Abstract
Despite the increasing use of lung ultrasound (LUS) in the evaluation of respiratory disease, operators’ competence constrains its effectiveness. We developed a deep-learning (DL) model for multi-label classification using LUS and validated its performance and efficacy on inter-reader variability. We retrospectively collected LUS and labeled as normal, B-line, consolidation, and effusion from patients undergoing thoracentesis at a tertiary institution between January 2018 and January 2022. The development and internal testing involved 7580 images from January 2018 and December 2020, and the model’s performance was validated on a temporally separated test set (n = 985 images collected after January 2021) and two external test sets (n = 319 and 54 images). Two radiologists interpreted LUS with and without DL assistance and compared diagnostic performance and agreement. The model demonstrated robust performance with AUCs: 0.93 (95% CI 0.92–0.94) for normal, 0.87 (95% CI 0.84–0.89) for B-line, 0.82 (95% CI 0.78–0.86) for consolidation, and 0.94 (95% CI 0.93–0.95) for effusion. The model improved reader accuracy for binary discrimination (normal vs. abnormal; reader 1: 87.5–95.6%, p = 0.004; reader 2: 95.0–97.5%, p = 0.19), and agreement (k = 0.73–0.83, p = 0.01). In conclusion, the DL-based model may assist interpretation, improving accuracy and overcoming operator competence limitations in LUS.
Similar content being viewed by others
Introduction
In recent years, lung ultrasound (LUS) has emerged as a valuable diagnostic tool for various respiratory diseases, including pneumonia, pulmonary edema, embolism, and pneumothorax1,2. It offers a non-invasive, cost-effective, and quick alternative to traditional methods without the need for radiation exposure3. LUS exhibits superior sensitivity compared to chest radiography (CXR)4, while maintaining a similar level of specificity5. Furthermore, it has proven effective in identifying conditions such as pleural effusion, pneumonia, pneumothorax, and pulmonary edema6. Nevertheless, the accuracy of LUS findings is highly reliant on the operators’ experience. In real-world clinical scenarios, LUS is performed by clinicians with varying degrees of expertise. A study assessing inter-reader agreement for LUS findings revealed only moderate to substantial agreement7. Ensuring precise and consistent interpretation among operators is crucial for guiding appropriate management in clinical practice.
In the field of medical imaging, deep learning (DL) techniques are widely used for classification, detection, segmentation, and noise reduction. Past research has developed DL-based models to aid physicians in diagnosing LUS-specific conditions8,9,10. For instance, Arntfield et al.8 created a DL model to distinguish A-line and B-line patterns, and Shang et al.9 enhanced radiologists' performance in diagnosing COVID-19 pneumonia via ultrasound images. Recently, multiclass DL models have been developed for LUS11,12,13. Ebadi et al.11 achieved 90% accuracy in classifying A-line, B-line, and pleural effusion/consolidation. Shea et al.12 further classified LUS images into five categories: A-line, B-line, confluent B-lines, consolidation, and pleural effusion. However, these studies did not assess whether the models enable consistent diagnostic performance among operators. Additionally, abnormal patterns like consolidation and B-lines can coexist in a single LUS image. Therefore, this study aimed to develop a DL model for multi-label classification of LUS images into normal, B-lines, consolidation, and effusion, and to evaluate its potential in assisting diagnostic performance and inter-reader agreement.
Results
Baseline characteristics of patients
Table 1 shows the distribution of LUS findings in each dataset. For model development (training, tuning, internal set), 7,580 LUS images from 1645 patients (mean age, 62.2 ± 16.6; 1018 male) were used. The temporally separated test set consists of 985 images from 146 patients, whereas 319 images from 54 patients and 53 LUS were collected as the first external test set and second external test set, respectively. Pleural effusion is the most common finding in developmental dataset (71.4%) and temporally separated test set (30.4%), while ‘normal’ was the most frequently observed in the first external test set. In the second external test set (n = 54), all four categories showed similar frequencies ranging from 29.6 to 35.2%.
