Abstract
Alzheimer's disease is the predominant form of dementia, and disulfidptosis is the latest reported mode of cell death that impacts various disease processes. This study used bioinformatics to analyze genes associated with disulfidptosis in Alzheimer's disease comprehensively. Based on the public datasets, the differentially expressed genes associated with disulfidptosis were identified, and immune cell infiltration was investigated through correlation analysis. Subsequently, hub genes were determined by a randomforest model. A prediction model was constructed using logistic regression. In addition, the drug-target affinity was predicted by a graph neural network model, and the results were validated by molecular docking. Five hub genes (PPEF1, NEUROD6, VIP, NUPR1, and GEM) were identified. The gene set showed significant enrichment for AD-related pathways. The logistic regression model demonstrated an AUC of 0.952, with AUC values of 0.916 and 0.864 in validated datasets. The immune infiltration analysis revealed significant heterogeneity between the Alzheimer's disease and control groups. High-affinity drugs for hub genes were identified. Through our study, a disease prediction model was constructed using potential biomarkers, and drugs targeting the genes were predicted. These results contribute to further understanding of the molecular mechanisms underlying Alzheimer's disease.
Similar content being viewed by others
Introduction
Alzheimer's disease (AD) is a chronic neurodegenerative disorder with an insidious onset, characterized by neurotic plaques associated with the accumulation of amyloid β protein (Aβ) in brain tissue and neurofilament tangles derived from hyperphosphorylation of microtubule-associated tau proteins1, along with synaptic dysfunction, neuronal loss, and various other pathological manifestations2. Despite extensive research, a cure for this remains elusive. Alzheimer's disease, which accounts for 50–60% of dementia cases, significantly affects cognitive abilities, memory, and independence, posing a substantial challenge to individuals' lives. The prevalence of AD is increasing worldwide due to the aging population, becoming an increasingly globalized health issue3. Estimates indicate that 75% of patients with AD remain undiagnosed globally, and this percentage rises to as high as 90% in certain underdeveloped regions4. Moreover, in 2019, AD was even ranked as the 6th leading cause of death in the United States5. The global burden on public health is immense, as AD poses a significant challenge worldwide. Currently, there are four primary hypotheses proposed to elucidate the pathogenesis of AD: The Aβ amyloid protein cascade theory6, the tau protein hyperphosphorylation theory7, the mitochondrial dysfunction8 and oxidative stress theory9, as well as the neuroinflammatory response10. These hypotheses have provided valuable insights into AD's pathogenesis, and drugs that target removing amyloid plaques from the brain are already being utilized in clinical practice11. However, current clinical drugs face challenges in reversing the pathological processes of AD and have certain limitations. As a result, traditional Chinese medicines have garnered significant attention. Unlike Western medicines that typically target a single pathway, Chinese medicines, with their multi-component, multi-target, and multi-pathway efficacies, have demonstrated greater advantages in the treatment of AD. To identify more effective herbal medicines for AD, Zhang et al. established the Integrated Traditional Chinese Medicine (ITCM) platform, the largest herbal ingredient-based pharmacotranscriptomic database12. Alongside this, they developed the COIMMR computational framework13, which facilitates the rapid screening of active ingredients in traditional Chinese medicine. This approach significantly enhances the efficiency of drug discovery compared to traditional pharmacological experiments. Despite significant progress in understanding AD, the complexity of its pathogenesis has led to a limited understanding of its specific mechanisms. Therefore, elucidating the molecular mechanisms, identifying biomarkers for diagnosis and treatment, and developing precise diagnostic approaches are imperative for addressing the challenges posed by AD.
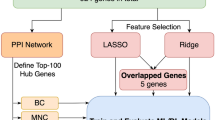
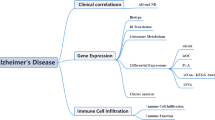
Disulfidptosis, an emerging mode of cell death, is triggered by disulfide stress. It is characterized by the accumulation of intracellular disulfides, resulting in the collapse of cytoskeletal proteins and F-actin, as supported in recent studies14. Notably, disulfidptosis has been strongly linked to tumor progression and has been implicated in various cancers, including bladder cancer15, breast cancer16, and hepatocellular carcinoma17,18, which contribute to the identification of new potential therapeutic targets. However, little research has been conducted to investigate the potential association between disulfidptosis and neurological disorders, particularly AD. Ma et al. explored genes and their subgroups associated with disulfidptosis in AD and constructed a predictive model19. Building on this work, we further identified key related genes, developed a model with higher diagnostic accuracy, and searched for drugs targeting these genes using an affinity prediction model. Additionally, we conducted in-depth studies on these key genes to elucidate their mechanisms of action and evaluate potential therapeutic targets. We also performed the immune infiltration analysis to investigate the interactions between these hub genes and immune cells, thereby examining the immune characteristics of AD. Figure 1 illustrates the flow chart outlining the study.
