Abstract
A series of 20 novel gefitinib derivatives incorporating the 1,2,3-triazole moiety were designed and synthesized. The synthesized compounds were evaluated for their potential anticancer activity against EGFR wild-type human non-small cell lung cancer cells (NCI-H1299, A549) and human lung adenocarcinoma cells (NCI-H1437) as non-small cell lung cancer. In comparison to gefitinib, Initial biological assessments revealed that several compounds exhibited potent anti-proliferative activity against these cancer cell lines. Notably, compounds 7a and 7j demonstrated the most pronounced effects, with an IC50 value of 3.94 ± 0.17 µmol L−1 (NCI-H1299), 3.16 ± 0.11 µmol L−1 (A549), and 1.83 ± 0.13 µmol L−1 (NCI-H1437) for 7a, and an IC50 value of 3.84 ± 0.22 µmol L−1 (NCI-H1299), 3.86 ± 0.38 µmol L−1 (A549), and 1.69 ± 0.25 µmol L−1 (NCI-H1437) for 7j. These two compounds could inhibit the colony formation and migration ability of H1299 cells, and induce apoptosis in H1299 cells. Acute toxicity experiments on mice demonstrated that compound 7a exhibited low toxicity in mice. Based on these results, it is proposed that 7a and 7j could potentially be developed as novel drugs for the treatment of lung cancer.
Similar content being viewed by others
Introduction
Cancer remains a critical illness that poses a significant threat to global public health. The treatment and prevention of cancer have consistently been formidable challenges for the medical community. Researchers have conducted comprehensive studies on oncogenes and tumor suppressor genes at the molecular level, leading to the development of novel drugs that target specific molecular markers in cancer cells. This approach has become a key focus in the development of anti-cancer therapeutics1. Tyrosine kinases are essential in cellular signal transduction, and there is increasing interest in developing drugs that target this pathway. Lung cancer is the most common malignant tumor in the world, with the epidermal growth factor receptor (EGFR) gene playing a crucial role in the development of lung cancer2,3,4. In recent times, targeted drugs against EGFR (such as gefitinib and erlotinib) have been launched sequentially (Fig. 1), offering fresh hope to individuals suffering from lung cancer5,6.
Protein tyrosine kinases are closely related to the occurrence and development of tumors. Overactive tyrosine kinases lead to abnormal activation of downstream signaling pathways, resulting in cell transformation, proliferation, resistance to apoptosis, and promotion of cell growth, ultimately leading to the formation of tumors7,8,9. The clinical trial results indicate that gefitinib belongs to the first generation of targeted therapeutic drugs, which can bind to the epidermal growth factor receptor, and has a good therapeutic effect on non-small cell lung cancer, breast cancer, etc6.
EGFR-TKIs exhibit a robust inhibitory effect on EGFR mutant lung cancer cells; however, their inhibitory activity against wild-type lung cancer cells is comparatively weaker. The objective of this study is to explore structural modifications of EGFR-TKIs in order to augment their inhibitory activity specifically against wild-type lung cancer cells.
The 1,2,3-triazole moiety a nitrogenous heterocyclic compound consisting of a five-membered ring. It has found widespread use in various fields such as medicine, pesticides, and more, owing to its ability to serve as a bioisosteric replacement for diverse structures including acyl groups, carboxylic acid groups, and heterocyclic groups. These compounds can bind to tumor cells through various non-covalent interactions such as hydrogen bonding, van der Waals forces, and metal chelation. They induce tumor cell apoptosis through multiple mechanisms, including blocking the growth cycle, inhibiting cell proliferation, and reducing mitochondrial transmembrane potential. These compounds have potential anti-tumor activity.
Furthermore, utilizing 1,2,3-triazole as a substitute structure has emerged as an effective strategy to overcome patent limitations, fostering imitation innovation. Numerous compounds incorporating the 1,2,3-triazole motif have been developed or investigated for their antitumor, antiviral, and antibacterial biological activities10,11,12. The copper-catalyzed azide-alkyne cycloaddition (Cu-AAC, “click chemistry”) has been employed for the synthesis of 1,2,3-triazole compounds13. Researchers have successfully introduced 1,2,3-triazole into the terminal alkyne groups of erlotinib (Fig. 2, 1) or icotinib (Fig. 2, 2), yielding derivatives with significant IDO1 inhibition activity14,15. These compounds possess two or more pharmacophores, enabling them to interact with multiple targets in tumor cells simultaneously. This characteristic enhances their anti-tumor activity, reduces side effects, overcomes drug resistance, and improves pharmacokinetics.
For instance, the CDK4/6 inhibitor with a 1,2,3-triazole-quinoline-2,4-dione structure (Fig. 2, 3) has been found to exhibit significant inhibitory activity against breast cancer cell lines, including MCF-7, MDA-MB-231, BT-474, and SK-BR3, comparable to that of abemaciclib. This compound arrests the G1/S cell cycle transition by blocking Rb phosphorylation. Pharmacokinetic studies demonstrate a longer plasma half-life and lower plasma clearance compared to abemaciclib, effectively controlling tumor growth volume while minimizing side effects16. In addition, further research has been conducted on the synthesis of acylamide derivatives of 1,2,3-triazole-benzofuran compounds (Fig. 2, 4). These derivatives have demonstrated potential anti-cancer activity against various types of cancer cell lines such as prostate cancer (PC3), lung cancer (A549), breast cancer (MCF7), and ovarian cancer (A2780). Molecular docking studies have supported the predicted binding affinity, indicating enhanced inhibitory effects17. Similarly, a novel pyrido[2,3-d]pyrimidine-thiazole-1,2,3-triazole compound (Fig. 2, 5) has been designed and synthesized, showing inhibitory effects on EGFR kinase and anti-proliferative activity in different cell lines18.