Model performance
The results of model performance across datasets are summarized in Table 2. In the temporally separated test set, the AUCs for normal, B-lines, consolidation, and effusion were 0.93 (95% CI 0.92–0.94), 0.87 (95% CI 0.84–0.89), 0.82 (95% CI 0.78–0.86), and 0.94 (95% CI 0.93–0.95) respectively. The AUCs for normal, B-lines, consolidation, and pleural effusion were 0.89 (95% CI 0.87–0.91), 0.78 (95% CI 0.74–0.81), 0.87 (95% CI 0.83–0.89), and 0.89 (95% CI 0.87–0.91) in the first external test set, and 0.802 (95% CI 0.78–0.83), 0.81 (95% CI 0.80–0.83), 0.77 (95% CI 0.75–0.80), and 0.77 (95% CI 0.74–0.79) in the second external test set (Fig. 3).
Results of reader test
Both readers demonstrated improved accuracy in binary discrimination (normal vs. other than normal) and B-line detection when aided by the DL model. Especially, reader 1 exhibited a significant improvement in sensitivity (75.4% vs. 86.2%, p = 0.002) and accuracy (87.5% vs. 95.6%, p = 0.004) for binary discrimination (Table 3).
Table 4 delineates the outcomes of inter-reader agreements with or without the DL model. Regarding each classification, there were no significant differences in detecting consolidation and B-lines (p > 0.05), and the agreement for pleural effusion decreased with the DL model (k = 0.71 vs. 0.58, p = 0.03). However, the agreement for binary discrimination (normal vs. abnormal) displayed a substantial improvement (k = 0.73 vs.0.83, p = 0.01) with use of the DL model.
Discussion
The previous studies have primarily focused on multi-class classification11,12 and have applied LUS deep learning models in specific diseases such as sepsis, dengue, and COVID-1910,11,14,16. However, this study deployed a multi-label deep learning model to distinguish multiple findings or common patterns in LUS into four categories: normal, B-lines, consolidation, and pleural effusion, and assessed its performance across multiple test sets. The model identified LUS findings with AUCs of 0.80–0.93 for normal, 0.78–0.87 for B-lines, 0.77–0.87 for consolidation, and 0.77–0.94 for pleural effusion, though performance decreased in external test sets. In the observer performance test, both readers demonstrated improved accuracy in distinguishing abnormal from normal (reader 1: 87.5–95.6%, p = 0.004; reader 2: 95.0–97.5%, p = 0.19) with the DL model's assistance. Additionally, the DL model improved inter-reader agreement for determining abnormal from normal (k = 0.73–k = 0.83, p = 0.01) and detecting B-lines (k = 0.70–0.76, p = 0.22). Our results demonstrate the feasibility of using a DL model in LUS for patients with respiratory symptoms and support the potential for overcoming the limitations due to variability between readers.
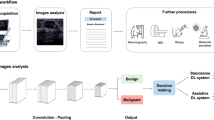
Although several DL-based models based on plain radiographs or CT scans have been developed for classifying respiratory disease, research on DL models applicable to LUS remains scarce. Ebadi et al.14 formulated a DL model utilizing LUS data from COVID-19 patients to classify LUS findings into normal, B-lines, and consolidation/effusion, achieving a model accuracy score of 0.90. Our study aligns with their work, also categorizing LUS findings into four distinct classes (Fig. 1). However, unlike the prior study, we developed a model that can distinguish between pleural effusion and consolidation individually.
In this study, the model exhibited strong performance in classifying LUS into normal, B-line, consolidation, and effusion, displaying AUCs ranging from 0.82 (consolidation) to 0.94 (effusion), and sensitivities ranging from 73.9% (consolidation) to 93.3% (normal) in the temporally separated test set. Nevertheless, its performance experienced a slight decrement in the external test set, with AUCs ranging from 0.78 (B-line) to 0.89 (normal) in the first external test set and 0.77 (effusion) to 0.81 (B-line) in the second external test. This disparity may be due to the use of different machines, the acquisition of LUS by various examiners, and differences in the distribution of LUS abnormalities. In reality, pleural effusion was the most common finding in the training and temporally separated test set. However, normal and consolidation were the most common findings in the first and second external test sets, respectively.