The workflow of the study. This study analyzed data from GSE33000 and GSE122063 for identifying DEGs. Intersecting these DEGs revealed 136 DEGs, 90 of which were associated with disulfidptosis-related genes. These 90 DEGs were subjected to enrichment analysis. Five hub genes were identified by randomforest for machine learning model construction and validation. A GAT_GCN model predicted affinities, validated by molecular docking.
Methods
Data acquisition and pre-processing
The Alzheimer's disease-related datasets were retrieved from the GEO database using the GEOquery package20. Three microarray datasets were obtained: GSE3300021, GSE12206322, and GSE528123, along with their corresponding gene annotation files. The GSE33000 dataset contains 310 AD samples and 157 control samples, the GSE122063 dataset contains 28 AD samples and 22 control samples, and the GSE5281 dataset contains 87 AD samples and 74 control samples. The samples were pre-processed using R, and the non-Alzheimer’s disease and abnormal control samples were filtered out24. Additionally, null gene probe counts were eliminated, and duplicate probe expression data were averaged. Furthermore, a set of 10 genes highly related to disulfidptosis was identified by Liu et al.25, which includes SLC7A11, SLC3A2, RPN1, NCKAP1, NUBPL, NDUFA11, LRPPRC, OXSM, NDUFS1, and GYS1.
Identification of disulfidptosis-related differentially expressed genes (DEGs)
The pre-processed expression data from two datasets, GSE33000 and GSE122063, were analyzed to identify DEGs between the AD group and the control group using the limma package26 (version 3.54.2) in R [screening criteria: p.adjust < 0.05, |log2fold change (FC)| > 1]. The up-regulated and down-regulated gene groups were intersected respectively and visualized in the Venn diagram. The resulting genes with consistent differential trends were considered the final DEGs for subsequent analyses. The Spearman correlation analysis was performed to screen out the DEGs associated with disulfidptosis (screening criteria: pvalue < 0.05, correlation > 0.75).
Identification of hub genes
The randomforest model is an integrated machine learning algorithm with decision trees based learner that provides variable importance scores during data analysis27. In this study, a randomforest model was constructed using the Randomforest package (version 4.7-1.1), and the feature importance metrics generated by the model were used to identify the hub genes.
Enrichment analysis
To explore the potential functions and biological mechanisms, the DEGs associated with disulfidptosis were subjected to gene ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analyses using the ClusterProfiler package (version 4.6.2)28. The GO analysis encompassed three levels: Biological Process (BP), Cellular Component (CC), and Molecular Function (MF). Furthermore, Gene Set Enrichment Analysis (GSEA) was performed on the final selected hub genes, using the correlation coefficients between hub genes and other genes to form the analysis list. The results of these analyses were visualized through histograms, network diagrams, and GSEA plots, where a significance level of p < 0.05 was considered indicative of significant enrichment.
Machine learning model construction and independent validation analysis
Logistic regression is a commonly employed statistical method for analyzing associations between diseases and causative factors, particularly in dichotomous classification problems29. In this study, a logistic regression model was constructed and validated on external datasets. The performance of the model in distinguishing between AD and non-AD samples was assessed using receiver operating characteristic (ROC) Curve analysis and by calculating the area under the ROC curve (AUC). The pROC package (version 1.18.0)30 in R was used to perform ROC analysis and obtain AUC values.
Evaluating the immune cell infiltration
CIBERSORT employs linear support vector regression to deconvolve the expression matrix of immune cell subtypes, which can estimate the abundance of immune cells and their characteristics of different populations31. The CIBERSORT package (version 0.1.0) was used to calculate the relative abundance of 22 immune cells in each sample of the gene expression matrix.
Drug affinity prediction
Graph Attention Network and Graph Convolutional Network (GAT_GCN) model, a graph neural network-based model for drug-target binding affinity prediction32, was applied to predicting the affinity between hub genes and drugs. For training the model, the KiBA dataset was used as the baseline dataset. The simplified molecular-input line-entry system (SMILES) of the drug compound was derived from the DrugBank database33. The targets were transformed into the amino acid sequences of the corresponding proteins, and each SMILES string strand was matched with each amino acid sequence to predict their binding affinity.
Molecular docking validation
The protein crystal structures in PDB format were retrieved from UniProt34. The three-dimensional (3D) structures of the drug compounds were obtained from PubChem. CB-DOCK235 was utilized to predict the binding cavity to which the small molecule binds. Global docking of the compounds and targets was performed using AutoDock Vina36, and binding energy scores were used to assess the binding ability of the drug-target interactions. 3D docking plots of the docking results were generated using Pymol.
Statistical analysis
All statistical analyses were performed using R version 4.1.2. Spearman's correlation analysis was used to determine the correlation between two variables. The Wilcox rank-sum test was utilized to analyze the difference between the two groups. Statistical significance was defined as pvalue < 0.05.
