Abstract
Common ragweed is an invasive alien species causing severe allergies in urban residents. Understanding its urban invasion pathways is crucial for effective control. However, knowledge is limited, with most studies focusing on agricultural and natural areas, and occurrence record-based studies exhibiting uncertainties. We address this gap through a study in East China cities, combining population genetics and occurrence records. Leaf samples from 37 urban common ragweed populations across 15 cities are collected. Genomic and chloroplast DNA extraction facilitate analysis of spatial genetic patterns and gene flows. Additionally, international grain trade data is examined to trace invasion sources. Results indicate spatial genetic patterns impacted by multiple introductions over time. We infer the modern grain trade between the United States and China as the primary invasion pathway. Also, cities act as transportation hubs and ports of grain importation might disperse common ragweed to urban areas. Invasive species control should account for cities as potential landing and spread hubs of common ragweed.
Similar content being viewed by others
Introduction
Common ragweed (Ambrosia artemisiifolia L., Asteraceae) is native to North America but is now an invasive alien species (IAS) on all continents except Antarctica1. Besides causing significant losses in agricultural production, common ragweed pollen is a primary allergen source, causing seasonal allergic rhinitis and asthma, which result in substantial health costs and labor losses1,2. The negative health impact of common ragweed pollens is worsened in urban areas because the production of pollens and allergenicity of pollens are both enhanced in urban environments3,4. Therefore, control of the common ragweed is a priority in many cities worldwide5.
Cities need information on the invasion pathway of common ragweed to control it more effectively. So far, this information has been primarily inferred from occurrence records. For example, the introduction of common ragweed was speculated to be related to grain shipment based on surveying the distribution of common ragweed in Israel6. In Magdeburg, Germany, researchers used the occurrence records of common ragweed populations in the urban area to link the introduction of common ragweed to seed trade7. Based on the occurrence data, communication lines were found to play a vital role in the dispersal of common ragweed in urban areas of Croatia8. While those inferences are helpful, they also contain notable limitations. The study in Israel and Magdeburg could not further trace the source of common ragweed to countries exporting the contaminated seeds7. It is difficult to determine the real dispersal hub of common ragweed in urban areas of Croatia8.
Some typical features of occurrence records contributed to the limitations in the above studies. First, like many other IAS, the data quality and availability of occurrence records for common ragweed are often inadequate9,10. The quality and availability of occurrence data for IAS in cities are particularly uneven10,11. Records with spatial and temporal errors may lead to an incomplete or even wrong interpretation of invasion pathways in the IAS. Second, interpretation of the occurrence data is challenging even when the quality of the data is not an issue. The spatial distribution of a species is not always a good indication of its spread pathway. For instance, plant populations with distinct ancestors in a city may be incorrectly judged as close relatives due to their spatial proximity12.
Population genetic methods provide a viable way to address the above limitations. The phylogenetic relationship among populations of IAS can be reconstructed using various genetic methods to interpret the invasion pathway. For example, in Argentina, the genetic analysis showed that the common sunflower (Helianthus annuus L.) was first introduced to Buenos Aires and then spread to Mendoza13. The population structure of slender false brome (Brachypodium sylvaticum (Huds.) P. Beauv.) in Corvallis, the United States, showed that the species was introduced from multiple locations in Europe14. Studies have already shown the potential of population genetic methods for revealing the invasion pathways of common ragweed. The method has been used to trace the source of common ragweed populations in Asia, South America, and Western and Eastern Europe15,16,17. It has also been adopted to identify the invasion pathways in Australia18 and quantify the genetic patterns of common ragweed populations in North America and China19,20. The invasion sources of common ragweed populations in East China was explored in an earlier study21. However, several key questions have not been answered. How did the propagules of common ragweed enter China? Are invasion sources different for common ragweed populations in East China cities? Which city(cities) is the dispersal centers of common ragweed? The answers to the above questions are crucial for devising effective and targeted management measures for common ragweed.
In this study, we intend to disclose the invasion pathways of common ragweed in urban areas. We selected cities in East China as the sites for this case study. China first recorded common ragweed in the 1930s22. Common ragweed currently occurs in more than 23 provinces, and most of the occurrences are in East China23,24. East China accounts for 43% of China’s land area but nearly 94% of China’s population25. The highly urbanized region contains key port cities for international trade and a well-developed transportation network26. Because existing studies found that ragweed seed is a common contaminant of the seed trade and international trade can accelerate the introduction of common ragweed27,28, we hypothesized that the international trade would contribute to the introduction of common ragweed into urban areas in East China cities. To test this hypothesis, we employed a combination of population genetic data, occurrence records, and international grain trade data. Our findings added new evidence of the facilitating role of international trade in spreading common ragweed.
Results
Genetic patterns of common ragweed populations in East China cities
We identified six nuclear microsatellite loci from the leaf samples. The results of exact tests for genotypic disequilibrium showed that none of the microsatellite loci presented significant linkage equilibrium. Also, the estimates of null alleles were highly variable (Supplementary Fig. 1). Therefore, these loci could be used for further analysis. Deviation from Hardy-Weinberg proportions was detected in most common ragweed populations (Table 1), which suggested heterozygote deficiency. Bottleneck effects caused by the population size reduction were found in six populations.
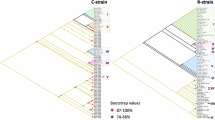
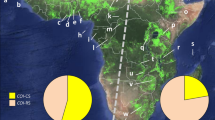
We identified ten cpDNA haplotypes based on indels and polymorphisms observed within the 836-bp concatenated alignment in chloroplast intergenic spacers. The values of haplotype diversity (HD) ranged from 0 to 0.68, and the values of nucleotide diversity (π) ranged from 0 to 0.00122 (Table 1). Common haplotypes (e.g., Hap 22 and Hap 21) and rare haplotypes (e.g., Hap 23, Hap 30, Hap 32, and Hap 33) co-existed within populations in coastal cities such as Qingdao, Guangzhou, and Fuzhou (Fig. 1). Populations in inland cities such as Chongqing and Guiyang only had rare haplotypes like Hap 30 and Hap 31. Three clades could be identified from the rooted neighbor-joining tree and the median-joining network based on the haplotypes (Supplementary Fig. 2).
a Study area. b Genetic patterns, the color proportions of the pie indicated the frequency of each cpDNA haplotype in the population. Abbreviations: Mudanjiang (MDJ), Changchun (CC), Shenyang (SY), Fushun (FS), Qinhuangdao (QHD), Beijing (BJ), Qingdao (QD), Shanghai (SH), Nanjing (NJ), Wuhan (WH), Changsha (CS), Fuzhou (FZ), Guangzhou (GZ), Chongqing (CQ), Guiyang (GY). Numbers are ids of the populations collected in the city.
