Abstract
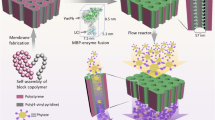
The identification or design of biocatalysts to mitigate the accumulation of plastics, including sub-micro- and nano-sized polyethylene terephthalate (nPET), is becoming a global challenge. Here we computationally incorporated two hydrolytic active sites with geometries similar to that of Idionella sakaiensis PET hydrolase, to fragaceatoxin C (FraC), a membrane pore-forming protein. FraCm1/m2 could be assembled into octameric nanopores (7.0 nm high × 1.6–6.0 nm entry), which deconstructed (40 °C, pH 7.0) nPET from GoodFellow, commodities and plastic bottles. FraCm1 and FraCm2 degrade nPET by endo- and exo-type chain scission. While FraCm1 produces bis(2-hydroxyethyl) terephthalate as the main product, FraCm2 yields a high diversity of oligomers and terephthalic acid. Mechanistic and biochemical differences with benchmark PET hydrolases, along with pore and nPET dynamics, suggest that these pore-forming protein catalytic nanoreactors do not deconstruct macro-PET but are promising in nanotechnology for filtering, capturing and breaking down nPET, for example, in wastewater treatment plants.

Similar content being viewed by others
Main
The World Economic Forum forecasted that by 2050 the production and use of plastics will grow at a rate of 4% per year1,2, making them the most common waste material in the world, accounting for >380 million tons. Their cumulative greenhouse gas emissions from production to disposal may be equivalent to 10–13% of the total carbon budget; 10.2% are predicted to correspond to polyethylene terephthalate (PET) plastics3,4. Approximately 80% of waste objects are composed of macroplastics, with PET accounting for 14.4% of total plastic waste1,2,3,4, which can reach the oceans from land2,5, and be degraded over time to microplastics (5 mm to 1 µm), sub-microplastics (100 nm to 1 µm), and nanoplastics (1 nm to 100 nm)1,2,6. Recent studies confirmed that PET particles comprise at least 5% of the total identified plastic particles1,2, which can be found in concentrations equivalent to trillions of particles in the air of some cities7, from 2,649 to 6,292 particles per litre in mineral water8, and from 5 to 52.3 ng ml−1 in mountain and Antarctica ice samples9. The ubiquity of plastic debris is causing an unprecedented ecological crisis1,2.
PET recycling is theoretically attainable10,11, compared with other extremely challenging plastics, such as thermosets6. While there are more widely used and better established chemical recycling methods for PET10, enzymatic PET recycling and upcycling strategies have been demonstrated as necessary alternatives10,11. Over the past 18 years, only a few benchmark enzymes from cultured and uncultured microorganisms have been shown to degrade macro-PET12, some of which have been engineered to increase the stability (up to +37.5 °C) and activity at temperatures close to or exceeding the PET glass transition temperature (TG), ∼70 °C. This temperature is recommended to access amorphous regions of PET polymer12, as it is physically impossible to enzymatically degrade crystalline PET, having a melting temperature (Tm) of 260 °C (Supplementary Note 1). Notable engineered examples are ThermoPETase13 and HotPETase14 from Idionella sakaiensis, and LCCICCG and LCCWCCG from leaf-branch compost15. Although challenging12, the degradation of PET below TG has also been reported, notably, for the machine learning-engineered variant FAST-PETase that completely depolymerizes at 50 °C untreated postconsumer-PET16, and for the I. sakaiensis PETase (IsPETase), active at 30 °C but labile at 37 °C after 24 h of incubation12,17, and its redesigned variant DuraPETase that is long-term active at 37 °C (ref. 18).
Micro-, sub-micro- and nano-PET, hereinafter referred to as nPET, pollution has also been targeted for enzymatic deconstruction. DuraPETase degrades PET microplastics (10–50 µm) and nanoplastics (50–100 nm) at concentrations (0.2–0.3 g l−1) 100-fold higher than those in wastewater plants, within 20 days at 37 °C (micro-) and within 1 h at 37 °C (nano-)18. The PETase from Thermobifida fusca, TfCut2, degrades nPET (100–164 nm) at 60 °C (refs. 19,20). These studies demonstrated the potential of PETases to also ameliorate nPET pollution, a line that we want to explore in this study, by designing a protein capable of filtering and degrading nPET. Within this context, actinoporins constitute a group of small and basic α pore-forming toxins produced by sea anemones as a defence mechanism21. In water, these easily produced and purified non-catalytic proteins remain perfectly soluble and stably folded; however, upon interaction with lipid membranes of a specific composition, they spontaneously become oligomeric integral membrane structures and make a pore22. Because of their physicochemical properties and configuration, most efforts towards designing pore-forming toxins have been directed at sensing molecules and sequencing23,24.
In this Article, we generated a pore-forming catalytic enzyme that could be further assembled into catalytic nanopores/nanoreactors that act as a reaction chamber to degrade nPET. This advance can be addressed due to recent modelling developments in designing de novo active sites25,26. As a target, we selected fragaceatoxin C (FraC) from Actinia fragacea, the only actinoporin membrane pore with a three-dimensional structure solved with atomic resolution21. FraC is a stable 7-nm-high octameric V-shaped pore formed by spontaneous self-assembly of the protein monomers and with access sizes on the order of 6.7 nm in the cis entry and on the order of 1.9 nm in the trans exit. A computational structure-based modelling method, the PluriZyme strategy26,27, was applied to add an artificial catalytic serine–histidine–aspartic triad and an oxyanion hole capable of ester hydrolysis to the non-catalytic FraC. These catalytic elements match the active sites of PETases12. When two newly engineered catalytic proteins (FraCm1 and FraCm2) were assembled as nanopores (npFraCm1 and npFraCm2) they became pore-based nanoreactors for the depolymerization of nPET. We discovered different degradation product profiles between them and in comparison with benchmark PETases, namely LCCWT, LCCWCCG and IsPETase12,15.
Results
In silico design of catalytic pore-forming proteins
The entire inner surface of the wild-type FraC pore structure, FraCWT, was explored using Protein Energy Landscape Exploration (PELE) software26,27, using as probes three esters (Supplementary Fig. 1) commonly hydrolysed by most esterases and lipases26. As in PluriZyme designs26,27, the initial goal is applying PELE to identify substrate binding sites for designing artificial hydrolase active sites. Energetic profiles of FraCWT from these simulations (Supplementary Data 1) are shown in Fig. 1a–c, in which the binding energy minima are split into three regions based on the distance to the trans exit of the pore. Notably, the three substrates found good pockets at similar distances, denoting a suitable ester stabilizing environment. As shown in Fig. 1d, from the cis side to the trans side, the outer cluster corresponds to the globular domain of the monomer at 70 Å from the bottom side (a more solvent-exposed side). The second cluster, located at a distance between 35 Å and 55 Å, corresponds to the hinge region of the monomer and is characterized by the presence of highly conserved residues involved in the conformational change forming the nanopore. We discovered several minima in this region but rejected them to avoid a loss of pore-forming activity28. At a distance of 20 Å, we discovered the transmembrane region configured by the N-terminal α-helices, where pockets are formed by non-conserved polar side chains facing the inner part of the channel. Both the first cluster and third cluster fulfilled our goal of introducing hydrolytic sites into the main pore, the dimensions/shapes of which can assist in channelling and holding substrates once they are assembled.
a, Energy profile of the global exploration with the branched ester glyceryl tripropionate and poses highlighted with the colour from each binding site (poses are highlighted only when the interaction energy is equal to or below −15 kcal mol−1). b, Energy profile of the global exploration with the aromatic small-sized phenyl acetate and poses highlighted with the colour from each binding site (poses are only highlighted when the interaction energy is equal to or below −12.5 kcal mol−1). c, Energy profile of the global exploration with the short chain alkenyl ester vinyl acetate and poses highlighted with the colour from each binding site (poses are highlighted only when the interaction energy is equal to or below −10 kcal mol−1). d, Cross-sectional representation of FraC with two opposing chains to visualize the localization of the different indicated binding sites (Protein Data Bank (PDB) ID: 4TSY). Computational data were collected and analysed with PELE. Calculations and raw data are shown in Supplementary Data 1.
