Abstract
Computational analysis of drug solubility was carried out using machine learning approach. The solubility of Decitabine as model drug in supercritical CO2 was studied as function of pressure and temperature to assess the feasibility of that for production of nanomedicine to enhance the solubility. The data was collected for solubility optimization of Decitabine at the temperature 308–338 K, and pressure 120–400 bar used as the inputs to the machine learning models. A dataset of 32 data points and two inputs (P and T) have been applied to optimize the solubility. The only output is Y = solubility, which is Decitabine mole fraction solubility in the solvent. The developed models are three models including Kernel Ridge Regression (KRR), Decision tree Regression (DTR), and Gaussian process (GPR), which are used for the first time as a novel model. These models are optimized using their hyper-parameters tuning and then assessed using standard metrics, which shows R2-score, KRR, DTR, and GPR equal to 0.806, 0.891, and 0.998. Also, the MAE metric shows 1.08E−04, 7.40E−05, and 9.73E−06 error rates in the same order. The other metric is MAPE, in which the KRR error rate is 4.64E−01, DTR shows an error rate equal to 1.63E−01, and GPR as the best mode illustrates 5.06E−02. Finally, analysis using the best model (GPR) reveals that increasing both inputs results in an increase in the solubility of Decitabine. The optimal values are (P = 400, T = 3.38E + 02, Y = 1.07E−03).
Similar content being viewed by others
Introduction
Production of nanomedicine has been a topic of first-rate hobby in pharmaceutical vicinity due to their importance significance for improving the drug efficacy. Nanosized drug production is one of the strategies to decorate drug solubility due to the expanded surface area to nanosized which therefore outcomes in an enhancement in drug solubility1–3. Given that, maximum newly discovered drugs are poorly soluble in aqueous media, underpinning studies is required to secorate the solubility of drugs thru one of the kind strategies such as nanonization, amorphous solid dispersion, crystallization, and salt formation4,5,6,7,8.
Drug nanonization has been used as an appealing technique for optimization of drugs thru solubility enhancement withinside the body. Supercritical solvents are extra appealing techniques due to the advanced characteristics of this process for preparation of nanosized drug particles3,6,9,10,11,12. The use of superficial CO2 as the secure solvent in pharmaceutical has been accredited by authorities. Moreover, there are some advantages of supercritical CO2 method like low price, easy operation, and moderate supercritical factor as compared to different gases. Therefore, supercritical CO2 is a great choice to be used as green solvent for preparation of nanomedicine in pharmaceutical area7,13,14,15,16.
Prior to implementing the nanonization process based on supercritical technology, first drug dissolution has to be analyzed in the solvent. Determination of drug solubility in supercritical process can be done via either experimental approach or computational, by which the computational method is more attractive17,18. Applying the experimental technique needs extensive time and cost for the analysis, while computational methods can save time and cost of the experiments, and they can be used for interpolation and extrapolation of the data19,20,21,22,23.
Different computational techniques have been utilized for the modeling of drug solubility, but the thermodynamic model and machine learning have shown better performance. The thermodynamic model establishes equilibrium between the solid phase and the solvent phase to determine the value of solubility19,24,25,26,27,28. On the other hand, machine learning models need measured data for training and validation of the algorithms29,30. The methods of machine learning have shown to be easier to implement and offer better accuracy for prediction of drug solubility in solvents.
Machine learning (ML) as a subject in artificial intelligence is a set of techniques to understand the patterns of data with no any suppostions regarding to the structure. One of these strategies' strengths is creating a relation among data and, then estimate some interaction. An important application of machine learning is regression, that could be defined as a specific type of problem in this study31,32,33,34. In this research, three approaches have been chose to make approaches on the drug solubility. Accepted methods in this research are Kernel Ridge Regression (KRR), Decision Tree Regression (DTR), and Gaussian Process Regression (GPR). Indeed, we implemented these efficient ML models for the first time for simulation of decitabine solubility in supercritical CO2 as the solvent. The results can help to assess the applicability of supercritical process for this drug candidate to be prepared in nanosized scale35.