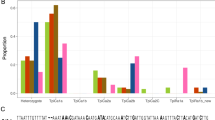
The results from GENELAND and STRUCTURE jointly indicated that common ragweed populations in East China cities could be classified into seven genetic clusters (Fig. 2). The posterior probability density calculated in GENELAND showed mixing around an apparent plateau at the value k = 7 (Supplementary Fig. 3). The result from STRUCTURE showed that the highest Delta K value (Delta K = 31) was obtained at K = 5, followed by K = 2, 4, and 3 (Supplementary Fig. 4). The membership of each population is shown in Supplementary Table 1. The classification of common ragweed populations into the seven clusters was supported by the moderate within-cluster pairwise FST values and significant or notable pairwise FST values among most genetic clusters (Supplementary Fig. 5). The pairwise FST values between Cluster D and E were moderate. Nevertheless, the two clusters were separable based on the result of STRUCTURE.
Bar heights represent the relative probability of assignment of individual plants to varying numbers of genetic clusters. Genetic clusters identified by GENELAND were indicated by letters. Abbreviations are names of sampled cities and populations: Mudanjiang (MDJ), Changchun (CC), Shenyang (SY), Fushun (FS), Qinhuangdao (QHD), Beijing (BJ), Qingdao (QD), Shanghai (SH), Nanjing (NJ), Wuhan (WH), Changsha (CS), Fuzhou (FZ), Guangzhou (GZ), Chongqing (CQ), Guiyang (GY).
The migration rates, the fraction of individuals in a population that are migrants derived from another population per generation, indicated that the metapopulations from Nanjing and Shenyang exported more current gene flows than the metapopulations from other cities (Fig. 3). The strength values were also much higher in Nanjing and Shenyang than in the remaining cities. Migration rates between paired cities can be found in Supplementary Table 2.
The similarity of common ragweed populations in East China cities to those in other countries
The common ragweed populations in East China cities were compared to those in North America and two European countries, France and Italy, where the species is an alien invasive plant (i.e., the clusters in Table 2). According to the result of GENECLASS2, at least 74% of individual plants of the genetic clusters in cities in East China were assigned to the modern North American cluster 2. The remaining individuals were assigned to the modern North American cluster 1 (no more than 21%), and the historical North American cluster (no more than 10%) in each genetic cluster. All individuals in cluster C belong to the modern North American cluster 2 while cluster F has the highest proportion of individuals from historical North American cluster (10%). No individual plant was assigned to the French and Italian clusters.
The expected heterozygosities (HE) of the populations in East China cities were not significantly different from those of the modern North American populations (p = 0.716, df = 81, F = 0.134, Supplementary Fig. 6) and the French and Italian populations (p = 0.137, df = 39, F = 2.305, Supplementary Fig. 6). The insignificant difference indicated no reduction of genetic diversity in the populations in East China cities. The inbreeding coefficients (FIS) of the populations in East China cities were significantly lower than those of the modern North American populations (p = 0.027, df = 81, F = 5.11; Supplementary Fig. 6). The low FIS value might suggest that the populations in East China cities had lower inbreeding levels than the modern North American populations.
Patterns inferred from the occurrence records of common ragweed
About 157 prefecture-level regions in China have reported the occurrences of common ragweed since 1930 (Fig. 4). The occurrence records showed that common ragweed first appeared in the Yangtze River region and then in other parts of China. Records from 1935 to 1949 were mainly along the Yangtze River. From 1970 to 1989, common ragweed appeared in regions further away from the Yangtze River, northeastern China, and the Taiwan Province. Hangzhou (first recorded in 1935), Wuhan (1946), Jiujiang (1948), Nanjing (1958), Qingdao (1964), Mudanjiang (1973), Qinhuangdao (1978), Laibin (1985), and Fuzhou (1991) could be identified as regions that have a high potential for dispersal.
Grain trade between the United States and China
From 1999 to 2009, the trade data showed that the states in the modern North American cluster 2 exported more grain products to China than those in the modern North American cluster 1 (Table 3). Importation of grains in the 15 cities showed that Nanjing and Qingdao received the most grain products from 2000 to 2006 (Fig. 5). From 1935 to 1939, the trade data revealed that Austrila, French indo-China and Thailand were the primary source countries for grain product imports to China (Supplementary Table 3).
Horizontal black bars represent the median value. Box limits indicate 75% and 25% quartiles. Whiskers indicated the minimum and maximum value. The white point indicated the mean value. The sample size for each city is 7 (n = 7). Abbreviations are names of sampled cities: Mudanjiang (MDJ), Changchun (CC), Shenyang (SY), Fushun (FS), Qinhuangdao (QHD), Beijing (BJ), Qingdao (QD), Shanghai (SH), Nanjing (NJ), Wuhan (WH), Changsha (CS), Fuzhou (FZ), Guangzhou (GZ), Chongqing (CQ), and Guiyang (GY). (Source: EPS CHINA DATA).
Discussion
The assignment tests suggested that the modern North American cluster 2 was likely the primary source of common ragweed populations in East China cities. The states within the distribution range of the modern North American cluster 2 exported far more grain products to China than other states. Studies showed that grain products in the U.S. retained many common ragweed seeds29,30. More than 90% of the grains imported by China are for consumption31. They were processed into food, feed, or alcohol in factories in urban areas32. Thus, grains contaminated by common ragweed seeds were primarily transported into urban areas. During transportation, grains can leak from vehicles33,34. The airflow created by vehicles can also facilitate the dispersal of common ragweed-contaminated grains35,36,37. Those processes can allow common ragweed seeds to escape into urban environments. Consequently, the propagule pressure of common ragweed in urban areas is high, contributing to the successful colonization of urban areas. Only a small portion of the imported grain in China is used as planting seeds31. These seeds often undergo careful screening before cultivation to ensure the desired species are planted, with non-target plants being removed during the growing process38. As a result, it is unlikely that seeds for planting are the primary source of common ragweed propagules in cities in East China. The impact of grain trade on the introduction source of common ragweed is also reflected in genetic clusters. For example, all individuals in cluster C belong to the modern North American cluster 2. This may be because all individual plants in cluster C were collected from Guangzhou, which is a major port for grain import. Our results corroborate the conclusion that introduction through contaminated food commodities is one of the presumed routes of IAS invasion39.
