Abstract
The promising next-generation probiotic Faecalibacterium prausnitzii is one of the most abundant acetate-consuming, butyrate-producing bacteria in the healthy human gut. Yet, little is known about how acetate availability affects this bacterium’s gene expression strategies. Here, we investigated the effect of acetate on temporal changes in the transcriptome of F. duncaniae A2-165 cultures using RNA sequencing. We compared gene expression patterns between two growth phases (early stationary vs. late exponential) and two acetate levels (low: 3 mM vs. high: 23 mM). Only in low-acetate conditions, a general stress response was activated. In high-acetate conditions, there was greater expression of genes related to butyrate synthesis and to the importation of B vitamins and iron. Specifically, expression was strongly activated in the case of the feoAABC operon, which encodes a FeoB ferrous iron transporter, but not in the case of the feoAB gene, which encodes a second putative FeoAB transporter. Moreover, excess ferrous iron repressed feoB expression but not feoAB. Lastly, FeoB but not FeoAB peptides from strain A2-165 were found in abundance in a healthy human fecal metaproteome. In conclusion, we characterized two early-stationary transcriptomes based on acetate consumption and this work highlights the regulation of feoB expression in F. duncaniae A2-165.
Similar content being viewed by others
Introduction
Bacteria in the genus Faecalibacterium (phylum Bacillota1, class Clostridia, family Oscillospiraceae) are commensal, strictly anaerobic bacteria, that are ubiquitous and abundant in the gastrointestinal tracts of humans and animals2,3. This genus is currently recognized as one of the most important commensals for human health4,5. According to the latest phylogenetic analysis6, the genus Faecalibacterium is composed of six species, of which prausnitzii is the best known for its effects on human health. The most extensively studied strain of F. prausnitzii, A2-165, has been recently reclassified and placed into a new species, duncaniae sp. nov.6. Interest in Faecalibacterium has increased over the last decade, mostly in response to the pioneering study of Sokol et al. reporting a depletion of F. duncaniae A2-165 in Crohn’s disease patients7. Since this report, many studies have confirmed reduced abundances of Faecalibacterium in patients with inflammatory bowel disease and metabolic diseases8,9, which has led to consider members of this genus into candidates for the development of diagnostic, prognostic, preventive or therapeutic approaches10.
In the healthy human colon, Faecalibacterium are among the main bacteria responsible for the consumption of acetate and the production of butyrate. Butyrate is the preferred energy source for colonocytes and has anti-inflammatory properties that are generally considered to be beneficial to intestinal health11,12. Recently, we demonstrated that dact3, an host gene linked to the Wnt/JNK pathway, mediates these F. prausnitzii anti-inflammatory effects13. In addition, several peptides originating from a single 15-kDa protein (Microbial Anti-inflammatory Molecule, MAM) have been identified in the supernatant of F. duncaniae A2-165 cultures14,15,16, and these have been shown to alleviate chemically induced colitis in mice17,18. The key enzyme for butyrate production is butyryl-CoA:acetate CoA-transferase. It generates butyrate and acetyl-CoA from extracellular acetate and intracellular butyryl-CoA2,19,20,21. In addition to generating butyrate, this process also promotes growth, thus explaining the growth-stimulating effects of culture supplementation with a high amount of acetate (33–50 mM)2,21,22. In co-culture models, acetate cross-feeding between Faecalibacterium and acetate-producing bacteria such as Bifidobacteria adolescentis23 or Blautia hydrogenotrophica21 has been observed to enhance butyrate formation. A positive correlation between acetate consumption and butyrate production was also established in vivo between F. duncaniae A2-165 and Bacteroides thetaiotaomicron24.
In the human gut, acetate is the most abundant short-chain fatty acid largely generated by the bacterial fermentation of dietary fibers. Its concentration fluctuates depending on various parameters, including gut transit time, diet, age, health status, and microbiota composition25,26,27. To cope with such changes, Faecalibacterium have presumably developed regulatory systems that sense and respond to acetate levels. These adaptive strategies would necessarily involve a well-coordinated gene expression network that takes into account both the physiological state of the bacteria (latency, exponential, stationary phases) and the environmental conditions (availability of acetate and other nutrients, physicochemical signals such as oxygen tension, pH, temperature). Although several recent studies have investigated the transcriptome of F. duncaniae21,28,29, none has examined the effects of acetate across growth phases. In light of its proposed vital role in intestinal health, this information is critical for understanding the metabolism of Faecalibacterium and how this might affect human health.
Here, we investigated the effect of acetate on temporal changes in the transcriptome of F. duncaniae A2-165 cultures using RNA sequencing. Next, we characterized the regulation of two feoB genes—which encode well-known ferrous iron transporter—in response to the availability of acetate and ferrous sulfate. Finally, using an novel integrated approach, we searched for and identified FeoB peptides specific to the A2-165 strain in the healthy human fecal metaproteome. The general workflow of our study can be found as Supplementary Fig. S5 online.
Results
Experimental design for acetate-dependent transcriptome analysis
Acetate consumption is the main driver of butyrate production by genus Faecalibacterium, as shown in Fig. 1A. To gain insight into the effect of acetate on the transcriptome of F. duncaniae A2-165, two sets of cultures were established in media that contained low (3 mM) or high (23 mM) concentrations of acetate (see “Methods” section). The resulting growth kinetics are shown in Fig. 1B. In both acetate conditions, there was no difference in growth kinetics over the first 7 h, and the stationary growth phase was reached after 9 h of culture in both treatments. In the early stationary phase, the biomass in high-acetate conditions (Sa) was about 1.5-fold higher than in low-acetate conditions (S) (2.30 ± 0.13 OD600 vs. 1.50 ± 0.18 OD600, N = 4, p < 0.05). Moreover, the growth rate was 1.25-fold higher in high-acetate conditions (Fig. 1C), suggesting that the general metabolism of F. duncaniae A2-165 was highly active in these cultures. Indeed, the generation time was reduced by about 25 min in this group compared to that of low-acetate cultures (1.67 h vs. 2.08 h). In high- and low-acetate conditions, we quantified butyrate production and acetate consumption in both the late exponential growth phase (Ea and E, respectively) and the early stationary growth phase (Sa and S, respectively) (Fig. 1D). In both acetate conditions, butyrate production was significantly higher in the early stationary phase than in the late exponential phase. Moreover, in the late exponential phase (7 h of growth) there was no difference between acetate conditions in either butyrate production or acetate consumption. In contrast, in the early stationary phase (10 h of growth), butyrate production and acetate consumption were significantly different between acetate conditions (butyrate: 11.94 mM ± 0.89 for Sa, 6.37 mM ± 1.07 for S, p < 0.01). Moreover, after 10 h of growth, almost all acetate had been consumed in the low-acetate cultures (− 2.16 mM ± 0.38), whereas only one-third of the acetate had been consumed in the high-acetate cultures (− 7.66 mM ± 0.58). The acetate limitation experienced in the low-acetate conditions strongly affected the growth of F. duncaniae A2-165 (Fig. 1B,C). We thus investigated the acetate-growth effect at the transcriptional level in both acetate-limited and acetate-saturated conditions using RNA-Seq. For this, gene expression profiles were analyzed at both E/Ea and S/Sa time points (Fig. 1B).
