Abstract
Oral squamous cell carcinoma (OSCC) is a common malignant tumor worldwide that is characterized by abnormal lesions or malignant hyperplasia of soft and hard tissues in the oral cavity. Previous research has found that HDAC6 may be a potential therapeutic target for cancer patients and has the ability to regulate immune cells. However, the mechanism of HDAC6 in OSCC pathogenesis is unclear. We collected clinical samples and analyzed the level of HDAC6 in OSCC patients. The results showed that in the high HDAC6 expression group, HDAC6 expression was positively correlated with the grade of OSCC (R = 0.182, P = 0.036) and that this group had a 3.248-fold increase in the mortality risk compared with the low HDAC6 expression group (P = 0.003). Survival analysis also identified a correlation between the expression of HDAC6 and overall survival in OSCC patients, and it was found that the expression of HDAC6 was inversely correlated with survival (P ≤ 0.001). In addition, we found that HDAC6 induced IL-13 expression through AP-1, resulting in M2 polarization of macrophages. Together, these results demonstrate that the level of HDAC6 may be a useful prognostic biomarker and offer a novel immune cell-related therapeutic strategy of targeting IL-13 in OSCC.
Similar content being viewed by others
Introduction
Oral squamous cell carcinoma (OSCC) is the seventh most common malignant tumor worldwide1 and is characterized by abnormal lesions or malignant hyperplasia of oral tissues, including the lips, the buccal mucosa (the lining of the lips and cheeks), the upper two-thirds of the tongue, the front part of the roof of the mouth (hard palate), the teeth, the sublingual floor of the mouth, the gums, and the small area behind the molars. The most common sites of OSCC in Taiwan are the tongue and buccal mucosa. OSCC mostly occurs in elderly people aged 55 to 64 years old. The main risk factors for OSCC are smoking, drinking, tobacco use, and betel nut chewing. Among these risk factors, smoking and drinking have a synergistic influence; other factors, including bad eating habits, viral infections (especially HPV infection) and poor oral hygiene, can also affect OSCC carcinogenesis2,3. According to cancer statistics, approximately 3% of cancers diagnosed in 2020 were OSCC, and more men than women have OSCC1. The most common type of OSCC is squamous cell carcinoma (SCC), which accounts for more than 90% of all OSCC cases. Other malignant tumors may arise from epithelial tissue, connective tissue, salivary glands, lymphoid tissue, and melanocytes or distant metastasis of the primary tumor4,5. Macrophages are multifunctional immune cells of the human innate immune system; they act through tissue-specific and environment-dependent mechanisms to regulate immune homeostasis in the body and play important roles as mediators of inflammation and antitumor immunity6,7,8. When inflammation and infection occur, blood monocytes accumulate in tissues and undergo a series of steps to differentiate into macrophages9. Macrophages have a high degree of plasticity, and exhibit disparate morphological manifestations under exposure to different environmental stimuli. Under physiological conditions, macrophages (M0) are induced by lipopolysaccharide (LPS), IFN-γ or TNF-α to polarize into M1-type macrophages (M1), and M1 macrophages secrete inflammatory factors (such as TNF-α, IL-1β, IL-6, and IL-12), which activate the immune system and defend against exogenous bacteria and viruses10,11. In contrast, when M0 macrophages are exposed to different environmental stimuli (such as IL-4 and IL-13), they polarize into M2-type macrophages (M2), which cause an anti-inflammatory response12. In tumors, macrophages are usually present as the M2 phenotype by exposure to environmental factors. M2 macrophages can inhibit cytotoxicity, thereby weakening their antitumor ability and promoting tumor growth13,14,15. In contrast, if the activation of M2 macrophages is blocked, the processes of tumor growth and metastasis can be inhibited, and M2 macrophages have thus been used in clinical treatment16,17,18,19. Current studies have shown that M1 macrophages have the ability to enhance the activation of inflammatory factors and phagocytose invading pathogens, while M2 macrophages have the opposite function, inhibiting immune functions and promoting tumor formation through IL-13. A previous study also found that an increased number of M2 macrophages