Abstract
Dormice (family Gliridae) are an ancient group of rodents. It was fully dominant in the Oligocene and Early Miocene, and its current diversity is represented by a few extant species. A Kazakhstani endemic, the desert dormouse Selevinia betpakdalaensis is one of the most enigmatic dormouse species. Lack of genetic data has not allowed Selevinia to be included in previous molecular phylogenetic analyses. In the current study, we report the first genetic data on S. betpakdalaensis as well as mitochondrial genomes of Myomimus roachi and Glirulus japonicus (retrieved from museum specimens) and a mitogenome of Graphiurus murinus (assembled from SRA data). The assembled mitochondrial genomes were combined with available mitochondrial data from GenBank to reconstruct the mitochondrial phylogeny of Gliridae. Taking into account a distortion of the phylogeny as a result of an analysis of the saturated third codon position, we obtained for the first time a resolved phylogeny of the family. The first split within Gliridae was estimated as an average of 34.6 Mya, whereas divergence time of subfamilies Graphiurinae and Glirinae was assessed at 32.67 Mya. The phylogenetic analysis confirmed the relationship (previously shown based on cranial and mandibular morphology) between Selevinia and the Myomimus.
Similar content being viewed by others
Introduction
Dormice (family Gliridae Muirhead, 1819) are an ancient group; current diversity is represented by 28 extant species. Nine genera of dormice (among which five are monotypic) are assigned to three subfamilies1: Glirinae (Glirulus Thomas, 1905 and Glis Brisson, 1762), Graphiurinae (Graphiurus Smuts, 1832) and Leithiinae (Chaetocauda Wang, 1985, Dryomys Thomas, 1905, Eliomys Wagner, 1840, Muscardinus Kaup, 1829, Myomimus Ognev, 1924, and Selevinia Belosludov and Bazhanov, 1939), which probably diverged ~ 50–27 million years ago (Mya)2. In the Oligocene and Early Miocene, approximately 16–15 Mya, the period of maximum Gliridae diversity, it was a fully dominant rodent family and may have represented up to 90% of rodent fauna, occupying almost all available niches3.
The desert dormouse Selevinia betpakdalaensis Belosludov and Bashanov, 1939, is one of the most enigmatic dormouse species. It is a rare rodent occurring in southern and eastern parts of Kazakhstan, especially in the deserts surrounding Lake Balkhash, and has been documented as far south as Karaungir, near the border with Kyrgyzstan1,4. Only a few specimens have been captured in the wild. Some Selevinia material has been found in bird pellets5.
This species was described by Belosludov and Bashanov6 based on specimens collected in 1938 by B.A. Belosludov and V.A. Selevin4 as a representative of a new monotypic family, Seleviniidae. The discovery of Selevinia was a real sensation. A find of a new mammalian species in the 20th century was an extremely rare event, not to mention a taxon of the genus or even family rank. Ognev7 has emphasised a similarity of this species to Myomimus (mouse-tailed dormouse) and placed the group as a subfamily in the family Gliridae. Subsequent studies have regarded Selevinia as a part of the family Gliridae8. On the basis of the presence of primitive characteristics, Storch9 has placed Selevinia closer to Myomimus and Chaetocauda and assigned all three genera to Seleviniidae, while others10 have considered Selevinia to be closer to genera Muscardinus and Glis. An analysis of the middle ear and of cranial and mandibular morphology11,12 has shown a clear resemblance between Selevinia and the small genus Myomimus. Wahlert et al.13 and Holden-Musser et al.14 have placed Selevinia and Myomimus in the tribe Seleviniini within the subfamily Leithiinae.
The lack of genetic data has prevented S. betpakdalaensis from being included in previous molecular phylogenetic analyses of Gliridae2,15,16. Therefore, the phylogenetic position of this species within the family still needs to be determined.
An analysis of relatively rapidly evolving mitochondrial DNA (mtDNA)17 is often used for phylogenetic purposes. Due to the maternal type of inheritance, this method has a number of limitations, such as discordance between mitochondrial and nuclear trees owing to hybridization with subsequent introgression of mtDNA or incomplete lineage sorting18,19. This risk is high when relationships are examined at the species or intraspecies rank, whereas intergeneric hybridization is an extremely rare phenomenon20. In the case of dormice, effects of introgression are unlikely because we are analysing taxa that diverged a long time ago. A distortion of phylogenetic results may also be a consequence of the adaptability of fast evolving mitochondrial genome; this adaptability has been shown in different groups of animals21,22,23,24,25. In contrast to these pitfalls, a big advantage of mtDNA, especially in the context of this study, is the ease of extraction because the number of mtDNA copies per cell greatly exceeds that of nuclear DNA. This state of affairs enables an analysis of old museum specimens when (especially in the case of a rare species) it is impossible to obtain fresh material.
Here we report the first genetic data from this enigmatic species, which were retrieved from a museum specimen. We assembled a mitochondrial genome of Selevinia and of several other species of the Gliridae family and combined this dataset with available mitogenomic sequences from GenBank to test our hypothesis about the phylogenetic position of Selevinia and other genera of the family.