LUS is widely used in diagnosing and monitoring respiratory diseases in ICU patients1. Dave et al.15 conducted a prospective study aiming to ascertain the DL model's ability to differentiate normal and abnormal patterns on bedside LUS in critically ill patients, demonstrating a 95% accuracy of the model, and advocating for its utility in ICU settings. In this study, LUS data from critically ill ICU-admitted patients were utilized, constituting a second external test set (Supplementary Table 1). Similar to the preceding research, the model exhibited an AUC of 0.82 (95% CI 0.79–0.94) for binary discrimination, affirming its applicability in immobilized patients. However, in terms of multi-label classification, the model's performance was relatively inferior when compared to recent advancements in the domain. This suboptimal performance might stem from several factors, including LUS examinations conducted on uncooperative patients with limited movement capabilities, poor sonic window, and a limited number of included cases. Consequently, further test employing a larger dataset appears essential to enhance the DL model's efficacy in multi-classification tasks.
Despite the numerous advantages of ultrasound imaging, its susceptibility to operator dependency presents a significant limitation. Previous research on inter-reader agreements indicated moderate to substantial concordance (k = 0.36–0.74), with a noted decrease in agreement when positive abnormalities such as consolidation or B-lines were encountered16. Consistent with these findings, our study observed substantial to moderate agreements for B-lines (k = 0.698), effusion (k = 0.710), and consolidation (k = 0.568), similar to prior investigations. Meanwhile, the integration of DL techniques has shown enhanced inter-reader agreements across various medical tasks and anatomical regions17,18,19. Our study also demonstrated that the utilization of the DL model significantly improved binary discrimination (k = 0.825, p = 0.01) and increased agreements for B-lines (k = 0.756, p = 0.22). Recognizing B-lines in LUS is important as they manifest in various pulmonary conditions, including pulmonary edema, interstitial fibrosis, and pneumonia20. Nhat et al.11 also demonstrated improved performance in interpreting LUS findings among non-expert clinicians, although they did not evaluate inter-reader agreement. However, our study indicated a lower level of agreement in the interpretation of consolidation and effusion. This observation may be attributed to the image-based analysis of LUS rather than video-based scrutiny. The observer performance test employed captured LUS images. Given LUS's established robust accuracy in diagnosing pleural effusion, direct implementation of LUS would likely distinguish effusion and consolidation with greater precision compared to image-based analysis.
The study indicated several limitations. First, it is a retrospective design study with a limited number of validated cases. Therefore, captured LUS was conducted with heterogeneous probe machines. However, extensive efforts were made to ensure comprehensive validation of the model through multiple test sets and observer performance tests. Second, the DL model developed for this study classified normal, B-line, consolidation, and pleural effusion based on representative captured frames rather than continuous video sequences. Consequently, evaluating the model's performance on video data remains a necessary avenue for exploration. Third, the reference standard for the training set was based on the interpretations of expert radiologists rather than the outcomes of CT scans. In clinical practice, LUS results are typically based on expert opinion without reliance on CT scans. Nevertheless, in this study, the observer performance test was conducted using selected cases that had undergone CT scans on the same day, and the reference standard was established based on CT findings.
Fourth, the observer performance test was executed with the participation of two radiologists. Generalizing the findings to other clinicians might pose challenges due to the limited scope of the participants. Further research is needed to develop models applicable to a wider range of scenarios, particularly in the implementation of optimization strategies for generalization and regulation purposes or deploying classifiers empowered by pre-trained weights from clinical images such as RadImageNet21 or RadFormer22. Additionally, the retrospective nature of our study presents inherent limitations, including potential biases in data selection and the inability to establish causal affinities. Therefore, further studies should focus on refining the model to handle a broader range of scenarios, such as video clip analysis, and improving its performance in multi-label classification tasks with large numbers of test datasets.
In conclusion, we developed multi-label DL model classifying lung ultrasound findings into normal, B-line, consolidation, and effusion, which enabled to enhance readers’ diagnostic accuracy and agreement between the readers.
Methods
This retrospective study obtained approval from the institutional review boards (IRBs) of [Seoul National University Hospital (SNUH)] and [Chung-Ang University Hospital (CAUH)] by the Ministry of Health and Welfare, Republic of Korea (approval number: H-2102-163-1199). Due to the retrospective nature of the study, IRB approval number H-2102-163-1199 waived the need of obtaining informed consent. All studies were performed according to relevant guidelines and regulations.
