Abstract
Wheat leaf rust, caused by Puccinia triticina (Pt), remains a constant threat to wheat production worldwide. Deployment of race-specific leaf rust (Lr) resistance genes in wheat provides effective protection against leaf rust, but often leads to selective pressures that drive the rapid emergence of new virulent Pt isolates in nature. However, the molecular mechanisms underlying the evasion of Lr-delivered resistance by leaf rust remain largely unknown. Here, we identify an avirulence gene AvrLr21 in Pt that triggers Lr21-dependent immune responses. BSMV (Barley stripe mosaic virus)-mediated host-induced gene silencing assay shows that silencing AvrLr21 compromises Lr21-mediated immunity. AvrLr21 interacts directly with Lr21 protein to induce a hypersensitive response in tobacco leaves. The evolved Lr21-breaking Pt isolates can suppress Lr21-mediated immunity. Our data provide a basis for studying the molecular determinants in Pt-wheat incompatible interaction and monitoring natural Pt populations to prioritize the deployment of Lr resistance genes in the field.
Similar content being viewed by others
Introduction
The exponentially growing world population demands increasing annual cereal production. However, wheat, the second most cultivated cereal, is under constant threat from fungal pathogens that constitute critical concerns for future food security1. Leaf rust, caused by Puccinia triticina, is one of the agronomically important wheat fungal diseases that dramatically constrains wheat production worldwide2. The introgression of leaf rust resistance (Lr) genes into commercial wheat cultivars is the most effective and sustainable approach to providing protection of wheat against leaf rust3,4. 83 Lr genes have been identified in common wheat, but only a limited number of Lr genes have been cloned, including Lr1, Lr9, Lr10, Lr13, Lr14a, Lr21, Lr22a, Lr34, Lr42, Lr47, and Lr675,6,7,8,9. Most Lr genes confer race-specific resistance against Pt isolates carrying the cognate avirulence genes following the gene-for-gene hypothesis10. Recently, the first avirulence gene AvrLr15 in Pt that can trigger Lr15-dependent immune response in wheat was identified, and overcoming Lr15-mediated resistance in wheat was caused by a deletion and point mutations of amino acid sequence of AvrLr15 in the Lr15-breaking Pt races11.
Leaf rust resistance gene Lr21 that was introgressed into bread wheat from Iranian Aegilops tauschii was the first cloned leaf rust resistance gene, encoding the typical nucleotide-binding leucine-rich repeat protein12,13,14. Recently, Lr21-mediated complete seedling resistance and partial adult plant resistance in field conditions were found to be associated with higher expression of Lr21 at the seedling stage and lower expression at the adult plants, indicating that the expression dynamics of the resistance gene of the host plant at different developmental stages can determine the degree of immune response15.
Owing to the co-evolutionary arms race between Pt pathogens and wheat, new virulent Pt isolates with the ability to overcome Lr-mediated resistance are continuously emerging16,17,18, which often results in the ineffectiveness of wheat resistance delivered by Lr genes after a few years of cultivation of wheat cultivars with specific Lr genes19. For example, hard red spring wheat cultivars carrying the combinations of Lr2a, Lr10, Lr13, Lr16, Lr21, Lr23, and Lr34 genes were released in 2004 in the North American regions, but virulent Pt isolates against Lr21 was first found in North America in 201020,21. Therefore, the rapid emergence of Lr-breaking Pt isolates demands the identification of more Pt avirulence (Avr) genes to study the molecular basis of Pt-wheat compatible and incompatible interactions, which will aid us in understanding the molecular mechanisms underlying Avr protein recognition by Lr proteins and Avr evasion by R gene and guiding the selective cultivation of Pt-resistant wheat cultivars in different wheat-growing regions.
Here, we identify an avirulence gene AvrLr21 with the ability to trigger its cognate resistance gene Lr21-dependent immunity in wheat. Functional analyses of the interaction between AvrLr21 and Lr21 uncover that AvrLr21 physically interacts with Lr21 protein and the evolved Lr21-breaking Pt isolates are able to inhibit Lr21-mediated immunity in the presence of AvrLr21. These findings can deepen our understanding of the molecular interaction in Pt-wheat incompatible interactions and expand the molecular markers used to monitor natural Pt populations in the field.