In summary, the integration of 1,2,3-triazole with other pharmacophores represents a highly effective approach in the development of novel anti-cancer drugs (Fig. 2, 6)19. The diverse studies presented demonstrate the remarkable potential of 1,2,3-triazole-modified compounds in cancer treatment, offering insights into their mechanisms and promising outcomes in cell-based studies.
To identify compounds with improved inhibitory effects on wild-type lung cancer cells, we modified gefitinib using a click reaction, introducing a 1,2,3-triazole moiety into its structure. We hoped that this modification would exhibit inhibitory effects on wild-type lung cancer cells. Using gefitinib as a positive control, we employed the CCK-8 assay to investigate the in vitro anti-tumor activity on wild-type human non-small cell lung cancer cells (NCI-H1299, A549) and human lung adenocarcinoma cells (NCI-1437). Furthermore, we studied the impact on tumor cell colony formation, migration ability, and apoptosis.
Chemistry
The synthetic strategy for the preparation of the target molecules is illustrated through a literature review in Scheme 120,21,22. Nitration of 1 was realized using HNO3/H2SO4 as the nitration agent, and the resulted compound 2 was readily converted to compound 3 under the conventional hydrogenation conditions. Cyclization reaction between 3 and formamide produced compound 4. The chlorination of hydroxyl group with phosphorus oxychloride produced compound 5. Compound 6 was obtained after reaction of 5 and 3-aminophenylacetylene. Finally, 1,2,3-triazole gefitinib derivatives 7a-7t was synthesized through a copper(I)-catalyzed azide-alkyne cycloaddition reaction. The reaction conditions of these operations were mild and easy to control. The structures of the key intermediates and all target compounds were confirmed through nuclear magnetic resonance spectroscopy (1H NMR and 13C NMR) and high-resolution mass spectrometry (HRMS). The detail and figures of NMR spectrums and HRMS were included in the supplementary files.
Results and discussion
Compounds 7a-7t suppressed cancer cells viability
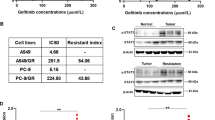
In this study, the CCK-8 assay was employed to investigate the cell inhibitory effects of compounds 7a-7t on human lung cancer cells NCI-H1299, NCI-H1437, and A549, with gefitinib used as a positive control. The results (Table 1) indicated that almost all compounds exhibited inhibitory effects on NCI-H1299, NCI-H1437, and A549 lung cancer cells, with compounds 7a and 7j showing the best performance. Specifically, in the NCI-H1299 cell line, the IC50 values for 7a and 7j were 3.94 ± 0.17 μM and 3.84 ± 0.22 μM, respectively. In the NCI-H1437 cell line, the IC50 values for 7a and 7j were 1.83 ± 0.13 μM and 1.69 ± 0.25 μM, respectively. In the A549 cell line, the IC50 values for 7a and 7j were 3.16 ± 0.11 μM and 3.86 ± 0.38 μM, respectively. These values were all superior to the inhibitory effects of gefitinib in the three cell lines (IC50 = 14.62 ± 0.90 μM, IC50 = 20.56 ± 2.45 μM, and IC50 = 14.62 ± 0.43 μM, respectively). The experimental results suggest that the gefitinib derivatives have higher cytotoxicity and play a pro-apoptotic role in lung cancer cells.
Compounds 7a and 7j exhibited IC50 values of 18.87 + 1.03 μM and 17.68 ± 0.52 μM respectively in the L02 cell line. Moreover, our study demonstrated that these compounds are less toxic to normal hepatocytes and more effective in killing lung cancer cells, with a survival rate of normal hepatocytes L02 treated with 7a and 7j at a concentration of 4 μM between 70% and 90%. These findings suggest that compounds 7a and 7j hold great potential as clinical therapeutic agents for lung cancer (Table 2). The introduction of phenyl or benzyl groups had a notable impact on the activity of the compounds. Notably, when the triazole linkage was benzyl, the activity was superior to phenyl. The study clearly demonstrates the successful structural modification of gefitinib, resulting in significant differences in compound activity. Furthermore, the addition of bromine atoms in the neighbouring position of the benzyl ring or the addition of fluorine or chlorine atoms in the interstitial position significantly improved the in vitro antitumour activity of the compounds.
Compounds 7a and 7j induce apoptosis in H1299 cells
To elucidate whether the inhibitory effects on cell proliferation and cytotoxicity of these compounds are associated with apoptosis, we conducted relevant experiments focusing on compounds 7a and 7j, which demonstrated good inhibitory effects on the proliferation of the three types of cancer cells. H1299 cells were treated with different concentrations of 7a and 7j for 48 h. The cells were stained with Annexin-V and PI, and the proportion of apoptotic cells was detected using flow cytometry.