Patients
Utilizing retrospective data, the study encompasses an internal set and incorporates wo additional external test sets to comprehensively evaluate the model's effectiveness. The developmental dataset for the LUS model was collated from patients who underwent diagnostic or therapeutic thoracentesis between 2018 and 2021 at SNUH. Following thoracentesis procedures, an assessment of lung parenchyma was conducted to detect any parenchymal anomalies or quantify residual pleural effusion. Patients with poor image quality were subsequently excluded, including those with severe blurring or darkness where A-line, B-line, and consolidation could not be distinguished, as well as those with inappropriate image depth, making it difficult to evaluate the ultrasound patterns, which might impact the generalizability of the model.
Acquisition of lung ultrasound and standard of reference
One of the two thoracic radiologists (H.C. and H.W.J., possessing 7 and 6 years of experience in LUS, respectively) conducted LUS examinations using four machines from three distinct vendors: LOGIQ E9 (GE Healthcare, Illinois, United States), EPIQ 5G and EPIQ 7G (Philips Healthcare, The Netherlands), and RS80A (Samsung Healthcare, Republic of Korea). The operators opted for either a high-resolution linear probe (7–11 MHz) or a convex probe (3–5 MHz) based on individual preferences. Following the protocol recommended by the bedside lung ultrasound in emergency (BLUE)-protocol1, the operators assessed three lung points— anterior upper, anterior lower, and posterolateral areas—thus acquiring a minimum of six frames or images per patient.
Post-LUS examination, the operators categorized LUS findings into four groups: normal, B-lines, consolidation, and pleural effusion. Normal LUS denotes a horizontal echogenic line parallel to the pleural line (A-line) without any irregularities. B-lines represented vertical low echogenic lines synchronized with respiration, while consolidation indicated a low echogenic area with an irregular border in aerated lung parenchyma. The categorization offered by the two thoracic radiologists served as the reference standard.
Data partitions
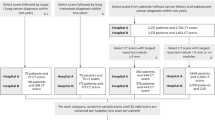
The patients included in the developmental dataset were randomly partitioned into training, validation, and internal test sets in an 8:1:1 ratio using a random number generator, ensuring that data from the same patient did not overlap across the sets. A manually fixed seed was implemented for reproducibility purposes. The test datasets were prepared in the following manner: (1) a temporally separated dataset at SNUH spanning from 2021 to 2022, serving as the temporally separated test set, (2) an external test set from 2021 to 2022 at CAUH, established as the first external test set, and (3) neurocritical ill patients admitted to an ICU at SNUH from 2017 to 2022, forming the second external test set (Fig. 2).
A total of 7580 images from 1645 patients were utilized in the training, validation and internal test stages. The temporally separated test set comprised 985 images obtained from 146 patients. Moreover, the first external test set consisted of 319 images from 54 patients, while the second external test set involved 53 LUS images. This methodology ensured the model's exposure to a diverse range of instances during training, test on distinct datasets to refine hyperparameters, and evaluation on a separate dataset to assess its overall performance and generalizability.
Data preprocessing
Convolutional Neural Networks (CNN) are typically applied in perceiving image classification to preserve spatial features. Moreover, the model selection has been made to EfficientNet-B023. The DICOM files underwent pseudonymization using an algorithm and were converted into PNG format during pre-processing. The input data have been converted from PNG format to tensors. For the classification task, we employed the EfficientNet-B0 model with ImageNet pre-trained weights, taking advantage of transfer learning to boost performance given our limited dataset. The images were resized to 224 × 224 pixels with three color channels (RGB) and further normalized to align with the distribution expected by the pre-trained model. The model was trained using a cross-validation approach, and its performance was assessed through accuracy, F1 score, and AUC-ROC metrics to ensure robustness and generalizability.
Deep learning-based model deployment in lung ultrasound
The integration of the Sigmoid activation function and BCE Loss in multi-label classification tasks is underpinned by a logical and effective approach. Augmentation techniques applied to random horizontal flips, controlled rotations up to 10 degrees, and random perspective distortion were employed to enhance the diversity of the data. Following augmentation, the images were converted into tensor format, an essential data structure in DL. The training process employed a learning rate of \(1{e}^{-5}\) and initialized model parameters using ImageNet pre-trained weights.The batch size was set to 128, and the model was trained for up to 2000 epochs using the Adam optimizer. Layers are not frozen to allow full utilization of pre-trained features and enable fine-tuning across the entire network. The classifier layer was modified to match the required output dimensions, incorporating a sigmoid activation function combined with BCE Loss to ensure correct output transformation for multi-label classification24,25. Additionally, a patience value of 100 was set for the early stopping mechanism. Optimal thresholds aimed at balancing sensitivity and specificity, were determined using an internal test dataset and subsequently applied across external datasets to evaluate model performance (Fig. 3). The experiments were conducted on hardware with an NVIDIA V100 32 GB GPU. The code is available at https://github.com/SeanPresent/iRAIL_LUS.