correlates with a higher histopathological grade of oral squamous cell carcinoma (OSCC)20. However, the detailed mechanism is unclear. Interleukins (ILs) are cytokines that play an important role in the activation, differentiation, proliferation, maturation, and migration of immune cells. Interleukins constitute a large family of proteins that can promote many reactions in cells and tissues by binding to high-affinity receptors on the cell surface21. Among ILs, IL-13 is synthesized by CD4 + T cells (Th2), NKT cells, and mast cells, and it acts on monocytes, fibroblasts, epithelial cells, and B cells. For example, it regulates the growth and differentiation of B cells, stimulates isotype conversion to IgE (immunoglobulin E), increases the amount of mucus in epithelial cells, promotes collagen synthesis by fibroblasts, and inhibits the production of inflammatory factors22. Studies indicate that IL-13 is regulated by two receptors—the IL-13Ra1/IL-4 Ra complex and IL-13Ra223; and IL-13Ra2 is a subunit of the IL-13R complex that has high affinity for IL-13. IL-13 induces the activity of the TGFβ1 promoter in an AP-1-dependent manner, which in turn leads to inflammation and fibrosis24. In addition, IL-13 transmits signals via the IL-13Rα2 and AP-1/ERK pathways to regulate tumor invasion and metastasis. According to a previous report, IL-13 regulates signal transduction through IL-13Rα2 in pancreatic cancer and ovarian cancer cells25. Histone deacetylase proteins (HDACs) constitute a family of enzymes that remove acetyl functional groups from histone proteins on DNA. HDAC6 has a unique structure among HDACs; it has two catalytic domains and a unique C-terminal zinc finger domain that binds to ubiquitin26. HDAC6 is located in the cytoplasm and mainly interacts with Non-histone proteins27; it is involved in many biological and pathological processes, such as cell migration, the DNA damage response and carcinogenesis, by regulating its substrates28,29. For example, the DNA mismatch repair protein MSH2 can be deacetylated and ubiquitinated by HDAC6 to regulate DNA mismatch repair and the DNA damage response30. Additionally, HDAC6 deacetylates the cell signaling regulators K-Ras and β-catenin, changing their carcinogenic activity and nuclear localization, respectively31,32. Related studies have shown that inhibition of HDAC6 may abolish the stability of oncogenic proteins, block tumor migration, and inhibit survival signals33. In addition, other studies have demonstrated that HDAC6 can regulate cancer-related signaling pathways, proving that HDAC6 has the potential to become a therapeutic target for cancer34,35,36 and has the capability to regulate immune cells. However, the understanding of the function of HDAC6 in OSCC is relatively unclear. Based on previous studies, we concluded that the tumor microenvironment, HDAC6, and IL-13 play a very important role in the carcinogenesis of OSCC. However, the correlation among them is unclear. Therefore, we hypothesized that HDAC6 is a key regulator of macrophage polarization and may be a therapeutic target to modulate IL-13. Our findings provide a scientific basis to evaluate the potential benefits of HDAC6 in human OSCC and indicate that future studies to understand the role of related molecular pathways are warranted.
Materials and methods
Clinical specimens and Statistical analysis
Prior to the collection of tissue samples, each subject or their legal guardian signed an informed consent form approved by the Institutional Review Board (IRB) of E-Da Hospital (EMRP-108-150). All of the study methodology satisfied the regulations issued and relevant guidelines, and approved by the IRB of E-Da Hospital. IRB provides instancedouble coded and delinked clinical specimens to us for experiments. Tissue samples were collected from 133 patients with OSCC. Detailed information on each patient, including stage, grade, tumor size, T stage, N stage, M stage, recurrence status and metastasis status, was recorded and is summarized in Table 1. Statistical analysis was performed using GraphPad Prism version 6.0 software (GraphPad, La Jolla, CA, USA). Correlations of high and low HDAC6 intensity with clinicopathologic parameters were analyzed through a univariate analysis. The significance of differences between the two independent groups was analyzed with a two-tailed Student’s t-test. P < 0.05 was considered to indicate a significant difference in all statistical analyses.