The assignment test results suggested that the historical North American clusters (collected during 1873 to 1939) were a minor source of common ragweed populations in cities in East China. Common ragweed was first recorded in China in 193522. Historical grain trade facilitated the introduction. Mainland China mainly imported grains from Australia, French Indo-China (including Vietnam, Cambodia, and Laos), Thailand, Kwangchowwan leased territory (a territory of China at the southern tip of the Liaodong Peninsula of Manchuria, leased to Japan in 1905), the U.S., Myanmar, Korea, Canada, and Argentina during 1935-193940,41 (Supplementary Table 3). The occurrences of common ragweed were first recorded in Vietnam, Cambodia, Laos, Thailand, Kwangchowwan leased territory, Myanmar, and Korea later than 19391,42. Therefore, those countries and regions are less likely the initial source. Common ragweed is native to the U.S., and Canada, and was introduced to Argentina before 182043. Also, the species was first recorded in Japan in 1877 and established in the country in the 1930s44, while it was first recorded in Australia in 190845. Common ragweed could be directly introduced into China from the U.S., Canada, and Argentina or indirectly from Japan and Australia. Since genetic data on common ragweed in Argentina, Japan, and Australia are unavailable, it is difficult to validate those possibilities. Nevertheless, the bridgehead introduction passage for common ragweed between Europe and Australia have already been identified18, our findings point to possible bridgehead introduction passages between China and other countries.
The minor role of the historical North American clusters as invasion sources could be attributed to the repeated modern introductions in East China cities and genetic exchange among cities. These findings support the possibility that modern genotypes have gradually replaced the genotypes introduced through historical introductions. Cluster F has the highest proportion of individual plants from the historical North American cluster. Individuals in cluster F came from Guiyang and Chongqing, which are in the inland region of China. The spread of modern genotypes of common ragweed to inland areas takes a longer time, resulting in a greater retention of historical genotypes. A previous study found that the proportion of historical North American clusters in the sources of non-urban common ragweed populations in China was three times that of modern North American clusters21. This can be attributed to the less developed transportation networks in non-urban areas, where populations of non-urban common ragweed interact less with populations from other areas. Thus, more historical genotypes retained. We offered an potential explanation to the early findings.
These findings support the hypothesis that international trade introduced common ragweed into urban areas in East China cities. We inferred that modern grain exportation from the United States to China dominated the introduction process. While economic variables such as grain importation are not directly related to the establishment of IAS, they integrate the effect of variables that directly affect the outcome of invasion, such as propagule pressure and pathways of introduction46,47. Although other studies have identified grain trade as an introduction pathway of IAS to cities through occurrence records48, our results provide new evidence from a population genetic perspective.
The existence of both the historical and modern North American clusters as invasion sources pointed to multiple introductions of common ragweed in East China cities that occurred in different periods. The spatial genetic pattern of common ragweed populations in East China cities showed that this possibility is highly likely. High genetic variability, low mean FIS, and rare bottleneck effects were observed in the populations in East China cities. Multiple introductions often lead to high genetic variability in invasive species17. The relatively low mean inbreeding coefficient (FIS = 0.342) suggests significant genetic differentiation among parental plants in most populations, which could be attributed to multiple introductions49. Rare bottleneck effects indicate that most populations did not experience a substantial reduction in population size. For species at the introduced range, the recovery of their gene pool may be replenished through multiple introductions15. These observations support the occurrence of multiple introductions.
Furthermore, we observed that populations from distant cities (e.g., Mudanjiang, Qinhuangdao, and Wuhan) were grouped together in the same genetic cluster. However, contemporary gene flows among these populations were low. The limited gene exchange between distant cities suggests that their assignment to the same genetic grouping is more likely a result of high genetic similarity among invasive source populations rather than recent gene flows50. In other words, the similar invasive source was introduced multiple times into those cities.
We also found that introductions occurred more frequently in coastal areas than inland. The co-existence of common and rare haplotypes from different clades in coastal cities indicated possible multiple introductions. The sole existence of the rare haplotype in inland cities suggested less frequent introductions. According to the literature, most areas of East China are considered suitable habitats for common ragweed23,51,52. Higher international trade activities in coastal areas than inland areas might explain this pattern because the trade increases the propagule pressure. Previous studies also found that common ragweed populations in Eastern and Western Europe were related to different regions of North America due to differences in trade patterns16,17.
Occurrence records also indicated a high likelihood of multiple introductions. For example, Qingdao first recorded common ragweed in 1964, and Fuzhou recorded the first observation in 199153,54. The two cities are far from each other (1118 km), making natural dispersal less likely. Also, the result of the genetic analysis showed that common ragweed populations from Qingdao and Fuzhou belonged to different genetic clusters. These findings support the possibility that common ragweed invaded East China cities through multiple introductions at different times.
In the spreading stage, the dispersal of common ragweed between cities in East China primarily might depend on stowaways associated with anthropogenic transportation. Studies have demonstrated that road networks and vehicle airflows promote the spread of common ragweed seeds36,55. In this study, we observed high levels of gene flow from Shenyang to Changchun (0.1544), Qingdao to Guiyang (0.1149), and Fuzhou to Guangzhou (0.1461). Those cities are connected by dense road networks.
Furthermore, the movement of soil and mowing machines may contribute to seed spread1. Common ragweed seeds can remain viable in the soil for up to 40 years45, and a single common ragweed plant can produce up to 6000 seeds56. Once established in an area, common ragweed can produce a large number of seeds that contaminate the soil over an extended period. If the invaded area is a center of horticulture or agriculture, common ragweed seeds can inadvertently be transported to other areas through the movement of soil and mowing machines. Among the studied cities in East China, Nanjing is one of the most significant horticulture centers in China, which may contribute to the high gene flow observed between Nanjing and other cities, e.g., Changsha (0.1721), Wuhan (0.1651), Shanghai (0.0889), Fuzhou (0.06), and Chongqing (0.0518).
Lastly, natural dispersal may have played a role in the spread of common ragweed seeds between cities. Barochory is the primary way of natural dispersal of common ragweed seeds1. However, the barochory distance of common ragweed seeds in terrestrial habitats was found to be less than 1 meter per year36. Hydrochory has been identified as a secondary mode of natural seed dispersal for common ragweed within its native range10,56. Its role in the dispersal of common ragweed seeds among East China cities worth further studies. We detected significant gene flow from Nanjing to Changsha (0.1721) and from Nanjing to Wuhan (0.1651). Those cities are connected by waterways. However, for other cities with significant gene flows, no direct waterway connections exist among them.