Acetate growth effect and butyrate production in F. duncaniae A2-165 cultures. (A) Schematic view of the importance of acetate in the butyrate biosynthetic pathway. The production of butyrate is dependent on glycolysis and the butyrate cycle. The key enzyme for butyrate production is butyryl-CoA:acetate CoA-transferase (ButCoAT), which consumes extracellular acetate to produce butyrate and acetyl-CoA from butyryl-CoA. (B) Growth kinetics of F. duncaniae A2-165 under high- and low-acetate conditions. Growth kinetics are shown for bacteria cultured in BHIS medium with 23 mM acetate (high-acetate, orange color) or with 3 mM acetate (low-acetate, blue color). Arrows indicate time points of sampling for the RNA-Seq transcriptome analysis. Sampling was performed after 7 h (late exponential phase, E and Ea for low- and high-acetate conditions, respectively) and 10 h (early stationary phase, S and Sa for low- and high-acetate conditions, respectively) of growth. N = 4, median with interquartile range (IQR) is shown. (C) Individual growth-rate measures in high/low-acetate conditions. N = 4, raw data, median with IQR are shown. (D) SCFA concentrations (expressed as molarity) of E/Ea and S/Sa culture supernatants. Butyrate production is presented as positive values while acetate consumption is presented as negative values. Acetate values were normalized with the acetate value of BHIS medium, which contains 3 mM of acetate. N = 4, median with IQR is shown. (E) Overview of RNA-Seq data. The number of (differentially expressed) DE genes for each comparison is presented in the form of a bar chart. Two-group comparisons were conducted between high- and low-acetate conditions in the late exponential phase (Ea vs. E) and early stationary phase (Sa vs. S). Additionally, two-group comparisons were performed between the early stationary and late exponential phases in high-acetate (Sa vs. Ea) and low-acetate (S vs. E) conditions. log2FC ≥ |2|, FDR-adjusted p-value ≤ 0.01.
Overview of transcriptomes in low- and high-acetate conditions
The RNA-Seq datasets contained a total of 2882 genes out of the 3013 predicted in the RefSeq database (i.e., over 95%). Of these, 1159 genes were found to be differentially expressed (DE) (log2 fold change (FC) ≥ |2| and FDR-adjusted p-value ≤ 0.01) between conditions. All DE genes are presented in Supplementary Tables S1–S4 online. Eight of these genes were differentially expressed in early exponential cultures, compared to 541 in the late stationary phase (339 and 202 transcripts up- and down-regulated, respectively, Fig. 1E), which is consistent with the different patterns of growth kinetics. This result clearly demonstrates that, under our experimental conditions, acetate concentration (3 vs. 23 mM) had little impact on transcriptomic profiles in F. duncaniae A2-165 for the first 7 h of growth (i.e., up to the late exponential growth phase), but had a significant impact when the cultures entered the stationary phase of growth. This prompted us to perform a detailed analysis of the adaptive changes that occur in the transcriptome in response to acetate availability between 7 and 10 h of growth. As shown in Fig. 1E, there was a large difference in the adaptive response between high-acetate (Sa/Ea, 118 DE genes) and low-acetate (S/E, 492 DE genes) conditions in this three-hour period. Specifically, we found that, under high-acetate conditions, only 67 and 51 genes were up- and downregulated, respectively, whereas under low-acetate conditions, 216 and 276 genes were up- and downregulated, respectively. When comparing the DE transcript lists in low- and high-acetate conditions (see Supplementary Fig. S6 online), it clearly appeared that the acetate-limiting condition triggered a larger adaptive transcriptional response at the onset of the stationary growth phase compared to the acetate-saturated conditions, in our growth model.
Using the COG and PATRIC databases, we classified DE genes into 12 functional categories (see Supplementary Table S5 online). As shown in Fig. 2A, for all transcriptomes (high-acetate, low-acetate, and early stationary), the largest category (27.2–58.8% of genes) was the superclass “Poorly characterized protein-genes”, which included the categories “Hypothetical proteins” and “General function prediction”. It is likely that these unknown DE genes contribute to the adaptive responses of F. duncaniae A2-165 as cells enter the stationary phase. In addition, clear differences between low- and high-acetate transcriptomes were observed with respect to the categories “Protein synthesis”, “Energy metabolism”, “Import system”, “Defense system”, “Stress response”, and “Transcription and Post-transcriptional regulation” (Fig. 2A). Interestingly, in acetate-limiting conditions (S vs. E), the highest proportions of up- and downregulated genes were related to “Stress response” (9.3%, i.e., 20 DE genes) and “Protein synthesis” (27.9%, i.e., 77 DE genes) respectively, whereas in acetate-saturated conditions (Sa vs. Ea), the highest proportions of affected genes were related to “Import system” (13.4%, i.e., 9 upregulated genes, and 23.5%, i.e., 12 downregulated genes).
Transcriptional responses of F. duncaniae A2-165 to acetate. (A) Pie charts of up- and downregulated genes showing the relative frequency (%) of each functional category. The number of genes in each category is shown in Table S5 in Supplementary file 2 and the total number of DE genes is shown below each pie-chart. The right part of the panel shows the representative color for each category. The category “Other biological processes” contained the metabolism subcategories “Amino acid”, “Amino sugar”, “Nucleotide”, “Vitamin and Cofactor”, “Lipid”, and “Cell wall”, as well as “Bacterial cell division processes”. The “Energy metabolism” category corresponds to the conversion of carbon sources and acetate to ATP and the carbon skeleton and includes glycolysis and the butyrate pathway. See Tables S2–S4 in Supplementary file 2 for data on DE genes (gene ID, COG number, protein annotation, log2FC, padj value, category). Bold characters indicate category selection in this study. (B) Description of relevant functional categories and import system subclassification in high, low and early stationary acetate transcriptomes. ABC (ATP-binding cassette) transport family. Others: other transport system family, FeoB (Ferrous) iron transport system family. PTS: phosphoenol pyruvate transferase system family, ECF (Energy Coupling Factor) transport family. Arrows indicate “Import system” category selection and FeoB transport system selection in this study.