Results
Identification of the disulfidptosis-related DEGs in AD
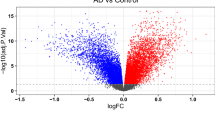
To identify the DEGs associated with AD, differential expression analysis was conducted in the GSE33000 and GSE122063 datasets following pre-processing. As a result, 377 DEGs were identified in the GSE33000 dataset, including 187 up-regulated genes and 190 down-regulated genes (Fig. 2A and Supplementary Table 1). Similarly, 716 DEGs were identified in the GSE122063 dataset, consisting of 235 up-regulated genes and 481 down-regulated genes (Fig. 2B and Supplementary Table 2). To visualize the expression patterns of the DEGs, a heatmap displaying the expression levels of 20 selected DEGs was generated (Fig. 2C,D). Upon comparing the two datasets, it was observed that 136 genes exhibited consistent expression trends, including 43 up-regulated genes and 93 down-regulated genes (Fig. 2E,F and Supplementary Table 3).
Identification of differentially expressed genes (DEGs) in AD. (A) Volcano plot of DEGs in the GSE3300 dataset. (B) Volcano plot of DEGs in the GSE122063 dataset. (C) Heatmap of selected DEGs in the GSE3300 dataset. (D) Heatmap of selected DEGs in the GSE122063 dataset. (E) Venn diagram of up-regulated DEGs. (F) Venn diagram of down-regulated DEGs.
Following that, 90 DEGs that exhibited strong associations with the disulfidptosis-related genes were screened by Spearman correlation analysis. The locations of partial genes on the chromosome and the correlations between them are displayed in Supplementary Fig. 1.
Functional enrichment analysis of the disulfidptosis-related DEGs in AD
To explore the potential functional mechanisms of 90 DEGs associated with disulfidptosis, enrichment analyses were conducted, including GO analysis at the BP, MF, and CC levels, as well as KEGG enrichment analysis.
The KEGG enrichment results are presented in Fig. 3A, revealing that these genes are primarily enriched in pathways such as "Alanine, aspartate, and glutamate metabolism," "GABAergic synapse," "Neuroactive ligand-receptor interaction," "Retrograde endocannabinoid signaling," and "Taurine and hypotaurine metabolism." These pathways play crucial regulatory roles in the nervous system, underscoring their potential significance in the processes related to AD.
Enrichment analysis of 90 DEGs associated with disulfidptosis. (A) Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analysis results show enriched items. (B) Gene Ontology (GO) enrichment analysis results at Biological Process (BP), Cellular Component (CC), and Molecular Function (MF) levels.
The GO enrichment results, depicted in Fig. 3B, highlight the top 6 enriched pathways across different levels. These pathways include "response to nerve growth factor," "neuropeptide signaling pathway," "neuronal cell body," and "neuropeptide receptor binding," among others. They encompass various aspects of neural signaling and metabolism, suggesting that their dysregulation may contribute to the onset and progression of neurological diseases, holding potential promise for the study of AD.
Identifying hub genes using a randomforest model
To identify the hub DEGs associated with disulfidptosis, a randomforest model was constructed, and the feature importance parameters provided by the model were utilized. Figure 4A displays the top 20 genes identified by the model. Ultimately, the top 5 genes were selected as hub genes: PPEF1, NEUROD6, VIP, NUPR1, and GEM. Additionally, the correlation patterns of these top 20 genes were investigated, revealing significant correlations between these five genes and other regulatory factors (Fig. 4B). Among these hub genes, PPEF1, NEUROD6, and VIP exhibited low expression levels in AD patients, whereas NUPR1 and GEM displayed high expression levels, as depicted in Fig. 4C.
Identification and analysis of hub genes associated with disulfidptosis. (A) Feature importance ranking in the randomforest model. (B) Correlation analysis of the top 20 ranked disulfidptosis-related DEGs. (C) Violin plot of the top 20 ranked disulfidptosis-related DEGs expressions (ns: nondifferential; *, **, ***, and **** indicates p < 0.05, < 0.01, < 0.001, and < 0.0001, respectively).
Functional annotation and enrichment analysis of hub genes
To further explore the potential functional mechanism of hub genes, the GSEA was performed. The results, depicted in Supplementary Fig. 2A–E, highlight several pathways associated with neurological diseases, including: "Alzheimer's disease," "PI3K-Akt signaling pathway," "JAK-STAT signaling pathway," "GABAergic synapse," "Retrograde endocannabinoid signaling," "NF-kappa B signaling pathway," "Notch signaling pathway," "Synaptic vesicle cycle," and "Pathways of neurodegeneration-multiple diseases." The results provide valuable insights into the potential involvement of these pathways in AD, offering potential targets for further investigation and therapeutic interventions.