Immunohistochemistry
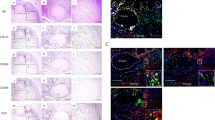
Patient tissues were sliced into 4-μm sections and stained with an automated Bond-Max immunostainer following the manufacturer’s protocol (Leica Microsystems). Slides were immunostained with a human anti-HDAC6 (Santa Cruz, sc-28386) antibody and counterstained with Mayer's hematoxylin (Sigma). Immunohistochemistry staining was assessed by 2 pathologist and clearly define for high and low intensity. The staining interpretation method was based on Iliopoulos D et al., 200537. The total score is calculated by multiplying the tumor cell range by the intensity. Representative images of HDAC6-positive and -negative tissues are presented in Fig. 1A.
Cell lines
The human oral squamous cell carcinoma (OSCC) cell lines OC-2, OECM-1 and human monocytic cell line (THP-1) were obtained from the American Type Culture Collection (ATCC) and maintained in DMEM/F12 culture medium (Gibco, France) supplemented with 1% penicillin/streptomycin and 10% fetal bovine serum (FBS) (Gibco) in a humidified incubator at 37 °C and 5% CO2.
Transfection and Luciferase assay
Cells (1X105) were seeded in 6-well plate. After 24 h, 2 µg of HDAC6 plasmid was transfected with TurboFect in vivo Transfection Reagent following the manufacturer’s protocol. For the luciferase assay, an IL-13 promoter sequence fragment (CTGGGAGTCAGAG) was synthesized and cloned into the pGL3-Basic expression vector (Promega). Luciferase activities were measured with a Dual-Luciferase Reporter Assay system and a GloMax 20/20 luminometer (Promega). In addition, luciferase assay data are presented as the mean ± SD of three independent replicates. P < 0.05 was considered to indicate a significant difference in all statistical analyses.
Cell growth assay
The cell growth assay was analyzed by Cell Counting Kit-8(CCK-8, Dojindo, Kumamoto, Japan). Brief, HDAC6-overexpring (5 × 103) cell were seeded into a 96-well plate 24 h before experimentation. Transient transgene expression occurs within 2–72 h after transfection according to TurboFect in vivo Transfection Reagent manufacturer’s protocol. Therefore, we monitoring the cell growth for 24, 48 and 72 h after HDAC6-overexpressing. The absorbances of each sample were measured at 450 nm in Microplate Reader.
Immunoblot analysis
Lysed cells were extracted with RIPA Cell Lysis Buffer, and proteins were transferred onto Amersham Hybond 0.45 PVDF membranes (GE Healthcare). The blots was cut prior and hybridization with antibodies. The following antibodies used for immunoblotting were purchased: anti-HDAC6(Santa Cruz, sc-28386), anti-IL13(Sigma, 3596M1), anti-p-AP1(Biovision, A1249), anti-AP1(Biovision, #3009) and anti-β-actin (Sigma, SAB5600204). The quantify of western blot bands was analyzed by Image J software in the Supplemental Materials.