Overall, our transcriptome analysis strongly suggests that F. duncaniae A2-165 is able to tightly regulate the expression of metabolic and stress-response genes according to acetate levels when cells enter the stationary phase, in our growth conditions. Remarkably, in all comparative transcriptional analyses, the CG447_03795 gene encoding the MAM protein did not appear among the DE genes; the transcription of this gene was quite high and unchanged between acetate conditions as well as between the growth phases examined here (see Supplementary Fig. S7 online).
We analyzed in detail the categories with the highest gene count belonging to the TOP 3, in low- and high- acetate transcriptomes (see Table S5 in Supplementary Information). These categories include ‘Protein synthesis’ and ‘Stress response’ in low acetate transcriptomes, while they encompass ‘Import system’ in high acetate transcriptomes. Differentially expressed transcriptional regulators may yield clues as to how the adaptive response is regulated. We concentrated on the ‘Transcription and Post-transcriptional Regulation’ category in low acetate growth conditions to associate putative transcriptional regulators with potential target genes.
A general stress response under acetate-limiting conditions
As depicted in Fig. 2B, we clearly observed significant differences in the transcriptional responses of F. duncaniae A2-165 under acetate-limiting condition compared with acetate-saturated condition. Interestingly, in low-acetate condition, we found a high number of downregulated transcripts in the “Protein synthesis” category (77 DE genes, log2FC range: 2–5, Table S3) that were not detected in high-acetate condition. These were mainly transcripts involved in translation machinery, such as translation initiation/elongation factors and aminoacyl-tRNA synthetases/transferases, but also included genes involved in ribosome biogenesis and/or stability. These findings likely indicated a major slowdown of translational processes under low-acetate growth conditions as F. duncaniae cells entered the stationary phase.
We also noted a drastic shift in the “Stress response” category (20 DE genes, Table S3) which again was not observed in high-acetate condition. Seven of these genes (log2FC range: 2.2–2.9) encoded chaperone proteins involved in protein remodeling, in the repair of proteins following stress damage, or even in the ubiquitin machinery and proteasome. The remaining 13 (log2FC range: 2.1–6.2) genes were associated with type II toxin/antitoxin (TA) systems30. More specifically, we observed the upregulation of four putative operons for RelE TA systems, one putative operon for a TA module in the Doc family, two genes encoding antitoxin proteins, and one gene encoding a toxin protein. Of these, the DinJ/YafQ system (CG447_14090/14095) was the most activated TA system (log2FC: 6.1–6.2). Overall, our data suggest that cells of F. duncaniae A2-165 in low-acetate condition experienced severe general stress upon entry to the early stationary growth phase.
Bacteria employ different transcriptional regulators and sigma transcription factors to respond to changing environments. In the low-acetate transcriptome examined here, we found a higher number of upregulated genes encoding transcriptional regulators and sigma transcription factors (18 DE genes, log2FC range: 2.1–5.5, see Supplementary Table S3 online) compared to high-acetate condition (4 DE genes, log2FC range: 3.0–5.0, see Supplementary Table S4 online). Moreover, in all bacteria studied thus far, the global regulator that mediates the general stress response is a specialized sigma factor31. In the low-acetate transcriptome, we specifically found five upregulated genes encoding sigma factors (CG447_02225/05485/06440/06875/08160, see Supplementary Table S3 online), while in high-acetate conditions there was only one, which was also upregulated in low-acetate conditions (CG447_02215, Tables S4-S5). It is thus possible that these five sigma factors could be involved in the general stress response described above when cells enter the stationary phase.
Deciphering import systems under high- and low-acetate conditions
Transcription of transporter genes is usually regulated in response to substrate availability32. As shown in Fig. 2B, we clearly observed two distinct import system transcriptomes of F. duncaniae A2-165, under acetate-limiting and saturated conditions. These findings likely indicated two major adaptation responses to substrate availability, according to acetate conditions. A large number of genes (46) belonging to the “Import system” category were found to be downregulated under low-acetate conditions (see Supplementary Table S3 online); this number was four-fold higher than under high-acetate conditions (see Supplementary Table S4 online). These were mainly transporter genes in the ABC superfamily (25 DE genes), in the sodium-dependent transporter family (5 DE genes), in the ECF subfamily (3 DE genes) and in the FeoB family (4 DE genes) (Supplementary Table S3 online, Fig. 2B). Within the ECF group, we noticed a reduction in gene expression related to B vitamins uptake (bioY gene, CG447_11745, involved in biotin import, log2 FC − 4.14; ribU gene, CG447_14300, log2 FC − 3.92, involved in riboflavine import). We noticed a considerable reduction in gene expression related to ferrous iron uptake (feoA1A2BC putative operon, CG447_12740-55, log2 FC range: − 4.9 to 5.9), and ferric iron uptake (fhu putative operon, CG447_03300-10, log2FC range: − 6.1 to 8.0). This suggested that the expression of major systems of iron transport was severely impaired in low-acetate conditions. Conversely, we found that expression of the feoA1A2BC operon was upregulated under high-acetate condition (log2FC: 1.5–2.1, Fig. 2B). This was particularly marked in early stationary transcriptome, in which we observed an even higher fold-change in expression (log2FC: 4.5–5.9, Sa vs. S, Supplementary Table S2 online, Fig. 2B). Many studies have explored the role of the FeoB high affinity transporters in ferrous iron uptake in iron-poor environments33,34. Our result suggests that in the early stationary phase in an acetate-saturated environment, ferrous iron is no longer abundant, and activation of the feoAABC system may allow F. duncaniae A2-165 cells to maintain iron homeostasis in these conditions.
Under high-acetate conditions, feoB is the most transcribed feo homolog
In order to confirm our hypothesis of limited iron availability in the high-acetate culture conditions, we analyzed feoB expression in the presence of excess ferrous sulfate (Fig. 3). Figure 3A illustrates ferrous iron uptake and iron homeostasis in F. duncaniae. As a control, we also analyzed gene expression in low-acetate conditions, as well as the expression of feoAB (CG447_08795, log2FC: − 4.3, see Supplementary Table S4 online) and butCoA, which encodes the terminal enzyme required for butyrate production (Fig. 3A). A new set of F. duncaniae A2-165 cultures was prepared with high or low levels of acetate and with or without ferrous sulfate, and sampling was performed as previously described (i.e., in the late exponential phase, indicated by E, and the early stationary phase, indicated by S, Fig. 3B). Figure 3B presents the results regarding the expression ratios (S vs. E) of feoB, feoAB, and butCoA obtained under the different conditions. These clearly demonstrated that the feoB expression was significantly triggered (log2FC: 6.0) during the early stationary phase, only in the high-acetate condition, and in the absence of ferrous sulfate supplementation. Moreover, its expression was totally repressed in the presence of ferrous sulfate (log2FC: − 2.0). Overall, the feoB regulation was dependent on the availability of an iron source in the medium, with strong activation and repression in the absence and presence of ferrous sulfate, respectively. For the feoAB gene, the expression ratios were similar in the absence or presence of an iron source in both acetate growth conditions, indicating that the regulation of feoAB is independent of iron availability in our growth condition. For the butCoA gene, we observed a positive growth phase effect in both acetate and iron conditions with stronger activation in the high-acetate conditions (Fig. 3B). Overall, these results demonstrate that regulation of the expression of feoAABC and feoAB differs significantly and suggest that the FeoB and FeoAB systems of F. duncaniae A2-165 play nonredundant roles in ferrous iron acquisition.
