Abstract
Our demand for electronic goods and fossil fuels has challenged our ecosystem with contaminating amounts of heavy metals, causing numerous water sources to become polluted. To counter heavy-metal waste, industry has relied on a family of physicochemical processes, with chemical precipitation being one of the most commonly used. However, the disadvantages of chemical precipitation are vast, including the generation of secondary waste, technical handling of chemicals and need for complex infrastructures. To circumvent these limitations, biological processes to naturally manage waste have been sought. Here, we show that yeast can act as a biological alternative to traditional chemical precipitation by controlling naturally occurring production of hydrogen sulfide (H2S). Sulfide production was harnessed by controlling the sulfate assimilation pathway, where strategic knockouts and culture conditions generated H2S from 0 to over 1,000 ppm (~30 mM). These sulfide-producing yeasts were able to remove mercury, lead and copper from real-world samples taken from the Athabasca oil sands. More so, yeast surface display of biomineralization peptides helped control for size distribution and crystallinity of precipitated metal sulfide nanoparticles. Altogether, this yeast-based platform not only removes heavy metals but also offers a platform for metal re-extraction through precipitation of metal sulfide nanoparticles.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Rucevska, I. et al. Waste Crime–Waste Risks: Gaps in Meeting the Global Waste Challenge (UNEP and GRID-Arendal, 2015); https://go.nature.com/38RZ1cY
Balde, C. P., Forti, V., Gray, V., Kuehr, R. & Stegmann, P. The Global e-Waste Monitor 2017: Quantities, Flows and Resources (United Nations University, International Telecommunication Union and International Solid Waste Association, 2017).
Statistics: All Mining (CDC, 2018); https://www.cdc.gov/niosh/mining/statistics/allmining.html
DeGraff, J. V. in Understanding and Responding to Hazardous Substances at Mine Sites in the Western United States (ed. DeGraff, J. V.) 1–8 (Geological Society of America, 2007).
Fu, F. & Wang, Q. Removal of heavy metal ions from wastewaters: a review. J. Environ. Manage. 92, 407–418 (2011).
Kang, D. H. P., Chen, M. & Ogunseitan, O. A. Potential environmental and human health impacts of rechargeable lithium batteries in electronic waste. Environ. Sci. Technol. 47, 5495–5503 (2013).
Song, X., Hu, S., Chen, D. & Zhu, B. Estimation of waste battery generation and analysis of the waste battery recycling system in China. J. Ind. Ecol. 21, 57–69 (2017).
Kurniawan, T. A., Chan, G. Y. S., Lo, W.-H. & Babel, S. Physico–chemical treatment techniques for wastewater laden with heavy metals. Chem. Eng. J. 118, 83–98 (2006).
Barakat, M. A. New trends in removing heavy metals from industrial wastewater. Arab. J. Chem. 4, 361–377 (2011).
Gupta, K. V., Ali, I., A. Saleh, T., Nayak, A. & Agarwal, S. Chemical treatment technologies for waste-water recycling—an overview. RSC Adv. 2, 6380–6388 (2012).
Gavrilescu, M., Demnerová, K., Aamand, J., Agathos, S. & Fava, F. Emerging pollutants in the environment: present and future challenges in biomonitoring, ecological risks and bioremediation. New Biotechnol. 32, 147–156 (2015).
Singh, A. & Prasad, S. M. Remediation of heavy metal contaminated ecosystem: an overview on technology advancement. Int. J. Environ. Sci. Technol. 12, 353–366 (2015).
Gadd, G. M. Microbial influence on metal mobility and application for bioremediation. Geoderma 122, 109–119 (2004).
Wiatrowski, H. A., Ward, P. M. & Barkay, T. Novel reduction of mercury (II) by mercury-sensitive dissimilatory metal reducing bacteria. Environ. Sci. Technol. 40, 6690–6696 (2006).
Silver, S. & Phung, L. T. Genes and enzymes involved in bacterial oxidation and reduction of inorganic arsenic. Appl. Environ. Microbiol. 71, 599–608 (2005).
