Abstract
Cockayne syndrome (CS) is a rare autosomal recessive disorder caused by mutations in ERCC8 or ERCC6. Most pathogenic variants in ERCC8 are single nucleotide substitutions. Structural variants (SVs) have been reported in patients with ERCC8-related CS. However, comprehensive molecular detection, including SVs of ERCC8, in CS patients remains problematic. Herein, we present three Japanese patients with ERCC8-related CS in whom causative SVs were identified using whole-exome-based copy number variation (CNV) detection tools. One patient showed compound heterozygosity for a 259-kb deletion and a deletion of exon 4 which has previously been reported as an Asia-specific variant. The other two patients were homozygous for the same exon 4 deletion. The exon 4 deletion was detected only by the ExomeDepth software. Intrigued by the discrepancy in the detection capability of various tools for the SVs, we evaluated the analytic performance of four whole-exome-based CNV detection tools using an exome data set from 337 healthy individuals. A total of 1,278,141 exons were predicted as being affected by the 4 CNV tools. Interestingly 95.1% of these affected exons were detected by one tool alone. Thus, we expect that the use of multiple tools may improve the detection rate of SVs from aligned exome data.
Similar content being viewed by others
Introduction
Cockayne syndrome (CS), first described in 19361, is a rare autosomal recessive (AR) disorder characterized by failure to thrive, developmental delay, and microcephaly, along with at least three of the following: cutaneous photosensitivity, pigmentary retinopathy/cataracts, sensorineural hearing loss, dental caries, and cachectic dwarfism. These symptoms result from pathogenic variants in either the Excision Repair Cross Complementing, Group 8 (ERCC8) or Group 6 (ERCC6) gene2,3, which encode proteins for DNA damage repair4. Most ERCC8 pathogenic variants reported until date are single-nucleotide variants (SNVs) or small insertion/deletions2. However, in rare cases, structural variants (SVs) have also been reported. A comprehensive molecular detection tool that can detect both types of variants (i.e., SNVs and SVs) is yet to be identified.
Various approaches have been utilized to detect SVs, including fluorescence in situ hybridization (FISH), array-based comparative genomic hybridization (aCGH)5, and multiplex ligation-dependent probe amplification (MLPA)6. Each approach has its own advantages and disadvantages related to the resolution and comprehensiveness of the genome coverage. For example, aCGH can screen the entire genome in a single assay, whereas FISH and MLPA can screen only the targeted regions. Recently, several tools have been developed to detect copy number variations (CNVs) from aligned whole-exome sequencing (WES) data, including ExomeDepth7, XHMM8, CODEX29, and EXCAVATOR210. Despite a number of studies having been conducted to compare the analytic performances of the CNV detection tools11,12,13,14,15,16,17, no “best practice” (gold standard) has been established until date.
Herein, we report three Japanese patients with CS due to bi-allelic SVs in ERCC8. In one patient, two causative SVs were detected using four WES-based CNV detection tools. This patient showed compound heterozygosity for a 259-kb deletion and deletion of exon 4. Two patients were homozygous for the same deletion of exon 4. The deletion of exon 4 was detected only by the ExomeDepth software. Intrigued by the discrepancy in the detection capability of various tools for the two variants, we conducted this study to evaluate the analytic performance of these WES-based CNV detection tools using an exome data set of 337 phenotypically normal parents of patients with neurodevelopmental disorders.
Results
We have been performing trio-WES on patients with rare and undiagnosed diseases18. Three patients were suspected as having a clinical diagnosis of CS, and SNVs and small indels were screened using WES. No such variants were found in the ERCC8 gene or ERCC6 gene in the three patients.
Patient 1—clinical summary
The patient was a 12-year-old Japanese boy with suspected CS. He was born at 39 weeks 5 days of gestation, with a birth weight of 2870 g (− 0.92 SD), body length of 48.4 cm (− 0.46 SD), and occipitofrontal circumference (OFC) of 33.0 cm (− 0.25 SD). He was the second child of unrelated healthy Japanese parents.
He showed failure to thrive and developmental delay. His neck became stable at 3 months, he could sit without support at 6 months, and he could walk by himself at 18 months. He uttered his first meaningful words at 2 years 6 months of age. Thereafter, he showed regression: he was unable to walk by himself and could utter no meaningful words at 5 years of age.