Expression analysis of feoB, feoAB and butCoA by RT-PCR. (A) Schematic representation of ferrous iron uptake, iron protein synthesis, glycolysis, and butyrate metabolism pathway. The FeoB membrane transporter, NFeoB domain with GTPase activity, and FeoA1, FeoA2, and FeoC cytosolic proteins are indicated. In the FeoB systems that have been described to date, cytosolic proteins are required for ferrous iron acquisition but the exact mechanism is not yet known33. The glycolysis pathway and Faecalibacterium’s genus-specific butyrate production pathway (shown as butyrate cycle) are shown. Pyruvate is converted to acetyl-CoA by pyruvate:ferredoxin oxidoreductase (PFOR) and the energy generated during butyrate production is conserved through the buildup of a proton gradient via the activity of a membrane-associated NADH:ferredoxin oxidoreductase encoded by two putative rnf operons (rnfDEA, CG447_09840-830, rnfCDGEAB CG447_10905-930)37. Ferredoxin proteins (orange color). See Tables S2–S4 in Supplementary file 2 for data on DE genes (gene ID, COG numbers, protein annotation, log2FC, padj value, category). (B) Effect of the addition of ferrous sulfate on the transcription of feoB, feoAB, and butCoA genes, quantified using RT-PCR. Growth kinetics of F. duncaniae strain A2-165 in the presence or absence of ferrous sulfate (50 µM) in high-acetate (23 mM acetate) and low-acetate (3 mM) growth conditions. Square: culture sampling at T6 (exponential (E) growth phase) and T10 (early stationary (S) growth phase) time points. Genetic organization of the putative feoA1A2BC operon is shown.
FeoB peptides from strain A2-165 are found in abundance in a healthy human fecal metaproteome
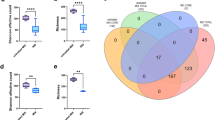
We were interested in studying feoB expression in the human gut microbiome. For this, we analyzed a fecal metaproteomic dataset obtained from eight healthy individuals, which included 123,425 peptides from the envelope fraction of the gut microbiota (Fig. 4)35. The metaproteomics/transcriptomics data can be found in Supplementary Tables S6–S8 online. Among these peptides, 236 matched with 10 of the 42 transporter genes that we had found to be upregulated in high-acetate conditions. Remarkably, the second highest degree of protein coverage (43.6%, 51 peptides) was found for the FeoB transporter. Of those 51 peptides, 9 were specific to a single protein in the metaproteomic dataset ‘a5.b59.a1’ (see Supplementary Table S7-2 ‘CG447_12750 FeoB’ and Supplementary Fig. S8 online), which was identified as FeoB of the A2-165 strain (K04759 in the KEGG orthology database). In addition, among the 573 identified proteins from A2-165 strain, FeoB belongs to the top 20 hits in term of spectra number (see Supplementary Table S8 online). On the other hand, FeoAB protein exhibited a fivefold lower number of spectra corresponding to the 284th position in the A2-165 proteins list (see Supplementary Table S8 online). Overall, this demonstrates that FeoB, but not FeoAB, from A2-165 is found in abundance in this healthy human fecal metaproteome35.
Workflow for the search for FeoB peptides in the healthy human gut metaproteome. RNA-Seq transcriptome of early stationary acetate cultures of F. duncaniae strain A2-165. (A) The general workflow employed for RNA-Seq in the current study: from identification of DE genes in the early stationary phase (Sa vs S comparison) to feoAABC operon selection. (B) Published human gut metaproteome. The general workflow employed in a published study of the gut metaproteome. The data were obtained from eight healthy subjects included in a study of intestinal bowel diseases35. (C) Integrating RNA-Seq and metaproteomic data. The general workflow employed to integrate RNA-Seq and metaproteomic data in the current study and the presentation of data for F. duncaniae A2-165.
Discussion
Two early stationary lifestyles of F. duncaniae A2-165
Here, we investigated the adjustments in the bacterial transcriptome when cells of F. duncaniae A2-165 entered the stationary growth phase in low- and high-acetate conditions. To our knowledge, this is the first physiological study based on transcriptomic and metabolic analyses that focuses on the entry into the stationary growth phase of F. duncaniae and, moreover, of a bacterium belonging to the Faecalibacterium genus.
Given that Faecalibacterium is a strictly anaerobic bacterium, cultivation presents challenges36. We have limited options for monitoring the growth of F. duncaniae, which include a semi-defined medium (“Yeast, Casitone, Fatty acid” named YCFA medium of Duncan laboratory2) with slow growth and low biomass, or rapid growth with high biomass in the rich complex BHIS medium. We opted for the latter method to enable accurate and precise tracking of stationary phase entry over a 12–14 h period.
Comparative analysis of the high- and low-acetate transcriptomes showed two different metabolic responses regarding the number of genes and their functions. Given that acetate is a major growth factor (i.e. major energy source)37, this seems consistent. Because of the lack of relevant data from the literature with which to compare our results, we chose to focus on the strongest and clearest effects, i.e., high fold-changes in expression. In the future, it will be necessary to study (i) the functions carried by the weaker adjustments in the transcriptome (i.e. lower fold change), (ii) this adaptive response in continuous culturing using chemostats, (iii) this adaptive response in co-culture models with acetate-producing bacteria.
