Abstract
Long extrachromosomal circular DNA (leccDNA) regulates several biological processes such as genomic instability, gene amplification, and oncogenesis. The identification of leccDNA holds significant importance to investigate its potential associations with cancer, autoimmune, cardiovascular, and neurological diseases. In addition, understanding these associations can provide valuable insights about disease mechanisms and potential therapeutic approaches. Conventionally, wet lab-based methods are utilized to identify leccDNA, which are hindered by the need for prior knowledge, and resource-intensive processes, potentially limiting their broader applicability. To empower the process of leccDNA identification across multiple species, the paper in hand presents the very first computational predictor. The proposed iLEC-DNA predictor makes use of SVM classifier along with sequence-derived nucleotide distribution patterns and physicochemical properties-based features. In addition, the study introduces a set of 12 benchmark leccDNA datasets related to three species, namely Homo sapiens (HM), Arabidopsis Thaliana (AT), and Saccharomyces cerevisiae (SC/YS). It performs large-scale experimentation across 12 benchmark datasets under different experimental settings using the proposed predictor, more than 140 baseline predictors, and 858 encoder ensembles. The proposed predictor outperforms baseline predictors and encoder ensembles across diverse leccDNA datasets by producing average performance values of 81.09%, 62.2% and 81.08% in terms of ACC, MCC and AUC-ROC across all the datasets. The source code of the proposed and baseline predictors is available at https://github.com/FAhtisham/Extrachrosmosomal-DNA-Prediction. To facilitate the scientific community, a web application for leccDNA identification is available at https://sds_genetic_analysis.opendfki.de/iLEC_DNA/.
Similar content being viewed by others
Introduction
Deoxyribonucleic acid (DNA) is comprised of billions of nucleotides and special arrangements of these nucleotides contain essential information for the development, functioning and inheritance of living organisms1,2. These nucleotides represent 25,000 protein-coding genes and various regulatory elements that control gene regulation2. In DNA sequence, these genetic components are organized in a structured manner and the sequence is wrapped around histone octamers also known as nucleosomes. Together around 30 million nucleosomes lead to the formation of chromosomes3. These chromosomes control vital biological processes like gene regulation, DNA replication, DNA damage response, and cell division1,4. However, aberrations within these processes produce additional genetic elements like extrachromosomal circular DNA (eccDNA)5.
To grasp the concept of eccDNA formation, one can examine the process of cell division5,6. During cell division, DNA replicates itself to ensure the transmission of chromosomes from parent to child cell. Within the replication process, DNA can incur damages, subsequently resulting in the fragmentation of chromosomes. DNA repair mechanisms reassemble these smaller segments and during the reassembling process, apart from chromosomes as a by-product eccDNAs are produced5,6. The lengths of these eccDNAs range from a few hundred to several thousand nucleotides because they are generated through random combinations of multiple segments6. Such fragments often harbor protein-coding genes, further complicating their impact on cellular processes such as gene expression, DNA replication, and DNA damage response
EccDNAs can be classified into two distinct categories based on their size and characteristics: short eccDNAs and long eccDNAs (leccDNA). EccDNAs with shorter lengths typically have tens to a few hundred nucleotides6,7. They are commonly found in the nucleus and cytoplasm of the cell and facilitate movement between genomic loci and drive genetic diversity along with adaptation. Moreover, they store genetic information and replicate independently as episomes. On the other hand, leccDNAs are longer and contain thousands of nucleotides8. LeccDNA are not formed only as the by-product of abnormal DNA replication process, but they can also form due to the recombination events in smaller eccDNA. Recent studies also provide similar evidence in agricultural weed systems9,10 and nuclear genomes11. LeccDNA are found only in the nucleus and contribute to genomic instability, gene amplification, cellular adaptation, and gene expression6. The presence of both types of eccDNA leads to excessive production of specific proteins, including oncogenes, enhancing the cell oncogenic potential and driving uncontrolled cell growth12. Futher, eccDNAs contribute to various diseases in multiple systems, such as glioblastoma, neuroblastoma, irregular immune response, and myocardial infarction6,13,14,15.
Identification of short eccDNA can reveal their roles in gene transfer, and genetic diversity. It is useful in understanding the molecular events of oncogene over-expression and therapeutic resistance. LeccDNA identification provides useful information about indications of genomic instability, genome organization as well as gene regulation. Furthermore, its identification is also useful for unveiling potential mechanisms responsible for the initiation and propagation of diseases such as cancer. Researchers are actively trying to explore its potential as a cancer biomarker and therapeutic resistance indicator7,16.
The identification of eccDNA is accomplished using a variety of wet-lab experimental methods, including pulsed-field gel electrophoresis (PFGE)17, southern blotting18, whole genome sequencing (WGS)19, fluorescence in situ hybridization (FISH)20, RT-PCR18, electron microscopy21, and rolling circle amplification (RCA)22. However, these methods often require prior knowledge or specific probes capable to bind with eccDNA which can limit their applicability to previously characterized eccDNAs. In addition, it is quite laborious, expensive, and time-consuming to identify eccDNAs at a larger scale across different organisms or cells.
The limitations of wet lab based methods and exceptional performance of AI based applications in natural language processing (NLP) tasks, have prompted a marathon of developing AI methods for DNA sequence analysis. Several AI models have been developed for various DNA analysis tasks such as enhancer identification23,24, DNA modification prediction25,26, promoter prediction27, DNA cyclizability prediction28, nucleosome position detection29 and so on. On the other hand, the identification of eccDNA is still being performed through wet lab-based methods due to the deficiency of AI applications for this particular task. According to the best of our knowledge, one predictor named DeepCircle30 is developed for the identification of short eccDNA sequences. There is currently no single predictor available for the identification of leccDNA, and also DeepCircle is not suitable for this specific purpose. The primary obstacle in utilizing DeepCircle for leccDNA identification lies in its reliance on the BERT model30, which can only handle sequence lengths of up to 512 tokens and leccDNA sequences exceed this token limit31.
In order to expedite and enhance research pertaining to the identification of leccDNA, there is an urgent necessity of a robust computational predictor. With an aim to develop a robust and precise computational predictor for leccDNA identification, the contributions of this study are manifold. Following the need for leccDNA identification datasets, it presents 12 benchmark datasets related to leccDNA sequences belonging to 3 different species i.e., Homo sapiens (HM), Arabidopsis thaliana (AT), and Saccharomyces cerevisiae (SC). It presents a robust and precise iLEC-DNA predictor that reaps the benefits of 2 different sequence encoding methods for transforming raw sequences into statistical vectors. Furthermore, to discriminate leccDNA and non-leccDNA sequences, it employs support vector machine (SVM) classifier that extracts more useful discriminative features from statistical vectors having nucleotide distribution patterns and physicochemical properties based information. Furthermore, it compares the performance of proposed predictor with more than 140 baseline predictors and 858 encoder ensembles that are developed by using 13 most widely used sequence encoding methods and 11 machine learning (ML) classifiers. It conducts extensive experimentation over 3 different species datasets to find important answers of the following research questions; I) Do leccDNA sequences exhibit any distinctive nucleotide patterns that distinguish them from non-leccDNA sequences? II) How can variable-length leccDNA sequences be effectively handled to train ML classifiers? III) Which sequence encoding method is more competent in transforming raw leccDNA sequences into statistical vectors by incorporating discriminatory information? IV) Which sequence encoding method demonstrates better performance with which ML classifier? V) Which specific ensemble of sequence encoding methods provide better classification performance? VI) Does the combined potential of the multiple sequence encoding methods enhance the classification efficacy? We believe answers to these questions will provide valuable guidance to the research community when it comes to selecting the optimal combination of encoding methods and classifiers. This will significantly contribute to the creation of an efficient end-to-end predictive pipeline.