Some studies suggest that pollen may play a role in gene exchange among cities. For instance, research conducted in Europe has demonstrated that common ragweed pollen can be dispersed over several hundreds or even thousands of kilometers through the air57,58,59. However, it is important to note that the pollination rate of common ragweed pollen originating from distant locations thousands of kilometers away would be extremely low, given that common ragweed populations were not widely distributed in the cities examined in our study.
Shanghai, Guangzhou, Qingdao, and Fuzhou emerge as potential main sites of introduction. These cities, being port cities, have a substantial grain importation volume. Previous studies have suggested that port cities are more likely to serve as entry points for the introduction of alien species60. The common ragweed populations in these cities exhibit a mixture of common and rare cpDNA haplotypes from different clades, and their inbreeding coefficients are low. However, as a result of effective control measures implemented in recent years2, the genetic diversity of Shanghai’s population is low. In contrast, the populations in Qingdao, Guangzhou, and Fuzhou maintain high levels of genetic diversity. Consequently, the potential risk posed by these latter populations are therefore greater.
Nanjing and Shenyang are identified as potential primary centers for the spread of common ragweed. Our findings demonstrate significant gene flows originating from the metapopulations of common ragweed in these cities to other cities in East China. This can be attributed to the prominent roles of Nanjing and Shenyang in grain importation. Nanjing ranks first in imported grain volume among all the studied cities, while Shenyang surpasses other cities within the same genetic cluster in terms of grain imports. Shenyang’s proximity to Yingkou, a historically crucial grain importation port61,62, further strengthens its influence. In addition, both Nanjing and Shenyang serve as key transportation hubs in China63,64, contributing to their dominance in gene flows along the transportation network. Notably, Nanjing’s floriculture adds another pathway for seed dispersal65, as soil containing common ragweed seeds can be inadvertently transported to major cities through floriculture products.
Previous studies listed Wuhan, Nanjing, and Shenyang as cities with a high spread potential of common ragweed24. The gene flow pattern did not support the designation of Wuhan as a spread center. Wuhan’s spread potential may have been significantly reduced due to effective biocontrol programs applied in recent years24,66. Another possibility is the small sample size in Wuhan because only one urban population was located in Wuhan. In addition to the cities mentioned above, we inferred that Hangzhou, Jiujiang, Qingdao, Mudanjiang, Qinhuangdao, Laibin, and Fuzhou had high spread potentials of common ragweed in China based on the temporal information of occurrence data. Among these cities, samples collected from Fuzhou, Qingdao, Mudanjiang, and Qinhuangdao were analyzed, but the metapopulations in Mudanjiang, and Qinhuangdao had low gene flows to nearby sampling cities in East China. The results of the genetic analysis did not corroborate with the inferences based on the occurrence records. The discrepancy might be due to the subjectivity of the inference method and the uncertainties associated with historical occurrence data.
Findings from this study can contribute to the control of common ragweed in the following aspects: (1) Our results show that urban areas can serve as landing points for and spread hubs of common ragweed. In order to control common ragweed, controlling the spread among urban areas should not be left out of the overall control program. (2) There is plenty of room for improvement in quarantine work at Chinese port cities because the primary source of common ragweed might be from the modern North American clusters. Also, coastal cities have experienced more introduction events than inland cities. Strict inspection and quarantine of grains imported from regions that are known sources of common ragweed seeds should be practiced in ports that handle grain importation. Monitoring programs and awareness campaigns should be set up in these cities to monitor common ragweed populations. Waste from factories that process imported grains must be treated before final disposal.
While our study contributes a new understanding of the invasion pathway of common ragweed in urban areas, several limitations should be addressed in future studies. First, although our field campaign covered the East China cities, more samples from more cities can further reduce uncertainty in the results. Second, we could only obtain genetic data for North America and two European countries after searching through the major international genetic databases and contacting authors who published papers on common ragweed’s genetics. Due to the lack of data, we could not explore the contribution of other potential sources to the genetic pattern of common ragweed populations in East China cities. For example, we could not confirm the bridgehead introduction passage from Argentina, Japan and Australia due to the lack of genetic data on common ragweed in these countries.
Knowledge of the invasion pathway of common ragweed in urban areas is crucial for controlling this IAS. However, limited information is available because studies have not adequately addressed this topic. This study examined the spatial genetic pattern of common ragweed plants in urban areas in East China cities. We also explored the possible role played by the grain trade between China and the U.S. in shaping the genetic pattern. The results indicated that common ragweed in East China cities had experienced multiple introductions at different times. We inferred that the modern-day grain trade between the U.S. and China could be the primary source of propagule. Finally, we identified cities that might serve as spread centers by examining the gene flows among the cities and the temporal information in occurrence records. Our findings contribute new knowledge on the invasion pathway of common ragweed in urban areas. Also, the information gained in this study is useful for controlling common ragweed in urban areas in China and other countries. Nevertheless, the entire study should be viewed only as the beginning of efforts to reveal invasion pathways of common ragweed and other IAS in urban areas. More studies combining the strength of observations with population genetic methods are needed to understand the pathways and driving mechanisms better.
Methods
Study area
The study area includes cities in East China (Supplementary Fig. 7). East China holds most of the Chinese population25 and is the main suitable habitat for common ragweed51. To locate the current distribution of common ragweed populations in cities, we referred to the literature15,67,68 and consulted local experts. Also, we checked the geographic locations of occurrence records of common ragweed contained in the Chinese Virtual Herbarium (CVH)69. Based on the compiled occurrence records, we selected 19 East China cities with reliable records of common ragweed to conduct field surveys and were able to find urban populations of common ragweed in 15 cities.
Data collection
We collected samples from 37 common ragweed populations in the 15 cities between 2017 and 2018 (Supplementary Table 1). The sampled populations were located in urban areas and were at least 3 km from each other. According to the varied population size of urban populations, 6–19 plants were sampled at least 2 m from each other inside each population. We collected at least five fresh leaves from each sampled plant and stored the leaves in sealed plastic bags filled with silica beads for desiccation. We recorded the longitudes and latitudes of sampling sites using a handheld GPS unit (AGM X2, with an accuracy of 10 m) during the field campaign.