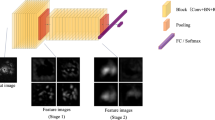
Basic feature extractor structure of EfficientNet-B0. The input data is converted from DICOM to PNG through a hand-made algorithm that goes through the model. The model combines EfficientNet-B0 with fully connected layers consisting of 4 nodes, fitting the purpose of classification. Afterward, the prediction scores are calculated using a sigmoid function and Binary Cross Entropy to enable multi-label classification.
Reader test
Arbitrarily selected 200 LUS images from the temporally separated test set and the first external test set was used for the observer performance test (OPT). For OPT, we arbitrarily selected patients among the patients who had taken chest CT and LUS on the same day to confirm the reference of the standard by comparing the results of the chest CT scan. Two board-certificated radiologists (W.H.L, and H.I.P., with 8- and 5- years of experience in thoracic radiology) independently read the test set two times over the 4-week intervals along with the DL model result and without the results (Figs. 1, 4).
Statistical analysis
Data presented with mean ± standard deviation for continuous variables and frequencies with percentages for categorical variables. Model performance is evaluated via AUC, sensitivity, specificity, and accuracy. Confidence intervals for the AUC values were calculated using the bootstrapping method, with predictions and true labels resampled 1000 times to estimate the variability and robustness of the model's performance. The optimal thresholds for determining LUS findings were determined based on the results of an internal test set, in which each classification (normal, B-lines, consolidation, and effusion) was balanced for sensitivity and specificity. The thresholds were applied across the first and second external test and evaluated the sensitivity, specificity, and accuracy in each test set. In the Observer Performance Test, accuracy, sensitivity, and specificity were assessed for both multi-label classification and binary discrimination (normal vs. abnormal [excluding normal, including B-line, consolidation, or pleural effusion]) by each reader. The inter-reader agreement was determined using Cohen's kappa coefficient: > 0.8 (almost perfect), 0.61–0.80 (substantial), 0.41–0.60 (moderate), 0.21–0.40 (fair), and 0–0.20 (slight). Statistical analyses were executed using Medcalc version 20.015 software (MedCalc, Mariakerke, Belgium), and a P-value below 0.05 was considered indicative of statistical significance.
Data availability
The datasets generated and/or analyzed during the current study are not publicly available due to the patient privacy regarding the IRB but are available from the corresponding author upon reasonable request.
Abbreviations
- AUC:
-
Area under the receiver operating characteristics curve
- DL:
-
Deep learning
- LUS:
-
Lung ultrasound
- ICU:
-
Intensive care unit
- OPT:
-
Observer performance test
References
Lichtenstein, D. A. & Mezière, G. A. Relevance of lung ultrasound in the diagnosis of acute respiratory failure: The BLUE protocol. Chest 134, 117–125 (2008).
Smit, M. R. et al. Lung ultrasound prediction model for acute respiratory distress syndrome: A multicenter prospective observational study. Am. J. Respir. Crit. Care Med. 207, 1591–1601 (2023).
Volpicelli, G. et al. International evidence-based recommendations for point-of-care lung ultrasound. Intens. Care Med. 38, 577–591 (2012).
Hew, M. & Tay, T. R. The efficacy of bedside chest ultrasound: From accuracy to outcomes. Eur. Respir. Rev. 25, 230–246 (2016).
Maw, A. M. et al. Diagnostic accuracy of point-of-care lung ultrasonography and chest radiography in adults with symptoms suggestive of acute decompensated heart failure: A systematic review and meta-analysis. JAMA Netw. Open 2, e190703 (2019).
Pivetta, E. et al. Lung ultrasound integrated with clinical assessment for the diagnosis of acute decompensated heart failure in the Emergency Department: A randomized controlled trial. Eur. J. Heart Fail. 21, 754–766 (2019).