Immune infiltration
To investigate potential differences in the immune system between the AD group and the non-AD controls, an immune infiltration analysis using the CIBERSORT algorithm was conducted. The proportions of 22 immune cells in the sample are depicted in Fig. 5A. The results in Fig. 5B reveal significant elevations in B cells naive, B cells memory, T cells CD4 memory resting, T cells gamma delta, NK cells resting, Monocytes, Macrophages M0, Macrophages M1, Macrophages M2, Dendritic cells activated and Neutrophils in AD patients. Conversely, AD patients exhibited significant reductions in Plasma cells, T cells CD8, T cells CD4 naive, T cells CD4 memory activated, T cells follicular helper, T cells regulatory (Tregs), NK cells activated, Dendritic cells resting, and Mast cells resting.
Results of immune infiltration analysis. (A) Percentage of 22 immune cells in AD and normal samples. (B) Boxplots illustrating the differences in immune infiltration between AD and Control groups. (C) Heatmap of the correlation between hub genes (GEM, NEUROD6, NUPR1, PPEF1, and VIP) and immune infiltrating cells (ns: nondifferential; *, **, ***, and **** indicates p < 0.05, < 0.01, < 0.001, and < 0.0001, respectively).
Furthermore, a correlation analysis between the hub genes and immune infiltrating cells was performed. The results showed significant correlations between the hub genes and Plasma cells, T cells CD8, T cells CD4 memory resting, NK cells activated, Monocytes, Macrophages M2, and Neutrophils (Fig. 5C).
Construction and validation of the predictive model
The ability of the five hub genes (PPEF1, NEUROD6, VIP, NUPR1, and GEM) to distinguish between AD and non-AD cases in the GSE33000 dataset was evaluated. The results demonstrated that the AUCs of the five hub genes on the GSE33000 dataset were all above 0.9, as shown in Fig. 6A. A logistic regression prediction model was constructed using the GSE33000 dataset, which exhibited strong discriminatory power, with an AUC of 0.952, as depicted in Fig. 6B. The model was further validated on the GSE122063 and GSE5281 datasets, yielding AUCs of 0.916 (Fig. 6C) and 0.864 (Fig. 6D). The logistic regression formula used for the prediction model is as follows: logit(p) = 0.180 − 0.964 × PPEF1 − 0.487 × NEUROD6 − 0.570 × VIP + 0.040 × NUPR1 + 1.074 × GEM.
Drug prediction
To investigate potential drugs targeting these hub genes, the GAT_GCN model was used for drug-target affinity prediction. The results, presented in Supplementary Table 4, show the top three drug compounds ranked by their affinity scores. To gain further insight into these findings, the top 50 drugs targeting each gene were selected to construct a network (Fig. 7A). The network depicted genes as substantial nodes and drugs as diminutive nodes, with the thickness of the connecting lines representing the level of affinity between them. Subsequently, a total of nine drugs that co-targeted these genes were identified (Fig. 7B). To further verify the binding capabilities of these drug-target pairs, molecular docking was conducted. The results revealed that all the binding affinities were less than − 5.0 kcal/mol, which showed strong interaction (Supplementary Table 5). The conformation of the core drug-target is depicted in Fig. 8A–E. Specifically, NEUROD6, VIP, and NUPR1 exhibited good binding affinity with Hypericin, PPEF1 showed a favorable binding affinity with Emodin, and GEM displayed a strong binding affinity with Rolitetracycline.
Discussion
The initial symptoms of Alzheimer's disease are mild and resemble normal age-related decline, making early diagnosis and identification notoriously challenging. Despite the growing knowledge and understanding of AD in recent years, the complex pathogenesis of the disease has hindered significant breakthroughs. Neurodegenerative diseases are characterized by progressive deterioration of neuronal function and structure, primarily attributed to the degeneration of synapses and axons, ultimately leading to neuronal cell death37. Therefore, it is crucial to identify the specific cell death mechanisms and signaling pathways affected in AD. Disulfidptosis, a novel type of cell death induced by intracellular disulfide accumulation due to SLC7A11 overexpression, has been identified.