Results and discussion
Pt_21 triggers Lr21-dependent cell death in the leaves of wheat and N. benthamiana
Our previous studies found that an effector gene Pt_21 was significantly up-regulated at 24 hpi during Pt infecting susceptible wheat cultivar and its product interacted with wheat TaTLP1 in the apoplast to suppress wheat immunity22. To identify the avirulence gene that genetically induces wheat resistance gene-mediated resistance, the produced recombinant protein ΔspPt_21 without signal peptide using the Escherichia coli (E. coli) expression system was purified and infiltrated into the leaves of 41 near-isogenic wheat lines of Thatcher carrying Lr resistance genes. Strikingly, cell death responses were specifically observed in the leaves of TcLr21 (RL6043). In contrast, no cell death responses were observed in the leaves of Thatcher and the other 40 near-isogenic lines of Thatcher carrying individual Lr gene as well as in the leaves infiltrated with buffer (Supplementary Fig. 1). We further validated these observations by infiltrating the purified ΔspPt_21 protein into the leaves of wheat TcLr21 and cultivar Fielder carrying non-functional lr21 allele (lr21-Fielder with 1 bp deletion in exon 2)15, respectively. As we expected, pronounced cell death responses appeared in the infiltrated region of Lr21 leaves, while no cell death responses were visible in the infiltrated leaves of Fielder and Thatcher (Fig. 1a). Leaves infiltrated with buffer did not display any cell death (Fig. 1a). In addition, a gene-based molecular marker of Lr21 confirmed the presence of Lr21 in the line TcLr21 and non-functional lr21 in Fielder (Supplementary Fig. 2). Taken together, these findings indicated that purified ΔspPt_21 protein induces an Lr21-dependent cell death response, indicating that Pt_21 might be AvrLr21. Protein-mediated cell death in wheat has been used to study the function of fungal effectors. For example, the infiltration of purified effector protein SnTox3 from Parastagonospora nodorum can trigger a robust necrotic response in the leaves of wheat carrying Snn323,24. Purified AvrSr35 protein was infiltrated into the leaves of wheat with Sr35 gene to induce cell death, but not in the leaves of wheat without Sr35 gene25.
a Infiltration of the ΔspPt_21 protein into the leaves of TcLr21, lr21-Fielder carrying a non-functional lr21 with a 1 bp deletion in exon 2 and Thatcher (Tc) wheat line without any leaf rust resistance genes. SDS-PAGE analysis of the recombinant ΔspPt_21 protein was conducted. Buffer was used as a negative control. b Co-infiltration of N. benthamiana leaves with Agrobacterium expressing the labeled Lr21 and ΔspPt_21 constructs. The same leaf was examined before (left) and after (right) being decolorized with the de-staining solution.
To further confirm that Pt_21 is AvrLr21, we constructed recombinant vectors carrying pCamA: ΔspPt_21 or pCamA: Lr21 (Supplementary Fig. 3) and co-infiltrated Agrobacterium tumefaciens carrying pCamA: ΔspPt_21 or pCamA: Lr21 on N. benthamiana leaves. Surprisingly, at 4 days after infiltration, strong cell death was observed in the co-infiltrated regions of N. benthamiana leaves (Fig. 1b). Another Pt effector, PTTG_27401, served as a negative control and failed to induce cell death when co-expressed with Lr21. To further validate this observed phenotype, co-infiltration was performed in Nicotiana tabacum leaves using Agrobacterium tumefaciens carrying ΔspPt_21 or Lr21 and the development of cell death was recorded at 2, 3 and 4 days after infiltration. Visible cell death appeared in co-infiltrated areas at 2 days, increased at 3 days, and fully occupied the infiltrated region at 4 days, but no cell death was observed in the negative control regions (Supplementary Fig. 4). Fluorescence microscopy observation confirmed the expression of GFP-tagged proteins, including Pt_21-GFP, Lr21-GFP and PTTG_27401-GFP (Supplementary Fig. 5). These results indicated that Pt_21 has an avirulence activity that induces an Lr21-dependent immune response in tobacco leaves. Previous studies showed that co-expression of wheat stem rust avirulence genes, such as AvrSr3525,26, AvrSr5027, and AvrSr2728, with its corresponding resistance gene in tobacco, triggers cell death.
Pt_21 is the avirulence gene AvrLr21
To further confirm that Pt_21 is the cognate avirulence gene of Lr21, we generated two BSMV constructs to knock down the expression of Pt_21 (Supplementary Fig. 6). The wheat PDS gene (TaPDS) served as a positive control to examine the silencing efficiency, and the empty BSMV:γ vector was used as a negative control. 10 days after the inoculation of BSMV by rubbing, the visible photobleaching phenotype was observed in BSMV:TaPDS-inoculated TcLr21 wheat leaves, whereas typical BSMV viral symptoms with mild chlorosis appeared in all leaves inoculated with BSMV:γ and BSMV:Pt_21. Next, the leaves of BSMV-inoculated TcLr21 wheat plants were challenged with urediniospores of Pt avirulent isolate PHNT. 14 days after inoculation of Pt, significantly increased rust pustules were observed in the leaves inoculated with BSMV: Pt_21 (Fig. 2a) compared to the leaves of TcLr21 without BSMV inoculation and the leaves of TcLr21 inoculated with BSMV:γ. The expression level of Pt_21 was significantly decreased in BSMV:Pt_21-treated TcLr21 plants compared with BSMV:γ-inoculated TcLr21 wheat at 24, 48 and 120 hpi (Fig. 2b). In addition, histological observation of Pt infection found that the hyphal length, infection areas, hyphal branching number and overall infection area of Pt at 24, 48 and 120 hpi were significantly increased following the silencing of Pt_21 by HIGS in TcLr21(Fig. 2c and Supplementary Fig. 7a–c). The areas of cell death and H2O2 accumulation were significantly lower at 120 hpi in BSMV:Pt_21 plants than those observed in BSMV:γ-inoculated and non-inoculated leaves (Fig. 2d and Supplementary Fig. 7d). Collectively, these findings indicate that BSMV-mediated silencing of Pt_21 is accurate, enabling us to conclude that Pt_21 is the avirulence gene AvrLr21. BSMV (Barley stripe mosaic virus)-mediated host-induced gene silencing assay has been widely used to study the functional role of wheat rust effectors during rust pathogen infection. For example, BSMV-mediated HIGS of the Pst_12806 effector from wheat stem rust Pst demonstrated that Pst_12806 is indispensable for Pst-wheat compatible interaction. Silencing of PstGSRE1 using BSMV-mediated HIGS impaired the virulence of Pst in wheat29,30,31,32,33.