Ridge regressions and the kernel approach are used in the Kernel Ridge Regression (KRR). KRR has the advantage of capturing nonlinear relationships, avoiding regression over-fitting problems through regularization and kernel techniques35.
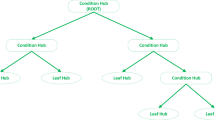
A decision tree regressor (DTR) is a straightforward, comprehensible, and efficient approach. The core principle of the decision tree algorithm has been distributed a large problem within multiple smaller sub-problems, it can be lead to an easier-to-interpret respond36,37. A DTR demonstrates a set of conditional queries ordered hierarchically and requested from the tree's root to the leaf38. DTRs are easy to understand and have a clear structure. DTRs produce a trained predictor, be able to express principles, and forecast new datasets using the splitting procedures, which is repeated39,40.
The other employed model of this study is based on the Gaussian process (GP) statistical concept, which refers to a group of random variables, as some of them are distributed with Gaussian distributions41,42. In geostatistics, the Gaussian process is the fundamental stochastic process. Gaussian processes directly represent Gaussian data and the base for non-Gaussian models such as linear regression models. As a result, Gaussian processes regression based on GPs is both accurate and straightforward for small datasets with high generality43,43,45. Additional target prediction studies of decitabine will be conducted in this research work to get better insights about the different plausible targets for this drug. We will use a hybrid approach in this study through combining both binding and ligand similarity analysis to predict other putative targets of decitabine. The purpose of this research is modifying the solubility of decitabine and GPR has been selected as the best model and reveals that increasing both inputs roughly increase the solubility of drug. So, the optimal is (P = 400, T = 3.38E + 02, Y = 1.07E-03).
Data set
To make models on solubility, we used a dataset with 32 data points identical to the reference46,47. Indeed, the experimental data have been collected from the reference and the machine learning models were fitted and implemented on the data. More detailed description of the method and experimental conditions can be found in the source published paper in46. Here, two inputs are considered, Pressure (bar) and Temperature (K), and a single output that shows the solubility of Decitabine drug in the supercritical carbon dioxide (CO2). The entire dataset is shown in Table 1.
Methodology
Kernel ridge regression (KRR)
The first machine learning (ML) method which is considered here for correlation of drug solubility values is the method of Kernel Ridge Regression (KRR). Suppose a data set \({\left\{({x}_{i},{y}_{i})\right\}}_{i=1}^{N}\) has been provided which is include \(N\) data points, and the goal is to estimate a function can analysis the mean squared error (MSE) of [\({(f\left(x\right)-y)}^{2}\)]. The conditional mean \({f}^{*}\left(x\right) : ={\mathbb{E}}[Y|X=x]\) has been illustrated as the best function48. In order to estimate the function \({f}^{*}\),
This equation can predict the kernel ridge regression35.
Decision tree regression (DTR)
A regression tree or decision trees regressor38 uses data from simulation inputs and outputs to create a structure that can be a leaf (terminal node), illustrating a estimation value, or an internal node (decision node), indicating some query to be performed on an input, with a branch and child for each possible output of the query. For continuous inputs, two options are available based on whether the condition is true or not. The structure of the data is declared at every node of the regression tree. To estimate the output for an unobserved data point, the inputs of that data point are employed to traverse the decision tree until a terminal node is seen. The estimated value is decided according to the output values from the training set ending up at that terminal node51.
An impurity measure for each node of the tree's test is decided by reviewing all input feature and obtaining an optimal split that maximizes the measure. MSE can be calculated by formulating the split A as follows for a particular input52:
Here, tL and tR denote the set of instances. Also, s(t) indicates the standard deviation of the N(t) data, ci, of instances within t:
Here, \(\overline{c(t)}\) is the average of the values in t. The split that minimizes mean square error across all input features for instances at each node of the regression tree is used at each node. Overfitting can occur in tree-based algorithms if the data is split too finely53,54,52.
Gaussian process regression (GPR)
Successor models, such as the Gaussian process (GP), provide predictions as well as the degree of uncertainty associated with those predictions. A GP is a group of random variables with the same Gaussian distribution for any quantity of variables42. GPs can be assumed as an infinite-dimensional buildup of multivariate Gaussian distributions. N- instance training data can be considered a singular data point taken from an N-variate Gaussian distribution; thus, it can be matched to the Gaussian process. Typically, the average of this Gaussian Process is reserved to zero.