Interestingly, two acidic residues, Asp17 and Glu24, lie in the vicinity of the identified transmembrane pocket. In general, the insertion of a negative charge into a structure is detrimental, so we typically take advantage of the presence (if any) of acidic residues. Following this approach, the Lys20His and Thr21Ser variant introduced intra- and interhelix catalytic triad hydrogen bonds. Regarding the outer pocket, we also identified several acidic residues, Asp38, Glu40 and Glu173. In this case, there is also a native histidine, His175; PELE simulations revealed that the Asp38Ser variant could generate a catalytic triad. Moreover, we wanted to add a mutation that could act as an oxyanion hole to stabilize the negative charge that appears during hydrolysis: Glu173Gln.
The potential catalytic poses involving an ester carbon-serine oxygen distance <4.5 Å concurrent with hydrogen bonds between Ser–His and His–Asp/Glu were quantified for the three exploration model esters and the two variants, FraCm1, including Lys20His and Thr21Ser, and FraCm2, including Asp38Ser and Glu173Gln. The computed PELE-normalized relative catalytic activity ranged from 19.01% to 10.98% for FraCm1 and from 2.05% to 1.42% for FraCm2 (Fig. 2a and Supplementary Data 1). Furthermore, the computed PELE-normalized relative catalytic activity for bis(2-hydroxyethyl)-terephthalate (or ETE, following the nomenclature by Schubert et al.29), and mono-(2-hydroxyethyl)-terephthalic acid (or TE), which are incomplete degradation products during the hydrolysis of PET by PETases12 (Extended Data Fig. 1), ranged from 8.79% to 3.61% for FraCm1 and from 0.47% to 1.10% for FraCm2 (Fig. 2a). FraCm1/m2 active sites can thus potentially accommodate and convert ester substrates, including TE and/or ETE. Note that although both our designs (FraCm1/m2) and IsPETase30 share an analogous active site geometry (Supplementary Fig. 2), as seen by clarifying the enzyme–substrate encounter complex by quantum/molecular mechanics (QM/MM) (Supplementary Data 1), the overall FraCm1/m2 pore is formed by eight different chains (Fig. 2b)21, thus introducing eight potential catalytic triads after pore assembly compared with the monomeric IsPETase30.
a, Correlation between computational PELE normalized relative activity (number of catalytic PELE poses/total number of accepted PELE steps and number of ester groups in the substrate) and experimental ester-hydrolysing activity (‘Exp. activity’ in the figure) for glyceryl tripropionate (labelled as GRP), vinyl acetate (VIN), phenyl acetate (PAE), TE and ETE. b, Schematic representation of PELE Local Exploration with ETE. Arrows indicate the random movement of the substrate in an N-terminal Å box during the PELE local sampling simulation. Calculations and raw data provided in Supplementary Data 1.
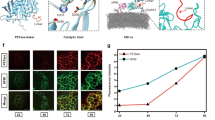
The stability of both the soluble form and the N-terminal region inserted into a membrane model by molecular dynamics (MD) was further confirmed (Supplementary Figs. 3–6). The analysis of catalytic distances for the inserted models indicated adequate hydrogen bond values, with the histidine–serine distance along the simulation being lower for the N-terminal mutant FraCm1 (Fig. 3a–d) than for the globular mutant FraCm2 (Fig. 3e–h).
FraCm1: a, N-terminal fragments are shown from residues 13 to 28, where the catalytic active site is located. Ser21 (introduced by a Thr21Ser mutation), His20 (introduced by a Lys20His mutation) and Glu24 (detected in the wild-type protein) formed the catalytic triad, and Asp17 and Glu24 (both detected in the wild-type protein) could act as the acid residue of the catalytic triad due to the proximity of the histidine residue. Hydrogen bond interactions between catalytic residues and their distances are displayed. b,c, Histogram (b) and violin plot (c) of the distances between two catalytic residues in the catalytic triad throughout the simulation. d, Distance between two catalytic residues over the simulation time. FraCm2: e, The active site in the globular region is shown. Ser38 (introduced by an Asp38Ser mutation), His175 (detected in the wild-type protein) and Glu40 (detected in the wild-type protein) conformed to the catalytic triad, and Gln173 (introduced by a Glu173Gln mutation) in the vicinity of the identified pocket conformed to the oxyanion hole. f,g, Histogram (f) and violin plot (g) of the distances between two catalytic residues in the catalytic triad throughout the simulation. h, Distance between two catalytic residues over the simulation time. MD simulations were collected and analysed with GROMACS (version 5.1.2) and OPENMM (version 7.3). Calculations and raw data are provided in Supplementary Data 1.
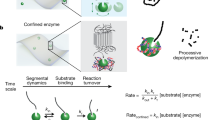
We further evaluated whether the two active centres, located inside the nanopores, could be accessed by nPET particles. Although the flexible nature of a single chain cannot directly be translated to that of a PET particle, we explored the flexibility of a PET chain of 200 units (which could mimic the molecular weights of various common PET samples, from 40 to 80 kDa) with MD simulations. The simulation shows that the PET chains in the nPET particle are dynamic entities and that their shape changes considerably over time (Fig. 4a), introducing multiple protuberances that could fit the pore dimensions, as observed in simple molecular docking (Fig. 4b). Moreover, one would expect that these protuberances could be stabilized by the pore shape (in an induced fit manner) and increase with partial hydrolysis. Moreover, one could also anticipate these particles to be formed from various chains, having multiple loose ends.
a, Solvent-accessible surface area of a nPET particle of 200 monomers over ~50 ns and 40 °C. Snapshots from the simulation are shown for different frames (where a frame is 0.1 ns). b, Graphical representation of a schematic coupling between the pore and the polymer. Calculations and raw data are provided in Supplementary Data 1.
npFraCm1 and npFraCm2 catalyse ester hydrolysis of TE and/or ETE
FraCWT, FraCm1 and FraCm2 were produced in an Escherichia coli expression system and purified (at least 95% pure; Supplementary Fig. 7 and Source Data). After evaluating that the intrinsic features and activities of the two mutants were indistinguishable from those of the wild type (Supplementary Fig. 8a–d and Supplementary Data 2), they were assembled as individual octameric pore water-soluble particles by adding them to empty nanodiscs to mimic the real situation encountered by actinoporins21 (Supplementary Fig. 9 and Source Data). See details in Supplementary Note 2. These preparations, referred to as npFraCWT, npFraCm1 and npFraCm2, were tested for their hydrolytic activity (Supplementary Note 3).
The measurement of p-nitrophenol (p-NP) release showed that npFraCm1 and npFraCm2 were capable of hydrolysing the three p-NP esters tested (pH 7.0, 40 °C), which are model esters for serine ester hydrolases (Extended Data Table 1 and Supplementary Data 3). Both engineered nanopores hydrolysed (pH 7.0, 40 °C) also the three model esters used for PELE exploration, ETE and TE, although npFraCm1 showed low specific activity for TE (Extended Data Table 1 and Supplementary Data 3). We observed notable correlations between the computed PELE-normalized relative catalytic activity and the experimental ester-hydrolysing activity (R2 = 0.72 for npFraCm1; R2 = 0.566 for npFraCm2; Fig. 2a). npFraCm1 and npFraCm2 hydrolysed ETE with kcat/Km values (Table 1, Supplementary Fig. 10 and Supplementary Data 4) that were directly benchmarked with those reported for PETases (550–17,000 M−1 s−1)31.