a BSMV-mediated host gene silencing of Pt_21 impairs Lr21-mediated resistance against Pt isolate PHNT that is avirulent on TcLr21 plant. The fourth leaves of TcLr21 wheat at 10 days after infection with BSMV:γ, BSMV:TaPDS, BSMV:Pt_21 were inoculated with urediospores. BSMV:γ-and BSMV:TaPDS-inoculated Thatcher and mock leaves treated with buffer were used as positive and negative control, respectively. At 14 days after urediospore inoculation, representative pictures were taken. b Relative expression levels of Pt_21 in Pt_21-silenced plants and negative control BSMV:γ plants at 24, 48, and 120 hpi after urediospore inoculation. The y-axis indicates the amounts of Pt_21 transcript normalized to the GAPDH gene. The x-axis indicates sampling times. Data are means ± standard errors (SE) of three independent experiments. Differences between Pt_21-silenced plants and control plants were assessed using Student’s t-tests (*p < 0.05). c Histological observations of hypha development and host cell death in BSMV-mediated HIGS of Pt_21 wheat leaves after inoculation with Pt PHNT at 24, 48, and 120 hpi. AP appresorium, SV substomatal vesicle, IH infection hypha, HMC haustorial mother cell, SH secondary hypha, NC necrotic cell, U urediospore. Scale bar, 50 μm. d H2O2 accumulation at infection sites was detected by staining with DAB and viewed under differential interference contrast microscopy. Scale bar, 20 µm.
AvrLr21 directly interacts with Lr21
AvrLr21 with the ability to induce the wheat Lr21-dependent cell death in planta suggests that the recognition specificity between AvrLr21 and Lr21 is likely mediated by direct interaction. To examine the interaction between AvrLr21 and Lr21, a yeast two-hybrid (Y2H) experiment was conducted using pGBKT7:ΔspAvrLr21 and pGADT7:Lr21 constructs (Supplementary Fig. 8). As shown in Fig. 3, the yeast Y2HGold transformant co-expressing pGBKT7:ΔspAvrLr21 and pGADT7:Lr21 was able to grow on the selective-HAWL medium and produced blue coloration, which was similar to the positive control ΔspTaPR1 and ΔspTaTLP1. In contrast, yeast co-transformed with Lr21 and the empty bait or ΔspAvrLr21 and the empty prey vector was unable to grow on the selective plates and failed to produce blue coloration (Fig. 3a). To further validate the interaction, a co-immunoprecipitation (Co-IP) assay was conducted. Figure 3 showed that recombinant Lr21-GFP proteins were co-purified with ΔspAvrLr21-GST protein using GST beads (IP), confirming that ΔspAvrLr21 directly interacts with Lr21 (Fig. 3b). In addition, RFP-tagged Lr21 and GFP-tagged ΔspAvrLr21 colocalized in the nucleus and cytosol of N. benthamiana leaves (Fig. 3c), implying that the direct interaction between ΔspAvrLr21 and Lr21 likely occurs in the plant nucleus and cytosol. Our results were consistent with previous studies. It has been documented that the rust avirulence protein is directly recognized by its corresponding resistance protein. The wheat stem rust avirulence protein AvrSr35 interacted directly with the Sr35 to induce Sr35 resistosome assembly25,26, and AvrSr50 interacted directly with the Sr50 protein in Y2H assay27. The flax rust avirulence protein AvrL567 specifically interacted with the corresponding R proteins in yeast34.
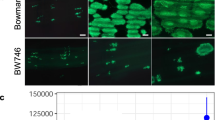
a Y2H analysis of the direct interaction between Lr21 and ΔspAvrLr21. Growth of yeast strains coexpressing ΔspAvrLr21 fused to the GAL4 DNA binding domain (BD) with Lr21 fused to the GAL4 activation domain (AD) on control media lacking leucine and tryptophan (SD-WL) or selective media lacking histidine and adenine supplemented with X-α-Gal (SD-HAWL-X-α-Gal). ΔspTaPR1 (BD) and ΔspTaTLP1 (AD) were used as positive controls. A ten-fold serial dilution is shown. b Confirmation of the interaction between ΔspAvrLr21 and Lr21 by Co-IP assays. Western blots of total proteins transiently expressing the marked constructs and proteins eluted from GST beads were detected using the anti-GFP or anti-GST antibody. The protein marker was labeled on the left. c Co-expressing GFP-tagged ΔspAvrLr21 and RFP-tagged Lr21 in the leaves of N. benthamiana, showing that ΔspAvrLr21 and Lr21 co-localize in plant nucleus and cytosol. The white arrows point to the nucleus. Scale bar, 50 μm.
