNA in organelle. (i) The number of organelles-associated mRNAs (P value = Wald test P-value. P > 0.05, foldchange > 1.5) was calculated in each classified GO terms. CC: cell component; TF: transcription factor. (j) The number of organelles-associated lncRNAs (P value = Wald test P-value. P > 0.05, foldchange > 1.5) was calculated in each classified GO terms. LncRNAs’ function was determined by the function of in cis associated neighbor genes (upstream or downstream 100 K from lncRNA genome location). CC: cell component; TF: transcription factor. (k) The number of lncRNAs that significantly enriched (Hypergeometric test, P < 0.05) in classified KEGG terms was shown. LncRNAs’ function was determined by the function of in cis associated neighbor genes (upstream or downstream 100 K from lncRNA genome location). CC: cell component; TF: transcription factor. (l) The number of organelles (lysosome, mitochondria and ER)-associated mRNAs that involved in significantly changed KEGG pathway (Hypergeometric test, P < 0.05).
Extended Data Fig. 2 Organelle-associated functional lncRNAs are identified and validated.
(a-c) The indicated candidates of mitochondria (a), lysosome (b) and ER (c) were verified by organelle purification coupled with RT-qPCR. The positive control of mitochondria (ATP8, MT-RNR1), lysosome (LAMP2) and ER (FGF2, TJP1) was shown as red column; and the negative control (GAPDH, U6, MALAT1, XIST) was shown as white column; the confirmed candidates were shown as blue column; other candidates were shown as grey column. The relative enrichment of each RNAs was calculated by 2^-(CtMito-CtTotal) followed by normalizing all ratio value to the GAPDH in control group (the first GAPDH column). Cut-off line was thereby determined by the relative GAPDH enrichment level. Data are presented as mean values ± S.D., n = 3 biologically independent experiments. (d-e) The knockdown efficiency of siRNA interference for the indicated mitochondria (d) and lysosome (e) candidates was determined by RT-qPCR. Data are presented as mean values ± S.D., n = 3 biologically independent experiments, Two-sided Student’s t-test, **P < 0.01, ***P < 0.001. (f) Immunoblotting detection of AMPK activation by measuring the p-AMPK ratio in HEK293T. Cells were interfered by lysosome candidate’s siRNA and treated with 2-hour glucose starvation. (g) Relative cellular ADP/ATP ratio was determined in HEK293T cells by mitochondrial candidate’s siRNA screening. Each candidate had two replicated siRNA, and three biological duplication for each siRNA. Data are presented as mean values ± S.D., n = 6 biologically independent experiments, Two-sided Student’s t-test, **P < 0.01, ***P < 0.001.
Extended Data Fig. 3 Mitochondria associated lncRNA GAS5 is identified and characterized.
(a) Lysosome fraction of HEK293T was isolated and the relative GAS5 enrichment was detected along with the cytosol marker GAPDH, lysosome marker LAMP1 and nucleus marker XIST, by RT-qPCR assay. Data are presented as mean values ± S.D., n = 3 biologically independent experiments. (b) Endoplasmic reticulum (ER) fraction of HEK293T was isolated and the relative GAS5 enrichment was detected along with the cytosol marker GAPDH, ER marker PD-L1, mitochondria marker ATP8 and nucleus marker XIST, by RT-qPCR assay. Data are presented as mean values ± S.D., n = 3 biologically independent experiments. (c) Cellular GAS5 expression