Our experimental design enabled the identification of a general stress response under low acetate growth conditions. A characteristic of the exponential-stationary transition is the induction of stress proteins adapted for general stress condition. The general stress response has been very well characterized in model bacteria such as Escherichia coli and Bacillus subtilis38, but this protective adaptive response is not known in Faecalibacterium. Notably, a general stress response was observed only in low-acetate conditions; concomitantly with the general stress response, a negative regulation of protein synthesis genes and a positive regulation of genes encoding type II toxin/antitoxin modules were also observed. As the majority of type II toxins inhibit translation30, we can hypothesize that the former observation may be the consequence of the latter. Sigma B is the sigma factor involved in the control of the general stress response38 in many bacteria including model bacteria and several anaerobic bacteria such as Clostridioides difficile39. Remarkably, the gene encoding Sigma B is not present in A2-165 genome. Our functional analysis led to a list of genes encoding Sigma factors, that could be involved in the control of this protective response in low-acetate conditions. To our knowledge, this study represents the first description of a general stress response, in the early stationary phase, in acetate-limiting conditions in Faecalibacterium.
FeoB transporters search strategy
In the second part of our study, we chose to study bacterial transporters of the Feo family because the transport function of ferrous iron and its regulation are well characterized32,33. For this, we focused on the fecal metaproteome of healthy individuals, previously published35. This metaproteome was of interest in our study of the F. duncaniae A2-165 import systems because it targeted bacterial envelope proteins. We combined high-resolution mass spectrometry of fecal specimens with RNA-seq of F. duncaniae A2-165 pure culture. Our metaproteomic analysis shows a significant difference in abundance between FeoB and FeoAB, suggesting variations in the regulation of feoAABC and feoAB expression in the context of an healthy gut microbiome. In conclusion, our search strategy allows us to capture the major A2-165 Feo import systems, namely the FeoB transporter. In the future, this search strategy can be extended to investigate Feo transporters in other bacteria of the genus Faecalibacterium using the same metaproteomic dataset.
Iron homeostasis and butyrate production
In many environments, including the human gut, iron is a limiting nutrient for growth, and high-affinity uptake systems play a central role in ferrous iron homeostasis40. Maintenance of ferrous iron homeostasis is crucial for the production of butyrate37 because this process requires two iron-binding ferredoxins as illustrated in Fig. 3A. Here, we demonstrated that butCoA expression was strongly upregulated regardless the supplementation of cultures with ferrous sulfate. This result suggests that, in the early stationary growth phase in high-acetate conditions, mechanisms of ferrous iron homeostasis were well established in F. duncaniae A2-165.
Interestingly, together with upregulation of feoAABC, we also detected upregulation of a flavodoxin-encoding gene (see Supplementary Table S4 online). This is consistent with previous studies reporting that iron-free flavodoxin replaces iron-sulfur ferredoxin under iron-limited conditions40,41. Because flavodoxin and ferredoxin both mediate electron transfer in redox processes, the use of flavodoxin could free up iron for utilization by other iron-dependent enzymes. In other word, flavodoxins are key players in maintenance of ferrous iron homeostasis in iron-limited environment40,41. For example, in C. difficile, the production of flavodoxin is tied to iron homeostasis in iron-limited conditions42. In addition, the switch to flavodoxin as electron transfer protein may have significant effects on the activity of numerous enzymes that use flavodoxin as a redox partner. For example, in B. subtilis, flavodoxin is the redox partner for the acyl lipid desaturase involved in fatty-acid desaturation43. To date, similar data are not available for Faecalibacterium, but the impact of flavodoxins on the activity of certain metabolic pathways, including butyrate pathway, as well as the link with iron availability, merits further investigation.
Methods
Strain used for experimental culture
This study used a stock of Faecalibacterium duncaniae A2-165 (DSM No. 17677, DSMZ collection, Braunschweig, Germany) originally isolated from human fecal stool by S. H. Duncan (University of Aberdeen, United Kingdom)2 and maintained by V. Robert, MICALIS Institute, INRAE, France.
Cultivation experiments of F. duncaniae
F. duncaniae A2-165 was grown under anaerobic conditions (anaerobic chamber: N2 = 90%, CO2 = 5%, and H2 = 5%) at 37 °C in BHIS medium (also named LYBHI in our previous studies), which is brain heart infusion broth (BHI, 37 g/L, Difco) supplemented with yeast extract (5 g/L, Difco), cellobiose (1 mg/mL, Sigma), maltose (1 mg/mL, Sigma), and cysteine (0.5 mg/mL, Sigma)17. The strain was thawed and grown from a 10% inoculation in 10 mL BHIS for 24 h at 37 °C, followed by a 1% inoculation in 10 mL BHIS for overnight growth (16–18 h). The experimental cultures were then created using 2% inoculation in 50 mL BHIS medium with (BHISAc) or without the addition of sodium-acetate solution (20 mM, Sigma2,37) at the time of inoculation. Growth kinetics were monitored by measuring the OD600 nm every hour. The low acetate condition (3 mM) corresponded to the basal amount of acetate that we measured in the BHIS growth medium. The high acetate condition (20 mM added to the medium, i.e. final concentration 23 mM), was based on the previous in vitro studies of Duncan et al.2 and Heinken et al.37, as well as our own measurements of acetate concentration in human feces (median value 20 mM, see Table 1 of Martin et al.27).
Addition of ferrous sulfate to F. duncaniae culture
A fresh solution of ferrous sulfate (FeSO4-7H2O, Sigma) was prepared with deionized water and filter-sterilized. The experimental cultures were created using 2% inoculations in 50 mL BHIS or BHISAc media. Ferrous sulfate (50 µM, Sigma) was added 3 h after inoculation.
Short-chain fatty acid (SCFA) production and consumption
Gas chromatography (7890B, Agilent Technologies) was used to identify the short-chain fatty acids present in culture supernatants. The SCFAs of interest in this study were acetate and butyrate. An internal standard of 2-ethylbutyrate was used to normalize the data in each run. Measurements of SCFA concentrations in the bacterial samples were normalized using the basal SCFA levels of BHISAc and BHIS media from the same experiment.
Collection of RNA samples
For both growth conditions, cells and culture supernatant were sampled in the late exponential growth phase (7 h after inoculation) and early stationary growth phase (10 h after inoculation) for RNA extraction (cells) and SCFA analysis (supernatant). At the times of sampling, dilutions were created of all cultures and plated on BHIS agar for quantification of colony-forming units (CFUs).