Results
Key idea
Over the newly developed 12 benchmark leccDNA datasets, we generate an effective statistical representation based on the gap-kmer distribution and physicochemical properties based information using two sequence encoding methods namely, complementary k-spaced nucleic acid pairs (CKSNAP), and pseudo electron-ion interaction pseudopotentials of trinucleotides (PseEIIP). Using discriminatory features from CKSNAP and PseEIIP we develop a novel predictor based on SVM for leccDNA identification namely, iLEC-DNA. In order to prepare fixed-length leccDNA and non-leccDNA sequences without losing information-rich regions, we perform a thorough intrinsic 2-mer distribution analysis. To validate the observations from the intrinsic analyses, an extrinsic performance analysis is conducted which affirms the information-rich regions that play a critical role in leccDNA identification. In addition, we compared the performance of the proposed predictor with more than 140 baseline 857 advanced predictive pipelines developed based on 13 commonly used sequence encoding methods and 11 ML classifiers. Extensive experimentation shows that the proposed predictor is able to achieve suitable performance across diverse benchmark datasets for leccDNA identification.
Summary of results
This section provides a comprehensive overview of the research objectives pertaining to the prediction of leccDNA. First, it investigates whether leccDNA sequences exhibit nucleotides distinctive patterns that can differentiate them from non-leccDNA sequences. It illustrates and compares the performance values of 143 baseline and 857 advanced predictors across 12 benchmark LeccDNA datasets. Finally, it illustrates the performance values of the proposed leccDNA predictor on 12 benchmark datasets.
RQ I: nucleotide patterns in LeccDNA and non-leccDNA sequences
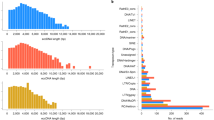
In order to perform DNA sequence classification, ML predictors require uniform length of DNA sequences and distinct nucleotide patterns across various classes. As depicted in Fig. 1, there is considerable variability in the lengths of both leccDNA and non-leccDNA sequences. However, for the purpose of training ML classifiers, these sequences must be of a fixed length. To tackle this issue, one solution involves the direct addition of a padding character ’P’ within sequences. However, the substantial variations in leccDNA sequence lengths, spanning from five to thirty thousand nucleotides necessitate more padding values which introduce noise and bias in data. This influx of padding values not only disrupts the original data distribution but also undermines the model’s potential to generalize effectively on unseen data. In an alternate strategy32, first information-rich regions are explored in the sequences, and padding values are added after truncating all sequences with a specific length threshold. This approach produces fixed-length DNA sequences without introducing substantial bias in data.
With the objective of delineating information-rich regions and obtaining uniform-length DNA sequences, an initial step involves the segmentation of DNA sequences into discrete subsequences. Further, to gain insights into the density and patterns of distinct nucleotide pairs, in 4 different steps an intrinsic analysis is performed. First, the occurrences of 16 unique 2-mers within each subsequence are calculated in leccDNA and non-leccDNA subsequences. Next, The subsequence-based densities of 2-mers are computed separately for leccDNA and non-leccDNA sequences across 10 different sequence lengths. In the subsequent step, the density-based values are normalized with a total number of sequences. Finally, the density differences of 2-mers among leccDNA and non-leccDNA sequences are computed to reveal distinctive nucleotide patterns. Details related to information-rich region analyses are provided in Supplementary Sect. S1.
Figure 2 shows the subsequence density-based differences of different 2-mers across 3 different leccDNA benchmark datasets namely, C4-2B, AT, and YS. The majority of density-based differences lie within the initial subsequences, whereas among the last subsequences, the densities of 2-mers are similar among leccDNA and non-leccDNA sequences. These differences in the densities of specific nucleotide pairs indicate that certain regions of the DNA sequences exhibit distinct nucleotide distribution patterns, which are characteristic of leccDNA sequences and can differentiate leccDNA from non-leccDNA sequences. Particularly certain 2-mers such as TT, TC, TA, AT, AC, GG, GC, and GA, have notably higher densities in leccDNA sequences, and 2-mers i.e., AG, GC, TG, CA, GA, and TC, show higher densities in non-leccDNA sequences. These distinguishing factors are captured with the help of specific sequence encoding methods and can be utilized for the identification of leccDNA sequences. In addition, similar patterns and 2-mer density-based differences across leccDNA and non-leccDNA sequences of other benchmark datasets are provided in the Supplementary File.
The intrinsic nucleotide pattern analysis affirms that leccDNA sequences display discernible nucleotide patterns that differentiate them from non-leccDNA sequences. These distinguishing features are predominantly situated in the initial regions of leccDNA sequences which are further investigated in the subsequent extrinsic performance analyses, as elaborated in the following subsection.
RQ II: handling variable length of LeccDNA sequences
With an aim to analyze the impact of 5 different regions (1000, 2000, 3000, 4000, and 5000) in leccDNA identification, using 13 encoding methods and 11 classifiers based 143 predictive pipelines an extrinsic performance outcome is discussed in this section.
Figure 3 illustrates top performing baseline predictors ACC values, across 5 different sequence lengths for 12 distinct benchmark datasets under independent test setting. Two different types of performance trends are observed across benchmark leccDNA datasets with respect to sequence lengths i.e., (I) steady linear performance increase with sequence length, (II) linear performance increase up to a specific sequence length. The leccDNA benchmark datasets namely, C4-2B, FAD, and SP lie under the trend category I as they have a gradual increase in performance with respect to the increase in the sequence length. These benchmark datasets have maximum performance value at the sequence length of 5000. In addition, the rest of the benchmark datasets fall under the trend category II, as the performance values increase up to a certain sequence length and afterward the performance decreases. For instance, OVCAR8, PC, and FaDu show a gradual increase in performance up to the 3000 nucleotides and a performance decline afterward. In addition, OP, UR, and AT show similar patterns until 2000 and PL, and MS show performance improvement up to 4000 nucleotides after which the performance deteriorates.
It is important to note that while dealing with leccDNA sequences, extracting meaningful information from subsequences that contain initial sequence nucleotides proves to be advantageous in achieving optimal classification efficacy. This highlights the potential benefits of breaking down complex sequences into manageable chunks, allowing for better classification performance and more efficient analysis. Furthermore, the superior performance of the baseline predictors for the initial sequence lengths reinforces the previously discussed observations that most of the discriminative patterns related to nucleotides are concentrated in the initial regions of leccDNA sequences.