In this study, microarray data from the brain tissues of patients with AD and healthy controls were utilized. Five hub DEGs associated with disulfidptosis, namely PPEF1, NEUROD6, VIP, NUPR1, and GEM, were identified by a randomforest model. The PPEF1 gene encodes a member of the serine/threonine protein phosphatase with the EF-hand motif family. It is believed to play a role in specific sensory neuron functions and development38. It has been shown that serine/threonine-specific protein phosphatase affects the function of plasma membrane ion channels in excitable tissues39. Additionally, aberrant phosphorylation of tau proteins, which is linked to the pathogenesis of AD, may be influenced by PPEF140. The role of the NEUROD6 gene in AD is well-established. It encodes a protein associated with the development and differentiation of the nervous system and has been shown to play a crucial role in sustaining the mitochondrial biomass and responding to oxidative stress41, both of which are implicated in the pathogenesis of AD42. Bioinformatics studies have demonstrated significantly reduced expression of NEUROD6 in Alzheimer's patients compared to normal subjects, suggesting its potential as a biomarker43,44. Vasoactive intestinal peptide (VIP), a neuropeptide that acts as a neuromodulator and neurotransmitter, with functions in vasodilation, smooth muscle relaxation, and immunomodulatory45,46, plays an essential role in various physiological activities. There is growing evidence linking VIP to the nervous system47,48,49. The neuropeptide exerts an effect on cAMP synthesis in the central nervous system50, and its variants have been associated with psychiatric disorders51. VIP-containing interneurons have been implicated in the pathology and treatment of neurological disorders, such as Alzheimer's disease49, Parkinson's disease52, and autism spectrum disorders53, among others. NUPR1 is a transcriptional regulator involved in various processes, including cell cycle regulation and apoptosis. It has been shown to play an important role in the progression of malignant tumors such as breast and ovarian cancers54. Relevant studies have also shown the involvement of NUPR1 in METH-induced neuronal apoptosis and autophagy55. GEM is a small GTP-binding protein in the Ras superfamily, and some studies have shown its role in neuronal morphological differentiation56. Less attention has been paid to the relevance of PPEF1, NUPR1, and GEM to AD as possible therapeutic targets, and further research is needed to determine their potential roles in the treatment.
To further explore the potential functions of the identified hub genes, the GSEA was performed to predict their associated signaling pathways. It has been suggested that altered signaling of 2-Arachidonoyl glycerol, an endocannabinoid, may contribute to synaptic silencing in AD57. Alterations in GABAergic circuits may also promote AD by disrupting overall neuronal network function58. Increased inflammatory signaling leads to upregulation of the transcription factor NF-kappa B, which plays a crucial role in AD pathogenesis by regulating the different disease molecules responsible for the promotion of AD59. These suggest that the identified hub genes could potentially serve as markers for therapeutic interventions in AD.
Our studies on immune infiltration analysis indicated that the activity of multiple immune cell types undergoes alterations during the onset and progression of AD. Increased levels of B cells and T cells align with previous findings suggesting that resident cells in the AD brain produce cytokines, reactive oxygen species (ROS), and inducible nitric oxide synthase (iNOS), thereby inducing a parenchymal neuroinflammatory response that leads to the infiltration of T cells into the brain60. Moreover, the upregulation of macrophages61 and neutrophils62 may also contribute to the neuroinflammatory response and neuronal damage process in AD, where due to the dysregulation of the brain microenvironment, microglial cells lose their functionality and release pro-inflammatory factors63, triggering neuroinflammation64, which influences the progression of AD.
Moreover, a disease prediction model constructed using these five hub genes based on logistic regression exhibited excellent performance on the test set (AUC = 0.952) and accurately predicted AD in two additional datasets (AUCs of 0.916 and 0.864, respectively), highlighting the potential value of these hub genes.
Finally, the drugs targeting the identified hub genes were predicted using the GAT_GCN model, and their binding affinity was verified through molecular docking. The results of molecular docking revealed that Hypericin, Emodin, and Rolitetracycline exhibited the strongest affinity for their respective targets among the tested drugs. Hypericin, a natural compound in Hypericum perforatum, possesses antitumor, antiviral, and antidepressant activities and induces apoptosis65. It has been shown to inhibit inflammatory responses induced by oligomeric amyloid β42 in microglia66 and is considered a potent anti-AD component. Emodin, an anthraquinone derivative, possesses antibacterial and anti-inflammatory properties67. Additionally, it exhibits potential antiviral activity68. A study has shown that it exerts neuroprotection against Alzheimer's disease through Nrf2 signaling in U251 cells and APP/PS1 mice69. Its ability to inhibit aggregation of amyloid-β peptide 1–42 makes it a promising candidate for AD treatment70. Rolitetracycline, a broad-spectrum tetracycline antibiotic, has been demonstrated to inhibit the formation of Aβ protofibrils71, thereby reducing the deposition of beta-amyloid peptide, which is one of the main pathological features of AD. These drugs hold promise as potential therapeutic agents for AD. However, it is important to note that further research and clinical trials are necessary to fully evaluate their safety and efficacy in treating the disease.
Conclusion
In conclusion, the genes associated with disulfidptosis were studied using bioinformatics, and the biological functions of these genes were explored. Potential biomarkers were identified in the study, and drugs targeting these biomarkers were predicted, shedding light on novel avenues for the treatment of Alzheimer's disease. Furthermore, the association between disulfidptosis and AD may provide valuable insights for the exploration of new therapeutic targets, opening up possibilities for innovative treatment strategies to be developed.
Data availability
The microarray data are sourced from the GEO database (https://www.ncbi.nlm.nih.gov/geo/). The drug compounds are derived from the DrugBank database (https://go.drugbank.com/).