Dissecting the molecular mechanisms underlying Lr21-breaking Pt isolates
The natural Lr21-breaking Pt isolates have been reported in America and China35. To understand the evolutionary mechanism underlying the emerged Pt isolates with the ability to overcome Lr21-mediated resistance, PCR amplification of the AvrLr21 genes from 6 natural Lr21-virulent isolates was conducted, and the resulting PCR products were sequenced. Surprisingly, all sequences of AvrLr21 genes cloned from Lr21-overcoming isolates are identical to the sequences of the AvrLr21 gene from the Lr21-avirulent isolate (Supplementary Fig. 9). We quantified the relative expression of AvrLr21 gene in our available Lr21-avirulent and -breaking isolates. The expression patterns of AvrLr21 in Lr21-avirulent and -breaking isolates at different time points upon infection of TcLr21 wheat line were similar, as evidenced by the observation that AvrLr21 was significantly induced and peaked at 72 hpi and subsequently decreased from 96 to 144 hpi. Notably, we found the expression level of AvrLr21 in Lr21-avirulent isolate both at 48 and 72 hpi was 2-fold higher than that in Lr21-virulent isolates, suggesting that expression variations of AvrLr21 at specific infection stage of Pt are likely attributed to the evasion of Lr21-mediated resistance in Lr21-breaking isolates (Supplementary Fig. 10). However, the protomer sequences of AvrLr21 in Lr21-breaking and Lr21-avirulent isolates are exactly same (Supplementary Fig. 11). Gain of virulence function through altering the expression level of avirulence genes by fungal pathogens has been reported. Wheat stem rust Puccinia graminis f. sp. tritici (Pgt) AvrSr27 can avoid the recognition by Sr27 gene via deletion, copy number and expression level variations at the AvrSr27 locus detected in the field Sr27-breaking Pst isolates28.
In addition, we found that pre-inoculation of Lr21-breaking Pt isolate THTT on TcLr21 leaves can completely inhibit AvrLr21-induced cell death in Tc·Lr21 leaves at 1d and 10d after infiltration of purified ΔspAvrLr21 protein (Fig. 4 and Supplementary Fig. 12). In contrast, ΔspAvrLr21 induces strong cell death in the leaves of TcLr21 without THTT inoculation as well as in leaves with Lr21-avirulent PHNT inoculation (Fig. 4 and Supplementary Fig. 12). Histological observation of the Pt hyphae at 24 hpi and 48 hpi indicated the successful inoculation of Pt isolates on wheat leaves (Supplementary Fig. 13). These observations indicated that Lr21-breaking Pt isolate evolved unknown mechanisms to suppress Lr21-mediated resistance, likely utilizing the secreted effector proteins. Previous studies have reported that fungal pathogens can employ secreted effectors to suppress plant resistance gene-delivered resistance. For example, xylem-invading fungus Fusarium oxysporum f.sp. lycopersici effector Avr1 can suppress tomato resistance genes I-2- and I-3-mediated immunity36. The effector AvrLm4-7 of Leptosphaeria maculans causing stem canker in oilseed rape can inhibit oilseed resistance genes Rlm3-, Rlm5- and Rlm9-mediated resistance37,38. In the future, we need to dissect the molecular mechanism underlying the inhibition of AvrLr21-induced cell death by Lr21-breaking Pt isolates.
Lr21-breaking Pt isolate THTT can specifically inhibit AvrLr21-induced cell death in the leaves of wheat line TcLr21. The leaves of TcLr21 were inoculated with urediospores of PHNT and THTT, respectively. At 12 h post-inoculation, the inoculated leaves were infiltrated with purified ΔspAvrLr21 protein. Representative pictures were taken at 1 day and 10 days after infiltration. The non-inoculated leaves and leaves inoculated with PHNT served as negative controls. PHNT (+): avirulent on TcLr21 wheat, THTT (+): virulent on TcLr21 wheat.
Methods
Plant materials, Pt race and sample collection
All near-isogenic lines of Thatcher, susceptible wheat Chinese Spring and tobacco are preserved in the Laboratory of Wheat Leaf Rust, Hebei Agricultural University. Thatcher line without any leaf rust resistance genes is susceptible to all leaf rust pathogens. The Fielder line contains the non-functional lr21 allele (lr21-Fielder with 1 bp deletion in exon 2). Second seedling leaves inoculated with PHNT (isolate 07-10-426-1) or distilled water (control) for RNA extraction were harvested at 0, 24, 48, 72, 96, 120, and 144 hpi. All samples were immediately frozen in liquid nitrogen and stored at −80 °C. Each treatment included three independent biological replicates.