Results
Mitochondrial genome structure and composition
For Selevinia betpakdalaensis, we de novo assembled the complete mitochondrial genome, which proved to be 16,608 bp in length (Fig. 1).
The map of the mitochondrial genome of Selevinia betpakdalaensis. Yellow ribbons mark annotations of protein-coding genes (CDSes); genes of rRNAs are highlighted in red, and genes of tRNAs in violet. The drawing of Selevinia is from Sokolov26.
The complete mitochondrial genome of S. betpakdalaensis contains the typical set of 13 protein-coding genes (PCGs), two ribosomal RNA genes (12S and 16S rRNA), 22 transfer RNA (tRNA) genes, and a putative control region (D-loop) (Fig. 1). The gene order and organisation in the S. betpakdalaensis mitogenome are consistent with those of other Gliridae representatives (Table S1). Nucleotide composition is significantly biased (levels of A, C, G and T are 31.5%, 23.8%, 13.6% and 31.1%, respectively) with a GC content of 37.4%. The GC skew of this genome was found to be −0.272: the highest among Gliridae values, ranging from −0.331 in Glirulus japonicus Schinz, 1845, to −0.283 in Muscardinus avellanarius Linnaeus, 1758 (Table S1). Nine genes (ND6 and seven tRNA genes) are oriented in the reverse direction, whereas the others are transcribed in the forward direction.
As is typical for most of mammalian species, the S. betpakdalaensis mitogenome harbours a total of 47 bp overlapping sequences in six regions. The longest overlap is 31 bp in length and located between ATP8 and ATP6. Initial codons in 13 PCGs of S. betpakdalaensis were found to be canonical putative start codons ATN (ATG for ND1, COX1-COX3, ATP8, ATP6, ND4L, ND4, ND6 and CYTB; ATT for ND2 and ND3; and ATA for ND5). The typical termination codon (TAA or TAG) was noted in eight PCGs. For ND1, ND2, ND4, ATP6 and COX3, the TAA stop codon is completed by the addition of 3′ A residues to mRNA. The CYTB stop codon is AGA as in other species (Table S2).
During the analysis of the Myomimus mitochondrial genome, it turned out that a species (Myomimus personatus Ognev, 1924) was recorded incorrectly in the voucher collection. According to our BLAST results, the similarity of the 12S gene of the analysed specimen with M. roachi Bate, 1937 (accession No. AJ536348) is 99.79%. Even in the absence of data on M. personatus in GenBank, with such a level of similarity, there is no doubt that our sample belongs to M. roachi. Additionally, our sample was trapped in Bulgaria, at a site within the geographic range of this species, whereas the geographic range of M. personatus is within mountain ranges of northeastern Iran and Turkmenistan (Kopet-Dag).
Mitochondrial genomes of Myomimus roachi and Glirulus japonicus were also found to be complete, 16,637 and 16,665 bp, respectively. The mitogenome of Graphiurus murinus Desmarest, 1822, assembled from SRA data proved to be 16,571 bp. Detailed information on the completeness of the 13 PCGs and their start and stop codons (for all the mitogenomes assembled in the current study) is given in Table S2.
Mitochondrial genomes sequenced and de novo–assembled in the current study were submitted to the NCBI GenBank database and have accession numbers PP971633–PP971635. The mitochondrial genome of G. murinus was uploaded to GitHub: https://github.com/ZaTaxon/Graphiurus_murinus.
Substitution saturation analysis
It is known that substitution saturation reduces a phylogenetic signal contained in sequences and complicates a phylogenetic analysis, especially if it involves deep branches. According to the analysis implemented in the DAMBE software (Table S3), the third codon position is either poorly suitable for phylogenetics (i.e. represents useless sequences) or shows substantial saturation. For all 13 PCGs, little saturation was revealed at the 1st codon position and 2nd codon position; thus, they were both suitable for the phylogenetic analysis.
Mitochondrial phylogeny of Gliridae
Gradual elimination of data subject to saturation
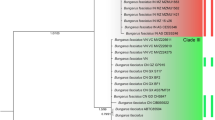
All types of the phylogenetic analysis (Fig. 2) indicated the monophyly of subfamilies Glirinae and Graphiurinae (groups highlighted in blue and orange, respectively) and that the monophyly of the subfamily Leithiinae is not stable. Selevinia was found to be a sister taxon to Myomimus (red branches) in all the cases.
Results of both Bayesian inference (BI) and maximum likelihood (ML) analyses involving all three codon positions (Fig. 2A, D) showed a sister position of Muscardinus to the Selevinia-and-Myomimus cluster, while garden and forest dormice proved to be sister to the Glirinae-and-Graphiurinae cluster. In both cases, in contrast to high Bayesian posterior probabilities (0.97 and 0.99), ML bootstrap support values turned out to be quite low (49% and 54%, respectively).