Using p-NP propionate as a substrate under different conditions, npFraCm1 and npFraCm2 showed maximum hydrolytic activity at a pH of approximately 9.0 and temperatures from 35 °C to 45 °C (Supplementary Fig. 11 and Supplementary Data 5). This is strongly consistent with the experimentally determined thermal melting temperature of FraCWT (Tm = 55.3 ± 0.5 °C), FraCm1 (Tm = 57.4 ± 0.5 °C) and FraCm2 (Tm = 61.6 ± 0.5 °C), as monitored by circular dichroism spectroscopy at 220 nm (Supplementary Fig. 12 and Supplementary Data 6).
Deconstruction of nPET by npFraCm1/m2
We further evaluated the possibility that npFraCm1 and npFraCm2 not only hydrolyse TE and ETE but also break down nPET particles of different types across the pores. Instead of using naturally occurring nPET particles, for example, those in wastewater treatment plants18, we chose to prepare, in vitro, particles of different nature. This allows us to assess their physical–chemical characteristics and their alterations during the degradation process, and the released products. nPET particles were produced (Supplementary Methods) from PET from a bottle of daily life (from a local shop, Granini brand, nPETb) and from a commodity thermoplastic polymer resin that is widely used for packaging (nonreheat PET resin RAMAPET N180, nPETc), as well as from two PET products from GoodFellow Cambridge: biaxially oriented crystalline PET film (GoodFellow crystalline, nPETGFc) and amorphous PET film (GoodFellow amorphous, nPETGFa). According to the manufacturers, PETc, PETGFa and PETGFc raw materials are of high purity (purity 99.9%), and the composition of PETb is according to legislation (Supplementary Note 4).
Using dynamic light scattering (DLS), the Z-average particle size (diameter) measured at 25 °C confirmed the presence of nPET particles with a stable, typical18,19,20 size distribution from 53.1 ± 0.1 nm to 108.0 ± 0.1 nm (Fig. 5a, Extended Data Table 2 and Supplementary Data 7), which is in the sub-micro to nanometre range. By differential scanning calorimetry, the degree of crystallinity (%), TG, melting temperature (Tm), cold crystallization temperature (Tc) and cold crystallization energy (ΔHc) of the bulk PET materials and the nPET particles could be also obtained (Extended Data Table 2 and Supplementary Fig. 13). The crystallinity, TG, Tm and Tc ranged from 1.3% to 34.5%, from 72.0 °C to 75.7 °C, from 245.9 °C to 251.9 °C, and from 192.8 °C to 215.1 °C, respectively, which is in the range of reported datasets12,32,33. For additional considerations, see Supplementary Note 5.
a,b, The distribution mean (Dv,50) size (±s.d. error bar) calculated by the scattered intensity for each particle size from three measurements (n = 3) is represented (a, for nPETx where x refers to GFa/GFc/b/c; b, for nPETb obtained at various transfer rates). Particles were obtained by predissolving PETb for 2 h at 150 rpm and 25 °C in 1,1,1,3,3,3-hexafluor-2-propanole (from Merck Life Science) and further transferred with the help of a burette into an ice-cold water-containing beaker (in an ice bath), which was strongly agitated (250 rpm). For a, the flow rate was fixed at 1 ml min−1, and for b, it was fixed at 0.05, 0.10, 0.25, 0.5, 1.0, 1.5 and 2.0 ml min−1 for producing 69.1, 78.8, 79.5, 85.4, 108.0, 125.9, 126.2 and 153.8 nm particles, respectively (Supplementary Methods). DLS datasets were collected at 25 °C with a Malvern Instrument (MALVERN ZETASIER NANO S ZEN 1600, IESMAT) with Dispersion Technology software 4.20 (Malvern Instrument). c, Evaluation of the degradation of nPETb particles of different sizes using npFraCm1/m2. Reaction conditions: [npFraCm1/m2], 1.5 μg ml−1; [nPETb], 1.1 mg ml−1; reaction volume, 100 μl; T, 40 °C; pH, 7.0 (20 mM HEPES buffer); reaction time, 48 h. The reactions were maintained in 2-ml safe-lock Eppendorf polypropylene tubes (ref. 0030 120.094) in a thermoshaker (model Thermomixer comfort, Eppendorf AG) at 1,000 rpm. The reactions were stopped by adding 900 μl dimethyl sulfoxide (from Merck Life Science), and the degradation products were immediately analysed by HPLC. Datasets were collected with a Varian Star LC workstation 6.41 (Varian). Quantifications of degradation products were performed by HPLC on the basis of calibrations with purified standards. Values in a and b are plotted as the mean of three independent replicates (n = 3) with the reported error ranges and s.d. calculated using Dispersion Technology software 4.20 (Malvern Instrument). Values in c and d are plotted as the mean of three independent replicates (n = 3) with the reported error ranges and s.d. calculated using the STDEV.S function in Excel 2019 (calculations and raw data are provided in Source Data). The figure was made using Excel 2019. Raw data are shown in Supplementary Data 7.
npFraCm1 or npFraCm2 were added at a concentration of 1.5 μg ml−1 (or 76 nM; Supplementary Note 6) to buffered suspensions (pH 7.0) containing 2.23 ± 0.06 mg ml−1 nPETGFa, nPETGFc, nPETb and nPETc, and the degradation products were observed by high-performance liquid chromatography (HPLC) (Fig. 6a,b) after 48 h of incubation at Topt, 40 °C. Topt refers to the temperature at which maximal ester-hydrolytic activity was observed using p-NP propionate (Supplementary Fig. 11 and Supplementary Data 5); at this temperature, npFraCm1 and npFraCm2 retained 54.7 ± 2.1% and 32.7 ± 6.3%, respectively, of their original activities after 48 h (Supplementary Fig. 14 and Supplementary Data 8). The chemical structures of all degradation products identified under our assay conditions are detailed in Extended Data Fig. 1. The identity was confirmed by high-resolution mass spectrometry (MS) analysis using products purified from reaction mixtures by semipreparative HPLC (Supplementary Fig. 15).
a,b, HPLC profiles of degradation products released by npFraCm1 (a) or npFraCm2 (b) under the experimental conditions: [npFraCm1/m2], 1.5 μg ml−1; [nPETGFa/GFc/b/c], 1.1 mg ml−1; reaction volume, 1,000 μl; T, 40 °C; pH, 7.0 (20 mM HEPES buffer); reaction time, 48 h. The reactions were maintained in 2-ml safe-lock Eppendorf polypropylene tubes (ref. 0030 120.094) in a thermoshaker (model Thermomixer comfort, Eppendorf AG) at 1,000 rpm. Aliquots of 100 μl were obtained, the reactions were stopped by adding 900 μl dimethyl sulfoxide (from Merck Life Science) and the degradation products were immediately analysed by HPLC. Datasets were collected with a Varian Star LC workstation 6.41 (Varian). All reactions and analyses were performed in triplicate (n = 3), with a representative chromatogram per enzyme and nPET particle shown. c,d, Concentration of degradation products found to be present in reaction mixtures with npFraCm1 (c) or npFraCm2 (d). Quantification was performed for products shown in a and b with unambiguous identification and for which enough material was recovered for HPLC calibration, namely from T to ETETE (Extended Data Fig. 1); purification was performed by semipreparative HPLC, and the molecular weights and structures were determined by mass spectrometry (Supplementary Fig. 15). Values are plotted as the mean of three independent replicates (n = 3) with the reported error ranges and s.d. calculated using the STDEV.S function in Excel 2019 (calculations and raw data are provided in Source Data). The figure was constructed using Excel 2019. Raw data are shown in Supplementary Data 9.
The HPLC chromatograms in Fig. 6a revealed four main degradation products when nPET particles were treated with npFraCm1: ETE, ETETE, TE and TETETE, in this order, with T being below the detection limit under our assay conditions. This substrate profile was markedly different from that of npFraCm2, which yields a higher diversity of oligomers, herein referred to as TET, TETE, TETET and TETETE, in addition to T, TE and ETE (Fig. 6b). Under the same experimental conditions, IsPETase, LCCWT and LCCWCCG released T, TE, ETE and the oligomers TET, TETE and/or TETETE, with T and TE being the main degradation products (Extended Data Fig. 2a–c); the presence of these products is in accordance with the literature and reaction mechanisms15,34. However, TET was not detected or suggested as a potential degradation product during the degradation of PET film by IsPETase34.