Picard, A., Gartman, A., Clarke, D. R. & Girguis, P. R. Sulfate-reducing bacteria influence the nucleation and growth of mackinawite and greigite. Geochim. Cosmochim. Acta 220, 367–384 (2018).
Gartman, A. et al. Microbes facilitate mineral deposition in bioelectrochemical systems. ACS Earth Space Chem. 1, 277–287 (2017).
Jong, T. & Parry, D. L. Removal of sulfate and heavy metals by sulfate reducing bacteria in short-term bench scale upflow anaerobic packed bed reactor runs. Water Res. 37, 3379–3389 (2003).
Kieu, H. T. Q., Müller, E. & Horn, H. Heavy metal removal in anaerobic semi-continuous stirred tank reactors by a consortium of sulfate-reducing bacteria. Water Res. 45, 3863–3870 (2011).
Neculita, C.-M., Zagury, G. J. & Bussière, B. Passive treatment of acid mine drainage in bioreactors using sulfate-reducing bacteria. J. Environ. Qual. 36, 1–16 (2007).
Lovley, D. R. Cleaning up with genomics: applying molecular biology to bioremediation. Nat. Rev. Microbiol. 1, 35–44 (2003).
Shiratori, T., Inoue, C., Sugawara, K., Kusano, T. & Kitagawa, Y. Cloning and expression of Thiobacillus ferrooxidans mercury ion resistance genes in Escherichia coli. J. Bacteriol. 171, 3458–3464 (1989).
Wang, C. L., Maratukulam, P. D., Lum, A. M., Clark, D. S. & Keasling, J. D. Metabolic engineering of an aerobic sulfate reduction pathway and its application to precipitation of cadmium on the cell surface. Appl. Environ. Microbiol. 66, 4497–4502 (2000).
Krämer, U. & Chardonnens, A. The use of transgenic plants in the bioremediation of soils contaminated with trace elements. Appl. Microbiol. Biotechnol. 55, 661–672 (2001).
Sayler, G. S. & Ripp, S. Field applications of genetically engineered microorganisms for bioremediation processes. Curr. Opin. Biotechnol. 11, 286–289 (2000).
Vieira, É. D., da Graça Stupiello Andrietta, M. & Andrietta, S. R. Yeast biomass production: a new approach in glucose-limited feeding strategy. Braz. J. Microbiol. 44, 551–558 (2013).
Worldwide Beer Production, 2016 (Barth-Haas Group, 2018).
Value of the Yeast Product Market Worldwide from 2016 to 2022 (in Billions U.S. Dollars) (PR Newswire, 2017).
Swiegers, J. H. & Pretorius, I. S. Modulation of volatile sulfur compounds by wine yeast. Appl. Microbiol. Biotechnol. 74, 954–960 (2007).
Linderholm, A. L., Findleton, C. L., Kumar, G., Hong, Y. & Bisson, L. F. Identification of genes affecting hydrogen sulfide formation in Saccharomyces cerevisiae. Appl. Environ. Microbiol. 74, 1418–1427 (2008).
Rickard, D. & Luther, G. W. Metal sulfide complexes and clusters. Rev. Mineral. Geochem. 61, 421–504 (2006).
Bolong, N., Ismail, A. F., Salim, M. R. & Matsuura, T. A review of the effects of emerging contaminants in wastewater and options for their removal. Desalination 239, 229–246 (2009).
Johnson, D. B. & Hallberg, K. B. Acid mine drainage remediation options: a review. Sci. Total Environ. 338, 3–14 (2005).
Table of Regulated Drinking Water Contaminants (EPA, 2019).
Secondary Drinking Water Standards: Guidance for Nuisance Chemicals (EPA, 2019).
Sparks, B. D., Kotlyar, L. S., O’Carroll, J. B. & Chung, K. H. Athabasca oil sands: effect of organic coated solids on bitumen recovery and quality. J. Pet. Sci. Eng. 39, 417–430 (2003).