Physical examination at 5 years of age showed microcephaly, sunken eyes, pigmentary retinopathy, sensorineural hearing loss, and joint contractures. Computed tomography (CT) of the head showed cerebellar degeneration and calcifications in the basal ganglia. Based on the above clinical features, the patient was diagnosed as having CS.
Patient 1—genetic analysis
Analysis based on WES data of the patient and his parents revealed no pathogenic SNVs and no short indels in the ERCC6 and ERCC8 gene; no candidate variants in other genes that could account for his phenotype could be identified either. Therefore, to identify pathogenic SVs, we performed CNV analysis using EXCAVATOR2, one of the CNV detection tools19. A 259-kb deletion (chr5:59,945,691_60,204,587) (GRCh37/hg19) partially overlapping the ERCC8 locus was detected in the patient and his mother by this tool (Fig. 1a, Supplementary Fig. 1, and Supplementary Table 1). Two flanking genes, ELOVL7 and DEPDC1B, were also deleted in addition to ERCC8. Junctional reads spanning the deletion was identified in the bam data and the both ends of the junction were confirmed by Sanger sequencing (Fig. 1a) using forward and reverse primers: 5-ACTGGCCAAGAGAACAAACCA-3 and 5-CCTTCCTTTGGCTTCACATTG-3, respectively.
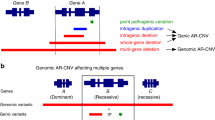
Identified compound-heterozygous SV in ERCC8. (a) The deletion spanning ERCC8 was detected in the patient and his mother by EXCAVATOR2. Sequencing analysis revealed a 259-kb deletion in chromosome 5 spanning intron 2 of DEPDC1B and intron 5 of ERCC8. (b) Schematic diagram of the exon 4 rearrangement. Note that the black dotted lines show two deletions consisting of a 3368-bp region in exon 4 (c.275+768 to c.399+346) and a 555-bp region in intron 4 (c.399+2003 to c.399+2557). The orange arrow indicates the 1656-bp inversion region in intron 4 (c.399+347 to c.399+2002) and the green bar shows the 8-bp insertion at the 5′ end of breakpoint. The figure below shows the results of validation by Sanger sequencing of SV breakpoints using primers at the positions shown in the schematic diagram. (c) Pedigree tree. The father and mother each had an identical complex SV of exon 4 rearrangement, and a large deletion, respectively.
In that CS is an AR disorder, the patient was inferred to have another pathogenic allele, other than the 259-kb deletion. In search of the second pathogenic variant in ERCC8, we applied the other three CNV detection tools (i.e., ExomeDepth, XHMM, and CODEX2). ExomeDepth detected a single exonic deletion in exon 4 of ERCC8 in both the patient and his father. This deletion was not detected by XHMM, CODEX2 or EXCAVATOR2 (Supplementary Fig. 2).
Detailed sequencing analysis of the deleted region showed that the deletion included an inversion of 1656 bp (chr5:60,212,090_60,213,745inv) from the deleted intron 4 segment and an insertion of 8 bp (chr5:60,211,535_60,212,089delinsATTAAGTA). This complex SV of the exon 4 rearrangement included a 3368-bp deletion of the exon 4 containing region from c.275+768 to c.399+346 (NM_000082.4), a 1656-bp inversion in intron 4 from c.399+347 to c.399+2002 with a 555-bp deletion from c.399+2003 to c.399+2557, and an 8-bp insertion (Fig. 1b). This complex SV of the ERCC8 exon 4 rearrangement has been reported multiple times, mainly in the Asian population (Table 1). The patient and his father were confirmed by Sanger sequencing to harbor this complex SV using forward and reverse primers: 5-CACCTAGGCCTGGAGATGTCA-3 and 5-TACCCACTCACCCCTGTCTTG-3, respectively.
Hence, the patient showed compound heterozygosity for two pathogenic SVs–a large deletion (259 kb) partially overlapping the ERCC8 locus, and a complex SV of the exon 4 rearrangement in ERCC8 (Fig. 1c).
Patient 2—clinical summary
The patient was a 4-year-old female, the first child of non-consanguineous Japanese parents. She was born at 37 weeks of gestation, with a birth weight of 2632 g (0.3 SD), body length of 46 cm (− 0.6 SD), and OFC of 35 cm (1.7 SD).
After birth, she often vomited, and her postnatal growth was delayed. Her motor development was delayed from infancy. Her neck became stable at 2 months, she sat without support at 11 months, and she could walk by herself at 2 years of age. She spoke no meaningful words and had severe intellectual disability (ID). There was spasticity of the lower extremities appeared. Later, she showed regression–she was unable to walk by herself at 10 years of age–and her motor ability gradually declined.