When the most saturated sites (transitions at the third codon position) were excluded from the analysis, the BI result (Fig. 2B) was similar to ML results both at the previous stage and in this case (Fig. 2E). It revealed a clustering of garden and forest dormice with Muscardinus, Selevinia and Myomimus: i.e. we observed monophyly of the subfamily Leithiinae, albeit poorly supported one (51%).
If the third codon position was entirely excluded from the analysis, then the BI method yielded trichotomy: garden and forest dormice represent a separate branch (Fig. 2C). The topology of the ML tree remained the same as at the previous step, but support for the subfamily Leithiinae was slightly greater (Fig. 2F).
Bayesian and Maximum Likelihood phylogenies based on mitochondrial PCG data. Three genes were used for Dryomys nitedula (CYTB, COX1 and ND1), and 13 PCGs for other species. (A–C): BI phylogenies; (D–F): ML phylogenies. (A ,D): all positions were used; (B,E): transitions at the 3rd codon position were excluded; (C,F): only 1st and 2nd positions were employed. The outgroup (Sciuridae) is hidden.
Examination of an extended dataset
Tree reconstruction based on the most complete taxonomic sample of 17 Gliridae species (based on partial 13 PCGs and 12S concatenated alignment) (Fig. S1) turned out to be the most supported. Both BI and ML analyses revealed a conventional taxon pattern – identical to that seen with ML applied to a smaller taxon set (Fig. 2E, F) – and identified three subfamilies: sister ones Glirinae and Graphiurinae as well as Leithiinae. Compared with the result of the analysis of the reduced dataset, the support of all nodes was significantly greater: the support of the subfamily Leithiinae was maximal (bpp = 1, bs = 94–96%), and within the subfamily, the sister position of Muscardinus toward Dryomys and Eliomys was rather well supported too (bpp = 0.91–0.92, bs = 65–69%) (Fig. S1). The cluster uniting Selevinia and Myomimus is basal within the subfamily. The presence of transversions of the 3rd position in the analysis (Fig. S1, part В) affected only the branching order within Graphiurus and slightly increased the support of several nodes.
Divergence dating
The first split within Gliridae (separation time of the subfamily Leithiinae) was estimated as an average of 34.6 Mya with 95% highest posterior density (HPD) of 41.04–28.03 Mya. Divergence time of subfamilies Graphiurinae and Glirinae was assessed at 32.67 Mya (95% HPD of 39.07–26.18 Mya). The first split within Leithiinae was estimated as an average of 31.24 Mya (95% HPD of 37.27–25.04 Mya) (Fig. 3).
Phylogenetic reconstruction of Gliridae as inferred from mitochondrial genomes (13 PCGs with the 3rd codon position excluded and 12S rRNA). Node labels show Bayesian posterior probability/ML bootstrap support; black circles denote nodes with 0.95–1.0 BI and 95–100 ML support. Blue bars represent 95% HPD intervals around mean estimates of divergence times. Plio: Pliocene, Plei: Pleistocene. Species names for which there are complete mitogenomes are black, and species for which mitogenomes were assembled as part of this work are marked with asterisks.
Discussion
In the current study, we for the first time sequenced and de novo assembled a complete mitochondrial genome of a rare rodent endemic to Kazakhstan – the desert dormouse S. betpakdalaensis – for which no genetic data have been obtained so far. Besides, we assembled complete mitochondrial genomes of three representatives of Gliridae: those of M. roachi and G. japonicus were derived from museum specimens, and the mitogenome of G. murinus was assembled from SRA data published previously.
The gene order and organisation of the mitochondrial genome of S. betpakdalaensis proved to be similar to those of other Gliridae representatives and typical for other vertebrates27. The mitochondrial genome of S. betpakdalaensis is characterised by an intermediate GC content (37.4%) compared with other Gliridae mitochondrial genomes (where it was found to be 34.6–38.6%).
Our results clearly indicate a strong influence of saturation on the topology; moreover, the topology may be erroneous but well supported (Fig. 2A). Saturation tests (Table S3) visualised the effect of saturation on the obtained phylogeny (Fig. 2). As a result of the BI analysis with complete exclusion of the 3rd codon position (Fig. 2C), trichotomy was observed, with an uncertain position of the cluster of garden and forest dormice. Meanwhile, the ML analysis of both types (with the exclusion of transitions at the 3rd codon position or with the exclusion of all 3rd positions) yielded the same result, identical to both ML and BI phylogenies obtained with the complete set of genes and species (Fig. 3). These findings are well consistent with modern classification1. At the same time, ML analysis was found to be more resistant to saturation.