We describe GP55 using a one-dimensional problem with an N-instance training set, [xi | i = 1,2,…,N] and the corresponding output values y = [y1, …, yN]. We use the same notation as in the previous sections to describe GP for a one-dimensional problem for ease of exposition. Two instances xi and xj in the training set are related to each other through the covariance function k(xi, xj). The squared-exponentiation function is employed here56:
where \({\sigma }_{f}^{2}\) the maximum allowable covariance and l is a length parameter that controls the extent of influence of each point. \({\sigma }_{f}^{2}\) Should be set to a large value for functions covering a broad range of values. In condition that data points xi and xj are close to each other, their output values are highly correlated, but if they are far away, then the value at one point does not influence the value at the other point. Accordingly, the hyper-parameter l determines the smoothness of the interpolation.
Assume we desire to employ the training data to estimate the output at an unseen data point x∗. Since the results be able toe depicted as an instance through a multivariate Gaussian distribution:
y denotes the output variable correlated to the N training data points, y∗ shows the estimated production at x∗ and the following sub-matrices:
And:
The probability of y∗, which is, the output at a data point, is formulates as:
The variance indicates the degree of uncertainty in the estimate:
The parameters l and σf of the Gaussian process regressor can be computed from the training subset using a maximum likelihood method. It is also feasible to incorporate a Gaussian noise component in the output variable, however we have supposed that the noise is zero in our current research.
Prediction of decitabine putative targets
Decitabine Smiles were generated via PubChem (https://pubchem.ncbi.nlm.nih.gov/compound/Decitabine) then we feed the smiles into The LigTMap server (https://cbbio.online/LigTMap/?action=home) to identify the plausible targets from seventeen target classes and more than six thousands of different types of proteins.
Results and discussion
Analysis of model outcomes
The three abovementioned models were implemented to the collected dataset to build the models for the drug solubility. The hyper-parameters of the models we introduced were optimized using Grid-Search57. More than 1000 distinct combinations were used to get these ideal parameters for each model. Then, the models were tested in their ideal configurations, and their performance was evaluated.
Three traditional statistical metrics will be used to assess and compare the efficiency of each model, such as R2 and Mean Absolute Error (MAE) and MAPE. In order to calculate each of the statistics, a mathematical equation must be used52:
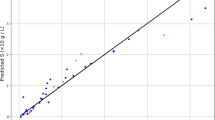
In these equations, n is size of data set, Yi,m is the estimated value, Yi,o indicates actual (observed) value. As well, \({\overline{Y}}_{i,m}\) is the average of estimated values and \({\overline{Y}}_{i,o}\) indicates average of actual values. A comparison among the estimated amounts and the real (observed) amounts in the model training is shown in Figs. 1, 2, and 3 for the methods of KRR, DTR, and GPR, respectively. The red dots indicate the test data, the blue dots are the training data (estimated amounts), and the green line represents the real amounts. Comparing these three shapes clearly shows the higher generality in the GPR method in comparison to other methods. The statistical results of the comparison for all methods have been also demonstrated in Table 2. As it is clear, all methods have great capability in fitting and correlating the experimental data which indicate that these models are of great choice for application in production of nanomedicine using supercritical based technology. The best outputs are illustrated for GPR through R2 higher than 0.99 in order to fit the solubility results.
The validated GPR method as the significant method has been applied to calculate the solubility data and find the influence of temperature and the pressure on the solubility of Decitabine in supercritical CO2. The results of 3D surface plot are explained in Fig. 4, the impact of temperature and pressure on the solubility of decitabine are significant, so that the highest value of solubility is observed at the maximum values of T and P in the 3D graph (see Fig. 4). The increase in the solubility with temperature and pressure could be attributed to the change of solvent density and consequently changing the solvation capacity of the solvent. Also, the 2D graphs of solubility versus temperature and pressure are indicated in Figs. 5 and 6, respectively. The optimum values calculated using the GPR model are listed in Table 3.