RQ III and IV: effectiveness of sequence encoding methods
This section briefly addresses research questions (III and IV) pertaining to the optimal sequence encoding methods and ML classifiers for effective leccDNA identification. To achieve this, two analyses are conducted here, first the performance rank scores of each sequence encoding method are calculated across diverse classifiers on 7 datasets. Additionally, the rank scores of classifiers are computed to identify consistently superior classifiers across all sequence encoding methods. Figure 4a shows the rank scores of 13 different sequence encoding methods with 11 ML classifiers across 7 datasets namely, AT, C4-2B, OP, OV, PL, UR, and YS. The rank scores are computed by determining the maximum performance of a sequence encoding method across a classifier for different datasets.
Three distinct categories of sequence encoders are established based on performance rank scores: (I) encoders with the lowest rank scores, (II) encoders with inconsistent performance across certain classifiers, (III) encoders with consistent performance across all classifiers. Among 13 sequence encoding methods, DNC, NAC, TNC, and EIIP fall under category I as these methods consistently show the lowest rank scores across 11 different ML classifiers. This attributes to the limited discriminatory power of these methods in capturing relevant patterns, nuanced variations, and characteristics in leccDNA sequences. Similarly, ANF, ENAC, NCP, PSEDNC and binary fall under category II as these methods show consistent ranks scores across few ML classifiers such as K-nearest neighbor (KNN), adaptive boosting (AB), decision tree (DT), Naive Bayes and linear discriminant analysis (LDA). Particularly, two sequence encoders namely, CKSNAP and SCPSEDNC lie under category III as they show consistent rank scores across majority of the classifiers. The consistent performance of these methods lies with their ability to efficiently capture nucleotide distribution and physicochemical properties that enable ML classifiers to identify leccDNA sequences with more efficacy as compared to the other sequence encoding methods.
Figure 4b shows the rank scores of 11 different ML classifiers with 13 distinct sequence encoding methods across AT, C4-2B, OP, OV, PL, UR, and YS datasets. Notably, the KNN classifier consistently exhibits the lowest predictive performance across all the sequence encoding methods. Similarly, AB classifier demonstrates inconsistent and comparatively lower performance ranks across all sequence encoding methods. Various classifiers including gradient boosting (GB), bagging, extra-trees (ET), DT, and NB exhibit relatively inconsistent performance rank scores, as they have marginal rank scores only with EIIP, NCP, and binary sequence encoding methods. On the other hand, SVM, LDA, random forest (RF) and logistic regression (LR) classifiers stand out with the highest rank scores, showing their effectiveness in the identification of leccDNA sequences with diverse sequence encoding methods.
(a) Unraveling the top-ranked scores of 13 sequence encoding methods across 11 classifiers over multiple datasets (AT, C4-2B, OP, OV, PL, UR, and YS) and sequence lengths: 1000, 2000, 3000, 4000, and 5000. (b) Exploring the ranking scores of 11 ML classifiers on 13 sequence encoding methods, shedding light on their comparative performance.
RQ V: the combined potential of multiple sequence encoding methods
To address research question V, we explore the potential of 11 ML classifiers in conjunction with 78 combinations of 13 sequence encoders. The primary aim of this analysis is to identify a combination of sequence encoders that consistently demonstrate high performance in leccDNA identification. Specifically, we examine 78 unique combinations of sequence encoders paired with the 11 ML classifiers which results in approximately 858 results for a single leccDNA benchmark dataset. Additionally, due to the substantial training time involved, the analysis spans 9 different leccDNA benchmark datasets: AT, YS, C4-2B, FaDu, FAD, PC, UR, OVCAR8, and OP. Notably, 3 datasets MS, SP, and PS have been excluded from this analysis due to their large number of samples and sequence sizes.
Table 1 summarizes the average performance measures for the top 10 sequence encoders and classifier combinations which are selected based on ACC. Among 858 total combinations, CKSNAP with physicochemical properties-based sequence encoders exhibit consistent performance across 9 diverse datasets. In general, the combination of CKSNAP and the PseEIIP encoder shows the maximum performance in terms of 5 out of 6 distinct evaluation measures i.e., ACC, SP, F1, MCC, and AUC-ROC. Particularly, CKSNAP-PseEIIP combination achieves superior performance as compared to 19 other top performing combinations with average performance margins of 4.583 % in terms of ACC, 5.436% over SP, 4.451% in terms of F1, 9.71% over MCC and 4.58% over AUC-ROC. It is important to mention that the combination of CKSNAP and PseEIIP produces maximum performance with SVM classifier. Hence, CKSNAP-PseEIIP combination and SVM are selected for the final performance analyses across all benchmark leccDNA datasets. Finally, 858 different results for 78 encoder combinations across 11 ML classifiers in terms of 6 distinct evaluation measures are provided in Supplementary Files.
RQ VI: the combined potential of CKSNAP and PseEIIP
To find the answer to research question VI, we explore the potential of CKSNAP and PseEIIP with SVM classifier. Here the objective is to analyze whether SVM produces better performance with statistical vectors of standalone encoders or with their combined statistical vectors. Figure 5a–i showcases the evaluation results based on performance values generated by standalone and combined statistical representations with SVM classifiers. It highlights the performance gains achieved through the utilization of diverse discriminatory features from CKSNAP and PseEIIP with SVM classifier across 9 different leccDNA benchmark datasets.
Out of 9 different benchmark leccDNA datasets, the combined potential of CKSNAP and PseEIIP shows performance improvements over 7 different datasets except AT and FaDu. Particularly, it shows average perfromance gains of 1.5 % in terms of ACC, 1.466 % and 1.97 % across SN and SP, 1.457% and 2.88% in terms of F1 and MCC, and 1.50% across AUCROC. Particularly, in terms of AT and FaDu datasets, CKSNAP along with SVM classifier manages to produce better performance as compared to CKSNAP and PseEIIP combination across 6 distinct evaluation measures i.e., ACC, SP, SN, MCC, F1, and AUC-ROC.
Overall, it is observed that both sequence encoding methods provide unique and discriminatory information to the classifiers for leccDNA identification. This discriminatory and unique information when presented to the ML classifiers in a concatenated way leads to signification performance gains which suggests the importance of using multiple sets of information while training a classifier for leccDNA identification. Therefore, the final experimentation over leccDNA benchmark datasets is performed by utilizing SVM, CKSNAP, and PseEIIP.
Performance analyses over 5-fold cross validation
Figure 6 illustrates the performance results of the proposed leccDNA predictor in terms of 5-fold cross-validation across 12 benchmark leccDNA datasets. The proposed predictor shows high-performance values over the dataset of SP, OVCAR8, and YS ranging from 90-94\(\%\) in terms of ACC, and AUC-ROC. For OVCAR8, SP and YS datasets there is an average gap of 3.73% in terms of SP and SN, which suggests that the proposed predictor is less prone to type I and type II error. This implies that the proposed leccDNA predictor is quite robust in predicting samples belonging to positive and negative classes. The high performance of the proposed predictor is due to the sufficient number of samples present to train the proposed predictor.