References
De-Paula, V. J., Radanovic, M., Diniz, B. S. & Forlenza, O. V. Alzheimer’s disease. Subcell. Biochem. 65, 329–352 (2012).
Ju, Y. & Tam, K. Y. Pathological mechanisms and therapeutic strategies for Alzheimer’s disease. Neural Regen. Res. 17, 543–549 (2022).
Lane, C. A., Hardy, J. & Schott, J. M. Alzheimer’s disease. Eur. J. Neurol. 25, 59–70 (2018).
Gauthier, S. et al. World Alzheimer Report 2022. Alzheimer’s disease International https://www.alzint.org/resource/world-alzheimer-report-2022 (2022).
2023 Alzheimer’s disease facts and figures. Alzheimers Dement. J. Alzheimers Assoc. 19, 1598–1695 (2023).
Hardy, J. A. & Higgins, G. A. Alzheimer’s disease: The amyloid cascade hypothesis. Science 256, 184–185 (1992).
Muralidar, S., Ambi, S. V., Sekaran, S., Thirumalai, D. & Palaniappan, B. Role of tau protein in Alzheimer’s disease: The prime pathological player. Int. J. Biol. Macromol. 163, 1599–1617 (2020).
Lin, M. T. & Beal, M. F. Mitochondrial dysfunction and oxidative stress in neurodegenerative diseases. Nature 443, 787–795 (2006).
Chen, Z. & Zhong, C. Oxidative stress in Alzheimer’s disease. Neurosci. Bull. 30, 271–281 (2014).
Preeti, K., Sood, A. & Fernandes, V. Metabolic regulation of glia and their neuroinflammatory role in Alzheimer’s disease. Cell. Mol. Neurobiol. 42, 2527–2551 (2022).
Neurology, T. L. Treatment for Alzheimer’s disease: Time to get ready. Lancet Neurol. 22, 455 (2023).
Tian, S. et al. Exploring pharmacological active ingredients of traditional Chinese medicine by pharmacotranscriptomic map in ITCM. Brief. Bioinform. 24, bbad027 (2023).
Tian, S. et al. COIMMR: A computational framework to reveal the contribution of herbal ingredients against human cancer via immune microenvironment and metabolic reprogramming. Brief. Bioinform. 24, bbad346 (2023).
Zheng, T., Liu, Q., Xing, F., Zeng, C. & Wang, W. Disulfidptosis: A new form of programmed cell death. J. Exp. Clin. Cancer Res. CR 42, 137 (2023).
Zhao, S. et al. Crosstalk of disulfidptosis-related subtypes, establishment of a prognostic signature and immune infiltration characteristics in bladder cancer based on a machine learning survival framework. Front. Endocrinol. 14, 1180404 (2023).
Wang, Y., Deng, Y., Xie, H. & Cao, S. Hub gene of disulfidptosis-related immune checkpoints in breast cancer. Med. Oncol. Northwood Lond. Engl. 40, 222 (2023).
Wang, T. et al. Disulfidptosis classification of hepatocellular carcinoma reveals correlation with clinical prognosis and immune profile. Int. Immunopharmacol. 120, 110368 (2023).
Zhang, L. et al. Development of tumor-evolution-targeted anticancer therapeutic nanomedicineEVT. Chem 10, 1337–1356 (2024).
Ma, S., Wang, D. & Xie, D. Identification of disulfidptosis-related genes and subgroups in Alzheimer’s disease. Front. Aging Neurosci. 15, 1236490 (2023).
Davis, S. & Meltzer, P. S. GEOquery: A bridge between the Gene Expression Omnibus (GEO) and BioConductor. Bioinforma. Oxf. Engl. 23, 1846–1847 (2007).
Narayanan, M. et al. Common dysregulation network in the human prefrontal cortex underlies two neurodegenerative diseases. Mol. Syst. Biol. 10, 743 (2014).
McKay, E. C. et al. Peri-infarct upregulation of the oxytocin receptor in vascular dementia. J. Neuropathol. Exp. Neurol. 78, 436–452 (2019).
Liang, W. S. et al. Gene expression profiles in anatomically and functionally distinct regions of the normal aged human brain. Physiol. Genom. 28, 311–322 (2007).
Liang, G. et al. Nanomedomics. ACS Nano 18, 10979–11024 (2024).
Liu, X. et al. Actin cytoskeleton vulnerability to disulfide stress mediates disulfidptosis. Nat. Cell Biol. 25, 404–414 (2023).
Ritchie, M. E. et al. Limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Altmann, A., Toloşi, L., Sander, O. & Lengauer, T. Permutation importance: A corrected feature importance measure. Bioinforma. Oxf. Engl. 26, 1340–1347 (2010).
Wu, T. et al. clusterProfiler 4.0: A universal enrichment tool for interpreting omics data. Innov. Camb. Mass 2, 100141 (2021).