Figure 7 aims to evaluate the impact of pressure on the solubility values of Decitabine at disparate temperatures. As indicated, there is a cross over area at each solubility figure. Indeed, the impact of temperature on drug solubility in SC-CO2 is paradoxical. Furthur, temperature's growth, influence on the sublimation pressure of drugs, causes the increment of solubility. In another side, increase the temperature results in decreasing the molecular compaction and as the result, the amount of SC-CO2 density, which has negative effect on the solubility of Decitabine. The pressure value of 18 bar is known as the cross over pressure. At the pressures between 12 and 18 bar, the negative effect of density deterioration entirely overcomes the desirable effect of vapor pressure increment. Moreover, at this range of pressures, temperature enhancement lead to a reduction in solubility. Above the cross over pressure (18 MPa), the solubility of Decitabine significantly enhances owing to the superiority of the positive effect of drug’s vapor pressure than the negative effect of density reduction. Therefore, at this pressure elevation of temperature improves the solubility.
Correlation of the solubility data with semi-empirical models
Figure 8a–d present the correlation outcomes of Decitabine-SC-CO2 system obtained by semi-empirical models. In this investigation, four principal semi-empirical density-based models (Sodeifian et al., K-J, Bartle et al. and Bian et al.) were pondered for the correlation of the experimental data of Decitabine solubility SC-CO258,56,57,58,62. Disparate values including settable parameters (a0, a1, a2, a3, a4 and a5), average absolute relative deviation (AARD%) and R2 are enlisted in Table 4. The AARD for developed models for Sodeifian et al., Bartle et al. and Bian et al. models were 12.15%, 11.61%, 14.46%, and 13.25%, respectively. Comparison of the results implies the fact that K-J model is the best model due to presenting the lowest value of AARD (11.61%).
Table 4 presents the correlation outcomes of Decitabine-SC-CO2 system obtained by semi-empirical models.
Additionally, through usage of LigTMap server we have found more than one hundred predicted targets for decitabine, these targets are classified according to disease target class into kinase 34 (29%), 30 (25%) transferase, 28 (24%) Hydrolase, 15 (13%) tuberculosis, 5 (4%) Hpyroli, 3 (2.5%) Influenza and 1 (0.8%) Beta secretase. Attached with this research work a supplementary data file that contains a list for the targets with docking scores in the binding sites of the specified proteins. Also, Pdb IDs for each specific protein are incorporated, the optimum binding of decitabine with these target proteins and binding mode, in addition to predicted affinity and docking scores all are obtained through the automated workflow of LigTMap. The obtained results (supplementary data) revealed that decitabine has ligand Similarity Score more than 0.6 with Deoxycytidine kinase and Thymidylate kinase TMK, target classes are kinase and tuberculosis with Pdb ID 3ipx and 1w2g respectively. Decitabine showed binding affinity more than 7 in disease target class Influenza, target name is Polymerase basic protein 2 (Pdb IDs: 5efc, 4or6 and 4q46). Decitabine showed the best docking score into Thymidylate kinase binding site with value equal −7.007 kcal/mol, Pdb ID: 1mrs in tuberculosis disease class. From these obtained results we can conclude that Thymidylate kinase (tuberculosis) and Polymerase basic protein 2 (Influenza) are plausible targets for decitabine, the following Fig. 9 illustrates the binding mode with Thymidylate kinase (tuberculosis).
Four semi-empirical models (Sodeifian et al., Bartle et al., K-J and Bian et al.) have been considered to make a correlation with the outputs of solubility experiments. The precision for all applied methods has analysed and measured through AARD% and R2. Comparison of the outputs implies the fact that K-J model is the best model due to presenting the lowest value of AARD (11.61%). Despite good efficiency of K-J model for the accurate prediction of drug solubility, the employed GPR model shows better performance compared to K-J owing to having higher value of R2.