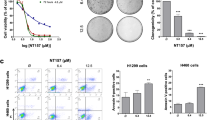
As shown in Fig. 3A, after treatment with compound 7a, the total apoptotic cell proportions in H1299 were 19.03 ± 2.10% (2 μmol/L), 28.73 ± 1.12% (4 μmol/L), and 50.1 ± 2.91% (8 μmol/L) (Fig. 3). Following treatment with compound 7j, the total apoptotic cell proportions in H1299 were 14.97 ± 1.54% (2 μmol/L), 27.07 ± 2.77% (4 μmol/L), and 65.77 ± 2.93% (8 μmol/L) (Fig. 3A). Compared to the control group, as the drug concentration increased, the apoptotic proportion gradually increased (P < 0.01, P < 0.05). These results suggest that both 7a and 7j significantly promote apoptosis in the H1299 lung cancer cell line and exhibit concentration-dependent effects.
In order to further verify the apoptotic activity induced by 7a and 7j in lung cancer cells, H1299 cells were treated with different concentrations of compounds 7a and 7j for 48 h, followed by DAPI staining. The cells were observed under a fluorescence microscope, and representative images are shown in Fig. 3B. Compared to the normal control group, H1299 cells treated with compounds 7a and 7j exhibited typical apoptotic features. As the drug concentration increased, nuclear staining intensified, and even nuclear condensation and fragmentation phenomena were observed. Therefore, the apoptosis-related results indicate that both 7a and 7j can significantly promote apoptosis in human lung cancer cells (H1299) in a concentration-dependent manner.
Compounds 7a and 7j suppress metastasis in H1299 cell
To investigate the impact of compounds on the migration of wild-type lung cancer cells, we utilized a wound healing assay and Transwell assay to assess the changes in migration ability of H1299 cells after treatment with compounds 7a and 7j. Figure 4 presents the results of the wound healing assay, indicating that after treatment with compound 7a, the migration closure rates of H1299 cells were 65.16 ± 7.88% (2 μmol/L), 50.12 ± 2.99% (4 μmol/L), and 23.54 ± 3.65% (8 μmol/L). After treatment with compound 7j, the migration closure rates of H1299 cells were 72.57 ± 3.43% (2 μmol/L), 60.91 ± 8.48% (4 μmol/L), and 48.88 ± 3.23% (8 μmol/L) (Fig. 4A). The migration closure rates in the compound-treated groups were significantly lower than the control group, showing a concentration-dependent trend (p < 0.01, p < 0.05).
Transwell chamber migration assay is another important method to assess cell migration ability. Tumor cells are seeded in the chamber, and the number of tumor cells that pass through the membrane is counted to evaluate the migration ability of tumor cells23. The results of the Transwell migration assay (shown in Fig. 4B) also indicate that compared to the normal control group, the number of cells passing through the membrane to the lower surface of the chamber in the compound-treated group significantly decreased (p < 0.01). Therefore, the experiments demonstrate that compounds 7a and 7j can inhibit the migration ability of non-small cell lung cancer cells in a concentration-dependent manner.
Compounds 7a and 7j inhibit the clonogenic ability in H1299 cells
The colony formation assay is an important method for evaluating the ability of adherent cells to proliferate and form colonies on a plate, and it is widely used to assess cell proliferation24. We also employed the colony formation assay to further confirm that compounds 7a and 7j can inhibit the proliferation of lung cancer cells, as shown in Fig. 5. After treating lung cancer cells with low, medium, and high concentrations (1, 2, 4 μmol/L) of compounds for 7 days, the number of cell colonies was counted to analyze the proliferation inhibition rate. The results showed a significant decrease in the number of cell colonies in the compound-treated groups, and this effect was concentration-dependent. Even at the lowest concentration of 1 μmol/L, the compounds began to markedly inhibit the formation of H1299 cell colonies. Particularly at a concentration of 4 μmol/L, colony formation was almost completely suppressed, indicating that compounds 7a and 7j exert a strong anti-proliferative effect on H1299 cells (Fig. 5).
Compounds 7a and 7j trigger apoptosis via the apoptosis signaling pathway
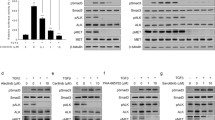
We conducted an in-depth investigation into the mechanism of apoptosis induced by compounds 7a and 7j. The Bcl-2 protein family is located upstream of the apoptosis signaling pathway. Bcl-2 proteins play an anti-apoptotic role by inhibiting the release of cytochrome C (cyt-c) from mitochondria. Previous literature reports that many anticancer drugs induce apoptosis by downregulating Bcl-2 expression25. During apoptosis, the expression of Bcl-2 is inhibited, cyt-c is released, and it binds with the apoptotic protease activation factor to form the apoptosome. This triggers a cascade reaction of Caspase family proteins, with apoptosome promoting the activation of Caspase9. Activated Caspase9 then activates Caspase3, forming the Caspase3 cleavage body, which further activates downstream PARP protein, ultimately promoting cell apoptosis26. Therefore, by detecting the expression levels of Bcl-2, caspase9, caspase3, and PARP proteins, we found that the treatment with compounds 7a and 7j resulted in a decrease in Bcl-2 and caspase9 protein expression levels, while the expression levels of Cleaved-Caspase3 and Cleaved-PARP proteins increased. These results suggest that compounds 7a and 7j may effectively induce apoptosis in non-small cell lung cancer cells through the Bcl-2/caspase3/PARP signaling pathway. After treating H1299 cells with different concentrations of compounds for 48 h, cell proteins were extracted, and western blot was used to detect the expression changes of apoptosis-related proteins. The results showed that the migration-related protein MMP9 did not show significant changes in the low drug concentration group but decreased significantly in the high drug concentration group. The apoptosis-related protein Bcl-2 exhibited a decrease in expression with increasing drug concentration, while Cleaved-PARP showed an increase in expression with increasing drug concentration. The expression of caspase3, PARP, and caspase9 proteins decreased with increasing drug concentration, indicating that apoptosis in the high drug concentration group was significantly stronger than that in the low drug concentration group and the control group (p < 0.01, p < 0.05).