RNA isolation and quantitative PCR
Total RNA was extracted using the TaKaRa MiniBEST Universal RNA Extraction Kit (TaKaRa, Beijing) according to the manufacturer’s instructions. Contaminating DNA was digested with amplification-grade DNase I (TaKaRa, Beijing) and cDNA was synthesized using the SuperScript II reverse transcriptase, oligo (dT)-18, and RNase Inhibitor (TaKaRa, Beijing). qPCR was performed with the LightCycler 96 (Roche, Shanghai). Expression of Pt_21 was investigated, and the β-actin gene was used to calibrate the expression level of the query genes, as previously described (Supplementary Table S1). Quantification of the target gene was assessed by relative standard curves. The 2−ΔΔCT method was employed to quantify the relative gene expression levels. Statistical analysis was conducted using Microsoft Excel to calculate mean values and standard errors (SE). The statistical significance of differences was calculated using Student’s t test by the SPSS 21.0 (IBM SPSS Statistic) to obtain the P value. Data were shown as mean ± SE of three biological replicates from one representative experiment. Significant differences between treatments and controls or between time-course points were represented by asterisk (**p < 0.01, *p < 0.05).
Expression and purification of the fusion protein
Pt_21 was individually cloned into pGEX-6p-3 (with GST tag) and expressed in E. coli. The recombinant plasmids were transformed into E. coli BL21 (DE3) chemically competent cells. The production of protein was induced by adding isopropyl-β-D-thiogalactopyranoside (IPTG, Mei5bio, Beijing). A GST-Agarose Label Kit (TRAN, Beijing) was used to bind GST-tagged protein following the manufacturer’s instructions. The purified protein products were separated by 15% SDS-PAGE and visualized using Coomassie Blue (Mei5bio, Beijing) staining.
Infiltration of wheat near-isogenic lines
We infiltrated the leaves of 21-day-old seedlings of all available 41 near-isogenic lines of Thatcher with ΔspPt_21 protein. This protein was diluted to 0.1 mg/mL in 1X PBS buffer (137 mM NaCl, 2.7 Mm KCl, 10 Mm Na2HPO4,and 2 Mm KH2PO4) and infiltrated into the abaxial side of the first leaf using a 1 ml needleless syringe. Buffer was infiltrated into a second leaf to test if the cell death was caused by a non-specific protein. The purified ΔspPt_21 protein was infiltrated into the leaves of TcLr21, and infiltrated seedlings were transferred to a growth chamber maintained at 22 °C with a photoperiod of 16 h. Cell death was evaluated at 24 h after infiltration. All these experiments were repeated three times. Genomic DNA of TcLr21 (RL6043) was extracted by a previously published method21.
Agrobacterium tumefaciens infiltration assays and confocal microscopy
The recombinant vectors pCamA:Lr21, pCamA:ΔspPt_21 and pCamA:PTTG_27401 were transformed into Agrobacterium tumefaciens strain GV3101 (Pyaest, Wuhan). Agrobacterium cultures containing the Lr21 and ΔspPt_21 or Lr21 and PTTG_27401 expression vectors were grown overnight at 28 °C in LB medium with appropriate antibiotic selections. The cells were pelleted and resuspended in MMAi buffer (5 g/L MS salts, 20 g/L sucrose, 10 mM MES, 200 μΜ AS) to an optical density (OD600) of 0.6, followed by incubation at room temperature for 1 hour. Cultures were infiltrated into leaves of 4-week-old tobacco with a 1 mL needleless syringe. Symptoms were monitored and recorded from 2 to 4 days after infiltration. Three independent biological replicates were conducted for each experiment. Protein expression was visualized using fluorescence microscopy with an excitation wavelength of 488 nm and an emission wavelength of 530 nm.
Y2H validation
Primers were designed to amplify the coding regions of AvrLr21 and Lr21 (Supplementary Table S1). The recombinant vectors pGBKT7:ΔspAvrLr21 and pGADT7:Lr21 were constructed. The Matchmaker Gold Y2H System was used to verify the interaction between ΔspAvrLr21 (bait) and Lr21 (prey). The bait and prey plasmids were co-transformed into yeast strain Y2HGold (Pyaest, Wuhan) according to the manufacturer’s instructions. Transformed yeast cells were assayed for growth on synthetic dropout SD/-Trp-Leu plates and SD/-Trp-Leu-His-Ade plates containing X-α-Galactosidase (X-α-Gal) and Aureobasidin A (AbA) (Pyaest, Wuhan).