Genomic DNA was extracted from the leaf samples using the CTAB method70,71. We also obtained the genomic DNA of four French and Italian populations and one North American population from Ciappetta et al.72. Six microsatellite markers were used to mark the genomic DNA of the common ragweed populations mentioned above. To ensure compatibility among Chinese, French and Italian, and North American populations, the microsatellite markers used in this research were the same as those used by Martin et al.19. Microsatellite primer sequences were Gen-Bank sequences FJ595149 (Ambart04), FJ5950 (Ambart06), FJ5952 (Ambart17), FJ5953 (Ambart18), FJ595155 (Ambart24), and FJ595156 (Ambart27) (Supplementary Table 4). In-house ROX-labeled size standards were used for genotyping73. GENEMarker version 1.5 was utilized for microsatellite loci fragment analysis and allele calling. Microsatellite loci fragment data of modern and historical North American populations collected from Martin et al.19 were re-read for data consistency19.
We sequenced chloroplast DNA (cpDNA) of the 37 East China populations. The two spacers (atpH–atpF and psbK–psbI) of chloroplast DNA spacer regions in common ragweed were chosen for analysis (Supplementary Table 5). Chloroplast DNA chromatograms of sequenced samples were aligned using MAFFT74 and Geneious version 11.0.4 (Biomatters Ltd., Auckland, New Zealand).
To further trace the invasion sources of common ragweed populations in East China cities, we obtained microsatellite loci fragment data of 45 North American populations and 426 historical herbarium individual plants (Supplementary Table 6) from Martin et al.19. The data was collected in the summer of 2009.
DNA extraction, amplification, and sequencing of nuclear microsatellites and cpDNA were completed by the Bio-ulab company (Beijing, China) (Details of the procedure were included in the supplementary materials).
Data analysis
Tests of genotypic linkage equilibrium and estimation of null allele frequency were performed in GENEPOP 4.7.075. Genetic diversity indices, including the number of different alleles (NA), expected heterozygosity (HE), and inbreeding coefficients (FIS) were estimated in GenAlEx 6.51 and GENEPOP 4.7.075,76 to represent diversity at nuclear microsatellite loci, respectively. Deviations from Hardy-Weinberg equilibrium of sampled populations were assessed using FIS77, and their significance was tested using GENEPOP 4.7.075. The potential bottleneck of sampled populations was detected using BOTTLENECK version 1.2.0.278.
We aligned the atpH–atpF and psbK–psbI sequences of common ragweed from CBOL Plant Working Group79 with our haplotypes to determine the correctness of our sequence. The number of haplotypes (h), haplotype diversity (HD), and nucleotide diversity (π) of sampled populations were computed using DnaSP 680. Except for software packages specifically noted, all statistical analyses were carried out using the basic R 4.0.281.
The Bayesian clustering algorithms contained in STRUCTURE version 2.3.382 and GENELAND version 4.0.383 were used to cluster common ragweed populations based on the nuclear microsatellite and the cpDNA haplotype (details of the parameters were in the supplementary materials). Pairwise population-specific fixation indices (FST) between sampled populations were calculated based on nuclear microsatellite markers. Pairwise population FST were calculated in the diveRsity package84.
The geographical distribution of cpDNA haplotypes was visualized using ArcGIS Desktop 10.2 (ESRI, Redlands, CA, USA). A rooted neighbor-joining tree with Ambrosia trifida L. as an outgroup was constructed using Geneious version 11.0.4 (Biomatters Ltd., Auckland, New Zealand) with a Tamura-Nei genetic distance model using 1000 replicates. A median-joining network of haplotypes was generated by using NETWORK 10.185. The tree and the network were used to explore the relationships among cpDNA haplotypes.
The gene flows among the sampled cities were estimated using BayesAss v3.0.486. A genetic graph was built using the gene flows among the sampled cities. The R packages POPGRAPH87 and IGRAPH88 were used to construct the graph and calculate graph centrality measures. More details of the analyses can be found in the supplementary materials.
Genetic diversity indices, including HE and FIS of French and Italian, and North American populations, were estimated in GenAlEx 6.51 and GENEPOP 4.7.075,76, respectively. Analysis of variance (ANOVA) was used to test differences among the geographic ranges in HE and FIS using the stats package81. In the previous study, the source populations of common ragweed in East China was identified21. This study further investigated the primary source of propagule and dispersal centers for common ragweed in cities in East China. Each plant was assigned to French and Italian, modern North American, and historical North American populations. The North American samples were grouped by the genetic clusters identified by Martin et al.19. The historical clusters are comprised of herbarium specimens collected between 1873 and 1939, while the modern clusters are consisted of samples collected in 200919. The assignment was performed in the GENECLASS2 software89.
Because this study focused on exploring the role of the grain trade in the introduction of common ragweed to East China cities, we collected exportation and importation data. Exports of grain products from states in the United States to China from 1999 to 2009 were provided by the Foreign Trade Division, the U.S. Census Bureau. The exports were adjusted by using the Consumer Price Index. This time range was chosen because the U.S. Census Bureau has provided data on grain trade to China since 1999, and the genetic analysis of the invasion source was based on modern North American common ragweed genetic data collected in 200919. Data collection and analysis were conducted using Excel.
Data on grain importation from 2000 to 2006 in studied cities were collected from EPS CHINA DATA (http://microdata.sozdata.com/home.html). The time range was chosen because China’s customs data on grain imports were available in EPS CHINA DATA from 2000, and China’s customs began to implement stricter quarantine measures for common ragweed seed in 2007. Data on China’s imports of grain products from 1935 to 1939 were collected from Chinese Maritime Customs Historical Material: 1859-194839. This time range was selected due to the historical North American genetic data being based on herbarium specimens collected between 1873 and 1939. In addition, common ragweed was first recorded in China in 193522, and importation data prior to 1935 utilized a different statistical scope compared to the later years. Data collection and analysis were performed using the stringi and the dplyr package90,91.
We searched occurrence data of common ragweed from databases such as the Global Biodiversity Information Facility (GBIF)92, CVH42, Plant Photo Bank of China93, and the Plant Specimen Database94 until June 07. 2021. The data was supplemented with survey data, anecdotal descriptions, and images from literature. We removed occurrence records in rural areas. We then generated the temporal and spatial distribution map of common ragweed in China using ArcGIS Desktop 10.2. We defined the first present time of common ragweed occurrence data in the prefecture-level city as the first observation time of common ragweed in that prefecture-level city. We inferred a city as a regional first landing point when common ragweed plants were observed in the city earlier than any other adjacent city. Bullock et al.10. assigned areas with more distribution records of common ragweed as the spread center because they had higher dispersal potential. Marchioro et al.95. assumed that if areas adjacent to an invasion area is suitable for the invasive species, the invasion area is designated as the spread center. Building upon the methods used in above studies, we classified a city as the potential spread center if it met three criteria: (1) The city first reported the occurrence of common ragweed in the region; (2) It has consistent occurrence records over a 20-year period; (3) Its adjacent cities are suitable habitats for common ragweed.