RNA extraction and qPCR
To confirm that HDAC6 regulates the polarization of macrophages, the mRNA levels of polarization genes, including M1 (iNOS and TNFα) and M2 (IL-10 and arginase I) polarization genes, were determined by reverse transcription-PCR (RT-PCR) and quantitative PCR (qPCR). Cellular RNA was extracted by TRIzol (Invitrogen) and transcribed into cDNA with a Reverse Transcription System Kit (Promega, Madison, WI). The primers (Genemessenger Scientific, Tainan, Taiwan) used in the experiment were as follows. iNOS: forward, 5′-CAG-CTG-GGC-TGT-ACA-AAC-CTT-3′ and reverse, 5′-CAT-TGG-AAG-TGA-AGC-GTT-TCG-3′. TNF-α: forward, 5′-AAA-ATT-CGA-GTG-ACA-AGC-CTG-TAG-3′ and reverse, 5′-CCC-TTG-AAG-AGA-ACC-TGG-GAG-TAG-3′. IL-10: forward, 5′-AGA-AGC-ATG-GCC-CAG-AAA-TC-3′ and reverse, 5′-CCA AGG-AGT-TGT-TTC-CGT-TAGC-3′; Arginase 1: forward, 5′-AGA-GCT-GAC-AGC-AAC-CCT-GT-3′ and reverse, 5′-GGA-TCC-AGA-AGG-TGA-TGG-AA-3′. QPCR was performed with SYBR Green PCR Master Mix (Applied Biosystems, Foster City, CA) in an ABI 7500 Real-Time PCR System (Applied Biosystems).
Results
Correlation between the HDAC6 expression profile and unfavorable clinicopathological parameters
To confirm whether the expression of HDAC6 is related to the prognosis and survival of OSCC patients, we initially collected clinical samples from 133 OSCC patients and recorded the patients’ clinical characteristics, including stage, grade, tumor size, T stage, N stage, M stage, recurrence status and metastasis status. We used the descriptive statistical analysis mode in SPSS statistical software and found that most of the patients had grade 1 (45.1%) or grade 2 (53.4%) OSCC (Table 1). Next, we analyzed HDAC6 expression in the clinical specimens of 133 OSCC patients and performed immunohistochemical staining on these specimens using an automatic staining machine. We classified patients into groups with high and low intensity (Fig. 1A) with the assistance of a pathologist and performed statistical analysis of the correlation between HDAC6 expression and the clinical features of OSCC. We found that the expression of HDAC6 was positively correlated with the grade of OSCC (R = 0.182) and that the correlation was statistically significant (P < 0.05) (Table 2). Furthermore, we used linear regression (Cox) analysis to evaluate correlations with clinicopathological features. The results showed that the mortality risk in the high HDAC6 expression group was 3.248-fold greater than that in the low HDAC6 expression group and that this difference was statistically significant (P = 0.003) (Table 3).
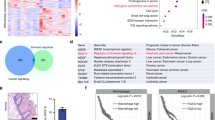
Finally, we analyzed the relationship between the expression of HDAC6 and the overall survival rate of OSCC patients through Kaplan–Meier analysis, and the results showed that the expression of HDAC6 was inversely correlated with the survival rate. When the expression of HDAC6 was high, the survival rate of patients with OSCC was decreased, and when the expression of HDAC6 was low, the survival rate was increased (Fig. 1B, P ≤ 0.001). The expression level of HDAC6 was thus correlated with the survival of OSCC patients.
HDAC6 promotes cell growth in OC-2 and OECM-1 cell lines
To analyze the effect of HDAC6 on cell growth in oral cancer cell lines, we evaluated cell growth with a CCK-8 assay in the OC-2 and OECM-1 cell lines. We transfected these cells with the HDAC6 overexpression plasmid, and the protein level of HDAC6 was analyzed by western blotting (Fig. 2A). The results showed that HDAC6 promoted cell growth in the OC-2 and OECM-1 cell lines (Fig. 2B and C).