Previous studies have reported that MMP9 plays a crucial role in angiogenesis and cell migration27. In non-small cell lung cancer tissues, the protein expression level of MMP9 is significantly higher than in normal adjacent tissues, suggesting a close association between the high expression of MMP9 and the malignant metastasis of lung cancer as well as poor prognosis28. To further investigate whether compounds regulate the migration ability of non-small cell lung cancer by modulating MMP9 protein levels, Western Blot results show a decrease in MMP9 protein expression with increasing drug concentration. Therefore, compounds 7a and 7j may inhibit the migration ability of non-small cell lung cancer by downregulating MMP9 protein expression (Fig. 6).
Assessment of apoptosis and migration induced by 7a and 7j. (A) Western blotting results of the protein levels in H1299 cells treated with 4, 8 or 16 μM. compounds 7a. Numbers below each lane indicated the relative expression level of the protein. (B) Western blotting results of the protein levels in H1299 cells treated with 4, 8 or 16 μM. compounds 7j. Numbers below each lane indicated the relative expression level of the protein.
Acute oral toxicity assessment
We further evaluated the safety of the compounds through acute toxicity experiments in mice. From the initial gavage to the 12th day, there was no statistical difference in mouse body weight compared to the control group. Observation of organ morphology during dissection revealed no pathological changes, and organ indices showed no statistical differences compared to the control group. Biochemical indicators, including serum alanine aminotransferase (ALT), aspartate aminotransferase (AST), blood urea nitrogen (BUN), and creatinine (CRE), were measured. ALT and AST are commonly used to assess liver function, reflecting physiological and pathological changes in liver function. Elevated levels of ALT and AST in the serum indicate damage to liver cells or mitochondria29. BUN and CRE are typically used to reflect renal function, and an increase in their levels in the serum suggests a decline in kidney function or kidney injury23. The results of these biochemical indicators showed no statistical differences compared to the control group and were within the normal range. HE staining of the mouse brain, heart, liver, spleen, lungs, kidneys, and stomach further confirmed that mice did not exhibit obvious toxic characteristics at the drug concentration tested. This additional evidence indicates a high level of safety for the compounds, providing experimental data reference for the clinical safety of the compounds (p > 0.05) as shown in Fig. 7.
Compound 7a in vivo toxicity of mice. (A) Acute toxicity experiments of compound 7a were conducted in mice. Mouse body weight changed. (B) The acute toxicity experiments examined the effects of compound 7a on mouse organs (C) Effect of acute toxicity experimental studies on blood biochemical indices in mice. (D) H&E staining were performed on various organs of mice treated with compound 7a.
From the day of self-administration until the 12th day, mice in all groups showed no apparent toxic reactions, and there were no deaths. When compared to the control group, there were no significant differences in body weight among the experimental groups. (p > 0.05) as shown in Fig. 7.
Conclusion
In summary, gefitinib derivatives containing a 1,2,3-triazole ring were designed, synthesized, and evaluated for their anti-tumor activity against wild-type lung cancer cells. The synthetic method was simple and efficient, and the compounds were structurally characterized and confirmed. Several of these compounds exhibited superior antitumor activity compared to gefitinib against one or more cancer cell lines employed in this study. Among them, compounds 7a and 7j exhibited strong anti-proliferative activity in three non-small cell lung cancer. Through cell function and mechanism studies, it was found that compounds 7a and 7j downregulated the expression of Bcl-2, Caspase9, and MMP9 proteins, upregulated the expression of Cleaved-Caspase3 and Cleaved-PARP proteins, inhibited tumor cell proliferation and migration, promoted cell apoptosis. Acute toxicity experiments in mice also confirmed their safety, providing foundational data for future in vivo experiments and clinical applications of these novel gefitinib derivatives, with the hope that these compounds may become effective and low-toxicity anti-cancer drugs.
Experimental work
Materials and Chemistry
The gefitinib-1,2,3-triazole derivative was synthesized in-house. All the reagents and solvents used were obtained from a commercially available source. The 1H and 13C NMR spectra were acquired in a DMSO-d6 solution using a Bruker 400 MHz NMR spectrometer. High-resolution mass spectra (HRMS) measurements were carried out using a Bruker Compact mass spectrometer.