On physical examination at 4 years 11 months of age, her height was 92.5 cm (− 3.2 SD), body weight was 10.5 kg (− 4.8 SD), and head circumference was 44.0 cm (− 5.0 SD). Subcutaneous fat loss, microcephaly, and sunken eyes were noted. Ophthalmologic examination revealed optic atrophy and pigmentary degeneration of the retina. She also suffered from photosensitivity. Routine laboratory investigations revealed elevated serum levels of the hepatic transaminases. Magnetic Resonance Imaging (MRI) and CT showed diffuse cerebral and cerebellar atrophy, demyelination, volume loss of the brain stem, and calcification of the bilateral basal ganglia. Based on the above clinical features, she was diagnosed as having CS.
Patient 2—genetic analysis
Karyotyping-banded chromosome analysis and array-CGH revealed no abnormalities in this patient. As mentioned above, WES analysis of the patient and her parents revealed no pathogenic SNVs and no short indels in the ERCC6 or ERCC8 gene, and there were no candidate variants in other genes that could account for her phenotype. To search for pathogenic SVs, we performed CNV analysis using EXCAVATOR2, ExomeDepth, XHMM, and CODEX2.
Among the four tools, only ExomeDepth detected the homozygous complex SV of the exon 4 rearrangement (Supplementary Fig. 2), which was the same as the complex SV of the exon 4 rearrangement detected in Patient 1.
Patient 3—clinical summary
The patient was a 17-year-old male, the first child of Japanese non-consanguineous parents. He was born at 37 weeks of gestation, with a birth weight of 2,360 g (− 0.9 SD), body length of 44 cm (− 1.5 SD), and OFC of 30.7 cm (− 1.4 SD).
He showed failure to thrive. Motor developmental delay had been observed from infancy. His neck became stable at 3 months, and he was able to sit without support at 12 months. He could crawl at 18 months. He could not walk independently. He spoke no meaningful words, and severe ID was noted. There was spasticity of the lower extremities. Subsequently, he showed regression: he was unable to crawl at 15 years of age. His motor ability also gradually declined.
On physical examination at 15 years 11 months of age, his height was 109.0 cm (− 5.3 SD), body weight was 14.1 kg (− 8.0 SD), and head circumference was 48.8 cm (− 3.5 SD). Subcutaneous fat loss, microcephaly, and sunken eyes were noted. Ophthalmologic examination revealed cataract, optic atrophy, and pigmentary degeneration of the retina. He also showed photosensitivity with frequent blister formation. Routine laboratory investigations revealed elevated serum levels of the hepatic transaminases and renal failure. MRI and CT revealed progressive cerebral and cerebellar atrophy, demyelination, volume loss of the brain stem, and calcification of the bilateral basal ganglia. Based on the above clinical features, he was diagnosed as having CS.
Patient 3—genetic analysis
Karyotyping-banded chromosome analysis and array-CGH in this patient revealed no abnormalities, so that WES was carried out. As mentioned above, WES analysis of the patient and his parents revealed no pathogenic SNVs and no short indels in the ERCC6 or ERCC8 gene, and there were no candidate variants in any other genes that could account for his phenotype. To search for pathogenic SVs, we performed CNV analysis using EXCAVATOR2, ExomeDepth, XHMM, and CODEX2.
ExomeDepth identified a homozygous complex SV rearrangement in exon 4, the same SV that was also detected in Patient 1 and Patient 2 (see Supplementary Fig. 2).
Genetic relatedness of Patient 1, Patient 2, and Patient 3
The same complex SV of the exon 4 rearrangement was identified in all three families. The genetic relatedness of the families was inferred from the genotype data. To determine the genetic closeness, the genotype similarities were analyzed (“Materials and methods”), which revealed that the three families were at least five degrees of relationship apart (Supplementary Fig. 3).
Comparing the features of the CNV detection tools using in-house WES data
Examination of the three patients with ERCC8-related CS in this study by the four CNV detection tools revealed observed differences in the analytical characteristics of the CNV detection tools. Of the four tools, only ExomeDepth successfully identified the complex SV of the exon 4 rearrangement in the current study. To further investigate the differences in the analytical characteristics of the four tools, we analyzed the WES data collected from the phenotypically normal parents (337 individuals) of patients with neurodevelopmental disorders.