In addition, across the datasets of OP, FaDu, PL, MS, UR, C4-2B, and FAD, the performance of the proposed predictor ranges from 75-82% in terms of ACC and AUC-ROC. The proposed predictor is not highly prone to type I and type II errors due to an average gap of 4% among SP and SN values. Moreover, the proposed predictor shows low performance on the PC and AT datasets with the performance values ranging from 69-73% in terms of ACC and AUC-ROC. In addition, over both of the datasets, the proposed predictor is prone to either type I or type II error due to an average gap of more than 5% in terms of SP and SN and lower AUC-ROC. The predictor is more prone to type II error as it is not able to successfully identify positive samples with a higher ratio as compared to the negative samples. The low performance of the proposed predictor on these datasets is due to the presence of a limited number of samples for positive and negative class samples (200 leccDNA sequences for AT and 400 leccDNA sequences in terms of PC).
Performance analyses over independent test set
Figure 7 illustrates the performance analyses over 6 distinct evaluation measures across 12 different benchmark datasets in terms of independent test sets. A closer look at the performance values at the 5-fold validation and independent test sets reveals that the performance of the proposed predictor either remains the same, increases, or decreases as compared to the 5-fold validation across various datasets. The performance of the proposed predictor remains the same across 6 different datasets such as FaDu, FAD, MS, YS, PL, and OVCAR8. Similarly, there is a slight decrease in the performance of the proposed predictor over C4-2B dataset, and an increase in the performance over 4 datasets namely, MS, PC, AT and OP.
Webserver
This article performs experimentation on 12 benchmark leccDNA datasets of 3 different species. To facilitate readers, we have provided all 12 benchmark datasets in the download section of our leccDNA prediction web application (https://sds_genetic_analysis.opendfki.de/iLEC_DNA/). In addition, users can train iLec-DNA on different datasets by using the training module of the web application.
Discussion
Intrinsic and extrinsic performance analyses of experimental results reveal that initial regions of leccDNA sequences carry significant discriminatory information for leccDNA identification. In addition, experimental results on 12 benchmark datasets from 3 different species, reveal that among 13 diverse types of encoding methods, two encoders CKSNAP and PseEIIP generate more comprehensive statistical vectors. A prime reason behind generating better statistical vectors is the extraction of both simple as well as gap-based nucleotide patterns. Specifically, CKSNAP encoder transforms raw DNA sequences into statistical vectors by computing occurrence distribution of simple as well as gap-based bimers. PseEIIP encoder makes used of predefined electron ionic pseudo potentials of trinucleotides, along with their frequency. In a nutshell, it can be concluded that those encoding methods are more suitable for transforming raw DNA sequences into statistical vectors that emphasize on gap-based nucleotides distribution. Furthermore, the ensembling of both encoders generates better statistical vectors. Primarily, the concatenation of statistical vectors of both encoders facilitates the extraction of two different types of features, gap based bimers distribution and gap-based nucleotides correlational information extracted through physicochemical properties. Among 11 different classifiers, SVM remains top performer because it finds optimal hyperplanes for discriminating sequences into leccDNA and non-leccDNA classes. Experimental results reveal that across all species benchmark datasets, it successfully designed optimal hyperplanes. Although several classifiers such as RF, LDA, and LR produce SVM comparable performance on a few datasets, overall they fail to produce consistent performance across all datasets.
Although the incorporation of distinct physicochemical, and nucleotide distribution information achieves a notable reduction in prediction errors across diverse leccDNA datasets, the robustness and efficacy of the proposed model are limited over the AT and PC datasets due to a bias towards type II errors. The proposed predictor suffers from this problem due to the availability of a lower number of leccDNA sequences across these two benchmark datasets. In the future, we intend to leverage additional sequence encoding methods, feature selection methods and incorporate certain deep learning models to enhance the classification efficacy and robustness across diverse leccDNA datasets. Moreover, by following the criteria of existing sequence analysis tools, hyperparameters of ML models can be optimized to further improve the predictive performance33,34,35,36,37.
Conclusion
The primary objective of this study is to introduce AI-based web application that can accurately predict leccDNA in various cell types and species. With an aim to develop a powerful web application, it presents 12 benchmark datasets that are utilized to evaluate the performance of leccDNA predictive pipelines and for the development of web application. In addition, the unique distribution of nucleotides is explored with an aim to decode the discriminatory potential in leccDNA sequences. To design a robust and precise leccDNA predictive pipeline, it explores the potential of 13 different sequence encoding methods in conjunction with 11 ML classifiers. Comprehensive experimentation across 12 benchmark datasets reveals that SVM classifier and 2 sequence encoding methods namely, PseEIIP and CKSNAP give superior and consistent performance across diverse leccDNA datasets. Furthermore, the concatenation of statistical vectors generated through CKSNAP and PseEIIP leads to significant performance gains. On top of the proposed predictor, the web application is developed that will facilitate biological researchers for conducting more comprehensive research for leccDNAs. In future, we will further enhance the scope of application by collecting data related to other species such as Drosophila (DM), Chimpanzee (CH), and Mouse (MM). We will perform cross-species analysis, where the model is trained on one species (AT, YS, DM, CH, MM) and evaluated on other species (HM), this will help in identifying leccDNA from other species at a larger scale. Based on various performance analysis, iLEC-DNA, a novel predictor for leccDNA, is proposed that captures discriminatory information through pseudo ionic potentials and nucleotide distribution information. iLEC-DNA is evaluated over 12 distinct benchmark datastes namely, MS, PS, SP, FaDU, FAD, PC, UR, CB, OV, OP, AT and YS. iLEC-DNA is a valuable tool for researchers examining intricate and lengthy eccDNA. Its capabilities can enable the exploration of leccDNA and their involvement in genomic instability and the onset of cancer.
Materials and methods
This section demonstrates comprehensive details of proposed and baseline predictors. It provides a comprehensive overview of benchmark datasets development process and characteristics of datasets. Finally, it presents evaluation measures that are used to evaluate and compare the performance of proposed and baseline predictors.
Graphical illustration of Benchmark datasets development process, proposed and baseline predictors. In datasets development process, leccDNA sequences are extracted from the eccDNA atlas and CD-HIT is utilized to remove redundant leccDNA sequences. Subsequently, USHUFFLE is applied to generate negative samples. In the second step, leccDNA sequences are converted into statistical vectors through baseline and proposed sequence encoding pipelines. In the classification and evaluation process, the performance of the proposed predictor is compared with the baseline predictor across all datasets.