Preparation of methyl 4-methoxy-5-(3-morpholinopropoxy)-2-nitrobenzoate (2)
To a mixture of compound 1 (15 g, 50 mmol) in acetic acid (100 mL) solution was added the mixture of nitric acid (3.92 g, 62 mmol) and sulfuric acid (6.07 g, 62 mmol) dropwise at 5 °C. After addition, the system gradually rose to room temperature, and the mixture was stirred at room temperature for 24 h. the reaction was poured into ice water (100 mL), The precipitate was collected by filtration, washed with water (20 mL) and recrystallized with isopropyl alcohol and dried successively to afford 2 (14 g, 80%). Mp:118–121 °C. 1H NMR (400 MHz, CDCl3): δ 7.45 (s, 1H),7.10 (s, 1H),4.18 (t, 2H), 3.94 (s, 3H), 3.93 (s, 3H), 3.73 (t, 4H), 2.55–2.47 (m, 6H), 2.06 (t, 2H).
Preparation of methyl 2-amino-4-methoxy-5-(3-morpholinopropoxy)benzoate (3)
A mixture of compound 2 (10 g, 28 mmol) and Pd/C (10%wt., 1 g) in methanol (150 mL) was stirred at 25 °C under H2 (15 psi) atmosphere for 24 h. The reaction mixture was filtered through celite and the filter cake washed with methanol (150 mL). The filtrate was concentrated to afford 3 (7.70 g, 85%) as a light yellow solid. Mp:89–90 °C. 1H NMR (400 MHz, DMSO-d6): δ 7.54(s,1H),7.14(s,1H), 6.43(s,2H), 4.28 (t, 2H), 3.97 (s, 3H), 3.91(s, 3H), 3.63(t, 4H), 2.57–2.52(m, 6H), 2.16–2.15 (m, 2H).
Preparation of 7-methoxy-6-(3-morpholinopropoxy)quinazolin-4(3H)-one (4)
A mixture of compound 3 (4.0 g, 10 mmol) and ammonium formate (2.0 g, 32 mmol) in DMF (30 mL) was stirred at 165 °C in nitrogen environment for 3–4 h. The reaction mixture was poured into ice-water (100 mL) and extracted with ethyl acetate. The combined extracts were washed with sat NaCl (50 mL) and then dried with anhydrous magnesium sulfate, concentrated to afford 4 (3.0 g, 75%) as a yellow solid. Mp:247–250 °C.1H NMR (400 MHz, DMSO-d6) δ 7.97 (s, 1H), 7.44 (s, 1H), 7.12 (s, 1H), 4.11 (t, J = 6.5 Hz, 2H), 3.90 (s, 3H), 3.58 (t, J = 4.6 Hz, 4H), 2.44 (t, J = 7.0 Hz, 2H), 2.37 (s, 4H), 1.99–1.80 (m, 2H).
4-(3-((4-chloro-7-methoxyquinazolin-6-yl)oxy)propyl)morpholine (5)
DMF (1.3 g, 18 mmol) was added dropwise over 20 min to a stirred solution of oxalyl chloride (2.28 g, 18 mmol) in DCM (12 mL) under N2 at 25 °C, resulting in an exotherm and gas evolution. When gas evolution ceased, compound 4 (2.6 g, 7.8 mmol) was added with mechanical agitation, the mixture was heated to reflux for 24 h and cooled to 25 °C, then the reaction was quenched with dilute aqueous Na2HPO4 solution (0.5 M, 25 mL). The resulting mixture was stirred on an ice bath for 2 h, and the solid was collected, rinsed with water (25 mL), and dried at 50 °C under vacuum to 5 (2.2 g, 85%) which was used directly. Mp:116–119 °C. 1H NMR (400 MHz, DMSO-d6) δ 8.86 (s,1H), 7.43 (s, 1H), 7.38 (s,1H), 4.23 (t, 2H), 3.95 (s, 3H), 3.57 (t, 4H), 2.43 (t, 2H), 2.37 (s, 4H), 2.03–1.92 (m, 2H).
Preparation of N-(3-ethynylphenyl)-7-methoxy-6-(3-morpholinopropoxy)quinazolin-4-amine (6)
Reaction of compound 5 (1.5 g, 4.5 mmol) and m-acetylenyl aniline (1.56 g, 13.4 mmol) was refluxed for 6 h in stirred 2-propanol (40 mL) under N2 for gave 6 (1.6 g, 65%). 1HNMR (400 MHz, DMSO-d6) δ 11.22 (s, 1H), 8.78 (s, 1H), 8.50 (s, 1H), 7.99 (s, 1H), 7.91 (d, J = 8.2 Hz, 1H), 7.48 (t, J = 7.9 Hz, 1H), 7.37 (d, J = 8.8 Hz, 2H), 4.40 (s, 2H), 4.28 (s, 1H), 3.96–3.79 (m, 7H), 3.39–3.160 (m, 6H), 2.35(s, 2H).
General procedure for the preparation of compound 7
Aryl-azido (150 mg, 0.7 mmol) and 6 (250 mg, 0.56 mmol) were added to 15 mL mixed solvent (water/tert-butanol/THF = 1:1:1). Copper sulfate pentahydrate (14 mg, 0.1 mmol) and sodium ascorbate (23 mg, 0.1 mmol) were added and the mixture was stirred at 85 °C for 12 h. After the completion of the reaction (monitored by TLC), the mixture was extracted with dichloromethane (15 mL × 3). The combined organic phase was washed successively with brine, dried over sodium sulfate and concentrated in vacuo. The residue was purified by through column chromatography (CH2Cl2/MeOH = 20:1) to give the desired compound 7a-7t as a crystalline powder.