levels under gradient glucose concentrations (0–25 mM) treatment was detected in HEK293T cells. Data are presented as mean values ± S.D., n = 3 biologically independent experiments, Two-sided Student’s t-test, *P < 0.05, **P < 0.01, ***P < 0.001. (d) GAS5 expression level in 12-hour FBS starved or not HEK293T cells was detected by RT-qPCR. Data are presented as mean values ± S.D., n = 3 biologically independent experiments, Two-sided Student’s t-test. (e) Confirmation of HEK293T GAS5 knockdown cell lines by RT-qPCR. Data are presented as mean values ± S.D., n = 3 biologically independent experiments, Two-sided Student’s t-test, **P < 0.01. (f) Oxygen consumption rate (OCR) profile was monitored in empty vector control (EV) or GAS5 overexpressed HEK293T cells with a Seahorse XF24 analyzer for 100 min. The indicated metabolic inhibitors were injected sequentially at different time points as indicated. Data are presented as mean values ± S.D., n = 3 biologically independent experiments. (g) Cellular NADH/NAD+ ratio was detected in GAS5 knocked down HEK293T under 8-hour glucose starvation. Data are presented as mean values ± S.D., n = 3 biologically independent experiments, Two-sided Student’s t-test, *P < 0.05, **P < 0.01. (h) Mitochondria were isolated from GAS5 knocked down HEK293T cells and mitochondrial NADH/NAD+ ratio was measured under 8-hour glucose starvation. Data are presented as mean values ± S.D., n = 3 biologically independent experiments, Two-sided Student’s t-test, **P < 0.01. (i) Electrophoresis of in vitro transcribed biotin labeled GAS5 sense (Sen.) and anti-sense (A.S.) transcripts. (j) RNA immunoprecipitation (RIP) assay was performed using FLAG empty vector (EV) or FLAG-MDH2 transduced HEK293T cells under 8-hour glucose starvation or not. Data are presented as mean values ± S.D., n = 3 biologically independent experiments, Two-sided Student’s t-test, *P < 0.05. (k-l) Generation of a standard curve for calculating GAS5 copy number. pGEM-T easy-GAS5 vector (a range of amounts from 10 copies to 108 copies) was used for standard curve generation in real-time PCR. The resultant CT values decreased linearly with increasing GAS5 copy number (log10) (k). GAS5 copy number was determined in HEK293T, MDA-MB-231 and MDA-MB-468 cell lines (l). Data are presented as mean values ± S.D., n = 3 biologically independent experiments. (m) The secondary structure of GAS5 was predicted and the stem loop region was separately marked as Loop1 (L1, 1–249 nt), Loop2 (L2, 269–465 nt) and Loop3 (L3, 546–634 nt). The GAS5 mutants D1, D2 and D3 were generated by deleted the corresponding loop region. (n) IB detection of His-MBP-MDH2 protein retrieved by in vitro-transcribed biotinylated GAS5 full-length (FL), GAS5-D2 (ΔL2, Δ269–465 nt) and GAS5-Loop2 (L2, 269–465 nt only). The input of biotin-RNAs was detected by dot blot using streptavidin-HRP. (o) Confirmation of HEK293T GAS5 knockout (GAS5-KO) cell lines by RT-qPCR. Data are presented as mean values ± S.D., n = 3 biologically independent experiments, Two-sided Student’s t-test, ***P < 0.001. (p) Confirmation of the GAS5 expression level in GAS5-FL, D1, D2 or D3 rescued GAS5-KO HEK293T cells, by RT-qPCR detection. Data are presented as mean values ± S.D., n = 3 biologically independent experiments.
Extended Data Fig. 4 GAS5 regulates the FH-MDH2-CS tandem association to modulate mitochondrial metabolism.