The average number of CNVs detected by the four CNV detection tools was the highest for ExomeDepth and lowest for CODEX2 (Fig. 2a). The average length of CNVs detected was shortest for ExomeDepth and longest for EXCAVATOR2 (Fig. 2b–e). The number of detected CNVs decreased with increasing CNV size. In summary, ExomeDepth and EXCAVATOR2 detected more CNVs than the other two tools; EXCAVATOR2 detected longer CNVs than the other tools.
Comparison of data obtained using four CNV detection tools. (a) Average numbers of CNVs per individual called by each tool. (b–e) Distribution of CNVs called by each tool in different size ranges. CNVs over 500 kb are aggregated to 500 kb. (f) Venn diagram describing the overlap in CNVs at the exon-level between the four CNV detection tools. The same individual exomes data were used for each algorithm.
We next investigated the degree of overlap of the CNVs detected by the four tools. In our in-house WES data set of 337 samples obtained from phenotypically normal individuals, there were 1,278,141 exons with CNVs detected by one or more of the four CNV detection tools. Of all the exons with CNVs, only 1,369 (0.1%) were detected by all the four tools (Fig. 2f); 15,042 exons (1.2%) and 45,881 exons (3.6%) were found by three and two tools, respectively; 1,215,849 exons with CNVs (95.1%) were detected by only one tool. Of the four tools, EXCAVATOR2 detected the highest number of exons with CNVs that could be detected by one tool alone. There was a slight overlap in the variety of CNVs detected by the four different tools.
Finally, although the accuracy and precision of CNV detection tools have been investigated in various papers11,12,13,14,15,16,17, we conducted a WES of the HG002 (NA24385) sample in our in-house platform and assessed the quality control in the data using a SV benchmark datasets registered with the Genome In A Bottle (GIAB) (see “Materials and methods”)20. Among the 176,564 exons of HG002, ExomeDepth exhibited the highest recall (sensitivity), precision, and F1 score. In contrast, XHMM exhibited the lowest recall value (lower true positive (TP)/false negative (FN) ratio), and CODEX2 exhibited the lowest precision value (lower TP/false positive (FP) ratio) (Supplementary Table 2). In the present study, assessments were carried out with the default parameters, but it should be noted that performance metrics may vary depending on the parameters used. Alternation of the adjustable parameters in XHMM or EXCAVATOR2 did not allow the detection of the ERCC8 exon 4 rearrangement. The exon 4 rearrangement identified in this study was exclusively detected by ExomeDepth as a 123 bp deletion.
Discussion
Herein, we report three patients who had biallelic SVs in the ERCC8 gene: the SV was a large 259-kb large deletion in one patient, and a complex SV of the exon 4 rearrangement in the three patients. These SVs were detected by applying software tools for detection of SVs to aligned exome data. Notably, the deletion of exon 4 was detected only by the ExomeDepth software. Intrigued by the different abilities of different tools to detect SVs in this case series, we evaluated the performance of 4 SV detection tools using a cohort of more than 300 samples and showed relatively limited conformity of the four tools with respect to predicting the CNV. Thus, we suggest that multiple tools used in combination may improve the detection rate of SVs from aligned exome data.
Determining the precise extent of deletion with respect to the neighboring genes is crucial to predicting the clinical impact of SVs. In Patient 1, the 259-kb deletion included ERCC8, ELOVL7, and DEPDC1B; NDUFAF2 was not included in this deletion. Biallelic loss-of-function of NUDFAF2 is known to lead to mitochondrial dysfunction, but Patient 1 is unlikely to develop mitochondrial dysfunction. On the contrary, at least one patient with ERCC8-related CS had a homozygous contiguous deletion of ERCC8 and NDUFAF2, and developed mitochondrial dysfunction21.
The complex SV of the exon 4 rearrangement in Patient 1, Patient 2, and Patient 3 (heterozygosity in Patient 1 and homozygosity in Patient 2 and Patient 3) is identical to that reported in previous patients. A review of the literature showed at least 29 cases in six previous study reports (Table 1)22,23,24,25,26,27. Almost all of the patients were East Asian patients (Chinese, 15; Japanese, 13; Australian, 1). Wang et al.24 reported that the complex SV of the exon 4 rearrangement accounted for 69.2% (18/26 variants) of all ERCC8 alleles in the 13 Chinese patients. Haplotype analysis of the complex SV of the exon 4 rearrangement detected in Patient 1 (heterozygous), Patient 2 (homozygous) and Patient 3 (homozygous) suggested that all of these mutated alleles with exon 4 rearrangement shared the mutation-bearing haplotype and mutation-associated haplotype24. Thus, the similarity of the complex SV of the exon 4 rearrangement would support the notion of a founder effect, as suggested by a previous report. This study confirmed that the three families with the exon 4 rearrangement were at least five degrees of relationship apart.