Conclusion
Computational simulation of Decitabine drug solubility in supercritical carbon dioxide was carried out in this study via three different machine learning models. We used a dataset of 32 data points and two inputs in this investigation to create solubility models (P and T). In this dataset, Y (solubility, mole fraction) is the lone result which is predicted by the models. Kernel Ridge Regression (KRR), Decision Tree Regression (DTR), and Gaussian process (GP) are the models which were employed in this work for correlation of the solubility data. Hyper-parameter tweaking was used to fine-tune these models, and standard metrics were used to assess their performance. KRR, DTR, and GPR have R2-scores of 0.806, 0.891, and 0.998. MAE's error rate is 1.08E−04, 7.40E−05, and 9.73E−06 in that sequence, too. The MAPE measure has a KRR error rate of 4.64E−01, a DTR error rate of 1.63E−01, and a GPR error rate of 5.06E−02 as the optimum option. As a conclusion, the best model (GPR) shows that increasing both inputs roughly raise the output. So, the best outcome is obtained as P = 400 bar, T = 3.38E + 02 °K, Y = 1.07E−03. Finally, LigTMap workflow revealed the promiscuity of decitabine to target Thymidylate kinase (disease class: tuberculosis) and Polymerase basic protein 2 (disease class: influenza). In this paper, the solubility value of Decitabine was evaluated at disparate values of pressure (120, 160, 200, 240, 280, 320, 360 and 400 bar) and temperatures (308, 318, 328, and 338 K). Four semi-empirical models (Sodeifian et al., Bartle et al., K-J and Bian et al.) have been considered to make a correlation with the outputs of solubility experiments. The precision of all applied methods has been evaluated through AARD% and R2. Comparison of the outputs implies the fact that K-J model is the best model due to presenting the lowest value of AARD (11.61%). Despite good efficiency of K-J model for the accurate prediction of drug solubility, the employed GPR model shows better performance compared to K-J owing to having higher value of R2.
Data availability
All data are available within the published paper.
Abbreviations
- ML:
-
Machine learning
- P:
-
Pressure
- T:
-
Temperature
- KRR:
-
Kernel ridge regression
- GPR:
-
Gaussian process regression
- DTR:
-
Decision tree regression
- MSE:
-
Mean squared error
- MAE:
-
Mean absolute error
- AARD:
-
Average absolute relative deviation
- SC-CO2 :
-
Supercritical CO2
references
Pishnamazi, M. et al. Evaluation of supercritical technology for the preparation of nanomedicine: etoricoxib analysis. Chem. Eng. Technol. 44, 559–564 (2021).
Khoshmaram, A. et al. Supercritical process for preparation of nanomedicine: Oxaprozin case study. Chem. Eng. Technol. 44, 208–212 (2021).
Sodeifian, G., Razmimanesh, F., Ardestani, N. S. & Sajadian, S. A. Experimental data and thermodynamic modeling of solubility of Azathioprine, as an immunosuppressive and anti-cancer drug, in supercritical carbon dioxide. J. Mol. Liq. 299, 112179 (2020).
Shaikh, R., Shirazian, S. & Walker, G.M. Application of artificial neural network for prediction of particle size in pharmaceutical cocrystallization using mechanochemical synthesis. Neural Comput. Appl. (2021).
Zabihi, S., Esmaeili-Faraj, S. H., Borousan, F., Hezave, A. Z. & Shirazian, S. Loxoprofen solubility in supercritical carbon dioxide: Experimental and modeling approaches. J. Chem. Eng. Data 65, 4613–4620 (2020).
Zabihi, S. et al. Thermodynamic study on solubility of brain tumor drug in supercritical solvent: Temozolomide case study. J. Mol. Liq. 321, 114926 (2021).
Hazaveie, S. M., Sodeifian, G. & Sajadian, S. A. Measurement and thermodynamic modeling of solubility of Tamsulosin drug (anti cancer and anti-prostatic tumor activity) in supercritical carbon dioxide. J. Supercrit. Fluids 163, 104875 (2020).
Zhuang, W., Hachem, K., Bokov, D., Javed Ansari, M. & TaghvaieNakhjiri, A. Ionic liquids in pharmaceutical industry: A systematic review on applications and future perspectives. J. Mol. Liq. 349, 118145 (2022).
Zhao, Z. et al. Multi support vector models to estimate solubility of Busulfan drug in supercritical carbon dioxide. J. Mol. Liq. 350, 118573 (2022).