Summary of the LeccDNA identification predictive pipeline
Figure 8 demonstrates different modules of the proposed iLEC-DNA predictor. It can be seen that after the datasets development process, DNA sequences are transformed into statistical vectors. The transformation of DNA sequences into statistical vectors is an essential task because AI predictors can only process numerical data and cannot operate directly on DNA sequences. While converting DNA sequences into statistical vectors the prime objective is to incorporate sequence order, semantic, nucleotide distribution, and positional features into the statistical vectors. First, the potential of 13 sequence encoding methods along with 11 different ML classifiers is explored to identify the most consistent sequence encoder and ML classifier from 143 total combinations. In the subsequent step, 858 ensembles of encoders (78 encoder combinations \(\times\) 11 ML classifiers) are created to reap the benefits of discriminative and unique information from multiple encoders. Our analysis shows that leccDNA sequences can be converted to discriminative statistical vectors by reaping the combined benefits of two different types of sequence encoding methods namely, CKSNAP and PseEIIP . Finally, the concatenated representations are passed to SVM classifier which shows superior performance as compared to any other combination. A comprehensive description of both sequence encoding methods and their concatenation is provided following subsections.
Benchmark datasets
In the pursuit of creating effective and reliable machine learning (ML) predictors for any biological sequence analysis tasks, the selection of appropriate data is a crucial task38. Inappropriate data can lead to the development of a biased and unreliable predictor that results in misleading insights and flawed decision-making.
EccDNA sequences are available across various databases such as eccDNA Atlas39, TeCD40, EccBase41, EccDB42, and EccDNADB43. Each database includes extrachromosomal DNA sequences of different species and cells. Among all databases, eccDNA Atlas39 offers a vast and comprehensive collection of eccDNA sequences derived from diverse organisms and experimental techniques. This extensive coverage ensures a broader representation of eccDNA diversity, enabling researchers to access a more complete picture of eccDNA characteristics across different species and experimental conditions.
To prepare leccDNA identification data, first necessary details such as specie, tissue, cell, isolate, genome version, and genomic coordinates, related to leccDNA sequences are acquired from eccDNA atlas database. Specifically, the genomic coordinates encompass information such as chromosome number, start and end positions of the leccDNA sequences. In addition, the genome versions/assemblies are downloaded from Refseq (https://www.ncbi.nlm.nih.gov/refseq/)44, namely araport11, S288C, and hrc38. In the subsequent step, the genomic coordinates and assemblies are utilized to retrieve relevant leccDNA sequences for diverse types of species. A summary of statistics related to obtained eccDNA sequences of 3 different species i.e., Saccharomyces cerevisiae (SC), Arabidopsis thaliana (AT), and Homo sapiens (HM) is presented in Table 2.
Table 2 provides statistics of 12 different benchmark leccDNA datasets. Among 12 benchmark leccDNA datasets, 10 datasets belong to different cell lines of HM namely, muscle (MS), plasma (PS), sperm (SP), FaDU, FaDU-DDP (FAD), placenta (PC), urine (UR), C4-2B (CB), ovcar-8 (OV), and ovary-primary (OP). Due to the low number of samples in terms of tissues/isolates of species AT and YS, only two datasets are formulated from them.
After the retrieval of leccDNA sequences, the leccDNA sequences may contain redundant or highly similar sequences. However, these similarities can introduce a bias when dividing the data into training and testing sets, leading to an overestimation of the model performance and the establishment of impractical benchmarks. To create reliable and comprehensive benchmark datasets, following previous studies45,46,47 we apply CD-HIT (sequence similarity >60\(\%\) ) to positive samples, where redundant or highly similar sequences are clustered together, resulting in a representative subset that encompasses the essential sequence variations. This process helps to prevent the over-representation of certain sequences, which could introduce biases during the training process of the ML classifier.
There are multiple ways to generate negative data samples for a DNA sequence classification task i.e., selection of sequences from genomic background48,49, and nucleotide shuffling49,50. For instance, sequences are randomly sampled from different positions of a genome to get a diverse pool of negative samples that are non-overlapping to the positive samples. In addition, sometimes negative samples are clustered with positive samples using psi-cd-hit to remove closely related positive and negative samples. In spite of its usage, this method has various disadvantages i.e., compositional bias, where the distribution of nucleotides in negative samples might differ completely as compared to positive samples which may lead to biased training of the ML models. In comparison, nucleotide shuffling tackles such problems by preserving various k-mers counts. Ushuffle is designed to preserve the statistical properties and local sequence features of the input sequences while removing specific sequence motifs and patterns. Following the existing work49, fasta\(\_\)ushuffle (k=2)(https://github.com/agordon/fasta_ushuffle) is utilized to shuffle nucleotides in positive samples to obtain suitable negative samples.
Complementary K-spaced nucleic acid pairs (CKSNAP)
CKSNAP encoder was proposed by Zhang et al.51 and has been widely used used in diverse types of DNA sequence classification predictors including, enhancer prediction52, DNA replication origin identification53, DNA modification prediction54 and promoter prediction55. The motivation behind the development of this encoder was to capture nucleotide occurrence distribution patterns with different gap values. CKSNAP56 generates gap based bimers such as for a hypothetical sequence GCTA, with gap value 1, gap-based bimers are be G-T and C-A. Similarly, for gap value 2, it first generates 1 gap-based bimers and then 2 gap-based bimers. Furthermore, for each gap value, it computes occurrence frequencies of bimers and normalizes them with total number of gap-kmers. Mathematically, CKNSAP can be written as;
where, N\(_{AA}\) represents the total occurrences of bimer AA in the DNA sequence and N\(_{total}\) denotes the total number of gap bimers. A detailed working paradigm of CKSNAP is shown in Fig. 9.
Electron-ion interaction pseudopotential of trinucleotides (PseEIIP)
Nair et al.,57 proposed electron ionic potential of 4 different nucleotides i.e., A, G, C, T (A: 0.1260, C: 0.1340, G: 0.0806, T:0.1335). These values represent the potential energy of the electrons and ions present in the atoms of the nucleotide. PseEIIP incorporates electron ionic potentials and nucleotide frequency of trinucleotides and converts a DNA sequence into a statistical representation.
To generate statistical representation through PseEIIP, first 3-mers dictionary is computed from a DNA sequence,
In the subsequent step, these values are normalized based on the total number of 3-mers in the DNA sequence,
where \(T_k\) denotes the total number of 3-mers in a DNA sequence. In addition, electron ionic potential values of 3-mers are summed and multiplied with corresponding normalized frequencies i.e.,
whereas, \(E_{AAA}\) represents the sum of ionic potentials for A, and \(E_{AAA}\) denotes the sum of ionic potentials for A, T, and A.
Feature fusion
Feature fusion involves the integration of diverse types of sequence information into a single vector which can improve the discriminative potential of statistical vectors and the efficacy of an AI predictor. Diverse types of feature fusion methods have been utilized to improve the performance of various sequence analysis tasks such as DNA hypersensitive site prediction58, DNA modification prediction59, promoter prediction60, and DNA binding proteins identification61.
In pursuit of harnessing the combined benefits of the two distinct sequence encoding methods, an early fusion strategy based on vector concatenation is adopted in the proposed iLEC-DNA predictor. Let \(\overrightarrow {X}\) and \(\overrightarrow {Y}\) be represented as statistical vectors of dimensions P and Q for a given sequence S:
Subsequently, the fused vector can be expressed as:
where, \(\overrightarrow {F}\) represents p\(+\)q dimensional fused vector.