Schober, P. & Vetter, T. R. Logistic regression in medical research. Anesth. Analg. 132, 365–366 (2021).
Robin, X. et al. pROC: An open-source package for R and S+ to analyze and compare ROC curves. BMC Bioinform. 12, 77 (2011).
Chen, B., Khodadoust, M. S., Liu, C. L., Newman, A. M. & Alizadeh, A. A. Profiling tumor infiltrating immune cells with CIBERSORT. Methods Mol. Biol. Clifton NJ 1711, 243–259 (2018).
Nguyen, T. et al. GraphDTA: Predicting drug-target binding affinity with graph neural networks. Bioinforma. Oxf. Engl. 37, 1140–1147 (2021).
Wishart, D. S. et al. DrugBank 5.0: A major update to the DrugBank database for 2018. Nucleic Acids Res. 46, D1074–D1082 (2018).
UniProt Consortium. UniProt: The universal protein knowledgebase in 2023. Nucleic Acids Res. 51, D523–D531 (2023).
Liu, Y. et al. CB-Dock2: Improved protein–ligand blind docking by integrating cavity detection, docking and homologous template fitting. Nucleic Acids Res. 50, W159–W164 (2022).
Trott, O. & Olson, A. J. AutoDock Vina: Improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 31, 455–461 (2010).
Andreone, B. J., Larhammar, M. & Lewcock, J. W. Cell death and neurodegeneration. Cold Spring Harb. Perspect. Biol. 12, a036434 (2020).
Montini, E. et al. A novel human serine-threonine phosphatase related to the Drosophila retinal degeneration C (rdgC) gene is selectively expressed in sensory neurons of neural crest origin. Hum. Mol. Genet. 6, 1137–1145 (1997).
Herzig, S. & Neumann, J. Effects of serine/threonine protein phosphatases on ion channels in excitable membranes. Physiol. Rev. 80, 173–210 (2000).
Srivastava, S., Ahmad, R. & Khare, S. K. Alzheimer’s disease and its treatment by different approaches: A review. Eur. J. Med. Chem. 216, 113320 (2021).
Uittenbogaard, M., Baxter, K. K. & Chiaramello, A. The neurogenic basic helix–loop–helix transcription factor NeuroD6 confers tolerance to oxidative stress by triggering an antioxidant response and sustaining the mitochondrial biomass. ASN Neuro 2, e00034 (2010).
Wallace, D. C. A mitochondrial paradigm of metabolic and degenerative diseases, aging, and cancer: A dawn for evolutionary medicine. Annu. Rev. Genet. 39, 359–407 (2005).
Satoh, J.-I., Yamamoto, Y., Asahina, N., Kitano, S. & Kino, Y. RNA-Seq data mining: Downregulation of NeuroD6 serves as a possible biomarker for Alzheimer’s disease brains. Dis. Markers 2014, 123165 (2014).
Fowler, K. D. et al. Leveraging existing data sets to generate new insights into Alzheimer’s disease biology in specific patient subsets. Sci. Rep. 5, 14324 (2015).
Wang, C. et al. Effect of vasoactive intestinal peptide (VIP) on NKG2D signal pathway and its contribution to immune escape of MKN45 cells. Sci. World J. 2013, 429545 (2013).
Villanueva-Romero, R. et al. Human CD4+CD45RA+ T cells behavior after in vitro activation: Modulatory role of vasoactive intestinal peptide. Int. J. Mol. Sci. 23, 2346 (2022).
Brenneman, D. E. et al. Vasoactive intestinal peptide. Link between electrical activity and glia-mediated neurotrophism. Ann. N. Y. Acad. Sci. 897, 17–26 (1999).
White, C. M., Ji, S., Cai, H., Maudsley, S. & Martin, B. Therapeutic potential of vasoactive intestinal peptide and its receptors in neurological disorders. CNS Neurol. Disord. Drug Targets 9, 661–666 (2010).
Deng, G. & Jin, L. The effects of vasoactive intestinal peptide in neurodegenerative disorders. Neurol. Res. 39, 65–72 (2017).
Nowak, J. Z., Jozwiak-Bebenista, M. & Bednarek, K. Effects of PACAP and VIP on cyclic AMP formation in rat neuronal and astrocyte cultures under normoxic and hypoxic condition. Peptides 28, 1706–1712 (2007).
Saus, E. et al. Comprehensive copy number variant (CNV) analysis of neuronal pathways genes in psychiatric disorders identifies rare variants within patients. J. Psychiatr. Res. 44, 971–978 (2010).
Gonzalez-Rey, E., Chorny, A., Fernandez-Martin, A., Varela, N. & Delgado, M. Vasoactive intestinal peptide family as a therapeutic target for Parkinson’s disease. Expert Opin. Ther. Targets 9, 923–929 (2005).
Goff, K. M. & Goldberg, E. M. A role for vasoactive intestinal peptide interneurons in neurodevelopmental disorders. Dev. Neurosci. 43, 168–180 (2021).