Statistics and reproducibility
Genetic diversity indices (NA, HE, FIS, h, HD and π) were estimated in GenAlEx 6.5176, GENEPOP 4.7.075 and DnaSP 680. The potential bottleneck was detected using BOTTLENECK version 1.2.0.278. STRUCTURE version 2.3.382 and GENELAND version 4.0.383 were used to cluster common ragweed populations. The rooted neighbor-joining tree was constructed using Geneious version 11.0.4. The median-joining network of haplotypes was generated by using NETWORK 10.185. The gene flows were estimated using BayesAss v3.0.486. The assignment was performed in the GENECLASS2 software89. Except for software packages specifically noted, all statistical analyses were carried out using the basic R 4.0.281. The specific details of the analyses and sample sizes are described in the methods above.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Microsatellite genotypes, chloroplast haplotypes, input files used in the analysis, and the source data behind the graphs in Figs. 1,2,4,5 are accessible through the Mendeley Data with identifier DOI:10.17632/kxgtk8r5kz.1.Chloroplast intergenic spacer locus sequences that support the findings of this study have been deposited in GenBank with the accession numbers MZ826750–MZ826763.The source data behind the graphs in Fig.3 are available in Supplementary Table 2.
Code availability
R codes used in the analysis are accessible through the Mendeley Data with identifier https://doi.org/10.17632/kxgtk8r5kz.1.
References
Montagnani, C., Gentili, R., Smith, M., Guarino, M. & Citterio, S. The worldwide spread, success, and impact of ragweed (Ambrosia spp.). CRC Crit. Rev. Plant Sci. 36, 139–178 (2017).
Zhou, Z. S., Wan, F. H. & Guo, J. Y. Common ragweed Ambrosia artemisiifolia L. in Biological Invasions And Its Management In China (eds. Wan, F. H., Jiang, M. X. & Zhan, A. B.) 99–109 (Springer Singapore, 2017).
Ziska, L. H. et al. Cities as harbingers of climate change: common ragweed, urbanization, and public health. J. Allergy Clin. Immunol. 111, 290–295 (2003).
Ghiani, A., Aina, R., Asero, R., Bellotto, E. & Citterio, S. Ragweed pollen collected along high-traffic roads shows a higher allergenicity than pollen sampled in vegetated areas. Allergy 67, 887–894 (2012).
Schaffner, U. et al. Biological weed control to relieve millions from Ambrosia allergies in Europe. Nat. Commun. 11, 1745 (2020).
Yair, Y. et al. Ragweed species (Ambrosia spp.) in Israel: distribution and allergenicity. Aerobiologia 35, 85–95 (2019).
Brandes, D. & Nitzsche, J. Biology, introduction, dispersal, and distribution of common ragweed (Ambrosia artemisiifolia L.) with special regard to Germany. Nachrichtenblatt des. Dtsch. Pflanzenschutzdienstes 58, 286–291 (2006).
Galzina, N., Barić, K., Šcepanović, M., Goršić, M. & Ostojić, Z. Distribution of invasive weed Ambrosia artemisiifolia L. in Croatia. Agric. Conspec. Sci. 75, 75–81 (2010).
Mang, T., Essl, F., Mosef, D. & Dullinger, S. Climate warming drives invasion history of Ambrosia artemisiifolia in central Europe. Preslia 90, 59–81 (2018).
Bullock, J. et al. Assessing and Controlling the Spread and the Effects of Common Ragweed in Europe. (European Commission, 2010).
Roy‐Dufresne, E. et al. Modeling the distribution of a wide‐ranging invasive species using the sampling efforts of expert and citizen scientists. Ecol. Evol. 9, 11053–11063 (2019).
Yakub, M. & Tiffin, P. Living in the city: Urban environments shape the evolution of a native annual plant. Glob. Chang. Biol. 23, 2082–2089 (2016).
Hernández, F., Presotto, A., Poverene, M. & Mandel, J. R. Genetic diversity and population structure of wild sunflower (Helianthus annuus L.) in Argentina: reconstructing its invasion history. J. Hered. 110, 746–759 (2019).
Rosenthal, D. M., Ramakrishnan, A. P. & Cruzan, M. B. Evidence for multiple sources of invasion and intraspecific hybridization in Brachypodium sylvaticum (Hudson) Beauv. in North America. Mol. Ecol. 17, 4657–4669 (2008).
Gaudeul, M., Giraud, T., Kiss, L. & Shykoff, J. A. Nuclear and chloroplast microsatellites show multiple introductions in the worldwide invasion history of common ragweed, Ambrosia artemisiifolia. PLoS ONE 6, e17658 (2011).
Gladieux, P. et al. Distinct invasion sources of common ragweed (Ambrosia artemisiifolia) in Eastern and Western Europe. Biol. Invasions 13, 933–944 (2011).
Genton, B. J., Shykoff, J. A. & Giraud, T. High genetic diversity in French invasive populations of common ragweed, Ambrosia artemisiifolia, as a result of multiple sources of introduction: genetic diversity in invasive common ragweed. Mol. Ecol. 14, 4275–4285 (2005).
van Boheemen, L. A. et al. Multiple introductions, admixture and bridgehead invasion characterize the introduction history of Ambrosia artemisiifolia in Europe and Australia. Mol. Ecol. 26, 5421–5434 (2017).
Martin, M. D. et al. Herbarium specimens reveal a historical shift in phylogeographic structure of common ragweed during native range disturbance. Mol. Ecol. 23, 1701–1716 (2014).
Li, F. F. et al. Patterns of genetic variation reflect multiple introductions and pre-admixture sources of common ragweed (Ambrosia artemisiifolia) in China. Biol. Invasions 21, 2191–2209 (2019).
Lu, S. et al. Genetic patterns reveal differences between the invasion processes of common ragweed in urban and non-urban ecosystems. Glob. Ecol. Conserv. 38, e02214 (2022).
Liu, J., Feng, S. & Mu, Y. Ecological characteristics of the ragweeds and countemeasure of the biological weed control. J. Northeast. Univ. 3, 61–67 (1997).
Qin, Z., DiTommaso, A., Wu, R. S. & Huang, H. Y. Potential distribution of two Ambrosia species in China under projected climate change. Weed Res. 54, 520–531 (2014).
Wan, F. H., Hou, Y. & Jiang, M. X. Invasion Biology (China Science Publishing & Media Ltd., 2015).