Co-IP assay
Agrobacterium strain GV3101 carrying the recombinant vector pCamA:Lr21 was infiltrated into the leaves of 4-week-old N. benthamiana. At 48 h post agroinfiltration, total protein was extracted using extraction buffer (50 mM Tris pH 7.5, 150 mM NaCl, 1% NP-40, 5 mM DTT, 100×Protease Inhibitor Cocktail, For Bacterial Cell, with EDTA (Songon, Shanghai). pGEX-6p-3:ΔspAvrLr21 was induced by IPTG with a final concentration of 0.5 mmol/L. For Co-IP assays, a mixture of 2 mL ΔspAvrLr21 and Lr21 containing 50 μl GST agarose beads were incubated at 4 °C for 4 h with gentle shaking. The precipitated proteins were washed five times with washing buffer. The precipitated proteins and crude proteins (input) were detected by immunoblotting with 5 μL anti-GST antibody (Solarbio, Beijing, K000478P) and 5 μL anti-GFP antibody (Solarbio, Beijing, K009323P) (The dilution ratio is 1:1000).
Co-localization assays
Red fluorescent fusion constructs were produced by cloning the coding sequence of Lr21 into the vector pGWB454. GFP fluorescent fusion constructs were produced by cloning the coding sequence of ΔspAvrLr21 into the vector pCamA (Supplementary Table S1). Agrobacterium strains GV3101 carrying pGWB454:Lr21 and pCamA:ΔspAvrLr21 were resuspended to a final OD600 of 0.8 and mixed in equal proportions immediately prior to infiltrations. Infiltrations were performed on 4-5-week-old of N. benthamiana plants. Fluorescence in leaves of N. benthamiana was monitored at 48 h after agroinfiltration, and then imaged directly using a confocal laser scanning microscopy with an excitation wavelength of 488 nm and emission of 540 nm (Olympus FluoView FV1000).
Sequence analysis of AvrLr21
AvrLr21 was amplificated from Pt race PHNT using 2×Es Taq MasterMix (CWBIO, Beijing). Sanger sequencing was used for sequencing of the AvrLr21 gene. The molecular weight of AvrLr21 protein was predicted by ProtParam (https://web.expasy.org/protparam/), and the localization of AvrLr21 protein was predicted by EffectorP-fungi 3.0 (http://effectorp.csiro.au/). The signal peptide of AvrLr21 was identified using SignalP 5.0 (http://www.cbs.dtu.dk/services/SignalP/). All primers used in this study are listed in Supplementary Table S1.
Host-induced gene silencing (HIGS)
To confirm the specificity of the silenced fragments, cDNA fragments of AvrLr21 with no or lower similarity to other genes were selected to prevent off-target effects. Next, the selected cDNA fragments, containing NotI and PacI restriction sites (Supplementary Table S1), were obtained by reverse transcription PCR and inserted into the empty barley stripe mosaic virus (BSMV) vector to generate BSMV:Pt_21. The wheat phytoene desaturase gene (PDS) was inserted into the empty BSMV vector to generate BSMV:TaPDS as a positive control for examining the silencing efficiency. BSMV:γ was used as a negative control. In vitro transcription was performed using the mMESSAGE mMACHINE Kit High Yield Capped RNA Transcription Kit (Ambion) with linearized plasmid as a template according to the manufacturer’s protocol. Then, 240 μl of the BSMV mixture (10 μl of linearized product, 90 μl of DEPC water, and 120 μl of GKP Buffer) was applied to the fully expanded leaves of 2-leave stage TcLr21 wheat plants by rubbing according to the method described previously. Leaf rust urediniospores were inoculated on the fourth and fifth leaves of TcLr21 and the susceptible wheat cultivar Thatcher. Wheat leaves were inoculated with sterile water as a negative control. 10 days after virus inoculation, the fresh urediniospores of PHNT were inoculated on the virus-inoculated leaves. The inoculated leaves were sampled at 24 h, 48 h, and 120 h after inoculation to examine the silencing efficiency of AvrLr21 using RT-qPCR and for histological observations. These experiments were repeated three times.
Histological observations of fungal growth and host responses
Auto-fluorescence of fungus-penetrated mesophyll cells was observed in necrotic areas by epifluorescence microscopy (excitation filter, 488 nm; dichromic mirror, 510 nm; and barrier filter, 520 nm). The 3,3-diaminobenzidine (DAB, Solarbio, Beijing) staining was conducted to detect H2O2 accumulation and viewed following the protocols described previously39. A minimum of 50 infection sites were examined on each of five randomly selected leaf segments for each treatment. The hyphal length, the branching of hyphae and H2O2 accumulation were observed with the Nikon Ti2-LAPP Ti2 Laser Application System (Nikon Corporation).
Inoculation of Lr21-breaking Pt isolate to inhibit cell death
Inoculation of the Lr21-breaking isolate THTT and the Lr21-avirulent isolate PHNT on the leaves of TcLr21 at the three-leaf stage were conducted following the methods previously described40. At 12 h after inoculation and moisturizing, the purified ΔspAvrLr21 protein with a concentration of 0.1 mg/mL with 1X PBS buffer (137 mM NaCl, 2.7 mM KCl, 10 mM Na2HPO4, and 2 mM KH2PO4) was infiltrated into the leaves of TcLr21 using a needleless syringe, and the infiltrated seedlings were transferred to a growth chamber and maintained at 22 °C with a photoperiod of 16 h. Development of cell death around the infiltration regions was observed, and representative pictures were photographed at 1 day and 10 days after protein infiltration. The experiment was repeated three times with similar results.