The choice of the genetic analysis method for the causative genetic variants in CS is crucial, and the focus should be on analyzing rearrangements in exon 4 of the ERCC8 gene. There was a founder effect of exon 4 rearrangement in East Asians, which will guide the development of a practical genetic diagnostic strategy for East Asian patients with CS. Moreover, we mention the issue of analyzing complex SVs of exon 4 rearrangement in CS patients. In our review of previous reports, we found that Cloney et al.27 reported only one case of this complex SV identified by WES, and in most cases, they identified it by exon-targeted qPCR, high-density aCGH, long-range PCR, or MLPA23,24,25,26. The small size of the complex SV of the exon 4 rearrangement often leads to its being overlooked in routine aCGH, due to insufficient probe coverage of the ERCC8 exon regions25. Thus, we believe that the complex SV of the exon 4 rearrangement is difficult to detect by normal aCGH and standard analysis of WES. Especially in suspected cases of CS among Asians, the causative mutation could not always be identified even by WES. In these cases, the exon 4 rearrangement SV might have been missed. Our study suggests that either use of several CNV detection tools in combination or a search for this founder mutation using specific primers by Sanger sequencing would undoubtedly be useful. These approaches are more effective for detecting the founder mutation.
The four CNV detection tools employed in the present study employ distinct strategies. ExomeDepth employs a beta-binomial model of read distribution7. XHMM employs principal component analysis to reduce noise and utilizes hidden Markov models8. CODEX2 employs a log-linear decomposition-based normalization approach9. EXCAVATOR2 predicts CNVs by considering not only the normalized read count of target regions but also that of off-target ones10. In the current study, 337 exome data from phenotypically normal parents of patients with neurodevelopmental disorders were evaluated. We characterized the size distribution of the CNVs detected by each CNV detection tool. Apparently, EXCAVATOR2 detects relatively large CNVs, while ExomeDepth detects relatively small CNVs. The overlap of CNVs detected by four tools were rather than restricted. Hence use of multiple tools and obtain union rather than intersection would be rational.
Effective detection of CNVs from WES data remains a challenge. Several studies have evaluated the recall and precision of various tools for detecting CNVs from exome data11,12,13,14,15,16,17. It has been demonstrated that the F1 scores achieved by each study can vary by several-fold in comparison to studies that utilize standard references11,13,14. The discrepancy in F1 scores is likely attributable to differences in the gold standard CNV data. Given that the majority of tools are capable of identifying a limited number of variations with an acceptable degree of precision, evaluating the union, rather than the intersection, of multiple CNV tools may be expected to increase the detection recall. The use of both the combination of multiple tools and parameter adjustment could further increase sensitivity at the expense of precision, as the detectable CNV features (e.g., length differences) are different with each tool or parameter adjustment. Clinical judgment of phenotype-gene correlation would overcome problems associated with the reduced precision (Fig. 3). In particular, if only a single pathological variant allele has been identified in a patient with suspected AR disorder and its counter-allele remains elusive, the possibility that the other allele may harbor a pathogenic SV should be borne in mind. We recommend the use of multiple tools to facilitate detection of such “unseen” SVs from aligned exome data.
Our study had several limitations. We only used four CNV detection tools, which is not an exhaustive list, and use of other tools could yield different results and observations. Due to the large number of regions and limited amount of DNA, validation of the CNVs by Sanger sequencing was not performed, and the specificities were not examined in this study. As the “true genome sequence” was not available for the 337 samples, it was not possible to evaluate the recall or precision of detection in-house data. Historically, CNVs have been detected by chromosomal microarray analysis in patients with suspected congenital neurodevelopmental disorders. However, it had become clear that chromosomal microarray analysis cannot detect SNVs/small indels. Since CNV detection tools can at least resolve relatively large CNVs, exome-based CNV analysis can be considered as a first-line test when multiple CNV detection tools are used in combination.