RNA extraction
The single-step RNA isolation method was used, with TRIzol reagent and the FastPrep-24™ 5G instrument. For RNA-Seq and RT-PCR analyses, 2 mL of each culture was harvested and centrifuged (30 s, 13,000 rpm) in the Freter chamber, and cell pellets were snap-frozen in liquid nitrogen. 1 mL of TRIzol reagent (phenol-based RNA extraction buffer, Ambion, ThermoFisher Scientific) and 0.4 g of glass beads (0.1 mm, Bühler, ThermoFisher Scientific) were added to the defrosted pellet, and cell lysis was performed with a FastPrep-24™ 5G instrument (MP Biomedicals) (speed 6, 40 s). RNA precipitation, washing, and solubilization was carried out following the TRIzol protocol. Residual DNA from the RNA preparations was enzymatically removed using TURBO DNA-free (Ambion, United Kingdom). RNA quality was assessed with the Agilent 6000 Nano kit using an Agilent 2100 bioanalyzer (Stratagene, Agilent Technologies, France); RIN values were in the 7.7–8.5 range. Total RNA was purified from three biological replicates for RNA-Seq experiments and for RT-qPCR analysis. Extracted RNA samples were stored in RNAase/DNAse-free water (Ambion, United Kingdom) at − 80 °C.
Library construction and Illumina sequencing
Total RNA quality was assessed using an Agilent Bioanalyzer 2100 and the RNA 6000 Pico kit (Agilent Technologies). Directional RNA-Seq libraries were constructed using the TruSeq Stranded Total RNA library prep kit, with bacterial Ribo-Zero reagents (Illumina), following the manufacturer’s instructions; 500 ng of total RNA were used (I2BC HTS platform, CNRS, Gif-sur-Yvettes, France). After the Ribo-Zero step, the samples were checked on the Agilent Bioanalyzer to verify rRNA depletion. The quality of the libraries was assessed on an Agilent Bioanalyzer 2100, using an Agilent High Sensitivity DNA Kit. Libraries were pooled in equimolar proportions and sequenced in a paired-end 2 × 75-pb run on an Illumina NextSeq500 instrument (I2BC HTS platform, CNRS, Gif-sur-Yvettes, France). Demultiplexing was performed with bcl2fastq2 v2.18.12. Adapters were trimmed with Cutadapt v1.15, and only reads longer than 10 pb were kept for further analysis.
Mapping and identification of differentially expressed genes
Reads were mapped on the genome of F. duncaniae strain A2-165 (GenBank: CP022479.1) with BWA 0.6.2-r126, and were counted using the subread feature of Counts v1.5.2. Analyses of differential expression were performed in R using DESeq2 (I2BC HTS platform, CNRS, Gif-sur-Yvettes, France). In total, 2900 genes—including 2819 protein-coding sequences, 63 tRNA genes, and 18 rRNA genes, were mapped. Only the first two groups were analyzed in this study, so that a total of 2882 genes were considered. These were further filtered to retain only ORFs that showed a four-fold change in expression (absolute value Log2FC > = 2) (up/down) between treatment conditions. RNA-Seq datasets can be found on the database Omics Dataverse under the DOI accession number BDNCT2 (https://doi.org/10.15454/BDNCT2).
RT-PCR analysis
We generated cDNA from 1 μg of total RNA using the High Capacity cDNA Reverse Transcription kit (Fisher Scientific, France) and random hexamers. The quality of cDNA was checked with an Agilent 6000 Pico kit and an Agilent 2100 Bioanalyzer (Stratagene, Agilent Technologies, France). We carried out qPCR in duplicate, in a reaction volume of 20 μL containing 500 pg of cDNA, 15 μL of SYBR® Green PCR Master Mix (Applied Biosystems, Courtaboeuf, France), and 300 nM of each gene-specific primer. The primers were designed with Primer Express® (version 3.0). We generated standard curves for each set of primers, using serial dilutions (four dilutions) of cDNA obtained from a mix of total RNA collected in BHISAc and BHIS. Amplification was carried out with an ABI® PRISM 7900 thermal cycler (Applied Biosystems), with the following thermal profile: 2 min at 50 °C, 10 min at 95 °C, and 40 cycles of 15 s at 95 °C and 60 s at 60 °C. The specificity of each PCR amplicon was checked by melting curve analysis. Here, the feoB (CG447_12750, F5′-tgatcttcaacctgctgtgc, R5′-gccacgatggtgaagaagtt), feoAB (CG447_08795, F5′-ataccgaagtcaccgacctg R5′-ctccatcagctgcatcgtta) and butCoA (CG447_01820, F5′-actttgttctgggcgcatac, R5′-ggtcagtcccttcaggttca) primers used in this study. The efp gene (CG447_05125), encoding the translation elongation factor P, was used for normalization with the primers efpF (5-gttgagttccagcacgtgaa-3) and efpR (5-aaagcctgagggaactttgc-3). The relative change in gene expression was recorded as the ratio of normalized target concentrations and was calculated with the comparative ΔΔCt method44. The mean values for three independent experiments are presented.
Protein sequence annotations, functional classification
Protein annotation data from the PATRIC database45 (PATRIC genome ID 853.173) were used for functional classification. As the functional subsystem coverage in PATRIC was low (29%), EggNOG database was used for COG-based functional classifications using the COG system (https://www.ncbi.nlm.nih.gov/research/COG).
We created a simplified classification scheme with 12 categories; the largest metabolism classes (protein synthesis, energy, and import system) were retained while smaller metabolism classes (cell wall metabolism, lipid metabolism, amino acid metabolism, amino sugar metabolism, nucleotide metabolism, vitamin and cofactor metabolism) were merged into an “Other metabolism” category. The “Import system” and “Export system” classes encompassed all membrane transport systems involved in the uptake/export processes of nutrients, micronutrients, ions, and unknown compounds. “Stress response” category encompassed toxin/antitoxin systems and chaperones. The TC-BLAST tool (TCD database https://www.tcdb.org/) was used for the determination of the number of TMS (putative TransMembrane Segment) and for the import system family assignment. To characterize gene similarity, homology, and gene context, NCBI BLAST was used, in particular for analysis of feo genes.
Fecal metaproteomics analysis
We conducted an analysis of 16 published available metaproteomes derived from the envelope fractions of fecal microbiota obtained from eight healthy donors without any family history of gastrointestinal disease and no medication use35. The details of the metaproteomic method has been described previously35. To note, a minimum of two distinct peptides identified across all samples in the dataset was set to validate a protein in order to exclude proteins of weak proof of presence.
Combined A2-165 transcriptomics and fecal metaproteomics analysis
Each of the 123,425 peptide sequences identified in the fecal metaproteomes was matched against each of the 42 query protein sequences identified in this RNA-seq analysis (see Supplementary Tables S6–S8 online) in the R environment.
Statistical analysis
Normally distributed data are presented as mean ± standard deviation (SD), skewed data as median with interquartile range (IQR), and categorical data as frequencies with proportions. T-tests for independent samples and one-way analysis of variance (ANOVA) (or Kruskal–Wallis when not normally distributed) were performed for continuous variables. Data were represented and analyzed with GraphPad Prism 8.