Statistics and reproducibility
Different statistical tests were conducted for different experiments. One-way ANOVA analysis was conducted with the Turkey’s post-hoc test using SPSS to determine the significance of differences between treatments or samples. Sample sizes and the number of replicates for each experiment are indicated in relevant experiments.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The gene sequence of AvrLr21 is available in the National Center for Biotechnology Information (NCBI) (http://www.ncbi.nlm.nih.gov/) with accession number OK033362.1. Source data underlying graphs can be obtained in Supplementary Data.
References
Fisher, M. C. et al. Emerging fungal threats to animal, plant and ecosystem health. Nature 484, 7393 (2013).
Dinh, H. X., Singh, D., Periyannan, S., Park, R. F. & Pourkheirandish, M. Molecular genetics of leaf rust resistance in wheat and barley. Theor. Appl. Genet. 133, 2035–2050 (2020).
Ellis, J. G., Lagudah, E. S., Spielmeyer, W. & Dodds, P. N. The past, present and future of breeding rust resistant wheat. Front. Plant Sci. 5, 641 (2014).
Liu, F. et al. Exome association analysis sheds light onto leaf rust (Puccinia triticina) resistance genes currently used in wheat breeding (Triticum aestivum L.). Plant Biotechnol. J. 18, 1396–1408 (2020).
Kumar, K. et al. An update on resistance genes and their use in the development of leaf rust resistant cultivars in wheat. Front. Genet. 13, 816057 (2022).
Li, H. N. et al. Cloning of the wheat leaf rust resistance gene Lr47 introgressed from Aegilops speltoides. Nat. Commun. 14, 6072 (2023).
Lin, G. F. et al. Cloning of the broadly effective wheat leaf rust resistance gene Lr42 transferred from Aegilops tauschii. Nat. Commun. 13, 3044 (2022).
Hewitt, T. et al. Wheat leaf rust resistance gene Lr13 is a specific Ne2 allele for hybrid necrosis. Mol. Plant. 14, 1025–1028 (2021).
Yan, X. C. et al. High-temperature wheat leaf rust resistance gene Lr13 exhibits pleiotropic effects on hybrid necrosis. Mol. Plant. 14, 1029–1032 (2021).
Bolton, M. D., Kolmer, J. A. & Garvin, D. F. Pathogen profile Wheat leaf rust caused by Puccinia triticina. Mol. Plant Pathol. 9, 563–575 (2008).
Cui, Z. C. et al. Evasion of wheat resistance gene Lr15 recognition by the leaf rust fungus is attributed to the coincidence of natural mutations and deletion in AvrLr15 gene. Mol. Plant Pathol. 25, e13490 (2024).
Huang, L. et al. Evolution of new disease specificity at a simple resistance locus in a crop–weed complex: reconstitution of the Lr21 gene in wheat. Genetics 182, 595–602 (2009).
Huang, L. et al. Map-based cloning of leaf rust resistance gene Lr21 from the large and polyploid genome of bread wheat. Genetics 164, 655–664 (2003).
Rowland, G. G. & Kerber, E. R. Telocentric mapping in hexaploid wheat of genes for leaf rust resistance and other characters derived from Aegilops squarrosa. Can. J. Genet. Cytol. 16, 137–144 (1974).
Naz, A. A. et al. Lr21 diversity unveils footprints of wheat evolution and its new role in broad-spectrum leaf rust resistance. Plant Cell Environ. 10, 1151–1158 (2021).
Zhao, J. & Kang, Z. S. Fighting wheat rusts in China: a look back and into the future. Phytopathol. Res. 5, 6 (2023).
Figlan, S. et al. Breeding wheat for durable leaf rust resistance in southern Africa: variability, distribution, current control strategies, challenges and future prospects. Front. Plant Sci. 11, 549 (2020).
Maor, R. & Shirasu, K. The arms race continues: battle strategies between plants and fungal pathogens. Curr. Opin. Microbiol. 8, 399–404 (2005).
McCallum, B. D. et al. A review of wheat leaf rust research and the development of resistant cultivars in Canada. Can. J. Plant. Pathol. 38, 1–18 (2016).
Oelke, L. M. & Komer, J. A. Genetics of leaf rust resistance in spring wheat cultivars alsen and norm. Genet. Resist. 95, 773–778 (2005).
Kolmer, J. A., Long, D. L. & Hughes, M. E. Physiologic specialization of Puccinia triticina on wheat in the United States in 2010. Plant Dis. 96, 1216–1221 (2012).
Wang, F. et al. Puccinia triticina effector protein Pt_21 interacts with wheat thaumatin-like protein TaTLP1 to inhibit its antifungal activity and suppress wheat apoplast immunity. Crop J. 11, 1431–1440 (2023).
Sung, Y. C. et al. PR1-mediated defence via C-terminal peptide release is targeted by a fungal pathogen effector. N. Phytol. 229, 3467–3480 (2021).