Phylogenetic reconstruction of 17 Gliridae species inferred from the 13 PCGs (3rd codon position excluded due to saturation) and 12S concatenated alignment revealed tree topology overall similar to the one previously published by Bover et al.2: three subfamilies (Glirinae, Graphiurinae and Leithiinae) were identified. Nevertheless, our result allowed us to resolve some complex nodes and obtain a supported topology. Clustering of Glirinae and Graphiurinae was demonstrated only by Bover et al.2, and the support of this cluster was low (bpp = 0.49). In the same paper, monophyly of Leithiinae was demonstrated, but the branching order within it was not resolved. Our findings resolved the topology within Leithiinae: the cluster of Myomimus and Selevinia turned out to be an early derivative, and Muscardinus is sister to the group of Eliomys and Dryomys. In the previous molecular study2, the genus Muscardinus was basal to the subfamily Leithiinae, probably as a consequence of saturation (see the section above). On the other hand, both the basal position of Muscardinus and the lack of support for Myomimus in their study may be explained by incomplete taxa sampling, namely the absence of Selevinia. It should be mentioned here that Montgelard et al.15 have also reported the basal position of Muscardinus within Leithiinae with good support in a study involving 12S and some nuclear genes.
It is worth noting that although BI analysis provided high posterior probability support for most of major nodes, the ML approach failed to recover the time course of divergence between several branches. In fact, such a pattern is seen often during construction of phylogenies28,29 Thus, it is necessary to carefully discuss the validity of nodes (such as the position of Muscardinus and the most recent common ancestor [MRCA] of subfamilies Glirinae and Graphiurinae) that are poorly supported by the ML bootstrap values.
Trying to combine morphological and molecular data for living and extinct Gliridae, Dalmasso et al.30 have performed Bayesian divergence dating including fossil species (as tip dates) alongside their living relatives in a tree-building procedure (fossilised birth–death models). As a result, Dryomys (Eliomys was not analysed) turned out to be phylogenetically closer to Glirulus than to Myomimus, in agreement with our most saturated results (Fig. 2A, B, D).
As for the primary goal of our study, the first molecular data obtained for S. betpakdalaensis strongly support its position as a sister taxon to M. roachi, as demonstrated earlier in studies based on middle-ear features and cranial and mandibular morphology11,12,31.
The divergence dates based on 13 PCGs and 12S rRNA as a whole do not differ much from the estimates made in previous articles (Table 1), except that we managed to significantly reduce the confidence intervals.
Thus, the age of the Gliridae family can be estimated as 34.6 (28.03–41.04) million years, which is slightly younger than Bover et al.’s2 estimate based on the same root calibration: 38.5 (26.91–50.08) million years. In the work of Dalmasso et al.30, the average age of the node uniting modern representatives of Glirulus, Dryomys and Myomimus (representatives of Graphiurus were not used in that study) was estimated at ~ 30 million years.
Our estimate for the MRCA of the subfamily Leithiinae is 31.24 (37.27–25.04) Mya, in good agreement with the average estimate 31.0 (41.4–20.6) made by Bover et al.2 by means of a 1330 bp fragment (CYTB and 12S). The age of the subfamily Glirinae, estimated in our study at 28.34 (35.05–21.56) million years, proved to be very close to the results of all previous projects (Table 1) based on nuclear genes16, mitochondrial genes2, or their combination15. The subfamily Graphiurinae’s mean age estimated here as 16.64 (11.43–21.97) million years is also similar to the previous estimate of Bover et al.2 at ~ 17 million years. A fundamentally new accomplishment in our study is the determination of time of divergence of Selevinia from the mouse-tailed dormouse: an average of 23.98 (30.44–17.9) Mya, that is, approximately the Oligocene–Miocene boundary.
Materials and methods
Sampling
We analysed three skin tissue samples from the theriology collection at the Zoological Institute of the Russian Academy of Sciences (RAS) (Saint Petersburg, Russia): S. betpakdalaensis (No. 70212), the masked mouse-tailed dormouse M. roachi (No. 44191) and the Japanese dormouse G. japonicus (No. 52705); see Table S1 for details.
In addition to the three museum specimens for which data were obtained in the present study, we downloaded raw reads of G. murinus from the NCBI SRA database (SRR7704813). Four complete mitochondrial genomes of Gliridae were downloaded from NCBI GenBank: those of Glis glis Linnaeus, 1766, Graphiurus kelleni Reuvens, 1890, Eliomys quercinus Linnaeus, 1766, and Muscardinus avellanarius. Besides, we included COX1, CYTB and ND1 partial sequences for Dryomys nitedula Pallas, 1778; CYTB and ND1 fragments for D. laniger Felten & Storch, 1968; partial CYTB sequences for Hypnomys morpheus Bate, 1918, and Eliomys melanurus Wagner, 1840; and 12S rRNA sequences for all available Gliridae species. The study sample in total included 17 Gliridae species and five Sciuridae species as an outgroup: Ratufa bicolor Sparrman, 1778, Sciurus vulgaris Linnaeus, 1758, Pteromys volans Linnaeus, 1758, Marmota himalayana Hodgson, 1841, and Tamias sibiricus Laxmann, 1769. See Table S4 for details.