Baseline predictors
This section summarizes 11 remaining encoders namely, nucleic acid composition (NAC)62, enhanced nucleic acid composition (ENAC)63, accumulated nucleotide frequency (ANF)64, dinucleotide composition (DNC)65, trinucleotide composition (TNC)66, nucleotide chemical property (NCP)67, binary68, electron ionic interaction potential (EIIP)57, series correlation pseudo dinucleotide composition (SCPseDNC),69, pseudo dinucleotide composition (PSEDNC)70,71, and pseudo k-tupler composition (PSEKNC)72.
Nucleic acid composition (NAC)62 computes the normalized frequency of each nucleotide across the DNA sequence. The normalization is done through the total length of the DNA sequence. Similarly, dinucleotide composition (DNC)65 and trinucleotide composition (TNC)66, use the pairs of nucleotides (k = 2, or k = 3) to compute normalized occurrence frequencies rather than taking into account individual nucleotides. Enhanced nucleic acid composition (ENAC)63 transforms raw sequences into statistical vectors by counting the number of different k-mers at a fixed sliding window. First, a dictionary of unique k-mers is created and then for each unique each k-mer, within each window its count is computed. This step is repeated by sliding over the DNA sequences with a step size of \(W_S\). In the end, all the count dictionaries are concatenated together to form a discriminative statistical vector.
Accumulated nucleotide frequency (ANF)64 encodes nucleotide frequency information in the statistical vectors. First, it computes the position-wise counts of nucleotides and then normalizes it with the position of nucleotides. Then, it represents each nucleotide with a 4-dimensional vector at each position, where the first three values indicate the presence or absence of a specific nucleotide, and the last value is the normalized positional density of that specific nucleotide. In the binary68 sequence encoding method, each nucleotide is represented by a vector of size 4. These vectors include ones and zeros with each one representing the presence of a specific nucleotide.
The nucleotides of the DNA have different chemical structures and chemical properties. Physicochemical properties-based sequence encoding methods make use of such information to capture discriminative information from the raw DNA sequences. Nucleotide chemical property (NCP)67, converts DNA sequences into statistical vectors based on the ring structure, functional group, and hydrogen group where each nucleotide is represented by a 3-dimensional vector. Electron-Ion Interaction Potential (EIIP)57 makes use of numerical values based on the average interaction potential between nucleotides constituent atoms, and electrons. It converts DNA sequences into statistical vectors by substituting each nucleotide with the predefined ionic potential value. Electron-ion interaction pseudopotentials of trinucleotide (PseEIIP)69 utilizes electronic ionic potential values of trinucleotides and their normalized occurrence frequency. For a trinucleotide, first the ionic potential is computed by summing up the individual pseudo-ionic potential values of three nucleotides which is multiplied by the normalized occurrence frequency of that specific trinucleotide.
Pseudo dinucleotide composition (PseDNC)70,71 makes use of six distinct DNA properties i.e., twist, roll, rise, tilt, shift, and slide, along with the frequencies of the nucleotide pairs. First, normalized occurrence frequencies of nucleotide pairs are computed which encode the contiguous local sequence-order information of the DNA sequence. To include the global sequence-order information, a set of correlation functions are computed among the neighboring nucleotides. These functions are computed by taking the mean over the difference among the nucleotide pairs property values. The output of pseDNC is a (16+\(\lambda\))-D vector, where the first 16 values represent the normalized frequencies of nucleotide pairs and the rest are higher-order correlation functions. Pseudo k-tupler Composition (PseKNC)72 works on a similar principle but the difference lies in K-tuple composition used in PseKNC. Rather than dealing only with dinucleotides or trinucleotides, PseKNC makes use of K = (1 \(\ldots\), L), to compute statistical vectors that contain higher and lower order features.
Evaluation measures
Following evaluation criteria of existing DNA sequence classification predictors23,24,25,27, we evaluate proposed and baseline predictors using five different evaluation measures namely, accuracy (ACC), sensitivity (SN), specificity (SP), Mathews correlation coefficeint (MCC), and area under the receiver operating curve (AUC-ROC).
Accuracy26 refers to the proportion of correct predictions with respect to the total predictions. Specificity or true negative rate (TNR)23 is the model’s ability to correctly predict the negative class samples. It is determined by dividing the number of correct negative predictions by the total number of true negatives. Sensitivity (or recall)26 measures the ability of the model to predict positive class samples by taking the ratio of correct positive predictions to the predictions on positive samples. MCC73 calculates the correlation between the model predictions and the true class, by taking into consideration true positives, true negatives, false positives, and false negatives. AUC-ROC74 computes the degree of separability of the model based on the true positive rate (TPR) and true negative rate (TNR) at various thresholds.
In the mathematical expression above, \(T^+\) and \(T^-\) denote the true predictions related to positive and negative classes, whereas \(F^+\) and \(F^-\) are the incorrect predictions related to the positive and negative classes respectively.
Experimental setup
To prepare benchmark datasets, we utilize two different APIs namely, Biopython75 and USHUFFLE50. The proposed and baseline predictive pipelines are developed on top of two libraries namely, iLearnPlus76 (https://ilearnplus.erc.monash.edu/) and Scikit-Learn77 v1.3.277 (https://scikit-learn.org/stable/ Following the evaluation criteria of existing DNA sequence classification predictors23,24,25,27, we perform experimentation in two different settings namely, 5-fold cross-validation and independent test set. All visualizations are generated using matplotlib v3.8.078 (https://matplotlib.org/). The parameter values for 11 different ML classifiers are provided in Table 3.
Data availability
The datasets generated and analysed during the current study are available in the following Github repository Extrachrosmosomal-DNA-Prediction, [https://github.com/FAhtisham/Extrachrosmosomal-DNA-Prediction].
References
Miller, O. J. & Therman, E. Human Chromosomes (Springer Science & Business Media, 2011).
Davis, L. Basic Methods in Molecular Biology (Elsevier, 2012).
Chaffey, N. et al. Molecular Biology of the Cell 4th edn. (Springer, 2003).
Sumner, A. T. Chromosomes: Organization and Function (Wiley, 2008).
Zuo, S. et al. Extrachromosomal circular dna (eccdna): From chaos to function. Front. Cell Dev. Biol. 9, 792555 (2022).
Zhao, Y., Yu, L., Zhang, S., Su, X. & Zhou, X. Extrachromosomal circular dna: Current status and future prospects. eLife 11, e81412. https://doi.org/10.7554/eLife.81412 (2022).
Ling, X. et al. Small extrachromosomal circular dna (eccdna): Major functions in evolution and cancer. Mol. Cancer 20, 1–15 (2021).
Paulsen, T., Kumar, P., Koseoglu, M. M. & Dutta, A. Discoveries of extrachromosomal circles of dna in normal and tumor cells. Trends Genet. 34, 270–278 (2018).