Outram, M. A. et al. The crystal structure of SnTox3 from the necrotrophic fungus Parastagonospora nodorum reveals a unique effector fold and provides insight into Snn3 recognition and pro-domain protease processing of fungal effectors. N. Phytol. 231, 2282–2296 (2021).
Salcedo, A. et al. Variation in the AvrSr35 gene determines Sr35 resistance against wheat stem rust race Ug99. Plant Sci. 358, 1604–1606 (2017).
Zhao, Y. B. et al. Pathogen effector AvrSr35 triggers Sr35 resistosome assembly via a direct recognition mechanism. Sci. Adv. 8, eabq5108 (2022).
Chen, J. P. et al. Loss of AvrSr50 by somatic exchange in stem rust leads to virulence for Sr50 resistance in wheat. Plant Sci. 358, 1607–1610 (2017).
Upadhyaya, N. M. et al. Genomics accelerated isolation of a new stem rust avirulence gene–wheat resistance gene pair. Nat. Plants 7, 1220–1228 (2021).
Panwar, V., McCallum, B. & Bakkeren, G. Host-induced gene silencing of wheat leaf rust fungus Puccinia triticina pathogenicity genes mediated by the Barley stripe mosaic virus. Plant Mol. Biol. 81, 595–608 (2013).
Bai, X. X. et al. A candidate effector protein PstCFEM1 contributes to virulence of stripe rust fungus and impairs wheat immunity. Stress Biol. 2, 12 (2022).
Wang, Y. Q. et al. A stripe rust fungal effector PstSIE1 targets TaSGT1 to facilitate pathogen infection. Plant J. 112, 1413–1428 (2022).
Xu, Q. et al. An effector protein of the wheat stripe rust fungus targets chloroplasts and suppresses chloroplast function. Nat. Commun. 10, 5571 (2019).
Qi, T. et al. Stripe rust effector PstGSRE1 disrupts nuclear localization of ROS-promoting transcription factor TaLOL2 to defeat ROS-induced defense in wheat. Mol. Plant. 12, 1624–1638 (2019).
Dodds, P. N. et al. Direct protein interaction underlies gene-for-gene specificity and coevolution of the flax resistance genes and flax rust avirulence genes. PNAS 103, 8888–8893 (2006).
Zhang, L., Zhang, L. Y., Meng, Q. F., Yan, H. F. & Liu, D. Q. Virulence and molecular genetic diversity, variation, and evolution of the Puccinia triticina population in Hebei Province of China from 2001 to 2010. Plant Sci. 14, 1095677 (2023).
Houterman, P. M., Cornelissen, B. J. C. & Rep, M. Suppression of plant resistance gene-based immunityby a fungal effector. PLoS Pathog. 5, e1000061 (2008).
Plissonneau, C. et al. A game of hide and seek between avirulence genes AvrLm4-7 and AvrLm3 in Leptosphaeria maculans. N. Phytol. 209, 1613–1624 (2016).
Ghanbarnia, K. et al. Leptosphaeria maculans AvrLm9: a new player in the game of hide and seek with AvrLm4-7. Mol. Plant Pathol. 19, 1754–1764 (2018).
Wang, F. et al. TaTLP1 interacts with TaPR1 to contribute to wheat defense responses to leaf rust fungus. PLoS Genet. 16, e1008713 (2020).
Roelfs, A. P. & Martell, L. B. Uredospore dispersal from a point source within a wheat canopy. Phytopathology 74, 1262–1267 (1984).
Acknowledgements
We thank Prof. R.A. McIntosh (Plant Breeding Institute, University of Sydney) for review and English editing of the manuscript; Prof. M. Hossein Borhan (Agriculture and Agri-Food Canada), for providing the gateway vectors; Prof. Zhimin Hao (Hebei Agricultural University) for providing the pGEX-6p-3 plasmids. This study was funded by the State Key Laboratory of North China Crop Improvement and Regulation (NCCIR2023ZZ-10), the National Natural Science Foundation of China (32172384, 31501623), Chunhui Talent Program of the Natural Science Foundation of Hebei (C2022204116) and the Hundred Talents Program for the introduction of high-level overseas talents in Hebei Province (E2020100004).
Author information
Authors and Affiliations
Contributions
H.Y.W., F.W., and S.T.Y. conceived the project and designed the experiments. S.S.S., F.W., Z.C.C., and L.S.M. performed the experiments and interpreted the results. F.W., S.S.S., Z.C.C., and L.S.M. analyzed the data. H.Y.W., S.S.S, and L.M. wrote and revised the manuscript. L. M., H.Y.W., and D.Q.L. supervised this experiment.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Anindya BANDYOPADHYAY, Ahmet Caglar Ozketen, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editors: Dr. Leena Tripathi and Dr. Ophelia Bu.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Shen, S., Wang, F., Cui, Z. et al. Puccinia triticina avirulence protein AvrLr21 directly interacts with wheat resistance protein Lr21 to activate wheat immune response. Commun Biol 7, 1170 (2024). https://doi.org/10.1038/s42003-024-06881-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-024-06881-4
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.