Operti, M. C. et al. PLGA-based nanomedicines manufacturing: Technologies overview and challenges in industrial scale-up. Int. J. Pharmaceut. 605, 120807 (2021).
Penoy, N., Grignard, B., Evrard, B. & Piel, G. A supercritical fluid technology for liposome production and comparison with the film hydration method. Int. J. Pharmaceut. 592, 120093 (2021).
Campardelli, R., Baldino, L. & Reverchon, E. Supercritical fluids applications in nanomedicine. J. Supercrit. Fluids 101, 193–214 (2015).
Sodeifian, G., Sajadian, S. A. & Ardestani, N. S. Determination of solubility of Aprepitant (an antiemetic drug for chemotherapy) in supercritical carbon dioxide: Empirical and thermodynamic models. J. Supercrit. Fluids 128, 102–111 (2017).
Sodeifian, G. & Sajadian, S. A. Solubility measurement and preparation of nanoparticles of an anticancer drug (Letrozole) using rapid expansion of supercritical solutions with solid cosolvent (RESS-SC). J. Supercrit. Fluids 133, 239–252 (2018).
Sodeifian, G., Razmimanesh, F. & Sajadian, S. A. Prediction of solubility of sunitinib malate (an anti-cancer drug) in supercritical carbon dioxide (SC–CO2): Experimental correlations and thermodynamic modeling. J. Mol. Liq. 297, 111740 (2020).
Pishnamazi, M. et al. Measuring solubility of a chemotherapy-anti cancer drug (busulfan) in supercritical carbon dioxide. J. Mol. Liq. 317, 113954 (2020).
Esfandiari, N. & Sajadian, S. A. Experimental and modeling investigation of Glibenclamide solubility in supercritical carbon dioxide. Fluid Phase Equilib. 556, 113408 (2022).
Sodeifian, G., Sajadian, S.A., Razmimanesh, F. & Hazaveie, S.M. Solubility of the Ketoconazole (an Antifungal Drug) in Supercritical Carbon Dioxide and Menthol as a Cosolvent (Ternary System): Experimental Data and Empirical Correlations. (2021).
Sodeifian, G., Sajadian, S. A., Razmimanesh, F. & Ardestani, N. S. A comprehensive comparison among four different approaches for predicting the solubility of pharmaceutical solid compounds in supercritical carbon dioxide. Korean J. Chem. Eng. 35, 2097–2116 (2018).
Sodeifian, G., Razmimanesh, F., Sajadian, S. A. & Hazaveie, S. M. Experimental data and thermodynamic modeling of solubility of Sorafenib tosylate, as an anti-cancer drug, in supercritical carbon dioxide: Evaluation of Wong-Sandler mixing rule. J. Chem. Thermodyn. 142, 105998 (2020).
Suleiman, D., Estévez, L. A., Pulido, J. C., García, J. E. & Mojica, C. Solubility of anti-inflammatory, anti-cancer, and anti-HIV drugs in supercritical carbon dioxide. J. Chem. Eng. Data 50, 1234–1241 (2005).
Pishnamazi, M. et al. Thermodynamic modelling and experimental validation of pharmaceutical solubility in supercritical solvent. J. Mol. Liq. 319, 114120 (2020).
Sodeifian, G., SaadatiArdestani, N., Sajadian, S. A., Golmohammadi, M. R. & Fazlali, A. Prediction of solubility of sodium valproate in supercritical carbon dioxide: Experimental study and thermodynamic modeling. J. Chem. Eng. Data 65, 1747–1760 (2020).
Sodeifian, G., SaadatiArdestani, N., Razmimanesh, F. & Sajadian, S. A. Experimental and thermodynamic analyses of supercritical CO2-solubility of minoxidil as an antihypertensive drug. Fluid Phase Equilibr. 522, 112745 (2020).
Sodeifian, G., Sajadian, S. A. & Derakhsheshpour, R. Experimental measurement and thermodynamic modeling of Lansoprazole solubility in supercritical carbon dioxide: Application of SAFT-VR EoS. Fluid Phase Equilibr. 507, 112422 (2020).