Ethics declaration
Our study was conducted using a voucher collection of the Zoological Institute of the RAS, and the research did not require fieldwork or live animal experimentation. The tissue samples used in the work are publicly deposited and accessible by others in a permanent repository at the Zoological Institute of the RAS. Methods are reported in accordance with ARRIVE guidelines.
DNA extraction, Library Preparation and sequencing
To reduce potential contamination, all manipulations with museum specimens were carried out in a separate laboratory room isolated from post-PCR facilities; it is predominantly employed for studies on historical samples from the collection at the Zoological Institute. All the working surfaces, instruments and plastics were sterilised with UV light. DNA from the museum skin samples was isolated by the phenol–chloroform extraction method32,33. DNA quality was checked with a Qubit 4.0 Fluorometer (Thermo Fisher Scientific, USA), and determination of the final library length distribution and checking for the absence of adapters were performed using Bioanalyzer 2100 (Agilent, Santa Clara, CA, USA). Sequencing was performed on Illumina Novaseq 6000 (Illumina, USA) in paired-end mode (2 × 150 bp) at the Core Sequencing Center of Kurchatov Center for Genome Research (national research center Kurchatov Institute, Russia).
Mitochondrial Genome Assembly, annotation and sequence analyses
The quality of raw reads was evaluated in FastQC ver. 0.11.934, then reads were rid of Illumina adapters, overrepresented sequences and low-quality reads (< Q20) using Trimmomatic v0.3935.
Clean reads were assembled in plasmidSPAdes version 3.10.136,37 with default settings. The contigs were annotated on the MITOS Web server38 with default settings. Gene boundaries were checked and refined by alignment against published Gliridae mitogenomes. Nucleotide composition and codon usage were calculated using Geneious Prime 2019.1 (Biomatters Ltd., Auckland, New Zealand). To calculate the GC skew, we utilized a known formula: GC skew = (G − C)/(G + C)39.
Sequence alignment
Overall, we investigated two taxon sets. The reduced one consisted of 14 species: eight Gliridae species with complete mitogenomes available and D. nitedula (COX1, CYTB and ND1 partial sequences); five Sciuridae species served as an outgroup. The complete taxon set (22 species) also included Gliridae species for which CYTB and/or 12S rRNA data were available in the NCBI (Table S2).
Taking into account the old age of the group and trying to avoid the phylogenetic reconstruction bias caused by saturation, we first used a concatenated alignment of 13 PCGs (excluding tRNAs and 12S and 16S rRNAs) for the reduced taxon set and 13PCGs + 12S rRNA for the complete one (simply to enlarge the taxon sampling). We performed the multiple alignments using MUSCLE40 implemented in Geneious Prime 2019.1 (Biomatters Ltd., Auckland, New Zealand).
Saturation tests
The third codon position is especially susceptible to saturation; this phenomenon is especially noticeable in the case of ancient groups. We performed the substitution saturation analysis on the reduced dataset consisting of 14 sequences by the Xia test41 implemented in the DAMBE 7.3.32 software42. This procedure is based on calculation of an index of substitution saturation (Iss) and of a critical Iss (Iss.c). We analysed 13 PCGs by examining the 1st, 2nd and 3rd codon positions. Additionally, we studied the same genes by focusing only on the 1st and 2nd codon positions. To interpret the obtained results, we conducted a comparison between the values of Iss and Iss.c and examined significant differences. According to the findings, we applied the method of RY-masking (R for purines and Y for pyrimidines) to transitions at the 3rd codon position to prevent saturation43.
Phylogenetic Reconstruction and Divergence Dating
To identify suitable substitution models, we utilised PartitionFinder v2.1.144. Due to possible saturation at the 3rd codon position of PCGs, three types of analyses were performed on both the complete and reduced taxon sets (see the Sequence Alignment section): (1) the 1st, 2nd and 3rd codon positions; (2) the 1st, 2nd and RY-masked 3rd codon positions; and (3) only the 1st and 2nd codon positions. Alignments were partitioned into 13 PCGs and 12S rRNA. The procedure was performed via the greedy algorithm45 in the PhyML software46. The ‘models’ option was set to ‘mrbayes’. The corrected Akaike Information Criterion (AICc) was used for model selection. All recommended models are listed in Table S5.
First, we performed preliminary analyses on three 14-species datasets with different degrees of the saturation influence by varying the involvement of the 3rd codon position in the analysis (Fig. 2). The extended taxon set consisted of 13 species with complete mitochondrial genomes available and all the mitochondrial genes’ data available in GenBank (Fig. 3).
Phylogenies were reconstructed by ML and BI methods. Trees were rooted by means of five Sciuridae species. The ML analysis was carried out on the IQ-TREE Web server47 with 10,000 ultrafast bootstrap replicates48. The BI analysis was performed in MrBayes 3.2.649. Each BI analysis started with random trees and was performed as two independent runs with four independent Markov Chain Monte Carlo (MCMC) algorithms for 10 million generations with sampling every 1,000th generation; the first 25% of the sampled trees were discarded as burn-in. Stationarity was examined in Tracer v1.7.250.