Data availability
The data that support the findings of this study are openly available in database Omics Dataverse at https://doi.org/10.15454/BDNCT2 [DOI accession number BDNCT2]. All of the material is owned by the authors and/or no permissions are required.
References
Oren, A. & Garrity, G. M. Valid publication of new names and new combinations effectively published outside the IJSEM. Int. J. Syst. Evol. Microbiol. 71, 005096. https://doi.org/10.1099/ijsem.0.005096 (2021).
Duncan, S. H., Hold, G. L., Harmsen, H. J. M., Stewart, C. S. & Flint, H. J. Growth requirements and fermentation products of Fusobacterium prausnitzii, and a proposal to reclassify it as Faecalibacterium prausnitzii gen. nov., comb. nov. Int. J. Syst. Evol. Microbiol. 52, 2141–2146. https://doi.org/10.1099/00207713-52-6-2141 (2002).
Miquel, S. et al. Faecalibacterium prausnitzii and human intestinal health. Curr. Opin. Microbiol. 16, 255–261. https://doi.org/10.1016/j.mib.2013.06.003 (2013).
Lopez-Siles, M., Duncan, S. H., Garcia-Gil, L. J. & Martinez-Medina, M. Faecalibacterium prausnitzii: From microbiology to diagnostics and prognostics. ISME J. 11, 841–852. https://doi.org/10.1038/ismej.2016.176 (2017).
Martin, R., Bermudez-Humaran, L. G. & Langella, P. Searching for the bacterial effector: The example of the multi-skilled commensal bacterium Faecalibacterium prausnitzii. Front. Microbiol. 9, 346. https://doi.org/10.3389/fmicb.2018.00346 (2018).
Sakamoto, M. et al. Genome-based, phenotypic and chemotaxonomic classification of Faecalibacterium strains: Proposal of three novel species Faecalibacterium duncaniae sp. nov., Faecalibacterium hattorii sp. nov. and Faecalibacterium gallinarum sp. nov. Int. J. Syst. Evol. Microbiol. 72, 005379. https://doi.org/10.1099/ijsem.0.005379 (2022).
Sokol, H. et al. Faecalibacterium prausnitzii is an anti-inflammatory commensal bacterium identified by gut microbiota analysis of Crohn disease patients. Proc. Natl. Acad. Sci. U.S.A. 105, 16731–16736. https://doi.org/10.1073/pnas.0804812105 (2008).
Sokol, H. et al. Low counts of Faecalibacterium prausnitzii in colitis microbiota. Inflamm. Bowel Dis. 15, 1183–1189. https://doi.org/10.1002/ibd.20903 (2009).
Tilg, H. & Moschen, A. R. Microbiota and diabetes: An evolving relationship. Gut 63, 1513–1521. https://doi.org/10.1136/gutjnl-2014-306928 (2014).
Langella, P., Guarner, F. & Martin, R. Editorial: Next-generation probiotics: From commensal bacteria to novel drugs and food supplements. Front. Microbiol. 10, 1973. https://doi.org/10.3389/fmicb.2019.01973 (2019).
Pryde, S. E., Duncan, S. H., Hold, G. L., Stewart, C. S. & Flint, H. J. The microbiology of butyrate formation in the human colon. FEMS Microbiol. Lett. 217, 133–139. https://doi.org/10.1111/j.1574-6968.2002.tb11467.x (2002).
Louis, P. & Flint, H. J. Diversity, metabolism and microbial ecology of butyrate-producing bacteria from the human large intestine. FEMS Microbiol. Lett. 294(1), 8. https://doi.org/10.1111/j.1574-6968.2009.01514.x (2009).
Lenoir, M. et al. Butyrate mediates anti-inflammatory effects of Faecalibacterium prausnitzii in intestinal epithelial cells through Dact3. Gut Microbes 12, 1–16. https://doi.org/10.1080/19490976.2020.1826748 (2020).
Quevrain, E., Maubert, M. A., Sokol, H., Devreese, B. & Seksik, P. The presence of the anti-inflammatory protein MAM, from Faecalibacterium prausnitzii, in the intestinal ecosystem. Gut 65, 882. https://doi.org/10.1136/gutjnl-2015-311094 (2016).
Quevrain, E. et al. Identification of an anti-inflammatory protein from Faecalibacterium prausnitzii, a commensal bacterium deficient in Crohn’s disease. Gut 65, 415–425. https://doi.org/10.1136/gutjnl-2014-307649 (2016).
Auger, S. et al. Intraspecific diversity of microbial anti-inflammatory molecule (MAM) from Faecalibacterium prausnitzii. Int. J. Mol. Sci. 23, 1705. https://doi.org/10.3390/ijms23031705 (2022).
Martin, R. et al. The commensal bacterium Faecalibacterium prausnitzii is protective in DNBS-induced chronic moderate and severe colitis models. Inflamm. Bowel Dis. 20, 417–430. https://doi.org/10.1097/01.MIB.0000440815.76627.64 (2014).
Martin, R. et al. Faecalibacterium prausnitzii prevents physiological damages in a chronic low-grade inflammation murine model. BMC Microbiol. 15, 67. https://doi.org/10.1186/s12866-015-0400-1 (2015).
Duncan, S. H. et al. Contribution of acetate to butyrate formation by human faecal bacteria. Br. J. Nutr. 91, 915–923. https://doi.org/10.1079/BJN20041150 (2004).
Charrier, C. et al. A novel class of CoA-transferase involved in short-chain fatty acid metabolism in butyrate-producing human colonic bacteria. Microbiology 152, 179–185. https://doi.org/10.1099/mic.0.28412-0 (2006).
D’Hoe, K. et al. Integrated culturing, modeling and transcriptomics uncovers complex interactions and emergent behavior in a three-species synthetic gut community. Elife 7, e37090. https://doi.org/10.7554/eLife.37090 (2018).
Lopez-Siles, M. et al. Cultured representatives of two major phylogroups of human colonic Faecalibacterium prausnitzii can utilize pectin, uronic acids, and host-derived substrates for growth. Appl. Environ. Microbiol. 78, 420–428. https://doi.org/10.1128/AEM.06858-11 (2012).
Rios-Covian, D., Gueimonde, M., Duncan, S. H., Flint, H. J. & de los Reyes-Gavilan, C. G. Enhanced butyrate formation by cross-feeding between Faecalibacterium prausnitzii and Bifidobacterium adolescentis. FEMS Microbiol. Lett. 362, fnv176. https://doi.org/10.1093/femsle/fnv176 (2015).
Wrzosek, L. et al. Bacteroides thetaiotaomicron and Faecalibacterium prausnitzii influence the production of mucus glycans and the development of goblet cells in the colonic epithelium of a gnotobiotic model rodent. BMC Biol. 11, 61. https://doi.org/10.1186/1741-7007-11-61 (2013).