Gao, X. D., Wang, X. X. & Zhu, B. Q. The distribution of Chinese minority populations and its change based on the study of the Hu Huanyong line. Int. J. Anthropol. Ethnol. 1, 2 (2017).
Hu, H. Y. The distribution, regionalization and prospect of China’s population. Acta Geogr. Sin. 45, 139–145 (1990).
Chapman, D. S. et al. Modelling the introduction and spread of non-native species: International trade and climate change drive ragweed invasion. Glob. Chang. Biol. 22, 3067–3079 (2016).
Chauvel, B., Dessaint, F., Cardinal-Legrand, C. & Bretagnolle, F. The historical spread of Ambrosia artemisiifolia L. in France from herbarium records. J. Biogeogr. 33, 665–673 (2006).
Simard, M. J. & Benoit, D. L. Potential pollen and seed production from early- and late-emerging common ragweed in corn and soybean. Weed Technol. 26, 510–516 (2012).
Schwartz-Lazaro, L. M. et al. Seed-shattering phenology at soybean harvest of economically important weeds in multiple regions of the United States. Part 1: broadleaf species. Weed Sci. 69, 95–103 (2021).
Zhong, Y. Quantitative interpretation of ‘basic self-sufficiency in cereals and absolute security in food rations’. Grain Issues Res. 2, 20–26 (2015).
Xie, Z. Y., Wang, R. Y. & Sun, R. Industrial cluster development of grain and oil. Cereals Oils Pro. 7, 14–21 (2015).
Alavi, H. R., Htenas, A., Kopicki, R., Shepherd, A. W. & Clarete, R. Trusting Trade and the Private Sector for Food Security in Southeast Asia (World Bank, 2012).
Pereira, P. S. X. et al. Wheat grain losses in highway transportation. J. Exp. Agric. Int. https://doi.org/10.9734/jeai/2019/v39i230328 (2019).
von der Lippe, M., Bullock, J. M., Kowarik, I., Knopp, T. & Wichmann, M. Human-mediated dispersal of seeds by the airflow of vehicles. PLoS ONE 8, e52733 (2013).
Lemke, A., Kowarik, I. & von der Lippe, M. How traffic facilitates population expansion of invasive species along roads: the case of common ragweed in Germany. J. Appl. Ecol. 56, 413–422 (2018).
Lemke, A. Interaction of traffic intensity and habitat features shape invasion dynamics of an invasive alien species (Ambrosia artemisiifolia) in a regional road network. NeoBiota 64, 155–175 (2021).
Lambert, D. Seed Production and Training Manual. (Federal Ministry of Food, Agriculture and Consumer Protection, 2012).
Harrower, C., Scalera, R., Pagad, S., Schönrogge, K. & Roy, H. Guidance for interpretation of the CBD categories of pathways for the introduction of invasive alien species. (European Commission, 2020). https://doi.org/10.2779/6172.
Mao, J. Q., Huang, S. Q. & Ma, Z. D. Chinese Maritime Customs Historical Material 1859-1948. (Jing Hua Press, 2001).
Xu, D. Statistics of agricultural production and trade in modern China. (Shanghai People′s Publishing House, 1983).
Institute of Botany, The Chinese Academy of Sciences. The Chinese Virtual Herbarium. https://www.cvh.ac.cn/index.php (2021).
Anton, A. M. & Zuloaga, F. O. Flora Argentina. Plantas vasculares de la Republica Argentina (2023).
National Institute for Environmental Studies. Invasive species of Japan. https://www.nies.go.jp/biodiversity/invasive/DB/detail/80400e.html (2021).
Essl, F. et al. Biological flora of the British Isles: Ambrosia artemisiifolia. J. Ecol. 103, 1069–1098 (2015).
Lockwood, J. L., Cassey, P. & Blackburn, T. The role of propagule pressure in explaining species invasions. Trends Ecol. Evol. 20, 223–228 (2005).
Pyšek, P. et al. Disentangling the role of environmental and human pressures on biological invasions across Europe. Proc. Natl Acad. Sci. USA 107, 12157–12162 (2010).
Ikeda, M. et al. The role of weed seed contamination in grain commodities as propagule pressure. Biol. Invasions 24, 1707–1723 (2022).
Tang, J. et al. Multiple introductions and genetic admixture facilitate the successful invasion of Plantago virginica into China. Biol. Invasions 24, 2261–2272 (2022).
Hagenblad, J. et al. Low genetic diversity despite multiple introductions of the invasive plant species Impatiens glandulifera in Europe. BMC Genet. 16, 103 (2015).
Chen, H., Chen, L. J. & Albright, T. P. Developing habitat-suitability maps of invasive ragweed (Ambrosia artemisiifolia.L) in China using GIS and statistical methods. in GIS for health and the environment (eds. Lai, P. C. & Mak, A. S. H.) 106–121 (Springer Berlin Heidelberg, 2007).
Liu, X. L. et al. The current and future potential geographical distribution of common ragweed, Ambrosia artemisiifolia in China. Pak. J. Bot. 53, 167–172 (2021).
Cui, H. Herbarium, Institute of Botany, CAS. https://pe.ibcas.ac.cn/index.html# (19640730).
Huang, K. & Guo, Q. The dangerous weed Common Ragweed was first found in Changle County. Fujian Prov. APQ 1, 45–45 (1993).
Lavoie, C. & Jodoin, Y. & De Merlis, A. G. How did common ragweed (Ambrosia artemisiifolia L.) spread in Québec? A historical analysis using herbarium records: Herbarium records and the spread of common ragweed. J. Biogeogr. 34, 1751–1761 (2007).
Fumanal, B., Chauvel, B., Sabatier, A. & Bretagnolle, F. Variability and Cryptic Heteromorphism of Ambrosia artemisiifolia Seeds: What Consequences for its Invasion in France? Ann. Bot. 100, 305–313 (2007).
Zink, K., Vogel, H., Vogel, B., Magyar, D. & Kottmeier, C. Modeling the dispersion of Ambrosia artemisiifolia L. pollen with the model system COSMO-ART. Int. J. Biometeorol. 56, 669–680 (2012).
Smith, M. et al. Geographic and temporal variations in pollen exposure across Europe. Allergy 69, 913–923 (2014).
Celenk, S. & Malyer, H. The occurrence of Ambrosia pollen in the atmosphere of Northwest Turkey: investigation of possible source regions. Int. J. Biometeorol. 61, 1499–1510 (2017).
Padayachee, A. L. et al. How do invasive species travel to and through urban environments? Biol. Invasions 19, 3557–3570 (2017).
Chi, Q. F. A study on the trade network of soybean in northeast China region in modern times (1861-1931). (Hebei University, 2013).