Koo, D.-H. et al. Extrachromosomal circular dna-based amplification and transmission of herbicide resistance in crop weed amaranthus palmeri. Proc. Natl. Acad. Sci. USA 115, 3332–3337 (2018).
Molin, W. T., Yaguchi, A., Blenner, M. & Saski, C. A. The eccdna replicon: A heritable, extranuclear vehicle that enables gene amplification and glyphosate resistance in amaranthus palmeri. Plant Cell 32, 2132–2140 (2020).
Spier Camposano, H., Molin, W. T. & Saski, C. A. Sequence characterization of eccdna content in glyphosate sensitive and resistant palmer amaranth from geographically distant populations. PLoS ONE 17, e0260906 (2022).
Li, R., Wang, Y., Li, J. & Zhou, X. Extrachromosomal circular dna (eccdna): An emerging star in cancer. Biomark. Res. 10, 1–13 (2022).
Wang, Y. et al. eccdnas are apoptotic products with high innate immunostimulatory activity. Nature 599, 308–314 (2021).
Møller, H. D. et al. Circular dna elements of chromosomal origin are common in healthy human somatic tissue. Nat. Commun. 9, 1069 (2018).
Rosswog, C. et al. Chromothripsis followed by circular recombination drives oncogene amplification in human cancer. Nat. Genet. 53, 1673–1685 (2021).
Yan, Y. et al. Current understanding of extrachromosomal circular dna in cancer pathogenesis and therapeutic resistance. J. Hematol. Oncol. 13, 1–16 (2020).
Mouakkad-Montoya, L. et al. Quantitative assessment reveals the dominance of duplicated sequences in germline-derived extrachromosomal circular dna. Proc. Natl. Acad. Sci. USA 118, e2102842118 (2021).
Wang, K. et al. Deciphering extrachromosomal circular dna in arabidopsis. Comput. Struct. Biotechnol. J. 19, 1176–1183 (2021).
Zhu, Y. et al. Whole-genome sequencing of extrachromosomal circular dna of cerebrospinal fluid of medulloblastoma. Front. Oncol. 12, 934159 (2022).
Decarvalho, A. C. et al. Discordant inheritance of chromosomal and extrachromosomal dna elements contributes to dynamic disease evolution in glioblastoma. Nat. Genet. 50, 708–717 (2018).
Lahey, J. & Chaudhry, M. A. Detection of Extrachromosomal Circular dna (eccdna) in Ionizing Radiation Exposed Cells (2014).
Diaz-Lara, A., Gent, D. H. & Martin, R. R. Identification of extrachromosomal circular dna in hop via rolling circle amplification. Cytogenet. Genome Res. 148, 237–240 (2016).
Zhang, T., Li, L., Sun, H. & Wang, G. Deepiteh: A deep learning framework for identifying tissue-specific ernas from the human genome. Bioinformatics 39, btad375 (2023).
Kleftogiannis, D., Kalnis, P. & Bajic, V. B. Progress and challenges in bioinformatics approaches for enhancer identification. Brief. Bioinform. 17, 967–979 (2016).
Nabeel Asim, M., Ali Ibrahim, M., Fazeel, A., Dengel, A. & Ahmed, S. Dna-mp: A generalized dna modifications predictor for multiple species based on powerful sequence encoding method. Brief. Bioinform 24, bbac546 (2023).
Zeng, W., Gautam, A. & Huson, D. H. Mulan-methyl-multiple transformer-based language models for accurate dna methylation prediction. bioRxiv 2023–01 (2023).
Oubounyt, M., Louadi, Z., Tayara, H. & Chong, K. T. Deepromoter: Robust promoter predictor using deep learning. Front. Genet. 10, 286 (2019).
Li, K., Carroll, M., Vafabakhsh, R., Wang, X. A. & Wang, J.-P. Dnacycp: A deep learning tool for dna cyclizability prediction. Nucleic Acids Res. 50, 3142–3154 (2022).
Fazeel, A., Agha, A., Dengel, A. & Ahmed, S. A Two-staged Bert Based Nucleosome Positioning Prediction Architecture for Multiple Species (Np-bert, 2023).
Chang, K.-L. et al. Short human eccdnas are predictable from sequences. Brief. Bioinform. 24, bbad147 (2023).
Devlin, J., Chang, M.-W., Lee, K. & Toutanova, K. Bert: Pre-training of deep bidirectional transformers for language understanding. arXiv preprint arXiv:1810.04805 (2018).
Asim, M. N., Ibrahim, M. A., Malik, M. I., Dengel, A. & Ahmed, S. Adh-ppi: An attention-based deep hybrid model for protein-protein interaction prediction. Iscience 25, 105169 (2022).
Ahmad, S. et al. Scorpion is a stacking-based ensemble learning framework for accurate prediction of phage virion proteins. Sci. Rep. 12, 4106 (2022).
Charoenkwan, P. et al. Amypred-frl is a novel approach for accurate prediction of amyloid proteins by using feature representation learning. Sci. Rep. 12, 7697 (2022).
Charoenkwan, P. et al. Sapphire: A stacking-based ensemble learning framework for accurate prediction of thermophilic proteins. Comput. Biol. Med. 146, 105704 (2022).
Hongjaisee, S., Nantasenamat, C., Carraway, T. S. & Shoombuatong, W. Hivcor: A sequence-based tool for predicting hiv-1 crf01_ae coreceptor usage. Comput. Biol. Chem. 80, 419–432 (2019).
Charoenkwan, P., Chotpatiwetchkul, W., Lee, V. S., Nantasenamat, C. & Shoombuatong, W. A novel sequence-based predictor for identifying and characterizing thermophilic proteins using estimated propensity scores of dipeptides. Sci. Rep. 11, 23782 (2021).
Libbrecht, M. W. & Noble, W. S. Machine learning applications in genetics and genomics. Nat. Rev. Genet. 16, 321–332 (2015).
Zhong, T. et al. eccdna atlas: A comprehensive resource of eccdna catalog. Brief. Bioinform 24, bbad037 (2023).
Guo, J., Zhang, Z., Li, Q., Chang, X. & Liu, X. Tecd: The eccdna collection database for extrachromosomal circular dna. BMC Genom. 24, 1–10 (2023).
Sun, H., Lu, X. & Zou, L. Eccbase: A high-quality database for exploration and characterization of extrachromosomal circular dnas in cancer. Comput. Struct. Biotechnol. J. 21, 2591–2601 (2023).
Yang, M. et al. eccdb: A comprehensive repository for eccdna-mediated chromatin contacts in multi-species. Bioinformatics 39, btad173 (2023).
Peng, L., Zhou, N., Zhang, C.-Y., Li, G.-C. & Yuan, X.-Q. eccdnadb: A database of extrachromosomal circular dna profiles in human cancers. Oncogene 41, 2696–2705 (2022).
O’Leary, N. A. et al. Reference sequence (refseq) database at ncbi: Current status, taxonomic expansion, and functional annotation. Nucleic Acids Res. 44, D733–D745 (2016).
Salem, M., Keshavarzi Arshadi, A. & Yuan, J. S. Ampdeep: Hemolytic activity prediction of antimicrobial peptides using transfer learning. BMC Bioinform. 23, 1–17 (2022).