In conclusion, we identified the disease-causing SVs in three patients of CS that could not be identified by standard SNV/indel analysis, using several CNV detection tools with WES. We believe that for WES analysis of cases of CS associated with variants of ERCC8, analysis using multiple CNV detection tools in combination, screening for panels of genes associated with a phenotype, and considering regional characteristics would be useful.
Materials and methods
WES analysis
This study was conducted with the approval of the Ethics Committee of Keio University (approval No. 20851) and Tokyo Medical and Dental University (TMDU) (approval No. O2015-502 and O2019-002). Samples were collected from Japanese individuals enrolled in a cohort study of neurodevelopmental disorders and families (patients with congenital dysneuroplasia enrolled in our consortium18). Informed consent was obtained from the patients and their parents for the molecular studies. To determine the molecular etiology, WES was performed. The methods of this study were carried out in accordance with approved guidelines and principles of the Declaration of Helsinki.
Genomic DNA was extracted from a peripheral blood sample using standard DNA extraction kits. A sequencing library was prepared using SureSelectXT Target Enrichment System for Illumina Paired-End Sequencing Library (Agilent). Targeted exonic DNAs were captured using SureSelect XT Human All Exon V4, V5 or V6 (Agilent) and Twist Exome 2.0 plus Comprehensive Exome spike-in (Twist Bioscience) probes, in accordance with the manufacturer’s instructions. WES was performed using the NovaSeq 6000 or HiSeq 4000 platform (Illumina, San Diego, CA). Mapping of the sequence reads to the human reference genome (GRCh37) was performed according to best practice guidelines for the Burrows-Wheeler aligner28 and Genome Analysis Tool Kit (GATK)29, as packaged in the integrated analysis suite variant tools30. The called variants were annotated using Annovar. Candidate pathogenic SNVs and short indels were narrowed down as previously reported18.
To further explore the molecular etiologies, WES-based CNV detection was performed. The aligned exome data in BAM format were used to screen for copy number aberrations using the ExomeDepth, XHMM, CODEX2, and EXCAVATOR2 tools in their default settings. When sequence reads spanning the breakpoints could be recognized by visual inspection of the bam files, the structural variants were further characterized by designing PCR primers and Sanger sequencing.
Analysis of kinship
To examine the blood relationship between two individuals, identity-by-descent (IBD) was estimated by calculating the identity-by-state (IBS) and PI_HAT values, using PLINK (v.1.9). The alleles used in the kinship analysis were SNPs with a minor allele frequency (MAF) of > 5% and Hardy–Weinberg equilibrium (HWE) of > 0.0001 in the Japanese population, based on the jMorp database31. Among the variants called in WES, SNPs with a MAF of > 5% in the Japanese population were analyzed.
Comparison of analytic performance of CNV detection tools
In view of the different performance characteristics of the 4 CNV detection tools, as exemplified in the CNV analysis of Patient 1, we sought to analyze a cohort of 337 Japanese samples (parents of patients with congenital dysneuroplasia collected by our consortium18) for SVs using the following 4 tools: ExomeDepth, XHMM, CODEX2, and EXCAVATOR2.
First, we directly compared the results of these four detection tools in terms of the number (average/sample), length (bp), and distribution of the CNVs detected by each of the tools. Second, we also compared the overlap of CNVs at the exon level among the four detection tools. An overlapping exon was defined as an exon that shared at least 50% of its total length with a CNV region detected by the different tools. Third, an in-house evaluation of the performance of four CNV tools was conducted using the HG002 (NA24385) sample, which is one of the most extensively characterized samples in Genome In A Bottle (GIAB, https://www.nist.gov/programs-projects/genome-bottle) of the National Institute of Standards and Technology (NIST). The genomic DNA of HG002 was procured from the Coriell Institute. A benchmark dataset for SVs of the HG002 samples was registered with the GIAB20. Four tools were run with their default parameters for our WES data of the HG002 sample. The regions selected for evaluation were the SV benchmark region (HG002_SVs_Tier1_noVDJorXorY_v.0.6.2.bed) for HG002 in the GIAB20 and the regions of exons that contain all bases in our WES targeted regions. When more than 50% of the bases on an exon were included in the CNV, the exon was considered to be an exon included in the CNV; other exons were considered to be exons not included in the CNV. The analysis of the overlap between the CNVs and the exons was performed using Bedtools (v.2.31.1, https://github.com/arq5x/bedtools2). A comparative analysis was conducted to evaluate the specificity, recall (sensitivity), precision, and F1 score of four CNV tools using our in-house data in comparison to the reference CNV dataset for the HG002. The benchmark SVs of GIAB (HG002_SVs_Tier1_v0.6.vcf) were employed to calculate values for evaluation using following formula:
In addition, we sought to determine whether adjusting the parameters to relax the detection criteria could further enhance sensitivity. To include smaller exons, the minimum target of XHMM was set to 1, and the Dnorm of EXCAVATOR2 was set to 103. CODEX2 lacked the functionality to adjust the parameters. A sample from patient 2 with an ERCC8 exon 4 rearrangement was analyzed to ascertain whether parameter adjustments enhanced detection.