Biological study
Lung cell lines A549, NCI-H1437, and NCI-H1299 were purchased from Procell Life Science &Technology Co., Ltd (Wuhan, China). Ham’s F-12 K Medium and Fetal bovine serum (FBS) were purchased from Procell Life Science &Technology Co., Ltd (Wuhan, China). RPMI 1640 Medium and diamidino-phenyl-indole (DAPI) were purchased from Solarbio Science Technology (Beijing, China). Cell Counting Kit-8 and Annexin V-EGFP Apoptosis Detection Kit were obtained from Beyotime Biotechnology (Shanghai, China). The primary antibodies against Caspase3 (1:500), PARP/cleaved-PARP (1:750), and MMP9 (1:1000) were obtained from Wan lei Biotechnology (Shenyang, China). Alpha Tubulin (1:2000), cleaved-Caspase3 (1:500), Caspase9(1:300) was purchased from Proteintech (Wuhan, China). Bcl-2(1:1000), HRP-conjugated affinipure goat anti-rabbit IgG (1:1000) was purchased from Cell Signaling Technology (MA, United States).
Cell culture
NCI-H1299 and NCI-H1437 cells were cultured in a humidified incubator at 37 °C with 5% CO2 in RPMI-1640 (No.31800, Solarbio) supplemented with 10% fetal bovine serum (FBS), and 1% Penicillin Streptomycin. A549 cells were cultured in a humidified incubator at 37 °C with 5% CO2 in Ham’s F-12 K (PM150910, Procell) supplemented with 10% FBS and 1% Penicillin Streptomycin.
CCK-8 assay for cell proliferation and cytotoxicity
Three types of lung cells were seeded into 96-well plate at a density of 5 × 103 cells per well at logarithmic growth phase and cultured in 37 °C. Then, the cells were treated initially with different concentrations of compounds (0, 2, 4, 8, 16 and 32 μmol/L) for additional 48 h, with three replica wells each. After that, cell viability was determined according to the instruction of the CCK-8 assay. Next, 10 μL of CCK-8 solution was added to each well of the plate and incubate the plate for 1–4 h in the incubator. The absorbance was measured at 450 nm using a microplate reader (Bio-Tek). The percentage of viable cells was measured using the following formula where three independent experiments were performed: [(A450 sample − A450blank)/(A450 control − A450blank)] × 100%.
Flow cytometry detection for cell apoptosis
H1299 cells were seeded in a 6-well plate at a density of 1 × 105 cells/well and incubated for 24 h. On the following day, the medium was replaced with fresh medium containing 7a or 7j (2, 4 and 8 μmol/L) and cells were incubated for an additional 48 h. Cell apoptosis was detected using Annexin V-EGFP/PI apoptosis detection Kit by flow cytometry. Subsequently, 5 µl of EGFR Annexin V and 10 µl of PI was added to the cell suspension, gently vortexed, and incubated at room temperature in the dark for 20 min, then single-cell suspension was prepared followed by flow cytometry (BD Accuri™ C6 Plus).
DAPI staining for cell apoptosis
H1299 (1.5 × 104 ~ 2 × 104 /well) cells were seeded in 24-well plates for 24 h, and then treated with 7a or 7j at the concentrations of 2,4 and 8 μmol/L for 48 h., the cells were fixed for 30 min at room temperature in 4% paraformaldehyde and washed 3 times in PBS. Finally, every well was stained with DAPI (10 µg/mL, C0065, Solarbio) at room temperature for 30 min, and washed at least three times in PBS. The fluorescence microscope was used to observe the morphological changes of cell nuclei.
Wound healing assay
H1299 cells were seeded in a 6-well plate, When the cell confluency reached 95%, a wound line was scratched using a 200 µl pipette tip, and then washed three times, fresh medium containing 7a or 7j (2, 4 and 8 μmol/L) was added, and the plate was incubated at 37 °C with 5% CO2. All the images were captured at 0 h and 24 h under an inverted microscope, and the quantification of scratches was analysed using Image J.
Transwell migration assay
Transwell assay was used to determine the migration ability of H1299 cells. Cells were starved in serum-free RPMI 1640 medium for 24 h, then detached and resuspended in serum-free RPMI 1640 medium. 5 × 105 cells/mL were inoculated into the upper chamber of a 24-well Transwell plate (Corning Inc., United States), with a volume of 100 µl per well. Then, 100 µL serum-free medium containing 7a or 7j (2, 4 and 8 μmol/L) was added to the upper chamber and the lower chamber was filled with 700 µL medium containing 20% FBS. After incubation for 24 h, H1299 cells on the upper membrane of the transwell were wiped off. The migrated cells were treated with 4% paraformaldehyde for 20 min, stained with 0.1% crystal violet for 20 min and washed three times with PBS. The number of H1299 cells that migrated to the underside of the membrane was counted under the inverted fluorescence microscope. Three randomly selected areas from each transwell were photographed and calculated using Image J.
Colony formation assay
H1299 cells were seeded in six-well plates at a density of 500 cells per well and treated with 7a or 7j (1, 2 and 4 μmol/L) for 7 days. Then, cells were per-fixed with 4% paraformaldehyde for 30 min, stained with 0.1% Crystal violet for 15 min, and then washed with pure water. Taking a picture after the plates were air-dry. The clone formations number was counted with Image J.