Estimates of divergence times among Gliridae taxa were calculated in the BEAST v2.7.4 software51. According to results obtained with PartitionFider, we chose appropriate substitution models. We specified the age of the Hypnomys morpheus specimen as a mean between 4,456 and 9,164 BP2. An optimised relaxed clock with the fossilised birth–death model was applied as tree priors. Because the convergence of the procedure was insufficient, we fixed two clades obtained earlier by the BI and ML methods. The first clade was represented by Gliridae species, and the second one by Dryomys, Eliomys, Hypnomys and Muscardinus species.
To properly compare our results with previous molecular studies, we calibrated the analysis according to Bover et al.2. On the basis of findings of Montgelard et al.15, Nunome et al.16 and Mouton et al.52 and the earliest known fossil representatives of Sciuridae and Gliridae53, those authors constrained the age of the divergence between these families according to a uniform distribution of 55–50 Mya. It is worth noting that there are controversies about such antiquity of the common ancestor of modern representatives of dormice. Freudenthal and Martín-Suárez54 doubt that representatives of Glirinae, Myomiminae and Dryomyinae that existed in the Miocene lost the caecum independently; those authors offer an alternative view: all modern glirids are descendants of a single Middle Miocene species; consequently, modern Gliridae’s MRCA age should be recalculated as 16 Mya. Although authors of subsequent studies have probably been familiar with this opinion, it has not been reflected in the reconstructions carried out in both molecular2,52 and morphological studies30,55.
The final maximum clade credibility time tree was summarised from two replicate runs (100 million MCMC generations each, sampling every 10,000 generations, discarding the first 10% as burn-in). The consensus tree was next visualised using FigTree v1.6 (http://tree.bio.ed.ac.uk/software/figtree/, accessed on 26 November 2021), and divergence time bars were generated automatically from the output with the help of 95% HPD of the ages for each node.
Data availability
Data is provided within the manuscript and in supplementary information files. Mitochondrial genomes sequenced and de novo assembled in the current study were submitted to the NCBI GenBank database under the accession numbers PP971633-PP971635. Mitochondrial genome of G. murinus was uploaded to github: https://github.com/ZaTaxon/Graphiurus_murinus.
References
Holden, M. E. Family Gliridae. in Mammal Species of the World: A Taxonomic and Geographic Reference (eds Wilson, D. E. & Reeder, D. M.) 819–841 (Johns Hopkins University, Baltimore, (2005).
Bover, P. et al. Ancient DNA from an extinct Mediterranean micromammal— hypnomys morpheus (Rodentia: Gliridae)—Provides insight into the biogeographic history of insular dormice. J. Zool. Syst. Evol. Res.58, 427–438 (2020).
Daams, R. & De Bruijn, H. A classification of the Gliridae (Rodentia) on the basis of dental morphology. Hystrix Ital. J. Mammal6, (1995).
Bashanov, B. S. & Belosludov, B. A. A remarkable family of rodents from Kasakhstan, U.S.S.R. J. Mammal. 22, 311–315 (1941).
Argyropulo, A. J. & Vinogradov, B. About the new wonderful rodent of our fauna (Selevinia Paradoxa gen. et spec. Nov). Priroda. 1, 81–83 (1939).
Belosludov, B. A. & Bashanov, B. S. A new genus and species of rodent from the Central Kasakhstan (USSR). Ucheniye Zap Kazakstanskogo Univiversiteta Alma-Ata Ser. Biol.1, 81–86 (1938).
Ognev, S. I. Mammals of the U.S.S.R. and Adjacent Countries. Vol. V. Rodents.Published for the Smithsonian Institution and the National Science Foundation by the Israel Program for Scientific Translations, Jerusalem, Washington, D.C., (1947).
von Koenigswald, W. V. Die Schmelzmuster in den Schneidezähnen Der Gliroidea (Gliridae Und Seleviniidae, Rodentia, Mammalia) und ihre systematische Bedeutung. Z. Für Säugetierkd. 58, 92–115 (1993).
Storch, G. Affinities among living dormouse genera. Hystrix. 6, 51–62 (1994).
Yachontov, E. L. & Potapova, E. G. On the position of dormice (Gliroidea) in the system of rodents. Proc. Zool. Inst. Acad. Sci. USSR 243, 127–147 (1991).
Potapova, E. G. Morphological patterns and evolutionary pathways of the middle ear in dormice (Gliridae, Rodentia). Trak Univ. J. Sci. Res. Ser. B. 2, 159–170 (2001).
Hennekam, J. J. et al. Cranial anatomy of the Desert Dormouse, Selevinia betpakdalaensis (Rodentia, Gliridae), revealed by Micro-computed Tomography. J. Mamm. Evol.28, 457–468 (2021).
Wahlert, J. H., Sawitzke, S. L. & Holden, M. E. Cranial anatomy and relationships of dormice (Rodentia, Myoxidae). Am. Mus. Novit. 3061, 1–32 (1993).