Hassan, R. I. et al. Lung ultrasound as a screening method for interstitial lung disease in patients with systemic sclerosis. J. Clin. Rheumatol. 25, 304–307 (2018).
Arntfield, R. et al. Automation of lung ultrasound interpretation via deep learning for the classification of normal versus abnormal lung parenchyma: A multicenter study. Diagnostics 11, 2049 (2021).
Shang, S. et al. Performance of a computer aided diagnosis system for SARS-COV-2 pneumonia based on ultrasound images. Eur. J. Radiol. 146, 110066 (2022).
Xue, W. et al. Modality alignment contrastive learning for severity assessment of COVID-19 from lung ultrasound and clinical information. Med. Image Anal. 69, 101975 (2021).
Nhat, P. T. et al. Clinical benefit of AI-assisted lung ultrasound in a resource-limited intensive care unit. Crit. Care 27, 45 (2023).
Shea, D. E. et al. Deep learning video classification of lung ultrasound features associated with pneumonia. In 2023 IEEE/CVF Conference on Computer Vision and Pattern Recognition Workshops (CVPRW) (2023).
Howell, L. et al. Deep learning for real-time multi-class segmentation of artefacts in lung ultrasound. Ultrasonics 140, 107251 (2024).
Erfanian Ebadi, S. et al. Automated detection of pneumonia in lung ultrasound using deep video classification for COVID-19. Inf. Med. Unlocked 25, 100687 (2021).
Dave, C. et al. Prospective real-time validation of a lung ultrasound deep learning model in the ICU. Crit. Care Med. 51, 301–309 (2023).
Herraiz, J. L. et al. Inter-rater variability in the evaluation of lung ultrasound in videos acquired from COVID-19 patients. Appl. Sci. 13, 1321 (2023).
Liu, Z. et al. Small lesion classification on abbreviated breast MRI: Training can improve diagnostic performance and inter-reader agreement. Eur. Radiol. 32, 5742–5751 (2022).
Upton, R. et al. Automated echocardiographic detection of severe coronary artery disease using artificial intelligence. JACC Cardiovasc. Imaging 15, 715–727 (2022).
Palmer, M. et al. The diagnostic accuracy of chest radiographic features for pediatric intrathoracic tuberculosis. Clin. Infect. Dis. 75, 1014–1021 (2022).
Dietrich, C. F. et al. Lung B-line artefacts and their use. J. Thorac. Dis. 8, 1356–1365 (2016).
Mei, X. et al. RadImageNet: An open radiologic deep learning research dataset for effective transfer learning. Radiol. Artif. Intell. 4, e210315 (2022).
Basu, S. et al. Radformer: Transformers with global-local attention for interpretable and accurate gallbladder cancer detection. Med. Image Anal. 83, 102676 (2023).
Tan, M. & Le, Q. V. EfficientNet: Rethinking model scaling for convolutional neural networks. ArXiv (2019). https://arxiv.org/abs/1905.11946
Durand, T., Mehrasa, N. & Mori, G. Learning a deep convnet for multi-label classification with partial labels. ArXiv (2019). https://arxiv.org/abs/1902.09720
Liu, W., Wang, H., Shen, X. & Tsang, I. W. The emerging trends of multi-label learning. IEEE Trans. Pattern Anal. Mach. Intell. 44, 7955–7974 (2022).
Acknowledgements
This work was supported by the National Research Foundation of Korea (NRF) grant funded by the Ministry of Science and ICT (MSIT) (grant number: NRF-2018R1A5A1060031)
Author information
Authors and Affiliations
Contributions
D.H., and Y.K. contributed to modeling and simulation. W.H., H.C., and T.J.K. contributed to the collecting data. C.M.P., H.C., D.H., Y.K., J.C., T.J.K. and S.B.K. discussed the results and commented on them. C.M.P., Y.K., and H.C. initiated the study, supervised all aspects of the study and D.H., H.C., C.M.P. and Y.K. wrote the manuscript. All the authors commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Hong, D., Choi, H., Hong, W. et al. Deep-learning model accurately classifies multi-label lung ultrasound findings, enhancing diagnostic accuracy and inter-reader agreement. Sci Rep 14, 22228 (2024). https://doi.org/10.1038/s41598-024-72484-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-72484-y
Keywords
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.