Western blot analysis
Firstly, H1299 cells were treated with different concentrations concentrations of 7a and 7j at 4, 8 and 16 μmol/L for 48 h, cells were harvested using RIPA lysate (R0010, Solarbio) and centrifuged at 12,000 rpm for 15 min at 4 °C, and then the concentration of total protein was measured and taken from the supernatant. Protein was separated by 12% SDS–polyacrylamide gel electrophoresis and transferred to nitrocellulose filter membrane. after blocking in 5% milk for 2 h, the membranes were incubated with specific primary antibody at 4 °C overnight. Next day, NC membranes were incubated for 1 h at room temperature with appropriate secondary antibodies, and then Protein bands was visualizing by chemiluminescence detection (ECL kit, Genview).
In vivo assay
Animals
SPF-grade healthy Kunming mice (weighing 20 ± 2 g) were purchased from Henan Skibbes Bio-technology Co., Ltd (Licence No.: SCXK (Yu) 2020–0005). The mice were provided free access to drinking water and adaptive feeding for 7 days.
Acute toxicity test
Twenty Kunming mice (half male and half female) were randomly allocate into 4 groups: 7a-treated group (male), 7a-treated group (female), control group (male) and control group (female). The control groups were given PBS intragastric administration, while the 7a groups were given the 7a solution (400 mg/kg, respectively), once a day for 2 consecutive days. Body weight and appearance were monitored for a total period of 12 days (from the first day after treating with 7a). The mice were killed by cervical dislocation on the 12th day, and blood samples were collected for the biochemistry test. Then, we measured liver and kidney function indicators such as alanine aminotransferase (ALT), aspartate aminotransferase (AST), serum creatinine (CRE) and blood urea nitrogen (BUN) (Nanjing Jiancheng Bioengineering Institute, China). The organs (brain, heart, liver, spleen, lungs, kidneys stomach) were harvested, and fixed in 10% formaldehyde and paraffin-embedded for histological examination. Finally, these sections were stained with haematoxylin and eosin for light microscopic examination.
Statistical analyses
We performed statistical analyses using GraphPad Prism 9.5 software these data were analyzed using one-way analysis of variance (ANOVA) followed by Dunnett’s tests. Data were obtained from no fewer than three independent experiments. A p-value less than 0.05 was considered statistically significant. A p-value more than 0.05 indicated no significant difference.
Ethical approval
The animal study was reviewed and approved by The Ethics Committee for the Care and Use of Laboratory Animals of Henan University of Science and Technology. All methods are reported in accordance with ARRIVE guidelines and in accordance with the relevant guideline and regulation.
Data availability
The data underlying this study are available in the published article and its Supporting Information.
References
Ganesh, K. & Massague, J. Targeting metastatic cancer. Nat. Med. 27, 34–44 (2021).
To, K. K. W., Fong, W. & Cho, W. C. S. Immunotherapy in treating EGFR-mutant lung cancer: Current challenges and new strategies. Front. Oncol. https://doi.org/10.3389/fonc.2021.635007 (2021).
Passaro, A., Janne, P. A., Mok, T. & Peters, S. Overcoming therapy resistance in EGFR-mutant lung cancer. Nat. Cancer 2, 377–391 (2021).
Mamdani, H., Matosevic, S., Khalid, A. B., Durm, G. & Jalal, S. I. Immunotherapy in lung cancer: Current landscape and future directions. Front. Immunol. https://doi.org/10.3389/fimmu.2022.823618 (2022).
Lurje, G. & Lenz, H.-J. EGFR signaling and drug discovery. Oncology 77, 400–410 (2009).
Cataldo, V. D., Gibbons, D. L., Perez-Soler, R. & Quintas-Cardama, A. Treatment of non-small-cell lung cancer with Erlotinib or Gefitinib. N. Engl. J. Med. 364, 947–955 (2011).
Ciardiello, F. & Tortora, G. A novel approach in the treatment of cancer: Targeting the epidermal growth factor receptor. Clin. Cancer Res. 7, 2958–2970 (2001).
Herbst, R. S. Review of epidermal growth factor receptor biology. Int. J. Radiat. Oncol. Biol. Phys. 59, 21–26 (2004).
Pao, W. & Chmielecki, J. Rational, biologically based treatment of EGFR-mutant non-small-cell lung cancer. Nat. Rev. Cancer 10, 760–774 (2010).
Liang, T., Sun, X., Li, W., Hou, G. & Gao, F. 1,2,3-triazole-containing compounds as anti-lung cancer agents: current developments, mechanisms of action and structure-activity relationship. Front. Pharmacol. 12, 661173 (2021).
Zhao, S. J., Liu, J., Lv, Z. S., Zhang, G. D. & Xu, Z. Recent updates on 1,2,3-triazole-containing hybrids with in vivo therapeutic potential against cancers: A mini-review. Eur. J. Med. Chem. 251, 115254 (2023).
Bozorov, K., Zhao, J. Y. & Aisa, H. A. 1,2,3-triazole-containing hybrids as leads in medicinal chemistry: A recent overview. Bioorg. Med. Chem. 27, 3511–3531 (2019).