Sodeifian, G., Detakhsheshpour, R. & Sajadian, S. A. Experimental study and thermodynamic modeling of Esomeprazole (proton-pump inhibitor drug for stomach acid reduction) solubility in supercritical carbon dioxide. J. Supercrit. Fluid 154, 104606 (2019).
Yamini, Y. et al. Solubility of capecitabine and docetaxel in supercritical carbon dioxide: Data and the best correlation. Thermochim. Acta 549, 95–101 (2012).
Xiang, S.-T., Chen, B.-Q., Kankala, R. K., Wang, S.-B. & Chen, A.-Z. Solubility measurement and RESOLV-assisted nanonization of gambogic acid in supercritical carbon dioxide for cancer therapy. J. Supercrit. Fluids 150, 147–155 (2019).
Zhu, H., Zhu, L., Sun, Z. & Khan, A. Machine learning based simulation of an anti-cancer drug (busulfan) solubility in supercritical carbon dioxide: ANFIS model and experimental validation. J. Mol. Liq. 338, 116731 (2021).
Sadeghi, A. et al. Machine learning simulation of pharmaceutical solubility in supercritical carbon dioxide: Prediction and experimental validation for busulfan drug. Arab. J. Chem. 15, 103502 (2022).
Senders, J. T. et al. Machine learning and neurosurgical outcome prediction: a systematic review. World Neurosurg. 109, 476-486e471 (2018).
Cherkassky, V. & Ma, Y. Comparison of model selection for regression. Neural Comput. 15, 1691–1714 (2003).
Carbonell, J. G., Michalski, R. S. & Mitchell, T. M. An overview of machine learning. Mach. Learn. 45, 3–23 (1983).
Goodfellow, I., Bengio, Y. & Courville, A. Machine learning basics. Deep Learn. 1, 98–164 (2016).
Rami, M. A. et al. Novel numerical simulation of drug solubility in supercritical CO2 using machine learning technique: Lenalidomide case study. Arab. J. Chem. 15(11), 104180 (2022).
Xu, M., Watanachaturaporn, P., Varshney, P. K. & Arora, M. K. Decision tree regression for soft classification of remote sensing data. Remote Sens. Environ. 97, 322–336 (2005).
Song, Y.-Y. & Ying, L. Decision tree methods: Applications for classification and prediction. Shanghai Arch. Psychiatry 27, 130 (2015).
Breiman, L., Friedman, J. H., Olshen, R. A. & Stone, C. J. Classification and Regression Trees (Routledge, 2017).
Ahmad, M. W., Mourshed, M. & Rezgui, Y. Trees vs neurons: Comparison between random forest and ANN for high-resolution prediction of building energy consumption. Energy Build. 147, 77–89 (2017).
Dumitrescu, E., Hue, S., Hurlin, C. & Tokpavi, S. Machine learning for credit scoring: Improving logistic regression with non-linear decision-tree effects. Eur. J. Oper. Res. 297, 1178–1192 (2022).
Grbić, R., Kurtagić, D. & Slišković, D. Stream water temperature prediction based on Gaussian process regression. Expert Syst. Appl. 40, 7407–7414 (2013).
Rasmussen, C. E. & Williams, C. K. Gaussian Processes for Machine Learning (MIT Press, 2006).
Rasmussen, C. E. Gaussian Processes in Machine Learning. Summer School on Machine Learning (Springer, 2003).
Daemi, A., Kodamana, H. & Huang, B. Gaussian process modelling with Gaussian mixture likelihood. J. Process Control 81, 209–220 (2019).
Wang, H., Guan, Y., & Reich, B. Nearest-neighbor neural networks for geostatistics. in 2019 International Conference on Data Mining Workshops (ICDMW). 196–205 (IEEE, 2019).
Pishnamazi, M. et al. Experimental and thermodynamic modeling decitabine anti cancer drug solubility in supercritical carbon dioxide. Sci. Rep. 11, 1–8 (2021).
Bader, H. et al. Solubility enhancement of decitabine as anticancer drug via green chemistry solvent: Novel computational prediction and optimization. Arab. J. Chem. 15(12), 104259 (2022).
Byrne, E. & Schniter, P. Sparse multinomial logistic regression via approximate message passing. IEEE Trans. Signal Process. 64, 5485–5498 (2016).