Kelly, E. N. et al. Oil sands development contributes elements toxic at low concentrations to the Athabasca River and its tributaries. Proc. Natl Acad. Sci. USA 107, 16178–16183 (2010).
Sakimoto, K. K., Wong, A. B. & Yang, P. Self-photosensitization of nonphotosynthetic bacteria for solar-to-chemical production. Science 351, 74–77 (2016).
Sweeney, R. Y. et al. Bacterial biosynthesis of cadmium sulfide nanocrystals. Chem. Biol. 11, 1553–1559 (2004).
DeSilva Tara, M., Gianluigi, Veglia, Fernando, Porcelli, Prantner Andrew, M. & Opella Stanley, J. Selectivity in heavy metal‐ binding to peptides and proteins. Biopolymers 64, 189–197 (2002).
Kim, J. et al. Effects of various pretreatments for enhanced anaerobic digestion with waste activated sludge. J. Biosci. Bioeng. 95, 271–275 (2003).
Harrison, R. G., Todd, P., Todd, P. W., Rudge, S. R. & Petrides, D. P. Bioseparations Science and Engineering (Oxford Univ. Press, 2015).
Hoek, P. V., Dijken, J. P. V. & Pronk, J. T. Effect of specific growth rate on fermentative capacity of baker’s yeast. Appl. Environ. Microbiol. 64, 4226–4233 (1998).
Yeasts, Yeast Extracts, Autolysates and Related Products: The Global Market (BCC Research, 2017).
Rugh, C. L. et al. Mercuric ion reduction and resistance in transgenic Arabidopsis thaliana plants expressing a modified bacterial merA gene. Proc. Natl Acad. Sci. USA 93, 3182–3187 (1996).
Arai, T. et al. Cu-doped ZnS hollow particle with high activity for hydrogen generation from alkaline sulfide solution under visible light. Chem. Mater. 20, 1997–2000 (2008).
Parveen, A., Agrawal, S. & Azam, A. Band gap tuning and fluorescence properties of lead sulfide Pb0.9A0.1S (A: Fe, Co, and Ni) nanoparticles by transition metal doping. Opt. Mater. 76, 21–27 (2018).
Bhattacharya, S. & Chakravorty, D. Electrical and magnetic properties of cold compacted iron-doped zinc sulfide nanoparticles synthesized by wet chemical method. Chem. Phys. Lett. 444, 319–323 (2007).
Selim, H., Gupta, A. K. & Al Shoaibi, A. Effect of reaction parameters on the quality of captured sulfur in Claus process. Appl. Energy 104, 772–776 (2013).
Minerals Information: Sulfur (USGS, 2018).
Little, B. J., Ray, R. I. & Pope, R. K. Relationship between corrosion and the biological sulfur cycle: a review. CORROSION 56, 433–443 (2000).
Seligman, A. M., Wasserkrug, H. L. & Hanker, J. S. A new staining method (OTO) for enhancing contrast of lipid-containing membranes and droplets in osmium tetroxide-fixed tissue with osmiophilic thiocarbohydrazide (TCH). J. Cell Biol. 30, 424–432 (1966).
Acknowledgements
We thank the Koch Institute Facilities, Center for Material Science Facilities (CMSE), the Whitehead Keck Microscopy Facility, as well as Y. Zhang (CMSE) and N. Watson (Keck) for assisting in TEM sample prep and imaging. We acknowledge the Amon Lab for helpful discussions and advice in setting up basic yeast experiments, specifically C. Brennan and S. Morrill. We also thank N. Eze for insightful discussions and paper editing. This work was supported by the Amar G. Bose Research Grant (G.L.S. and A.M.B.) and the NSF Graduate Fellowship (G.L.S.). Partial support for this research was provided by a core center grant P30-ES002109 from the National Institute of Environmental Health Sciences, National Institutes of Health.
Author information
Authors and Affiliations
Contributions
G.L.S., E.E.R. and A.M.B. conceived the study and designed experiments; G.L.S. performed and E.E.R. helped with experiments; G.L.S. analysed the data and assembled figures; G.L.S., E.E.R. and A.M.B. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
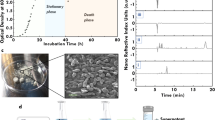
Extended Data Fig. 1 Measuring yeast H2S production.