(a) Immunofluorescence staining of MDH2 (Alexfluo594 red) and mitochondria marker Tom20 (Alexfluo488 green) in HEK293T cells revealed their consistent mitochondria colocalization. Scale bar for 20 μm. (b-c) Immunofluorescence staining of CS (b) or FH (c) (Alexfluo488 green) with mitochondria marker COX IV (Alexfluo594 red) in HEK293T revealed their consistent mitochondria colocalization. Scale bar for 20 μm. (d) Immunofluorescence staining of MDH2 (Alexfluo488 green) and CS (Alexfluo594 red) in HEK293T revealed their consistent colocalization. Scale bar for 20 μm. (e) Immunofluorescence staining of MDH2 (Alexfluo594 red) and FH (Alexfluo488 green) in HEK293T revealed their consistent colocalization. Scale bar for 20 μm. (f) Immunofluorescence staining of MDH2 (Alexfluo488 green) and CS (Alexfluo594 red) in mtagBFP2-FH transduced HEK293T revealed their consistent colocalization. Scale bar for 20 μm. (g) The percentage of mitochondria-colocalized MDH2, CS or FH per HEK293T cell was revealed by Manders’ Colocalization Coefficients (MCC) of MDH2-Tom20, CS-COX IV and FH-COX IV separately, corresponding to fluorescence images (a-c). Data are presented as mean values ± S.D., n = 10 biologically independent cells. (h) The percentage of MDH2-colocalized CS or FH per HEK293T cell was revealed by Manders’ Colocalization Coefficients (MCC) of MDH2-CS, MDH2-FH separately, corresponding to fluorescence images (d-e). Data are presented as mean values ± S.D., n = 10 biologically independent cells. (i) The percentage of mtagBFP2-FH-colocalized MDH2 or CS per HEK293T cell was revealed by Manders’ Colocalization Coefficients (MCC) of mtagBFP2-FH-MDH2, mtagBFP2-CS separately, corresponding to the fluorescence image (f). Data are presented as mean values ± S.D., n = 10 biologically independent cells. (j) SFB-MDH2 and Myc-CS or FH were co-overexpressed in HEK293T cells. Co-IP assay was performed using S-protein beads to detect MDH2-CS or MDH2-FH interaction. (k) The schematic for MDH2 deletion mutant generation. The substrate binding sites were contained in 99–251 amino acids. KRD for lysine (K) rich domain. MLS for mitochondria location signal peptide. (l) SFB-MDH2 WT and its deletion mutants (MDH2-D1, D2, D3) were co-overexpressed with Myc-CS in HEK293T cells. Co-IP was performed using S-protein beads, and the co-precipitated Myc-CS was detected by Myc-tag antibody immunoblot. (m) SFB-MDH2 and Myc-CS were co-overexpressed in HEK293T cells. MDH2-CS interaction was detected by co-IP coupled IB under the indicated gradient glucose concentration treatment for 8 hours. (n) FLAG-FH and Myc-MDH2 were co-overexpressed in HEK293T cells. MDH2-FH interaction was detected by co-IP coupled IB under indicated gradient glucose concentration treatment for 8 hours. (o) In vitro MDH2 enzyme activity assay was performed with additional in vitro- transcribed GAS5 sense (Sen.) or anti-sense (A.S.). Data are presented as mean values ± S.D., n = 3 biologically independent experiments, Two-sided Student’s t-test, nonsignificant. (p) SFB-MDH2 and Myc-CS or Myc-FH were co-overexpressed in empty vector (EV) control or GAS5 overexpressed HEK293T. Co-immunoprecipitation (Co-IP) assay was performed to detect the MDH2-CS and MDH2-FH interaction. (q) Confirmation of the GAS5-FL and its mutants (D1, D2, D3) overexpression levels in HEK293T cells by RT-qPCR detection. Data are presented as mean values ± S.D., n = 3 biologically independent experiments, Two-sided Student’s t-test, ***P < 0.001.
Extended Data Fig. 5 GAS5 regulates FH-MDH2-CS association by declining MDH2 acetylation.
(a) The 3D molecular model of MDH2 tetramer was established using PyMol software and the indicated acetylation sites were marked in red. (b) S-protein beads pulldown coupled IB detection of SFB-MDH2 acetylation level in HEK293T, under normal (25 mM) or high (50 mM) glucose concentration, using Pan-Acetylation antibody. (c) IB detection of the transduced SFB-MDH2 acetylation level under 8-hour glucose starvation or not using the Pan-Acetylation antibody in GAS5 or Scramble control knocked down HEK293T cells. (d) SFB-MDH2 was transduced in empty vector control (EV), full-length GAS5 (FL) or GAS5 mutant (D2) overexpressed HEK293T cells. Acetylation of SFB-MDH2 was detected by IP-IB using indicated antibodies. (e-f) FLAG-CS (e) or FLAG-FH (f) was overexpressed in HEK293T and the acetylation of CS or FH was detected by IP-IB using indicated antibodies under indicated gradient glucose concentration for 8 hours.