According to calibration with pure standards (T, TE, ETE, TET, TETE and ETETE), the concentration of degradation products at 48 h using npFraCm1 ranged from 1,718 to 5,743 µM, depending on the type of nPET (Fig. 6c and Supplementary Data 9), with nPETGFa and nPETGFc being degraded to a higher extent than nPETb and nPETc, despite their higher crystallinity. In all cases, ETE was the major degradation product, whose concentration ranged from 2.1- to 12.5-fold higher than that of TE, depending on the type of nPET. When npFraCm2 was used, the concentration of degradation products at 48 h ranged from 1,500 to 2,660 µM, depending on the type of nPET, with nPETb and nPETGFa being the preferred particles, and TE was produced at approximately 3-fold higher concentrations than ETE (Fig. 6d and Supplementary Data 9). IsPETase, LCCWT and LCCWCCG released degradation products at concentrations of 1,154–1,884 µM, 312–410 µM and 309–414 µM, respectively, depending on the type of nPET (Extended Data Fig. 2d–f and Supplementary Data 9). In all cases, T was observed, and TE was found to have a concentration 6- to 31-fold higher than ETE, depending on the nPET and the PETase. The higher efficiency of IsPETase compared with LCCWT and LCCWCCG could be associated with the higher activity of this enzyme at 40 °C compared with LCC variants17,18.
This finding confirmed the capacity of npFraCm1 and npFraCm2 to efficiently deconstruct nPET materials at 40 °C. The uniqueness of npFraCm1 compared with the npFraCm2, IsPETase and LCCWT/WCCG variants in degrading nPET is that ETE is the main degradation product compared with TE, with no appreciable formation of T under our assay conditions. This feature could represent an advantage over the other enzymatic systems that will produce a mixture of E, T, TE and ETE.
Time course and kinetic study of nPETb particle degradation
The time course of the released degradation products, determined by HPLC using npFraCm1 and npFraCm2, was followed and compared with that of LCCWT at 40 °C and pH 7.0. nPETb (diameter 108.0 ± 0.1 nm; Extended Data Table 2) was targeted as the closest example to real nPET substrates, and all three enzymatic preparations were able to efficiently degrade it. Although, IsPETase is currently the most active wild-type enzyme for PET hydrolysis at around 40 °C, LCCWT was selected for comparative purposes as a benchmark PETase given its higher stability compared with mesophilic IsPETase under our thermal assay conditions, of 40 °C (for details, see Supplementary Note 7; Supplementary Fig. 16 and Supplementary Data 11).
As shown in Fig. 7 (Supplementary Data 10), degradation products were constantly observed since the early stages for the three enzymatic preparations. Under the tested assay conditions, the time course of degradation product release was much steeper with npFraCm2 than with npFraCm1 in the first few reaction hours. Whereas npFraCm2 was only highly active until 5–6 h of incubation, hydrolysis by npFraCm1 continuously progresses up to 24 h where almost maximal depolymerization was achieved, as for LCCWT for which hydrolysis progressed beyond 24 h. This is consistent with the lower thermal stability of npFraCm2 (t1/2 at 40 °C: 5.3 ± 0.2 h) compared with npFraCm1 (t1/2 at 40 °C > 48 h) under the assay conditions (Supplementary Fig. 14 and Supplementary Data 8). Additionally, the better positioning of the long protuberances that conformed the nPET particles (Discussion) in the nanopore channel once the particle is seated at the pore entrance, could also explain the differences in the reaction kinetics and dynamics of npFraCm1 and npFraCm2. Regardless of the underlying reasons for a different kinetics, these data demonstrate that the access of nPET particles across the nanopores is not a limiting factor during depolymerization mediated by npFraCm1 and npFraCm2.
a–c, Time course degradation of nPETb particles using npFraCm1 (a), npFraCm2 (b) and LCCWT (c). Reaction conditions and dataset collections are detailed in Figs. 5 and 6, and Extended Data Fig. 2. In brief: 1.1 mg ml−1 nPETb, 1.5 μg ml−1 npFraCm1/m2, LCCWT, 40 °C, pH 7.0. Values are plotted as the mean of three independent replicates (n = 3) with the reported error ranges and s.d. calculated using the STDEV.S function in Excel 2019 (calculations and raw data are provided in Source Data). Raw data are provided in Supplementary Data 10.
To evaluate the effect of the particle size, we prepared nPETb particles of different mean sizes (diameter), namely from 69.1 ± 0.8 to 153.8 ± 1.4 nm (Fig. 5b and Supplementary Data 7). We discovered that smaller particles were hydrolysed by npFraCm1 more easily than larger particles (Fig. 5c), whereas npFraCm2 preferred particles from 85.4 to 108 nm (Fig. 5d), in accordance with the increased exposure of its active site to the particles (Fig. 4). These data suggest that, while particles as large as 153.8 nm were degraded (from 8.0- to 4.4-fold lower than that of the best performing size), there will be a particle size limit that will most likely no longer be degraded. Regardless of this finding, the analysed particle sizes are well above the npFraCm1 and npFraCm2 sizes, namely 6.7 nm in the cis entry and 1.9 nm in the trans exit21 (Fig. 4).
The results of the conventional (convMM) and inverse (invMM) Michaelis‒Menten kinetics35 are summarized in Supplementary Fig. 17a–f (Supplementary Data 12 and 13), and the parameters resulting from the fits (pH 7.0, 40 °C) are presented in Table 1. The results highlight that the affinity, deconstruction rates and reactive site density (Γattack) of the catalytic nanopores are comparable to those of soluble PETases, demonstrating that having a catalytic unit in a nanopore does not substantially affect either the affinity for, or the reaction rate towards, the nPET particles, as well as the concentration of reactive sites available per gram of substrate compared with a soluble enzyme with free access to the particles. For additional kinetic details, see Supplementary Note 7.
Morphological evolution during nPETb degradation by npFraCm1 and npFraCm2
The change in particle size distribution and morphology was analysed by field emission scanning electron microscopy (FE-SEM) in dried samples collected before, during and after hydrolysis. The FE-SEM micrographs in Supplementary Fig. 18 (Source Data) show the morphological degradation of the nPETb particles by npFraCm1 and npFraCm2. The as-synthesized spheres, with a relatively uniform size distribution, start aggregating and degrading after few minutes of hydrolysis, appearing covered by a film of insoluble degradation products in the samples incubated with npFraCm1/m2. Eventually, all particles disappeared and only large aggregates of degradation products are visible, as observed for PETases with a random-hydrolysis mechanism20.
Discussion
This study focused on the design of two pore-based biocatalytic nanoreactors for nPET particle depolymerization, using a non-catalytic pore-forming protein as the starting target. Through a combination of computational structure-based modelling tools and different experimental techniques, we provide robust datasets demonstrating that these biocatalytic nanopores can break down primary nPET of different types at 40 °C. The diameter of the nPET particles produced and tested ranged from approximately 53 to 154 nm, whereas the FraC pore has an inner diameter ranging from 1.9 nm (cis exit) to 6.7 nm (trans exit)21. Therefore, if we stick with this static image, one might suggest that nPET particles would not be able to access the artificial active centres introduced into the nanopores, although experimental evidence shows otherwise. Since the entry of each particle in its entirety into the pore would not be possible, we propose that the nPET particles may have protuberances and that it is not the particles that enter into the nanopores but the protuberances (Fig. 4), the size of which will be reduced during hydrolysis, and with it the size of the particles, that will be more accessible as the hydrolysis proceeds. This may agree with the fact that the molecular weight of sub-micro- and nano-sized PET particles may be lower compared with that of the pristine materials, and that the degradability of sub-micrometre particles and nanoparticles may not only be due to the increased surface area, but also to the lower chain lengths and more flexible polymer chain ends36.