Zhang, X. Modern northeast grain trade. (Harbin Normal University, 2014).
Wu, L. Y. Shenyang’s position and function in northeast. Asia. NEAF 4, 68–69 (1994).
Yang, T. & Yang, S. F. Strengthen the status of Nanjing as a transport center, accelerate the growth of the metropolis of Nanjing. Mod. Urban Stud. 1, 28–33 (2002).
Li, H. & Chen, C. The current situation and development countermeasures in the circulation of Nanjing’s flower industry. Jiangsu Agric. Sci. 39, 540–544 (2011).
Zhou, Z. S., Guo, J. Y. & Wan, F. H. Review on management of Ambrosia artemisiifolia using natural enemy insects. Chin. J. Biol. Control 31, 657–665 (2015).
Huang, J. et al. Genetic divergence of populations of Ambrosia artemisiifolia L. in Guangdong. Guangdong Agric Sci. 3, 135–138 (2012).
Wan, F. H., Wang, R. & Qiu, S. B. Control method of severe invasive weed common ragweed. 40 th Anniversary of Peking Insect Association 155–156 (1990).
Institute of Botany, The Chinese Academy of Sciences. The Chinese Virtual Herbarium. https://www.cvh.ac.cn/index.php (2017).
Lee, S. B., Milgroom, M. G. & Taylor, J. W. A rapid, high yield mini-prep method for isolation of total genomic DNA from fungi. Fungal Genet. Rep. 35, 23 (1988).
Zhang, Y. J., Zhang, S., Liu, X. Z., Wen, H. A. & Wang, M. A simple method of genomic DNA extraction suitable for analysis of bulk fungal strains: fungal DNA isolation by thermolysis. Lett. Appl. Microbiol. 51, 114–118 (2010).
Ciappetta, S. et al. Invasion of Ambrosia artemisiifolia in Italy: assessment via analysis of genetic variability and herbarium data. Flora 223, 106–113 (2016).
DeWoody, J. A. et al. Universal method for producing ROX-labeled size standards suitable for automated genotyping. BioTechniques 37, 348–352 (2004).
Katoh, K. & Toh, H. Parallelization of the MAFFT multiple sequence alignment program. Bioinformatics 26, 1899–1900 (2010).
Rousset, F. GENEPOP’007: a complete re-implementation of the genepop software for Windows and Linux. Mol. Ecol. Resour. 8, 103–106 (2008).
Peakall, R. & Smouse, P. E. GenAlEx 6.5: Genetic analysis in Excel. Population genetic software for teaching and research-an update. Bioinformatics 28, 2537–2539 (2012).
Weir, B. S. & Cockerham, C. C. Estimating F-statistics for the analysis of population structure. Evolution 38, 1358–1370 (1984).
Cornuet, J. M. & Luikart, G. Description and power analysis of two tests for detecting recent population bottlenecks from allele frequency data. Genetics 144, 2001–2014 (1996).
Hollingsworth, P. M. et al. A DNA barcode for land plants. Proc. Natl Acad. Sci. USA 106, 12794–12797 (2009).
Rozas, J. et al. DnaSP 6: DNA sequence polymorphism analysis of large data sets. Mol. Biol. Evol. 34, 3299–3302 (2017).
R. Core Team. R: A Language and Environment for Statistical Computing. (R Foundation for Statistical Computing, 2018).
Pritchard, J. K., Stephens, M. & Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 155, 945–959 (2000).
Guillot, G., Estoup, A., Mortier, F. & Cosson, J. F. A spatial statistical model for landscape genetics. Genetics 170, 1261–1280 (2005).
Keenan, K., McGinnity, P., Cross, T. F., Crozier, W. W. & Prodöhl, P. A.diveRsity: an R package for the estimation and exploration of population genetics parameters and their associated errors. Methods Ecol. Evol. 4, 782–788 (2013).
Bandelt, H., Forster, P. & Röhl, A. Median-joining networks for inferring intraspecific phylogenies. Mol. Biol. Evol. 16, 37–48 (1999).
Wilson, G. A. & Rannala, B. Bayesian inference of recent migration rates using multilocus genotypes. Genetics 163, 1177–1191 (2003).
Dyer, R. J. GeneticStudio: a suite of programs for spatial analysis of genetic-marker data. Mol. Ecol. Resour. 9, 110–113 (2009).
Csardi, G. & Nepusz, T. The igraph software package for complex network research (2006).
Piry, S. et al. GENECLASS2: a software for genetic assignment and first-generation migrant detection. J. Hered. 95, 536–539 (2004).
Gagolewski, M. stringi: Fast and Portable Character String Processing in R. J. Stat. Soft. 103, (2022).
Wickham, H. et al. dplyr: A Grammar of Data Manipulation. (2023).
GBIF.org (04 June 2021) GBIF Occurrence Download. https://doi.org/10.15468/dl.fjq5mm.
Institute of Botany, Chinese Academy of Sciences. Plant photo bank of China. http://ppbc.iplant.cn/ (2021).
College of Life Sciences, Sichuan University. The Plant Specimen Database. http://mnh.scu.edu.cn (2021).
Marchioro, C. A. & Krechemer, F. S. Potential global distribution of Diabrotica species and the risks for agricultural production. Pest. Manag. Sci. 74, 2100–2109 (2018).
Acknowledgements
We thank all people who assisted us in collecting samples during our field trips. We are grateful for the microsatellite loci fragment data of modern and historical North American populations shared by Dr. Micheal D. Martin from the Norwegian University of Science and Technology. We also want to thank the two anonymous reviewers for helping us to improve the paper. This work is supported by a grant from the National Natural Science Foundation of China (Grant no. 3217542).
Author information
Authors and Affiliations
Contributions
S.L. conceived the research idea, designed and conducted field surveys, analyzed data, and wrote the manuscript. X.L. conducted field surveys, interpreted the results, and designed figures. H.W. interpreted the results and revised the manuscript. R.G. and S.C. provided the DNA materials of common ragweed in Europe and revised the manuscript. J.Y. and J.J. conducted field surveys, analyzed data, and interpreted the results. J.L. provided information about grain trade and importation between China and USA. J.Y. conceived the research idea, secured the funding, designed field surveys, interpreted the results, and revised the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editor: Christina Karlsson Rosenthal.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lu, S., Luo, X., Wang, H. et al. China-US grain trade shapes the spatial genetic pattern of common ragweed in East China cities. Commun Biol 6, 1072 (2023). https://doi.org/10.1038/s42003-023-05434-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-023-05434-5
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.