Extended Data Fig. 6 GAS5 functions as a tumour suppressor by controlling TCA flux.
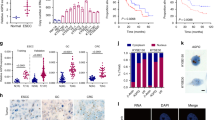
(a) RT-qPCR detection of GAS5 expression level in empty vector control (EV) or GAS5 overexpressed MDA-MB-231. Data are presented as mean values ± S.D., n = 3 biologically independent experiments, Two-sided Student’s t-test, ***P < 0.001. (b) Cellular NADH/NAD+ ratio was determined in empty vector control (EV) or GAS5 overexpressed MDA-MB-231 cells under 8-hour glucose starvation. Data are presented as mean values ± S.D., n = 3 biologically independent experiments, Two-sided Student’s t-test, **P < 0.01. (c-d) Mitochondria were isolated from 8-hour glucose starved (d) or normal (c) MDA-MB-231 cells and the mitochondrial NADH/NAD+ ratio level was measured. Data are presented as mean values ± S.D., n = 3 biologically independent experiments, Two-sided Student’s t-test, ***P < 0.001. (e) Mitochondria were isolated from MDA-MB-231 cells and the mitochondrial ATP generation was measured. Data are presented as mean values ± S.D., n = 3 biologically independent experiments, Two-sided Student’s t-test, *P < 0.05. (f) Oxygen consumption rate (OCR) profile was monitored in empty vector control (EV) or GAS5 overexpressed MDA-MB-231 cells with a Seahorse XF24 analyzer for 100 min. The indicated metabolic inhibitors were injected sequentially at different time points as indicated. Data are presented as mean values ± S.D., n = 3 biologically independent experiments. (g) RT-qPCR detection of GAS5 expression level in empty vector control (EV) or GAS5 overexpressed MDA-MB-468 cells. Data are presented as mean values ± S.D., n = 3 biologically independent experiments, Two-sided Student’s t-test, ***P < 0.001. (h) Cellular NADH/NAD+ ratio was determined in empty vector control (EV) or GAS5 overexpressed MDA-MB-468 cells. Data are presented as mean values ± S.D., n = 3 biologically independent experiments, Two-sided Student’s t-test, **P < 0.01. (i) Mitochondria were isolated from MDA-MB-468 cells and the mitochondrial NADH/NAD+ ratio level was measured. Data are presented as mean values ± S.D., n = 3 biologically independent experiments, Two-sided Student’s t-test, ***P < 0.001. (j) Cellular ATP was determined in empty vector control (EV) or GAS5 overexpressed MDA-MB-468 cells. Data are presented as mean values ± S.D., n = 3 biologically independent experiments, Two-sided Student’s t-test, *P < 0.05. (k) Mitochondria were isolated from MDA-MB-468 cells and the mitochondrial ATP level was measured. Data are presented as mean values ± S.D., n = 3 biologically independent experiments, Two-sided Student’s t-test, *P < 0.05. (l) Cellular malate level was determined in empty vector control (EV) or GAS5 overexpressed MDA-MB-468 cells. Data are presented as mean values ± S.D., n = 3 biologically independent experiments, Two-sided Student’s t-test, *P < 0.05. (m) Cellular citrate level was determined in empty vector control (EV) or GAS5 overexpressed MDA-MB-468 cells. Data are presented as mean values ± S.D., n = 3 biologically independent experiments, Two-sided Student’s t-test, **P < 0.01. (n) Cell proliferation viability was assessed by cell counting in empty vector control (EV) or GAS5 overexpressed MDA-MB-231 cells. Data are presented as mean values ± S.D., n = 3 biologically independent experiments, two-way ANOVA analysis, **P < 0.01. (o) Colony formation assay in empty vector (EV) control or GAS5 overexpressed MDA-MB-231 cells. Data are presented as mean values ± S.D., n = 3 biologically independent experiments, Two-sided Student’s t-test, ***P < 0.001. (p) Cell proliferation viability of GAS5 overexpressed MDA-MB-231 cells with 0.2 mM