Chowdhury, U. R., Samant, R. S., Fodstad, O. & Shevde, L. A. Emerging role of nuclear protein 1 (NUPR1) in cancer biology. Cancer Metastasis Rev. 28, 225–232 (2009).
Xu, X. et al. Nupr1 modulates methamphetamine-induced dopaminergic neuronal apoptosis and autophagy through CHOP-Trib3-mediated endoplasmic reticulum stress signaling pathway. Front. Mol. Neurosci. 10, 203 (2017).
Leone, A. et al. The Gem GTP-binding protein promotes morphological differentiation in neuroblastoma. Oncogene 20, 3217–3225 (2001).
Mulder, J. et al. Molecular reorganization of endocannabinoid signalling in Alzheimer’s disease. Brain 134, 1041–1060 (2011).
Li, Y. et al. Implications of GABAergic neurotransmission in Alzheimer’s disease. Front. Aging Neurosci. 8, 31 (2016).
Sun, E., Motolani, A., Campos, L. & Lu, T. The pivotal role of NF-kB in the pathogenesis and therapeutics of Alzheimer’s disease. Int. J. Mol. Sci. 23, 8972 (2022).
Chen, X. & Holtzman, D. M. Emerging roles of innate and adaptive immunity in Alzheimer’s disease. Immunity 55, 2236–2254 (2022).
Silvin, A. et al. Dual ontogeny of disease-associated microglia and disease inflammatory macrophages in aging and neurodegeneration. Immunity 55, 1448-1465.e6 (2022).
Zenaro, E. et al. Neutrophils promote Alzheimer’s disease-like pathology and cognitive decline via LFA-1 integrin. Nat. Med. 21, 880–886 (2015).
Liu, P. et al. Biomimetic dendrimer-peptide conjugates for early multi-target therapy of Alzheimer’s disease by inflammatory microenvironment modulation. Adv. Mater. 33, 2100746 (2021).
Thawkar, B. S. & Kaur, G. Inhibitors of NF-κB and P2X7/NLRP3/Caspase 1 pathway in microglia: Novel therapeutic opportunities in neuroinflammation induced early-stage Alzheimer’s disease. J. Neuroimmunol. 326, 62–74 (2019).
Kubin, A., Wierrani, F., Burner, U., Alth, G. & Grünberger, W. Hypericin—The facts about a controversial agent. Curr. Pharm. Des. 11, 233–253 (2005).
Zhang, M., Wang, Y., Qian, F., Li, P. & Xu, X. Hypericin inhibits oligomeric amyloid β42-induced inflammation response in microglia and ameliorates cognitive deficits in an amyloid β injection mouse model of Alzheimer’s disease by suppressing MKL1. Biochem. Biophys. Res. Commun. 481, 71–76 (2016).
Dong, X. et al. Emodin: A review of its pharmacology, toxicity and pharmacokinetics. Phytother. Res. 30, 1207–1218 (2016).
Ho, T.-Y., Wu, S.-L., Chen, J.-C., Li, C.-C. & Hsiang, C.-Y. Emodin blocks the SARS coronavirus spike protein and angiotensin-converting enzyme 2 interaction. Antiviral Res. 74, 92–101 (2007).
Li, Z. et al. Neuroprotective effect of emodin against Alzheimer’s disease via Nrf2 signaling in U251 cells and APP/PS1 mice. Mol. Med. Rep. 23, 108 (2021).
Wang, L. et al. Emodin inhibits aggregation of amyloid-β peptide 1–42 and improves cognitive deficits in Alzheimer’s disease transgenic mice. J. Neurochem. 157, 1992–2007 (2021).
Howlett, D. R., George, A. R., Owen, D. E., Ward, R. V. & Markwell, R. E. Common structural features determine the effectiveness of carvedilol, daunomycin and rolitetracycline as inhibitors of Alzheimer β-amyloid fibril formation. Biochem. J. 343, 419–423 (1999).
Funding
This work was supported by National Natural Science Foundation of China (82174531), the Fundamental Research Funds for the Central Universities (BH202439), National High Level Hospital Clinical Research Funding (XK2023-13) and Scientific and Technological Research Project of Xinjiang Production and Construction Corps (Grant number 2022AB022).
Author information
Authors and Affiliations
Contributions
CY Yu and YJ Li designed the concept of this manuscript. L Huang and ZT Li contributed to data analysis and drafted the manuscript. XY Zhang and YF Li supervised the research and revised the manuscript. YT Lv participated in molecular docking. All the authors approved the final version. All authors have given consent for publication.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Huang, L., Li, Z., Lv, Y. et al. Unveiling disulfidptosis-related biomarkers and predicting drugs in Alzheimer’s disease. Sci Rep 14, 20185 (2024). https://doi.org/10.1038/s41598-024-70893-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-70893-7
Keywords
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.