The data suggest that npFraCm1 performs endo-hydrolysis on the small protuberances that can access the internal area of the catalytic nanopore; this would result in the generation of E (EPET) and T (or TPET) terminal PET oligomers, from which smaller subproducts, preferentially ETE and ETETE (from EPET), and to a lesser extent TETETE (from TPET), will be formed by exo-type chain scission. In the case of npFraCm2, the active site is remarkably more exposed to the solvent and to different accommodations of the sub-micro- and nanoparticles (and of its protuberances), initiating a more random endo- and exo-type chain degradation that yields a high diversity of oligomers and monomers. The high accumulation of all such degradation products (Fig. 6) suggests that the cleavage of TPET and EPET by npFraCm2 would occur at similar rates. This finding contrasts with the PET degradation process proposed for IsPETase, where the terminal digestion step of TPET is preferred34.
Compared with benchmark PETases, namely LCCWT, LCCWCCG and IsPETase, we discovered remarkable differences in hydrolysis capacity and degradation products (Extended Data Fig. 3). Under our experimental conditions, npFraCm1 breaks down nPET to ETE as the main degradation product without appreciable production of T, which is a final PET-building block that is usually produced by the benchmark PETases12,13,14,15,16,17,18,19,20,34. Instead, npFraCm2 behaves similarly to benchmark PETases in the sense that it can fully convert nPET to T, but unlike them, it produces a greater diversity of degradation products. The possibility of using FraC to design two catalytic nanopores capable of degrading nPET at relatively low temperatures (40 °C) and with different degradation profiles among them and compared with benchmark PETases demonstrates the versatility of this protein as a scaffold for the design of pore-based catalytic nanoreactors to deconstruct nPET particles. Note that up to eight refined artificial biocatalytic sites could be incorporated due to the homo-octamer assembly. Therefore, the limitations in the access of the nPET particles to the active sites can be compensated by the presence of multiple catalytic groups, whose number would be higher than in conventional enzymes with PETase activity. Indeed, our results demonstrated that, using the same conditions and particles, and considering the limitations of access to the particles, the catalytic efficiency for nPETb of npFraCm1 and npFraCm2 is similar to that of soluble LCCWT15.
The above features, with the easy production, simple purification and stability of the engineered catalytic nanopores designed herein (Supplementary Note 2), can create new alternatives to degrade sub-micro- and nano-sized PET, for example, in wastewater treatment plants18, under sustainable conditions, for example, 40 °C. The proof of concept presented in this study may open up new lines of research. We envision, for example, the co-integration of the two artificial hydrolytic sites or designing thermostable variants by further protein engineering efforts to have robustness above 70 °C to degrade real-world plastic pollution at an industrially relevant scale18. Also, the co-integration of engineered pore-forming proteins and nanoscale materials or membranes to develop inorganic–organic hybrid catalysts that can act as both filtering systems to effectively capture sub-micro- and nano-sized PET, mimicking the well-known water desalination nanopores37, and reaction chambers to further degrade synthetic particles38 (Supplementary Note 9). The possibility of producing engineered transmembrane pore-forming proteins, which include FraC or other larger pore-forming proteins that can oligomerize in annular pores of more than 30 monomers and large diameters (for example, the 25–40 nm Perfringolysin O39), in target cell membranes may also enable the design of microbial reactors with integrated catalytic pores supporting multiple conversions, yet to be defined, as exemplified here by nPET degradation.
Methods
In silico protein and substrate preparation
We selected the homo-octamer biological assembly crystal structure of FraC (4TSY40) and its monomer soluble structure (3W9P41). Each system was prepared and optimized at pH 7.5 with a Protein Preparation Wizard42 from Schrodinger to fix the protonation states and correct alternative positions or other common problems. The three model esters (Supplementary Fig. 1), TE and ETE were modelled using the OPLS2005 force field43, and the atomic charges were obtained using single-point energy quantum mechanics calculations with density functional theory using the B3LYP-D3 exchange-correlation and CC-PVTZ basis set.
PELE, adaptive simulations and distance metrics
The engineering protocol starts by mapping possible substrate binding sites in the target protein. Interactions between the different ester-type substrates and the inner surface of the assembled pore-forming protein scaffold were thus sampled using PELE software44. Through a PELE Global Exploration, we obtained an energy landscape profile. Further, in PELE Local Exploration, we analysed the energy landscape profile with respect to the catalytic distance between substrate ester reactive atoms and the nucleophile oxygen of the catalytic serine. Moreover, we estimated the relative activity of the active site for each substrate, calculating the number of catalytic events (equation (1)). The number of catalytic events for each substrate was determined by multiple substrate-active site distance metrics. The FraCm1 contains eight possible combinations of serine–histidine interactions between α-helices and two equidistant acids that can complete the catalytic triad, increasing the possible combinations. To avoid underestimation of catalytic events, we generated a distance matrix for each PELE step for all active sites and selected the catalytic triad nearest the substrate. For FraCm2, we focused on only one monomer globular part. For extensive details, see Supplementary Methods.
MD simulations of FraCm1, FraCm2 and the polymer
To analyse the stability of the membrane-inserted N-terminal domains in FraCm1/m2, MD simulations were performed using the Bilayer Membrane Builder protocol with the replacement method provided by CHARMM-GUI45 and GROMACS 5.1.2 simulations46. To analyse the MD of the polymer, including its flexibility in the solvent of a 200-monomer PET fragment, MD simulations were performed using the Polymer Builder protocol provided by CHARMM-GUI45 and GROMACS 5.1.2 (ref. 46). To obtain the different metrics of all MD simulations, the MDTraj47 and MDAnalysis48 Python modules were employed. For the simulation of the polymer, we used GROMACS49 analysis tools to obtain its solvent-accessible surface area. For extensive details, see Supplementary Methods.
QM/MM minimizations
QM/MM minimizations of the reactant state in IsPETase, FraCm1 and FraCm2 mutants were performed using Qsite from the Schrödinger Suite50. The calculations were performed at the DFT/B3LYP and 6-31G* basis set level of theory. The QM region included ETE as a substrate and catalytic serine and histidine residues: Ser131 and His208 for IsPETase, Ser21 and His20 for FraCm1 and Ser38 and His175 for FraCm2. The IsPETase reactant state’s structure was based on the PDB code 5XH3 (ref. 51), and both FraC mutant structures were based on a catalytic PELE pose.
npFraCWT, npFraCm1 and npFraCm2 production and assembly
The complementary DNA encoding FraCm1, with Lys20His/Thr21Ser mutations, and FraCm2, with Asp38Ser/Glu173Gln mutations, were obtained via the overlap extension mutagenesis method22,52. FraCWT, FraCm1 and FraCm2 were produced in an E. coli expression system and purified as previously described21,22. Pore assembly was performed to mimic the actual situation encountered by actinoporins in nature, as detailed in Supplementary Methods.
LCCWT, LCCWCCG and IsPETase: source and purification
The genes coding for the cutinase LCC in its wild-type form (LCCWT)18, as well as the WCCG variant (LCCWCCG)19, were codon-optimized for E. coli K12 and synthesized by Biomatik in pET21a(+) (Novagen) between the restriction sites NdeI and SalI and were kindly donated by Pablo Pérez-García (Universität Hamburg, Germany). The construct containing the gene coding for IsPETase fused to a maltose-binding protein in the backbone of pMAL-p4x was kindly donated by Sebastian Weigert (University of Bayreuth, Germany)20. The proteins were produced and purified as reported and stored at 1.5 mg ml−1 in 40 mM HEPES buffer pH 7.0 until use.