Rostovtsev, V. V., Green, L. G., Fokin, V. V. & Sharpless, K. B. A stepwise Huisgen cycloaddition process: Copper(I)-catalyzed regioselective “ligation” of azides and terminal alkynes. Angew. Chem Int. Ed. 41, 2596–2599 (2002).
Xu, G.-Q. et al. Novel 1,2,3-triazole erlotinib derivatives as potent IDO1 inhibitors: Design, drug-target interactions prediction, synthesis, biological evaluation, molecular docking and ADME properties studies. Front. Pharmacol. 13, 854965 (2022).
Mao, L.-F. et al. Discovery of icotinib-1,2,3-triazole derivatives as IDO1 inhibitors. Front. Pharmacol. 11, 579024 (2020).
Praveenkumar, E. et al. Selective CDK4/6 inhibition of novel 1,2,3-triazole tethered acridinedione derivatives induces G1/S cell cycle transition arrest via Rb phosphorylation blockade in breast cancer models. Bioorg. Chem. 116, 105377 (2021).
Gariganti, N. et al. Design, synthesis, anticancer activity of new amide derivatives derived from 1,2,3-triazole-benzofuran hybrids: An insights from molecular docking, molecular dynamics simulation and DFT studies. J. Mol. Struct. 1273, 134250 (2023).
Bandi, S. R. et al. Synthesis of novel pyrido[2,3-d]pyrimidine-thiazolidine-1,2,3-triazoles: Potent EGFR targeting anticancer agents. J. Mol. Struct. 1294, 136451 (2023).
Othman, E. M., Fayed, E. A., Husseiny, E. M. & Abulkhair, H. S. Apoptosis induction, PARP-1 inhibition, and cell cycle analysis of leukemia cancer cells treated with novel synthetic 1,2,3-triazole-chalcone conjugates. Bioorg. Chem. 123, 105762 (2022).
Lipunova, G. N., Nosova, E. V., Charushin, V. N. & Chupakhin, O. N. Synthesis and antitumour activity of 4-aminoquinazoline derivatives. Russ. Chem. Rev. 85, 759–793 (2016).
Li, H.-X., Qian, Y.-M. & Xu, L.-S. Design, synthesis and anticancer activity evaluation of novel Quinazoline derivatives as EFGR inhibitors. Chin. J. Struct. Chem. 40, 933–941 (2021).
El-Azab, A. S. et al. Synthesis, antiproliferative and enzymatic inhibition activities of quinazolines incorporating benzenesulfonamide: Cell cycle analysis and molecular modeling study. J. Mol. Struct. 1278, 134928 (2023).
Katt, M. E., Placone, A. L., Wong, A. D., Xu, Z. S. & Searson, P. C. In vitro tumor models: Advantages, disadvantages, variables, and selecting the right platform. Front. Bioeng. Biotechnol. 4, 12 (2016).
Oike, T. et al. Reporting of methodologies used for clonogenic assays to determine radiosensitivity. J. Radiat. Res. 61, 828–831 (2020).
Tajuddin, W., Lajis, N. H., Abas, F., Othman, I. & Naidu, R. Mechanistic understanding of curcumin’s therapeutic effects in lung cancer. Nutrients 11, 2989 (2019).
Hu, Q. et al. Baicalin and the liver-gut system: Pharmacological bases explaining its therapeutic effects. Pharmacol. Res. 165, 105444 (2021).
Augoff, K., Hryniewicz-Jankowska, A., Tabola, R. & Stach, K. MMP9: A tough target for targeted therapy for cancer. Cancers 14, 1847 (2022).
Gong, L. et al. Prognostic impact of serum and tissue MMP-9 in non-small cell lung cancer: A systematic review and meta-analysis. Oncotarget 7, 18458–18468 (2016).
Sookoian, S. & Pirola, C. J. Liver enzymes, metabolomics and genome-wide association studies: From systems biology to the personalized medicine. World J. Gastroenterol. 21, 711–725 (2015).
Funding
This work was supported by Scientific Research Foundation for PhD (Henan Normal University, NO. 5101039170343 and Xinxiang University, NO. 1366020232). And the Key Scientific Research Projects of Universities in Henan Province (24A350006). The Henan Province Medical Science and Technology Research Project (LHGJ20230450). College Students’ Innovative Entrepreneurial Training Plan Program (202310464081). The Tackle Key Problems in Science and Technology Project of Henan Province, China (Grant No. 242102311222 and 242102311226).
Author information
Authors and Affiliations
Contributions
E.G. and L.F.M. are the principal two authors of this paper. They were responsible for the conceptualization of its idea and have holistically contributed to all parts of the work and manuscript. Y.W. and G.L.F. was also responsible for drafing. G.Q.X. and Z.Y.W. was also responsible for reviewing. Z.J.L. and J.C.L. was also responsible for proofreading the diferent successive versions of the manuscript, starting from the primary submission until the final publication. X.X.H. and S.H.L. was also responsible for reviewing. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Gao, E., Wang, Y., Fan, Gl. et al. Discovery of gefitinib-1,2,3-triazole derivatives against lung cancer via inducing apoptosis and inhibiting the colony formation. Sci Rep 14, 9223 (2024). https://doi.org/10.1038/s41598-024-60000-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-60000-1
Keywords
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.