HDAC6 induced the expression of IL-13 through AP-1
We next analyzed the ability of HDAC6 to regulate macrophage polarization into alternatively activated (M2) macrophages. A previous study found that IL-13 induced macrophage polarization from the M0 phenotype to the M2 phenotype38. In addition, treatment with an HDAC6 inhibitor was found to decrease the production of the Th2 cytokine Il-1339. Therefore, we speculated that HDAC6 may have the ability to induce the expression of IL-13. Transcription factor affinity prediction (TRAP) software was used to predict the transcription factor binding sites, and a binding motif for AP-1 (TGAGTCA) was identified in the regulatory region of the IL-13 promoter (Fig. 3A).To test the abovementioned hypothesis, luciferase reporter plasmids (pGL3) encoding wild-type (WT) and mutated (MT) fragments of the IL-13 5′-UTR were constructed and cotransfected with an AP-1 plasmid into HEK-293 T cells. We found that AP-1 overexpression increased luciferase reporter activity in a dose-dependent manner (Fig. 3B) in the WT group but not the mutated group (Fig. 3C). Next, to investigate the expression of IL-13, the OSCC cell lines OC-2 and OECM-1 were transfected with the HDAC6 overexpression plasmid, and the expression of IL-13 was determined by western blotting. The results showed that HDAC6 induced the expression of IL-13 in the OC-2 and OECM-1 cell lines (Fig. 3D). In addition, we used an AP-1 inhibitor (T-5224) to block the mechanism of AP-1, and these results showed that AP-1 phosphorylation was activated by HDAC6 overexpression but blocked by treatment with the AP-1 inhibitor (T-5224) in OSCC cell lines (Fig. 3E, F).
HDAC6 induced IL-13 expression via AP-1. (A) Transcription factor affinity prediction (TRAP) software predicted that the promoter region of IL-13 has an AP-1 transcription binding sequence. (B, C) pGL3-IL-13 luciferase plasmids containing wild-type (WT IL-13-Luc) or mutated (MT IL-13-Luc) 5′-UTRs were constructed. 293 T cells were cotransfected with the IL-13 5′-UTR plasmids and a plasmid expressing AP-1 at different dose ratios. (D) The HDAC6 plasmid (2 µg) was transfected into OECM-1 cells, and OC-2, HDAC6, IL-13, AP-1, p-AP-1 and B-actin were analyzed by western blotting. B-actin was used to normalize protein expression levels. (E) The AP-1 inhibitor T-5224 (40 µM) and (F) 2 µg of the HDAC6 plasmid were transfected into cells, and the level of IL-13 was analyzed by western blotting.
HDAC6 mediated THP-1 macrophage polarization into alternatively activated (M2) macrophages
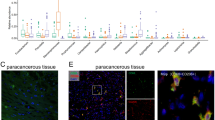
To explore whether HDAC6 can regulate the polarization of macrophages, we collected the supernatant of OSCC cells overexpressing HDAC6. The supernatant from untreated cells was called conditioned medium (CM), and the supernatant from the HDAC6-overexpressing cells was called HDAC6-induced CM. The mRNA levels of the M1 (iNOS and TNFα) and M2 (IL-10 and arginase I) macrophage markers were analyzed by qPCR after THP-1 cells were exposed to CM, HDAC6-induced CM, IL-13 antibody and IL-13 antibody + HDAC6-induced CM. We found that compared to CM, HDAC6-induced CM from THP-1 cells significantly increased the expression of IL-10 and arginase I, but blocked by IL-13 antibody. (Fig. 4A,B). Therefore, we hypothesized that HDAC6 activates M2 polarization of THP-1 macrophages through the AP-1/IL-13 signaling pathway.
HDAC6-induced CM promoted M1 polarization of macrophages. CM, HDAC6-induced CM, IL-13 antibody and IL-13 antibody + HDAC6-induced CM from OEC-M1 (A) or OC-2 (B) cells was used to treat THP-1 cells. After 24 h, THP-1 cell mRNA was extracted, and iNOS, TNFα, IL-10 and arginase I gene expression was analyzed by qPCR.