Data availability
The ClinVar accession numbers for the ERCC8 variants reported in this study are SCV004177229 (259 kb deletion) and SCV004177228 (exon 4 rearrangement). The datasets analyzed during the current study will be made available by the corresponding author upon reasonable request.
References
Cockayne, E. A. Dwarfism with retinal atrophy and deafness. Arch. Dis. Child 11, 1–8. https://doi.org/10.1136/adc.11.61.1 (1936).
Laugel, V. et al. Mutation update for the CSB/ERCC6 and CSA/ERCC8 genes involved in Cockayne syndrome. Hum. Mutat. 31, 113–126. https://doi.org/10.1002/humu.21154 (2010).
Nance, M. A. & Berry, S. A. Cockayne syndrome: Review of 140 cases. Am. J. Med. Genet. 42, 68–84. https://doi.org/10.1002/ajmg.1320420115 (1992).
de Boer, J. & Hoeijmakers, J. H. Nucleotide excision repair and human syndromes. Carcinogenesis 21, 453–460. https://doi.org/10.1093/carcin/21.3.453 (2000).
Duan, J., Zhang, J. G., Deng, H. W. & Wang, Y. P. CNV-TV: A robust method to discover copy number variation from short sequencing reads. BMC Bioinform. 14, 150. https://doi.org/10.1186/1471-2105-14-150 (2013).
Ceulemans, S., van der Ven, K. & Del-Favero, J. Targeted screening and validation of copy number variations. Methods Mol. Biol. 838, 311–328. https://doi.org/10.1007/978-1-61779-507-7_15 (2012).
Plagnol, V. et al. A robust model for read count data in exome sequencing experiments and implications for copy number variant calling. Bioinformatics 28, 2747–2754. https://doi.org/10.1093/bioinformatics/bts526 (2012).
Fromer, M. et al. Discovery and statistical genotyping of copy-number variation from whole-exome sequencing depth. Am. J. Hum. Genet. 91, 597–607. https://doi.org/10.1016/j.ajhg.2012.08.005 (2012).
Jiang, Y. et al. CODEX2: Full-spectrum copy number variation detection by high-throughput DNA sequencing. Genome Biol. 19, 202. https://doi.org/10.1186/s13059-018-1578-y (2018).
D’Aurizio, R. et al. Enhanced copy number variants detection from whole-exome sequencing data using EXCAVATOR2. Nucleic Acids Res. 44, e154. https://doi.org/10.1093/nar/gkw695 (2016).
Gordeeva, V. et al. Benchmarking germline CNV calling tools from exome sequencing data. Sci. Rep. 11, 14416. https://doi.org/10.1038/s41598-021-93878-2 (2021).
Tan, R. et al. An evaluation of copy number variation detection tools from whole-exome sequencing data. Hum. Mutat. 35, 899–907. https://doi.org/10.1002/humu.22537 (2014).
Samarakoon, P. S. et al. Identification of copy number variants from exome sequence data. BMC Genom. 15, 661. https://doi.org/10.1186/1471-2164-15-661 (2014).
Hong, C. S., Singh, L. N., Mullikin, J. C. & Biesecker, L. G. Assessing the reproducibility of exome copy number variations predictions. Genome Med. 8, 82. https://doi.org/10.1186/s13073-016-0336-6 (2016).
Yao, R. et al. Evaluation of three read-depth based CNV detection tools using whole-exome sequencing data. Mol. Cytogenet. 10, 30. https://doi.org/10.1186/s13039-017-0333-5 (2017).
Zhao, L., Liu, H., Yuan, X., Gao, K. & Duan, J. Comparative study of whole exome sequencing-based copy number variation detection tools. BMC Bioinform. 21, 97. https://doi.org/10.1186/s12859-020-3421-1 (2020).