Holden-Musser, M. E., Juškaitis, R. & Musser, G. M. Gliridae. in Handbook of the Mammals of the World - Volume 6. Lagomorphs and Rodents I (eds Wilson, D. E., Lacher, T. E. & Mittermeier, R. A.) 838–889 (Lynx Edicions, Barcelona, (2016).
Montgelard, C., Matthee, C. A. & Robinson, T. J. Molecular systematics of dormice (Rodentia: Gliridae) and the radiation of Graphiurus in Africa. Proc. R. Soc. Lond. B Biol. Sci. 270, 1947–1955 (2003).
Nunome, M., Yasuda, S. P., Sato, J. J., Vogel, P. & Suzuki, H. Phylogenetic relationships and divergence times among dormice (Rodentia, Gliridae) based on three nuclear genes. Zool. Scr.36, 537–546 (2007).
Brown, W. M., George, M. & Wilson, A. C. Rapid evolution of animal mitochondrial DNA. Proc. Natl. Acad. Sci. 76, 1967–1971 (1979).
Hassanin, A. et al. Evolutionary history of Carnivora (Mammalia, Laurasiatheria) inferred from mitochondrial genomes. PLOS ONE. 16, e0240770 (2021).
Bodrov, S. Y. et al. Evolutionary history of mountain voles of the subgenus Aschizomys (Cricetidae, Rodentia), inferred from mitochondrial and nuclear markers. Integr. Zool.15, 187–201 (2020).
Franco-Trecu, V. et al. Sex beyond species: the first genetically analyzed case of intergeneric fertile hybridization in pinnipeds. Evol. Dev.18, 127–136 (2016).
Bondareva, O. et al. Signatures of adaptation in mitochondrial genomes of Palearctic Subterranean Voles (Arvicolinae, Rodentia). Genes. 12, 1945 (2021).
Shen, Y. Y. et al. Adaptive evolution of energy metabolism genes and the origin of flight in bats. Proc. Natl. Acad. Sci. 107, 8666–8671 (2010).
Ramos, E. K. D. S., Freitas, L. & Nery, M. F. The role of selection in the evolution of marine turtles mitogenomes. Sci. Rep.10, 16953 (2020).
Da Fonseca, R. R., Johnson, W. E., O’Brien, S. J., Ramos, M. J. & Antunes, A. The adaptive evolution of the mammalian mitochondrial genome. BMC Genom.9, 119 (2008).
Wang, Y., Xiong, F. & Song, Z. Molecular phylogeny and adaptive mitochondrial DNA evolution of Salmonids (Pisces: Salmonidae). Front. Genet.13, 903240 (2022).
Sokolov, V. E. Rare and Endangered Animals. Mammals (Vysshaya Shkola, 1986).
Anderson, S. et al. Complete sequence of bovine mitochondrial DNA conserved features of the mammalian mitochondrial genome. J. Mol. Biol.156, 683–717 (1982).
Suzuki, Y., Glazko, G. V. & Nei, M. Overcredibility of molecular phylogenies obtained by bayesian phylogenetics. Proc. Natl. Acad. Sci.99, 16138–16143 (2002).
Douady, C. J. Comparison of bayesian and maximum likelihood bootstrap measures of phylogenetic reliability. Mol. Biol. Evol.20, 248–254 (2003).
Dalmasso, A., Peláez-Campomanes, P. & López‐Antoñanzas, R. Relative performance of bayesian morphological clock and parsimony methods for phylogenetic reconstructions: insights from the case of Myomiminae and Dryomyinae glirid rodents. Cladistics. 38, 702–710 (2022).
Rossolimo, O. L., Potapova, E. G., Pavlinov, I. Y., Kruskop, S. V. & Voltzit, O. V. Dormice (Myoxidae) of the World (Moscow University, 2001).
Barnett, R., Larson, G. A. & Phenol–Chloroform Protocol for extracting DNA from ancient samples. in Ancient DNA (eds Shapiro, B. & Hofreiter, M.) vol 840 13–19 (Humana, Totowa, NJ, (2012).
Green, M. R. & Sambrook, J. Isolation of High-Molecular-Weight DNA from Mammalian Tissues Using Proteinase K and Phenol. Cold Spring Harb. Protoc. pdb.prot093484 (2017). (2017).
Andrews, S. & FastQC: A Quality Control Tool for High Throughput Sequence Data. (2010).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics. 30, 2114–2120 (2014).
Bankevich, A. et al. SPAdes: a New Genome Assembly Algorithm and its applications to single-cell sequencing. J. Comput. Biol.19, 455–477 (2012).
Antipov, D. et al. plasmidSPAdes: assembling plasmids from whole genome sequencing data. Bioinformatics btw493 doi: (2016). https://doi.org/10.1093/bioinformatics/btw493
Bernt, M. et al. Improved de novo metazoan mitochondrial genome annotation. Mol. Phylogenet Evol. 69. MITOS, 313–319 (2013).