Zhang, Y., Duchi, J. & Wainwright, M. Divide and conquer kernel ridge regression. in Conference on Learning Theory, PMLR. 592–617 (2013).
Hoerl, A. E. & Kennard, R. W. Ridge regression: Biased estimation for nonorthogonal problems. Technometrics 12, 55–67 (1970).
King, W. E. et al. Laser powder bed fusion additive manufacturing of metals; physics, computational, and materials challenges. Appl. Phys. Rev. 2, 041304 (2015).
Seyed, A. S. et al. Solubility of favipiravir (as an anti-COVID-19) in supercritical carbon dioxide: An experimental analysis and thermodynamic modeling. J. Supercrit. Fluids. 183, 105539 (2022).
Kamath, C., & Cantu-Paz, E. Creating Ensembles of Decision Trees Through Sampling. (Lawrence Livermore National Lab (LLNL), 2001).
Kamath, C. Scientific Data Mining: A Practical Perspective (SIAM, 2009).
Ebden, M. Gaussian Processes: A Quick Introduction. arXiv preprint arXiv:1505.02965 (2015).
Kamath, C. Data mining and statistical inference in selective laser melting. Int. J. Adv. Manuf. Technol. 86, 1659–1677 (2016).
Lerman, P. Fitting segmented regression models by grid search. J. R. Stat. Soc. Ser. C (Appl. Stat.) 29, 77–84 (1980).
Kumar, S. K. & Johnston, K. P. Modelling the solubility of solids in supercritical fluids with density as the independent variable. J. Supercrit. Fluids 1, 15–22 (1988).
Bartle, K., Clifford, A., Jafar, S. & Shilstone, G. Solubilities of solids and liquids of low volatility in supercritical carbon dioxide. J. Phys. Chem. Ref. Data 20, 713–756 (1991).
Esfandiari, N. & Sajadian, S. A. Solubility of lacosamide in supercritical carbon dioxide: An experimental analysis and thermodynamic modeling. J. Mol. Liq. 26, 119467 (2022).
Bian, X.-Q., Zhang, Q., Du, Z.-M., Chen, J. & Jaubert, J.-N. A five-parameter empirical model for correlating the solubility of solid compounds in supercritical carbon dioxide. Fluid Phase Equilib. 411, 74–80 (2016).
Sodeifian, G., Razmimanesh, F. & Sajadian, S. A. Solubility measurement of a chemotherapeutic agent (Imatinib mesylate) in supercritical carbon dioxide: Assessment of new empirical model. J. Supercrit. Fluids 146, 89–99 (2019).
Acknowledgements
The authors extend their appreciation to the Deputyship for Research & Innovation, Ministry of Education in Saudi Arabia for funding this research work through the project number (IF-PSAU-2021/03/18826). The authors would like to thank the Deanship of scientific research at Umm Al-Qura University for supporting this work by grant code (22UQU4290565DSR93). Authors would like to introduce appreciated thanks to Taif University Researchers Supporting, Project number (TURSP-2020/35), Taif University, Taif, Saudi Arabia. The authors extend their appreciation to the Deanship of Scientific Research at King Khalid University for funding this work through Large Groups Project under grant number (R.G.P. 2/50/43).
Author information
Authors and Affiliations
Contributions
S.M.A: supervision, writing the original draft, software, validation B.K.A: data analysis, writing the original draft, optimization M.M.A: investigation, optimization, writing the original draft, data analysis A.B: Supervision, writing the original draft, validation, modeling, software M.A.S.A: writing the original draft, modeling, software A.A.S: editing, data analysis, writing, investigation A.S.A: editing, writing, investigation, data analysis K.V: editing, writing, investigation, data analysis A.M.A: editing, writing, investigation, data analysis M.P: supervision, writing the original draft, software, validation.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Alshahrani, S.M., Almutairy, B.K., Alfadhel, M.M. et al. Computational simulation and target prediction studies of solubility optimization of decitabine through supercritical solvent. Sci Rep 12, 18875 (2022). https://doi.org/10.1038/s41598-022-21233-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-21233-0
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.