Sub-micro- and nano-sized PET degradation tests: time course and kinetics
For the nPETb, nPETc, nPETGFa and nPETGFc degradation tests, the following conditions were utilized, if otherwise not indicated: [npFraCm1/m2, IsPETase and LCCWT/WCCG], 1.5 μg ml−1; [nPETGFa/GFc/b/c], 1.1 mg ml−1; reaction volume, 1,000 μl; T, 40 °C; pH, 7.0 (20 mM HEPES buffer); reaction time, 48 h. For conventional Michaelis‒Menten kinetic analysis, reaction conditions are listed as follows: [npFraCm1], 15 µg ml−1; [npFraCm2], 30 µg ml−1; [LCCWT], 132 µg ml−1; [nPETb], 0.38–4.4 mg ml−1; reaction volume, 50 μl; reaction time, 30 min (for npFraCm1 and npFraCm2) or 60 min (for LCCWT); T, 40 °C; pH, 7.0 (20 mM HEPES buffer); 1,000 rpm. For inverse Michaelis‒Menten kinetic analysis, the following reaction conditions were used: [npFraCm1], 0–73 µg g−1 nPETb; [npFraCm2], 0–118 µg g−1 nPETb; [LCCWT], 0–118 µg g−1 nPETb; [nPETb], 1.1 g l−1; reaction volume, 50 μl; T, 40 °C; pH, 7.0; agitation, 1,000 rpm; time of reaction, 30 min (for npFraCm1 and npFraCm2) or 60 min (for LCCWT). Evaluation of the degradation of nPETb particles of different sizes using npFraCm1/m2 was performed using the following reaction conditions: [npFraCm1/m2], 1.5 μg ml−1; [nPETb], 1.1 mg ml−1; reaction volume, 100 μl; T, 40 °C; pH, 7.0 (20 mM HEPES buffer); reaction time, 48 h. All reactions were maintained in 2-ml safe-lock Eppendorf polypropylene tubes (ref. 0030 120.094; Eppendorf AG) in a thermoshaker (model Thermomixer comfort, Eppendorf AG). The reactions were stopped, at the indicated times, by diluting ten times with dimethyl sulfoxide (from Merck Life Science) and the degradation products were immediately analysed by HPLC (Supplementary Methods). In all cases, datasets were collected with a Varian Star LC workstation 6.41 (Varian) and analysed using Excel 2019 and SigmaPlot 14.5. Quantifications of degradation products were performed by HPLC on the basis of calibrations with purified standards, whose identity was confirmed by high-resolution MS (see raw MS data in Supplementary Data 14). All reactions were performed in triplicate (n = 3) with the following control reactions set up using the same amount of material: (1) soluble FraCWT, FraCm1 and FraCm2; (2) npFraCWT; (3) no protein; and (4) nPETb, nPETc, nPETGFa and nPETGFc, without protein added.
Ester hydrolysis assays
The hydrolysis of glyceryl tripropionate (ref. W328618), vinyl acetate (ref. V150-3), phenyl acetate (ref. 108723), ETE (ref. 465151), from Merck Life Science, and TE (purified as detailed in Supplementary Methods) was assayed using a pH indicator assay at 40 °C and pH 8.0 by monitoring the absorbance at 550 nm (extinction coefficient (ε) of phenol red, 8,450 M−1 cm−1), as reported26,27. The hydrolysis of the model esters p-NP acetate (ref. N-8130; Merck Life Science), propionate (Santa Cruz Biotechnology, ref. sc-256813) and butyrate (ref. N-9876; Merck Life Science) was assessed by monitoring the continuous production of 4-nitrophenol at 348 nm (pH-independent isosbestic point, ε = 4,147 M−1 cm−1)26. In all cases, a Synergy HT Multi-Mode Microplate Reader with Gen5 2.00 software (Biotek Instruments) was used. All reactions were performed in triplicate (n = 3) with control reactions and background signals considered, as reported26,27. For extensive details, see Supplementary Methods.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The authors declare that the main data supporting the findings of this study are available within the paper and related Supplementary Information, Supplementary Data and Source Data files. The molecular simulations, the MD simulations and the quantum mechanical minimizations have been deposited at Zenodo (https://zenodo.org/deposit/7755566)53 under the identifier https://doi.org/10.5281/zenodo.7755566. To use the archive, download the file and extract its contents to a local directory using appropriate software. The directory contains separate folders for each type of simulation, along with input, output and README files. Source data are provided with this paper (the mass spectrometry files can be analysed with the software MassLynx V4.1). All other data are available from the authors upon reasonable request.
References
York, A. Adapting to plastic.Nat. Rev. Microbiol. 18, 362–363 (2020).
Rosenboom, J. G., Langer, R. & Traverso, G. Bioplastics for a circular economy. Nat. Rev. Mater. 7, 117–137 (2022).
Geyer, R., Jambeck, J. R. & Law, K. L. Production, use, and fate of all plastics ever made. Sci. Adv. 3, e1700782 (2017).
Cabernard, L. et al. Growing environmental footprint of plastics driven by coal combustion. Nat. Sustain. 5, 139–148 (2022).
Jambeck, J. R. et al. Marine pollution. Plastic waste inputs from land into the ocean. Science 347, 768–771 (2015).
Gigault, J. et al. Current opinion: what is a nanoplastic? Environ. Pollut. 235, 1030–1034 (2018).
Allen, S. et al. Atmospheric transport and deposition of microplastics in a remote mountain catchment. Nat. Geosci. 12, 339–344 (2019).
Schymanski, D. et al. Analysis of microplastics in drinking water and other clean water samples with micro-Raman and micro-infrared spectroscopy: minimum requirements and best practice guidelines. Anal. Bioanal. Chem. 413, 5969–5994 (2021).
Materić, D. et al. Nanoplastics measurements in Northern and Southern polar ice. Environ. Res. 208, 112741 (2022).
Wei, R., Bertling, J., O’Connor, K., Bank, L. M. & Bornscheur, U. T. Possibilities and limitations of biotechnological plastic degradation and recycling. Nat. Catal. 3, 867–871 (2020).
Ellis, L. D. et al. Chemical and biological catalysis for plastics recycling and upcycling. Nat. Catal. 4, 539–556 (2021).
Wei, R. et al. Mechanism-based design of efficient PET hydrolases. ACS Catal. 12, 3382–3396 (2022).
Son, H. F. et al. Rational protein engineering of thermo-stable PETase from Ideonella sakaiensis for highly efficient PET degradation. ACS Catal. 9, 3519–3526 (2019).
Bell, E. L. et al. Directed evolution of an efficient and thermostable PET depolymerase. Nat. Catal. 5, 673–681 (2022).
Tournier, V. et al. An engineered PET depolymerase to break down and recycle plastic bottles. Nature 580, 216–219 (2020).
Lu, H. et al. Machine learning-aided engineering of hydrolases for PET depolymerization. Nature 604, 662–667 (2022).
Yoshida, S. et al. A bacterium that degrades and assimilates poly(ethylene terephthalate). Science 351, 1196–1199 (2016).
Cui, Y. et al. Computational redesign of a PETase for plastic biodegradation under ambient condition by the GRAPE strategy. ACS Catal. 11, 1340–1350 (2021).
Barth, M. et al. Effect of hydrolysis products on the enzymatic degradation of polyethylene terephthalate nanoparticles by a polyester hydrolase from Thermobifida fusca. Biochem. Eng. J. 93, 222–228 (2015).
Vogel, K. et al. Enzymatic degradation of polyethylene terephthalate nanoplastics analyzed in real time by isothermal titration calorimetry. Sci. Total Environ. 773, 145111 (2021).
Tanaka, K., Caaveiro, J. M. M., Morante, K., González-Mañas, J. M. & Tsumoto, K. Structural basis for self-assembly of a cytolytic pore lined by protein and lipid. Nat. Commun. 6, 1–11 (2015).