citrate rescue or not was analyzed by CCK8 assay in the indicated time points. Data are presented as mean values ± S.D., n = 3 biologically independent experiments, two-way ANOVA analysis, **P < 0.01. (q) In vivo analyses of tumors’ weight in xenograft mouse model were shown. Data are presented as mean values ± S.D., n = 5 biologically independent samples of xenograft tumors per group. Two-sided Student’s t-test, **P < 0.01, ***P < 0.001. (r) RT-qPCR detection of CCND1 (gene of cyclin D1) expression level in the indicated xenograft tumors. Data are presented as mean values ± S.D., n = 5 biologically independent samples of xenograft tumors per group, Two-sided Student’s t-test, *P < 0.05, ***P < 0.001. (s) RT-qPCR detection of SIRT3 expression in adjacent normal tissues (N) and malignant breast cancer (T) (SYSUCC Cohorts. n = 48 biologically independent samples of tumors with matched adjacent normal tissues. Two-sided Student’s t-test, **P < 0.01. (t) Analysis of SIRT3 expression in adjacent normal tissues (N) and malignant breast cancer (T), in other cohorts’ microarray data sets from Oncomine database. Data are presented as box plots with median (horizontal line) and 25%/75% quartiles with whiskers to the last point; Two-sided Student’s t-test, **P < 0.01. (u-w) Immunohistochemical (IHC) detection of MDH2 (u), FH (v) or CS (w) was performed in adjacent normal tissues and matched malignant breast cancer tissues (Bio MAX, n = 100 biologically independent samples of patients, Gehan-Breslow test, **P < 0.01). Scale bar, 100 µm.
Supplementary information
Supplementary Table 1
List of identified subcellular RNAs.
Supplementary Tables 2–4
Table S2: protein identification results for biotinylated GAS5 pull-down assay. Table S3: the correlation between clinicopathological parameters and GAS5 expression. Table S4: sequences of oligonucleotides for qPCR and RNA interference.
Source data
Source Data Fig. 1
Unprocessed western blots and gels.
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Unprocessed western blots and gels.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Unprocessed western blots and gels.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Unprocessed western blots and gels.
Source Data Fig. 5
Unprocessed western blots and gels.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 2
Unprocessed western blots and gels.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Unprocessed western blots and gels.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 4
Unprocessed western blots and gels.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 5
Unprocessed western blots and gels.
Source Data Extended Data Fig. 6
Statistical source data.
Rights and permissions
About this article
Cite this article
Sang, L., Ju, Hq., Yang, Z. et al. Mitochondrial long non-coding RNA GAS5 tunes TCA metabolism in response to nutrient stress. Nat Metab 3, 90–106 (2021). https://doi.org/10.1038/s42255-020-00325-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42255-020-00325-z
This article is cited by
-
A sustainable approach to universal metabolic cancer diagnosis
Nature Sustainability (2024)
-
ANT2 functions as a translocon for mitochondrial cross-membrane translocation of RNAs
Cell Research (2024)
-
LncRNA like NMRK2 mRNA functions as a key molecular scaffold to enhance mitochondrial respiration of NONO-TFE3 rearranged renal cell carcinoma in an NAD+ kinase-independent manner
Journal of Experimental & Clinical Cancer Research (2023)
-
The ASH1L-AS1-ASH1L axis controls NME1-mediated activation of the RAS signaling in gastric cancer
Oncogene (2023)
-
LncRNA modulates Hippo-YAP signaling to reprogram iron metabolism
Nature Communications (2023)