Perna, N. T. & Kocher, T. D. Patterns of Nucleotide Composition at Fourfold Degenerate Sites of Animal Mitochondrial Genomes. J. Mol. Evol.41, 353–358 (1995).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res.32, 1792–1797 (2004).
Xia, X., Xie, Z., Salemi, M., Chen, L. & Wang, Y. An index of substitution saturation and its application. Mol. Phylogenet Evol.26, 1–7 (2003).
Xia, X. DAMBE7: New and Improved Tools for Data Analysis in Molecular Biology and Evolution. Mol. Biol. Evol.35, 1550–1552 (2018).
Abramson, N. I. et al. Phylogenetic relationships and taxonomic position of genus Hyperacrius (Rodentia: Arvicolinae) from Kashmir based on evidences from analysis of mitochondrial genome and study of skull morphology. PeerJ. 8, e10364 (2020).
Lanfear, R., Frandsen, P. B., Wright, A. M., Senfeld, T. & Calcott, B. PartitionFinder 2: new methods for selecting partitioned models of evolution for molecular and morphological phylogenetic analyses. Mol. Biol. Evol.msw260https://doi.org/10.1093/molbev/msw260 (2016).
Lanfear, R., Calcott, B., Ho, S. Y. W., Guindon, S. & PartitionFinder Combined selection of partitioning schemes and substitution models for phylogenetic analyses. Mol. Biol. Evol.29, 1695–1701 (2012).
Guindon, S. et al. New algorithms and methods to Estimate Maximum-Likelihood Phylogenies: assessing the performance of PhyML 3.0. Syst. Biol.59, 307–321 (2010).
Trifinopoulos, J., Nguyen, L. T., von Haeseler, A. & Minh, B. Q. W-IQ-TREE: a fast online phylogenetic tool for maximum likelihood analysis. Nucleic Acids Res.44, W232–W235 (2016).
Hoang, D. T., Chernomor, O., Von Haeseler, A., Minh, B. Q. & Vinh, L. S. UFBoot2: improving the Ultrafast bootstrap approximation. Mol. Biol. Evol.35, 518–522 (2018).
Ronquist, F. et al. MrBayes 3.2: efficient bayesian phylogenetic inference and model choice across a large Model Space. Syst. Biol.61, 539–542 (2012).
Rambaut, A., Drummond, A. J., Xie, D., Baele, G. & Suchard, M. A. Posterior summarization in bayesian phylogenetics using Tracer 1.7. Syst. Biol.67, 901–904 (2018).
Bouckaert, R. et al. BEAST 2: a Software platform for bayesian evolutionary analysis. PLoS Comput. Biol.10, e1003537 (2014).
Mouton, A. et al. Evolutionary history and species delimitations: a case study of the hazel dormouse, Muscardinus avellanarius. Conserv. Genet.18, 181–196 (2017).
Hartenberger, J. L. Description de la radiation des Rodentia (Mammalia) Du Paléocène supérieur Au Miocène; incidences phylogénétiques. Comptes Rendus Académie Sci. - Ser. IIA - Earth Planet. Sci.326, 439–444 (1998).
Freudenthal, M. & Martínez-Suárez, E. New ideas on the systematics of Gliridae (Rodentia, Mammalia). Span. J. Palaeontol.28, 239 (2020).
Lu, X., Costeur, L., Hugueney, M. & Maridet, O. New data on early oligocene dormice (Rodentia, Gliridae) from southern Europe: phylogeny and diversification of the family. J. Syst. Palaeontol.19, 169–189 (2021).
Acknowledgements
The authors are thankful to Olga Makarova and Eugenya Maksimova (curators of a small-mammal collection in the Laboratory of Theriology, Zoological Institute RAS in St. Petersburg) for their assistance and Dr. Leonid Voyta for fruitful discussions during the work on this manuscript. Special thanks go to Dr. Maria Logacheva and Alexei Penin for inspiring the study. Authors thankful to two anonymous referees for taking time and fruitful comments that help to improve the initial version of the manuscript.The English language was corrected by shevchuk-editing.com.
Funding
Funding for this study was provided by the Ministry of Science and Higher Education of the Russian Federation, project No. 075-15-2021-1069 and state research assignment No. 122031100282-2.
Author information
Authors and Affiliations
Contributions
T.V.P., prepared figures, wrote the main text, methodology, carried formal analysis,; V.A.P., genome assembly, Formal analysis, Writing—original draft, Writing—review and editing; S.Yu.B., Data curation, Methodology, Writing—review and editing; N.I.A., Conceptualization, Methodology, Resources, Writing—original draft, Writing—review and editing, Project administration, Funding acquisition. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Petrova, T.V., Panitsina, V.A., Bodrov, S.Y. et al. The mitochondrial genome of the critically endangered enigmatic Kazakhstani endemic Selevinia betpakdalaensis (Rodentia: Gliridae) and its phylogenetic relationships with other dormouse species. Sci Rep 14, 22259 (2024). https://doi.org/10.1038/s41598-024-73703-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-73703-2
Keywords
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.