Illustrations left of the images represent H2S detection columns with tick marks indicating the level of sulfide measured in ppm. a, Sulfide detection using 200 pm columns for mutants ΔCYS4, ΔHOM2, ΔMET17, and ΔHM217. b, Sulfide detection using 60 ppm columns for ΔMET17 in cultures of YPD, CSM, and CSM with the addition (+) of methionine (M) or cysteine (C). c, Sulfide detection using 2000 ppm columns for ΔMET17 in CSM cultures lacking (-) methionine or cysteine, or both.
Extended Data Fig. 2 Strain, culture density (OD600), and media composition effects on metal precipitation.
a, Precipitation of copper, zinc, cadmium, lead, and mercury with mutants ΔCYS4, ΔHOM2, ΔMET17, and ΔHM217, and WT as a control, in CSM. b, Effects of removing methionine (M) and/or cysteine (C) from CMS on precipitation efficacy using ΔMET17. Columns represent removal of M while rows represent removal of C from CSM. 1X stands for 100% removal (that is 0.2X = 20% and 0.5X=50%). Annotated values per cell grid represent the percent cadmium removed and standard error. c, Optimal culture density (marked within grey bounds) was determined by titrating cultures of ΔMET17 at different OD600 with copper, zinc, cadmium, lead, and mercury. Metal color coding matches those used in the main text. For all data, the mean ±s.d. of three replicates were taken for each data point.
Extended Data Fig. 3 Elemental mapping of precipitated metal sulfide particles.
a, Elemental mapping of HRTEM images of cadmium sulfide nanoparticles deposited on the cell wall of ΔMET17. Cadmium was false colored as red, sulfide as blue. Scale bars represent 50 nm. b, Elemental dispersive X-ray (EDX) spectroscopy was performed on purified precipitated copper, cadmium, lead, mercury, and zinc sulfide particles under TEM. Elemental Kα peaks were colored and highlighted as areas under the curve for qualitative comparisons. Metal color coding of spectral plots match those used in the main text.
Supplementary information
Supplementary Information
Supplementary Figs. 1–11 and Tables 1 and 3.
Supplementary Table
Supplementary Table 2.
Source data
Source Data Fig. 1
Growth curve and sulfur production values for several strains at various time points
Source Data Fig. 2
Metal removal of Cu, Zn, Cd, Pb, and Hg with respects to media composition in YPD, CSM, CSM-M, and CSM-C
Source Data Fig. 3
Metal concentrations at different rounds of metal removal taken from the Athabasca oil sands
Source Data Fig. 4
Measured sizes of metal sulfide particles examined under TEM
Source Data Fig. 5
Excitation and emission spectra of several purified metal sulfide nanoparticles
Source Data Extended Data Fig. 2
Effects on strain and culture density on metal precipitation efficiency
Source Data Extended Data Fig. 3
Raw EDX signal of purified metal sulfide nanoparticles examined under TEM
Rights and permissions
About this article
Cite this article
Sun, G.L., Reynolds, E.E. & Belcher, A.M. Using yeast to sustainably remediate and extract heavy metals from waste waters. Nat Sustain 3, 303–311 (2020). https://doi.org/10.1038/s41893-020-0478-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41893-020-0478-9
This article is cited by
-
Upcycling waste sewage sludge into superior single-atom Fenton-like catalyst for sustainable water purification
Nature Water (2024)
-
Self-doping active sites in microbe-derived carbonaceous electrocatalysts for the oxygen reduction reaction performance
Nano Research (2024)
-
Ultrasensitive and highly selective detection of strontium ions
Nature Sustainability (2023)
-
Solar-driven waste-to-chemical conversion by wastewater-derived semiconductor biohybrids
Nature Sustainability (2023)
-
Metallurgical pathways of lead leaching from brass
npj Materials Degradation (2023)