Alegre-Cebollada, J. et al. Silent mutations at the 5′-end of the cDNA of actinoporins from the sea anemone Stichodactyla helianthus allow their heterologous overproduction in Escherichia coli. J. Biotechnol. 127, 211–221 (2007).
Wloka, C., Mutter, N. L., Soskine, M. & Maglia, G. Alpha-helical Fragaceatoxin C nanopore engineered for double-stranded and single-stranded nucleic acid analysis. Angew. Chem. Int. Ed. Engl. 55, 12494–12498 (2016).
Versloot, R. C. A. et al. Quantification of protein glycosylation using nanopores. Nano Lett. 22, 5357–5364 (2022).
Roda, S., Robles-Martín, A., Xiang, R., Kazemi, M. & Guallar, V. Structural-based modeling in protein engineering. A must do. J. Phys. Chem. B 125, 6491–6500 (2021).
Alonso, S. et al. Genetically engineered proteins with two active sites for enhanced biocatalysis and synergistic chemo- and biocatalysis. Nat. Catal. 3, 319–328 (2020).
Roda, S. et al. A Plurizyme with transaminase and hydrolase activity catalyzes cascade reactions. Angew. Chem. Int. Ed. Engl. 61, e202207344 (2022).
Mesa-Galloso et al. Disrupting a key hydrophobic pair in the oligomerization interface of the actinoporins impairs their pore-forming activity. Protein Sci. 26, 550–565 (2017).
Schubert, S. et al. Reaction pathways for the enzymatic degradation of poly(ethylene terephthalate): What characterizes an efficient PET-hydrolase? ChemBioChem 24, e202200516 (2023).
Jerves, C. et al. Reaction mechanism of the PET degrading enzyme PETase studied with DFT/MM molecular dynamics simulations. ACS Catal. 11, 11626–11638 (2021).
Bååth, J. A., Borch, K., Jensen, K., Brask, J. & Westh, P. Comparative biochemistry of four polyester (PET) hydrolases. ChemBioChem 22, 1627–1637 (2021).
Menzel, T. et al. Impact of enzymatic degradation on the material properties of poly(ethylene terephthalate). Polymers 13, 3885 (2021).
Bashir, Z., Al-Aloush, I., Al-Raqibah, I. & Ibrahim, M. Evaluation of three methods for the measurement of crystallinity of pet resins, preforms, and bottles. Polym. Eng. Sci. 40, 2442–2455 (2000).
Joo, S. et al. Structural insight into molecular mechanism of poly(ethylene terephthalate) degradation. Nat. Commun. 9, 382 (2018).
Erickson, E. et al. Comparative performance of PETase as a function of reaction conditions, substrate properties, and product accumulation. ChemSusChem 15, e202101932 (2022).
Pfaff, L. et al. Multiple substrate binding mode-guided engineering of a thermophilic PET hydrolase. ACS Catal. 12, 9790–9800 (2022).
Heiranian, M., Farimani, A. & Aluru, N. Water desalination with a single-layer MoS2 nanopore. Nat. Commun. 6, 8616 (2015).
Gigault, J. et al. Nanoplastics are neither microplastics nor engineered nanoparticles. Nat. Nanotechnol. 16, 501–507 (2021).
Johnstone, B. A. et al. Cholesterol-dependent cytolysins: the outstanding questions. IUBMB Life 74, 1169–1179 (2022).
4TSY: crystal structure of FraC with lipids. NCBI https://www.ncbi.nlm.nih.gov/Structure/pdb/4TSY
3W9P: crystal structure of monomeric FraC (second crystal form). NCBI https://www.ncbi.nlm.nih.gov/Structure/pdb/3W9P
Sastry, G. M., Adzhigirey, M., Day, T., Annabhimoju, R. & Sherman, W. Protein and ligand preparation: parameters, protocols, and influence on virtual screening enrichments. J. Comput. Aided Mol. Des. 27, 221–234 (2013).
Banks, J. L. et al. Integrated Modeling Program, Applied Chemical Theory (IMPACT). J. Comput. Chem. 26, 1752–1780 (2005).
Lecina, D., Gilabert, J. F. & Guallar, V. Adaptive simulations, towards interactive protein–ligand modeling. Sci. Rep. 7, 8466 (2017).
Lee, J. et al. CHARMM-GUI input generator for NAMD, GROMACS, AMBER, OpenMM, and CHARMM/OpenMM simulations using the CHARMM36 additive force field. J. Chem. Theory Comput. 12, 405–413 (2016).
Bauer, P., Hess, B., & Lindahl, E. GROMACS 2022 Manual. Zenodo https://doi.org/10.5281/ZENODO.6103568
McGibbon, R. T. et al. MDTraj: a modern open library for the analysis of molecular dynamics trajectories. Biophys. J. 109, 1528–1532 (2015).
Michaud-Agrawal, N., Denning, E. J., Woolf, T. B. & Beckstein, O. MDAnalysis: a toolkit for the analysis of molecular dynamics simulations. J. Comput. Chem. 32, 2319–2327 (2011).
Abraham, M. J. et al. GROMACS: high performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1–2, 19–25 (2015).
Murphy, R. B., Philipp, D. M. & Friesner, R. A. A mixed quantum mechanics/molecular mechanics (QM/MM) method for large-scale modeling of chemistry in protein environments. J. Comput. Chem. 21, 1442–1457 (2000).
5XH3: crystal structure of a novel PET hydrolase R103G/S131A mutant in complex with HEMT from Ideonella sakaiensis 201-F6. NCBI https://www.ncbi.nlm.nih.gov/Structure/pdb/5XH3
Sambrook, J. & Russell, D. W. In vitro mutagenesis using double-stranded DNA templates: selection of mutants with DpnI. Cold Spring Harb. Protoc. https://doi.org/10.1101/pdb.prot097766 (2018).
Robles-Martín, A. et al. Sub-micro and nano-sized polyethylene terephthalate deconstruction with engineered pore-forming protein nanopores. Zenodo https://zenodo.org/deposit/7755566 (2023).
Acknowledgements
This study was conducted under the auspices of the FuturEnzyme Project funded by the European Union’s Horizon 2020 Research and Innovation Programme under the auspices of the FuturEnzyme Project (grant agreement no. 101000327) and the PlasticsFatE project (grant agreement no. 95921), and Horizon Europe Research and Innovation Programme under grant agreement no. GA101060625 (Nymphe project). We also acknowledge financial support under grants PID2020-112758RB-I00 (M.F.), PDC2021-121534-I00 (M.F.), TED2021-130544B-I00 (M.F.), PID2019-106370RB-I00 (V.G.) and PID2019-105838RB-C31 (F.J.P.) from the Ministerio de Ciencia e Innovación, Agencia Estatal de Investigación (AEI) (Digital Object Identifier MCIN/AEI/10.13039/501100011033), Fondo Europeo de Desarrollo Regional (ERDF) A way of making Europe and the European Union NextGenerationEU/PRTR, UCM-Banco Santander Grants PR87/19-22556 and PR108/20-26896 and UnaEuropa (Unano) SF2106 (to A.M.P.). S.G.-L. was supported by a Real Colegio Complutense Postdoctoral Fellowship for Distinguished Junior Scholars. S.R. thanks the Spanish Ministry of Science and Innovation for a PhD fellowship (FPU19/00608). D.H.-M. thanks Complutense University of Madrid and Banco Santander for a PhD fellowship (CT82/20/CT83/20). A.R.-M. thanks the Spanish Ministry of Science and Innovation for a PhD fellowship (PRE2020-091825) and the project PID2019-106370RB-I00. We thank M. J. Vicente for the ESI–MS analysis, performed at the Servicio Interdepartamental de Investigación (SIDI) from the Autonomous University of Madrid, Spain.