Uchiyama, Y. et al. Efficient detection of copy-number variations using exome data: Batch- and sex-based analyses. Hum. Mutat. 42, 50–65. https://doi.org/10.1002/humu.24129 (2021).
Miya, F. et al. A combination of targeted enrichment methodologies for whole-exome sequencing reveals novel pathogenic mutations. Sci. Rep. 5, 9331. https://doi.org/10.1038/srep09331 (2015).
Suzuki, H., Yamada, M., Uehara, T., Takenouchi, T. & Kosaki, K. Parallel detection of single nucleotide variants and copy number variants with exome analysis: Validation in a cohort of 700 undiagnosed patients. Am. J. Med. Genet. A 182, 2529–2532. https://doi.org/10.1002/ajmg.a.61822 (2020).
Zook, J. M. et al. A robust benchmark for detection of germline large deletions and insertions. Nat. Biotechnol. 38, 1347–1355. https://doi.org/10.1038/s41587-020-0538-8 (2020).
Janssen, R. J. et al. Contiguous gene deletion of ELOVL7, ERCC8 and NDUFAF2 in a patient with a fatal multisystem disorder. Hum. Mol. Genet. 18, 3365–3374. https://doi.org/10.1093/hmg/ddp276 (2009).
Ren, Y. et al. Three novel mutations responsible for Cockayne syndrome group A. Genes Genet. Syst. 78, 93–102. https://doi.org/10.1266/ggs.78.93 (2003).
Ting, T. W. et al. Cockayne Syndrome due to a maternally-inherited whole gene deletion of ERCC8 and a paternally-inherited ERCC8 exon 4 deletion. Gene 572, 274–278. https://doi.org/10.1016/j.gene.2015.07.065 (2015).
Wang, X. et al. Molecular spectrum of excision repair cross-complementation group 8 gene defects in Chinese patients with Cockayne syndrome type A. Sci. Rep. 7, 13686. https://doi.org/10.1038/s41598-017-14034-3 (2017).
Xie, H. et al. A complex intragenic rearrangement of ERCC8 in Chinese siblings with Cockayne syndrome. Sci. Rep. 7, 44271. https://doi.org/10.1038/srep44271 (2017).
Calmels, N. et al. Functional and clinical relevance of novel mutations in a large cohort of patients with Cockayne syndrome. J. Med. Genet. 55, 329–343. https://doi.org/10.1136/jmedgenet-2017-104877 (2018).
Cloney, T. et al. Lessons learnt from multifaceted diagnostic approaches to the first 150 families in Victoria’s Undiagnosed Diseases Program. J. Med. Genet. 59, 748–758. https://doi.org/10.1136/jmedgenet-2021-107902 (2022).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760. https://doi.org/10.1093/bioinformatics/btp324 (2009).
McKenna, A. et al. The Genome Analysis Toolkit: A MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303. https://doi.org/10.1101/gr.107524.110 (2010).
San-Lucas, F. A., Wang, G., Scheet, P. & Peng, B. Integrated annotation and analysis of genetic variants from next-generation sequencing studies with variant tools. Bioinformatics 28, 421–422. https://doi.org/10.1093/bioinformatics/btr667 (2012).
Tadaka, S. et al. jMorp: Japanese Multi-Omics Reference Panel update report 2023. Nucleic Acids Res. https://doi.org/10.1093/nar/gkad978 (2023).
Acknowledgements
We appreciate the patients’ family members for permission to publish this report. The present research was supported by JSPS KAKENHI Grant Number 16K07211 (F.M.), 20K08236 (M.K. and F.M.), 20H03646 (S.S. and F.M.), the Japan Agency for Medical Research and Development (AMED) under grant numbers grant number JP23ek0109549 and JP23ek0109672 (K.K.).
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. N.O., H.S., To.T., M.Y. and K.K. performed the clinical evaluation of the patient. Data collection was performed by N.O., M.K., S.S., Y.K., Ta.T., and F.M. Genetic analysis and interpretated the results were performed by D.W., Y.K., H.S., M.Y., D.N., M.S., K.K., and F.M. D.W. wrote the original draft; all the authors revised the final manuscript. F.M. supervised the work.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Watanabe, D., Okamoto, N., Kobayashi, Y. et al. Biallelic structural variants in three patients with ERCC8-related Cockayne syndrome and a potential pitfall of copy number variation analysis. Sci Rep 14, 19741 (2024). https://doi.org/10.1038/s41598-024-70831-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-70831-7
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.