Ullah, W. et al. Splicing sites prediction of human genome using machine learning techniques. Multimed. Tools Appl. 80, 30439–30460 (2021).
Zhang, Y. & Hamada, M. Deepm6aseq: Prediction and characterization of m6a-containing sequences using deep learning. BMC Bioinform. 19, 1–11 (2018).
Lee, D. et al. A method to predict the impact of regulatory variants from dna sequence. Nat. Genet. 47, 955–961 (2015).
Krützfeldt, L.-M., Schubach, M. & Kircher, M. The impact of different negative training data on regulatory sequence predictions. PLoS ONE 15, e0237412 (2020).
Jiang, M., Anderson, J., Gillespie, J. & Mayne, M. ushuffle: A useful tool for shuffling biological sequences while preserving the k-let counts. BMC Bioinform. 9, 1–11 (2008).
Zhang, W. et al. Prediction of methylation sites using the composition of k-spaced amino acid pairs. Protein Peptide Lett. 20, 911–917 (2013).
Basith, S., Hasan, M. M., Lee, G., Wei, L. & Manavalan, B. Integrative machine learning framework for the identification of cell-specific enhancers from the human genome. Brief. Bioinform. 22, bbab252 (2021).
Manavalan, B., Basith, S., Shin, T. H. & Lee, G. Computational prediction of species-specific yeast dna replication origin via iterative feature representation. Brief. Bioinform 22, bbaa304 (2021).
Liu, Q. et al. Deeptorrent: A deep learning-based approach for predicting dna n4-methylcytosine sites. Brief. Bioinform. 22, bbaa124 (2021).
Zhang, P., Zhang, H. & Wu, H. ipro-wael: A comprehensive and robust framework for identifying promoters in multiple species. Nucleic Acids Res. 50, 10278–10289 (2022).
Bi, Y. et al. An interpretable prediction model for identifying n7-methylguanosine sites based on xgboost and shap. Mol. Ther.-Nucleic Acids 22, 362–372 (2020).
Nair, A. S. & Sreenadhan, S. P. A coding measure scheme employing electron-ion interaction pseudopotential (eiip). Bioinformation 1, 197 (2006).
Liang, Y. & Zhang, S. Identifying dnase I hypersensitive sites using multi-features fusion and f-score features selection via chou’s 5-steps rule. Biophys. Chem. 253, 106227 (2019).
Cai, J. et al. A bioinformatics tool for the prediction of dna n6-methyladenine modifications based on feature fusion and optimization protocol. Front. Bioeng. Biotechnol. 8, 502 (2020).
Wang, M., Li, F., Wu, H., Liu, Q. & Li, S. Predpromoter-mf (2l): A novel approach of promoter prediction based on multi-source feature fusion and deep forest. Interdiscip. Sci. Comput. Life Sci. 14, 697–711 (2022).
Zhang, J., Gao, B., Chai, H., Ma, Z. & Yang, G. Identification of dna-binding proteins using multi-features fusion and binary firefly optimization algorithm. BMC Bioinform. 17, 1–12 (2016).
Li, L. et al. Sequence-based identification of recombination spots using pseudo nucleic acid representation and recursive feature extraction by linear kernel svm. BMC Bioinform. 15, 1–9 (2014).
Zhu, H., Ao, C.-Y., Ding, Y.-J., Hao, H.-X. & Yu, L. Identification of d modification sites using a random forest model based on nucleotide chemical properties. Int. J. Mol. Sci. 23, 3044 (2022).
Xu, H., Jia, P. & Zhao, Z. Deep4mc: Systematic assessment and computational prediction for dna n4-methylcytosine sites by deep learning. Brief. Bioinform. 22, bbaa099 (2021).
Park, S., Wahab, A., Nazari, I., Ryu, J. H. & Chong, K. T. i6ma-dnc: Prediction of dna n6-methyladenosine sites in rice genome based on dinucleotide representation using deep learning. Chemom. Intell. Lab. Syst. 204, 104102 (2020).
Tahir, M., Hayat, M. & Kabir, M. Sequence based predictor for discrimination of enhancer and their types by applying general form of chou’s trinucleotide composition. Comput. Methods Prog. Biomed. 146, 69–75 (2017).
Nguyen-Vo, T.-H. et al. ipseu-ncp: Identifying rna pseudouridine sites using random forest and ncp-encoded features. BMC Genom. 20, 1–11 (2019).
Liu, B., Gao, X. & Zhang, H. Bioseq-analysis 2.0: An updated platform for analyzing dna, rna and protein sequences at sequence level and residue level based on machine learning approaches. Nucleic Acids Res. 47, e127 (2019).
Alam, W., Tayara, H. & Chong, K. T. Xg-ac4c: identification of n4-acetylcytidine (ac4c) in mrna using extreme gradient boosting with electron-ion interaction pseudopotentials. Sci. Rep. 10, 20942 (2020).
Chen, W., Feng, P.-M., Lin, H. & Chou, K.-C. irspot-psednc: Identify recombination spots with pseudo dinucleotide composition. Nucleic Acids Res. 41, e68–e68 (2013).
Liu, B., Liu, F., Fang, L., Wang, X. & Chou, K.-C. repdna: A python package to generate various modes of feature vectors for dna sequences by incorporating user-defined physicochemical properties and sequence-order effects. Bioinformatics 31, 1307–1309 (2015).
Guo, S.-H. et al. inuc-pseknc: A sequence-based predictor for predicting nucleosome positioning in genomes with pseudo k-tuple nucleotide composition. Bioinformatics 30, 1522–1529 (2014).
Chicco, D. & Jurman, G. The advantages of the matthews correlation coefficient (mcc) over f1 score and accuracy in binary classification evaluation. BMC Genom. 21, 1–13 (2020).
Davis, J. & Goadrich, M. The relationship between precision-recall and roc curves. In Proceedings of the 23rd International Conference on Machine Learning 233–240 (2006).
Chapman, B. & Chang, J. Biopython: Python tools for computational biology. ACM Sigbio Newsl. 20, 15–19 (2000).
Chen, Z. et al. ilearnplus: A comprehensive and automated machine-learning platform for nucleic acid and protein sequence analysis, prediction and visualization. Nucleic Acids Res. 49, e60–e60 (2021).
Pedregosa, F. et al. Scikit-learn: Machine learning in Python. J. Mach. Learn. Res. 12, 2825–2830 (2011).
Tosi, S. Matplotlib for Python Developers (Packt Publishing Ltd, 2009).
Author information
Authors and Affiliations
Contributions
A.F.A and M.N.A conceived and conducted the experiments, S.A. and D.A. analysed the results. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Abbasi, A.F., Asim, M.N., Ahmed, S. et al. Long extrachromosomal circular DNA identification by fusing sequence-derived features of physicochemical properties and nucleotide distribution patterns. Sci Rep 14, 9466 (2024). https://doi.org/10.1038/s41598-024-57457-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-57457-5
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.