Author information
Authors and Affiliations
Contributions
A.R.-M., R.A.-S., and L.F.-L. contributed equally to this work. V.G., S.G.-L. and M.F. lead this contribution. V.G., A.M.-P. and M.F. wrote the original manuscript, which was further prepared through the contributions of A.R.-M., S.R., S.G.-L., R.P. and M.A.B. All the authors have given approval to the final version of the manuscript. R.A.-S., D.H.-M., A.M.-P. and S.G.-L. contributed to the production, purification and haemolytic and structural characterization of the FraC pore-forming proteins. J.L.G.-A. and F.J.P. contributed to the HPLC analysis. L.F.-L., C.C., D.A. and M.F. performed the ester and nPET hydrolysis assays and determinations. A.R.-M., S.R. and V.G. contributed to the computational analysis. V.A.-R., R.P. and M.A.B. contributed to the FE-SEM analyses. V.G. originally conceived the computational analysis, which was further experimentally conceived by A.M.-P., S.G.-L. and M.F.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Catalysis thanks Pedro Alexandrino Fernandes, Bian Wu, Ren Wei and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Chemical structures of the degradation products identified in the present study.
The figure was constructed using ChemDraw 18.2. E refers to ethylene glycol, T refers to terephthalic acid, TE refers to MHET, ETE refers to BHET, TETE refers to (MHET)2 as reported by Joo et al. (2018)34, and ETETE refers to 2-HE(MHET)2 as reported by Joo et al. (2018)34. Nomenclature as reported by Schubert et al.29.
Extended Data Fig. 2 Degradation profiles of nPETGFa/GFc/b/c treated with IsPETase and LCCWT/WCCG.
a-c HPLC profiles of degradation products released under the experimental conditions: [IsPETase, LCCWT/WCCG], 1.5 μg ml-1; [nPETGFa/GFc/b/c], 1.1 mg ml-1; reaction volume, 1000 μl; T, 40 °C; pH, 7.0 (20 mM HEPES buffer); reaction time, 48 h. The reactions were maintained in 2-ml safe-lock Eppendorf® polypropylene tubes (ref. 0030 120.094) in a thermoshaker (model Thermomixer comfort, Eppendorf AG, Hamburg, Germany) at 1000 rpm. Aliquots of 100 μl were obtained, the reactions were stopped by adding 900 μl dimethyl sulfoxide (from Merck Life Science S.L.U., Madrid, Spain), and the degradation products were immediately analysed by HPLC. Datasets were collected with a Varian Star LC workstation 6.41 (Varian Inc., Palo Alto, California, USA). All reactions and analyses were performed in triplicate (n = 3), with a representative chromatogram per enzyme and nPET particle shown. d-f Concentration of degradation products from T to ETETE corresponding to a-e. Quantification was performed for products with unambiguous identification and for which enough material was recovered for HPLC calibration, namely, from T to ETETE (Extended Data Fig. 1); purification was performed by semipreparative HPLC, and the molecular weights and structures were determined by mass spectrometry (Supplementary Fig. 15). Values are plotted as the mean of three independent replicates (n = 3) with the reported error ranges and SD calculated using the STDEV.S function in Excel 2019 (calculations and raw data are provided in Source Data). The figure was constructed using Excel 2019. Raw data are shown in Supplementary Data 9.
Extended Data Fig. 3 Schematic of the degradation product profiles.
The degradation products identified after nPET hydrolysis by npFraCm1, npFraCm2, LCCWT, LCCWCCG and IsPETase are shown.
Supplementary information
Supplementary Information
Supplementary methods, Figs. 1–18, Notes 1–9 and references.
Supplementary Data 1–13
Supplementary Data 1. Bioinformatic/computational raw data obtained from the different software simulations related to Figs. 1-4, Supplementary Figs. 2-6. Supplementary Data 2. Raw data for Supplementary Fig. 8 (Structural and functional comparison of FraCWT and FraCm1 and FraCm2. Supplementary Data 3. Raw data (absorbance per min) and calculations for Extended Data Table 1 (Ester-hydrolysing activity of npFraCm1 and npFraCm2). Supplementary Data 4. Raw data and calculations for Table 1 (Kinetic hydrolysis parameters of ETE and nPETb for npFraCm1 and npFraCm2) and Supplementary Fig. 10 (Kinetic hydrolysis parameters of ETE for npFraCm1 and npFraCm2). Supplementary Data 5. Raw data and calculations for Supplementary Fig. 11 (pH and thermal profiles of the npFraCm1 and npFraCm2). Supplementary Data 6. Raw data for Supplementary Fig. 12 (Thermal denaturation of FraCWT, FraCm1 and FraCm2 as monitored by circular dichroism at 220 nm). Supplementary Data 7. Raw data and calculations for Fig. 5 (Particle size distribution of nPETGFa/GFc/b/c as determined by DLS). Supplementary Data 8. Raw data and calculations for Supplementary Fig. 14 (Thermal stability of npFraCm1 and npFraCm2 as monitored by activity measurements). Supplementary Data 9. Raw data and calculations for Fig. 6 (degradation profiles of nPETGFa/GFc/b/c treated with npFraCm1/m2) and Extended Data Fig. 2 (degradation profiles of nPETGFa/GFc/b/c treated with IsPETase and LCCWT/WCCG). Supplementary Data 10. Raw data and calculations for Fig. 7 (time course study of nPETb degradation using npFraCm1/m2 compared with LCCWT). Supplementary Data 11. Raw data and calculations for Supplementary Fig. 16 (nPETc and PETGFa hydrolysis by LCCWT at 40 °C, 50 °C and 60 °C). Supplementary Data 12. Raw data and calculations for Supplementary Fig. 17a–c (conventional Michaelis–Menten (MM) kinetic study of nPETb particle degradation using npFraCm1/m2 in comparison with the benchmark LCCWT) and the corresponding data in Table 1 (kinetic hydrolysis parameters of ETE and nPETb for npFraCm1 and npFraCm2). Supplementary Data 13. Raw data and calculations for Supplementary Fig. 17d–f (inverse Michaelis–Menten (MM) kinetic study of nPETb particle degradation using npFraCm1/m2 in comparison with the benchmark LCCWT) and the corresponding data in Table 1 (Kinetic hydrolysis parameters of ETE and nPETb for npFraCm1 and npFraCm2).
Supplementary Data 14
Raw MS data for Supplementary Fig. 15. The data in the .zip file can be analysed with the software MassLynx V4.1 (Waters) to obtain the mass spectra of the oligomers ETE, TET, ETETE, TETE, TETET and TETETE.
Source data
Source Data Supplementary Fig. 7
Uncropped scan of the gel shown in Supplementary Fig. 7.
Source Data Supplementary Fig. 9
Original unprocessed negative stain electron microscopy micrographs, as shown in Supplementary Fig. 9. Original unprocessed negative stain electron microscopy micrograph of MSP1E3D1-assembled nanodiscs in the presence of FraCWT, FraCm1 and FraCm2, in this order. Electron microscopy datasets have been collected with JEOL JEM1400 transmission electron microscope and processed using a DigitalMicrograph software (Gatan).
Source Data Supplementary Fig. 18
Original unprocessed negative stain electron microscopy micrographs, as shown in Supplementary Fig. 18. In the order of appearance: non-treated nPETb (Supplementary Fig. 18a), treated nPETb with npFraCm1 during 3 min, 15 min, 30 min and 24 h (Supplementary Fig. 18b), and treated nPETb with npFraCm2 during 3 min, 15 min, 30 min and 24 h (Supplementary Fig. 18c). Magnification, 100,000×. Scale bar, 500 nm. FE-SEM was performed with an FEI Nova NanoSEM 230 microscope.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Robles-Martín, A., Amigot-Sánchez, R., Fernandez-Lopez, L. et al. Sub-micro- and nano-sized polyethylene terephthalate deconstruction with engineered protein nanopores. Nat Catal 6, 1174–1185 (2023). https://doi.org/10.1038/s41929-023-01048-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41929-023-01048-6
This article is cited by
-
Designer catalytic nanopores meet PET nanoparticles
Nature Catalysis (2023)