Macfarlane, S. & Macfarlane, G. T. Regulation of short-chain fatty acid production. Proc. Nutr. Soc. 62, 67–72. https://doi.org/10.1079/PNS2002207 (2003).
Miquel, S. et al. Ecology and metabolism of the beneficial intestinal commensal bacterium Faecalibacterium prausnitzii. Gut Microbes 5, 146–151. https://doi.org/10.4161/gmic.27651 (2014).
Martin, R. et al. Functional characterization of novel Faecalibacterium prausnitzii strains isolated from healthy volunteers: A step forward in the use of F. prausnitzii as a next-generation probiotic. Front. Microbiol. 8, 122. https://doi.org/10.3389/fmicb.2017.01226 (2017).
Lebas, M., Garault, P., Carrillo, D., Codoner, F. M. & Derrien, M. Metabolic response of Faecalibacterium prausnitzii to cell-free supernatants from lactic acid bacteria. Microorganisms 8, 1528. https://doi.org/10.3390/microorganisms8101528 (2020).
Kang, D. et al. Functional dissection of the phosphotransferase system provides insight into the prevalence of Faecalibacterium prausnitzii in the host intestinal environment. Environ. Microbiol. 23, 4726–4740. https://doi.org/10.1111/1462-2920.15681 (2021).
Harms, A., Brodersen, D. E., Mitarai, N. & Gerdes, K. Toxins, targets, and triggers: An overview of toxin-antitoxin biology. Mol. Cell 70, 768–784. https://doi.org/10.1016/j.molcel.2018.01.003 (2018).
Gottesman, S. Trouble is coming: Signaling pathways that regulate general stress responses in bacteria. J. Biol. Chem. 294, 11685–11700. https://doi.org/10.1074/jbc.REV119.005593 (2019).
Saier, M. H. Jr. A functional-phylogenetic system for the classification of transport proteins. J. Cell. Biochem. Suppl 32–33, 84–94. https://doi.org/10.1002/(sici)1097-4644(1999)75:32+%3c84::aid-jcb11%3e3.0.co;2-m (1999).
Sestok, A. E., Linkous, R. O. & Smith, A. T. Toward a mechanistic understanding of Feo-mediated ferrous iron uptake. Metallomics 10, 887–898. https://doi.org/10.1039/c8mt00097b (2018).
Lau, C. K., Krewulak, K. D. & Vogel, H. J. Bacterial ferrous iron transport: The Feo system. FEMS Microbiol. Rev. 40, 273–298. https://doi.org/10.1093/femsre/fuv049 (2016).
Henry, C. et al. Modern metaproteomics: A unique tool to characterize the active microbiome in health and diseases, and pave the road towards new biomarkers-example of Crohn’s disease and ulcerative colitis flare-ups. Cells 11, 1340. https://doi.org/10.3390/cells11081340 (2022).
Martin, R. et al. Faecalibacterium: A bacterial genus with promising human health applications. FEMS Microbiol. Rev. 47, fuad039. https://doi.org/10.1093/femsre/fuad039 (2023).
Heinken, A. et al. Functional metabolic map of Faecalibacterium prausnitzii, a beneficial human gut microbe. J. Bacteriol. 196, 3289–3302. https://doi.org/10.1128/JB.01780-14 (2014).
Hecker, M., Schumann, W. & Volker, U. Heat-shock and general stress response in Bacillus subtilis. Mol. Microbiol. 19, 417–428. https://doi.org/10.1046/j.1365-2958.1996.396932.x (1996).
Morvan, C., Folgosa, F., Kint, N., Teixeira, M. & Martin-Verstraete, I. Responses of Clostridia to oxygen: From detoxification to adaptive strategies. Environ. Microbiol. 23, 4112–4125. https://doi.org/10.1111/1462-2920.15665 (2021).
Pi, H. & Helmann, J. D. Ferrous iron efflux systems in bacteria. Metallomics 9, 840–851. https://doi.org/10.1039/c7mt00112f (2017).
Pisithkul, T. et al. Metabolic remodeling during biofilm development of Bacillus subtilis. mBio https://doi.org/10.1128/mBio.00623-19 (2019).
Berges, M. et al. Iron regulation in Clostridioides difficile. Front. Microbiol. 9, 3183. https://doi.org/10.3389/fmicb.2018.03183 (2018).
Chazarreta-Cifre, L., Martiarena, L., de Mendoza, D. & Altabe, S. G. Role of ferredoxin and flavodoxins in Bacillus subtilis fatty acid desaturation. J. Bacteriol. 193, 4043–4048. https://doi.org/10.1128/JB.05103-11 (2011).
Bustin, S. A. et al. The MIQE guidelines: Minimum information for publication of quantitative real-time PCR experiments. Clin. Chem. 55, 611–622. https://doi.org/10.1373/clinchem.2008.112797 (2009).
Wattam, A. R. et al. PATRIC, the bacterial bioinformatics database and analysis resource. Nucleic Acids Res. 42, D581–D591. https://doi.org/10.1093/nar/gkt1099 (2014).
Acknowledgements
We acknowledge the sequencing and bioinformatics expertise of the I2BC High-throughput sequencing facility. This work has benefited from the facilities and expertise of @BRIDGe (Université Paris-Saclay, INRAE, AgroParisTech, GABI, 78350 Jouy-en-Josas, France). We thank V. Robert, C. Mayeur, R. Martin and J.M. Chatel for helpful advices, F. Mazouz for RT-PCR analysis and L. Higgins for English correction service.
Funding
This work was supported by the French funding agency “Agence Nationale de la Recherche” under Grant (ANR-16-CE29-0020); under Grant (BP-Acetate, Qualiment-Institut Carnot); and under Grant (ANR-10-INBS-09).
Author information
Authors and Affiliations
Contributions
E.H., S.L. and M.T. conceptualized, conceived, and designed the study; E.H., S.V., S.C., and Y.J. performed research; E.H., S.V., S.A., S.L., Y.J., C.J., C.H. analysed data; E.H., S.L., C.J., and M.T. wrote the original draft; E.H., S.L., M.T., C.J., S.V., P.L., H.S., C.H., L.B., S.A. revised the manuscript; E.H., L.B. and M.T., funding acquisition. All approved the submitted version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Verstraeten, S., Layec, S., Auger, S. et al. Faecalibacterium duncaniae A2-165 regulates the expression of butyrate synthesis, ferrous iron uptake, and stress-response genes based on acetate consumption. Sci Rep 14, 987 (2024). https://doi.org/10.1038/s41598-023-51059-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-51059-3
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.