Discussion
Many previous studies have found that macrophages have both positive and negative effects on tumor growth in various cancers. The ratio of M1 to M2 macrophages plays a major regulatory role and affects the function of the immune system. It is currently known that an increase in the number of M2 macrophages is related to the developmental process of oral squamous cell carcinoma40, to angiogenesis and to more malignant histopathological features (i.e., higher grade)41. Our study found that HDAC6 expression was significantly associated with the tumor grade and mortality risk in OSCC. Therefore, HDAC6 expression and macrophage polarization are strongly correlated. According to our results, HDAC6 induced IL-13 through AP-1 and activated the polarization of M0 macrophages into M2 macrophages. In addition, silencing HDAC6 decreased macrophage adhesion to the extracellular matrix and significantly blocked LPS-induced activation of macrophages. Hyperacetylation of tubulin in microtubules may impair microtubule dynamics, leading to failure of LPS-induced M1 activation42. We also found that HDAC inhibition shifted macrophage polarization toward the M2 phenotype through the GSK3β/PTEN/Akt axis43. Therefore, we believe that HDAC6 has the ability to regulate M1/M2 polarization of macrophages. The transcription factor activator protein-1 (AP-1) is a downstream factor of the M-CSF/MAPK signaling pathway and is a major transcription factor that mediates proinflammatory and activated macrophages44,45. Some results indicate that IL-6 mediates OSCC cell migration through the SYK/JNK/AP-1 signaling pathway46. Previous studies have found that HDAC6 induces the expression of proinflammatory genes (such as IL-1β, IL-6 and TNF-α) via the ROS-MAPK-NF-κB/AP-1 signaling pathway in macrophages47. TNF-α, IL-1β, IL-6, and IL-12 are secreted from M1 macrophages and activate the immune system to defend against exogenous bacteria and viruses. Interestingly, another study also found that the production of the Th2 cytokine IL-13 was induced by AP-125 and that selective HDAC6 inhibition impaired IL-13 expression39. A previous study found that the salivary level of IL-13 was significantly different in OSCC patients compared to controls and that IL-13 could prove to be a potential biomarker for OSCC48. This finding is consistent with our experimental results showing that HDAC6 induced M2 polarization through the AP-1/IL-13 signaling pathway. Therefore, AP-1-mediated regulation of macrophage polarization still needs further in-depth study and confirmation of its true role.
Data availability
The datasets generated during the current study are available from the corresponding authors on reasonable request.
References
Sung, H. et al. Global Cancer Statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 Countries. CA Cancer J. Clin. 71, 209–249. https://doi.org/10.3322/caac.21660 (2021).
Lee, K. D., Wang, T. Y., Lu, C. H., Huang, C. E. & Chen, M. C. The bidirectional association between oral cancer and esophageal cancer: A population-based study in Taiwan over a 28-year period. Oncotarget 8, 44567–44578. https://doi.org/10.18632/oncotarget.17818 (2017).
Lin, Y. S., Jen, Y. M., Wang, B. B., Lee, J. C. & Kang, B. H. Epidemiology of oral cavity cancer in taiwan with emphasis on the role of betel nut chewing. ORL J. Otorhinolaryngol. Relat. Spec 67, 230–236. https://doi.org/10.1159/000089214 (2005).
Ling, Z., Cheng, B. & Tao, X. Epithelial-to-mesenchymal transition in oral squamous cell carcinoma: Challenges and opportunities. Int. J. Cancer 148, 1548–1561. https://doi.org/10.1002/ijc.33352 (2021).
Warnakulasuriya, S., Johnson, N. W. & van der Waal, I. Nomenclature and classification of potentially malignant disorders of the oral mucosa. J. Oral. Pathol. Med. 36, 575–580. https://doi.org/10.1111/j.1600-0714.2007.00582.x (2007).
Wynn, T. A., Chawla, A. & Pollard, J. W. Macrophage biology in development, homeostasis and disease. Nature 496, 445–455. https://doi.org/10.1038/nature12034 (2013).
Ginhoux, F., Schultze, J. L., Murray, P. J., Ochando, J. & Biswas, S. K. New insights into the multidimensional concept of macrophage ontogeny, activation and function. Nat. Immunol. 17, 34–40. https://doi.org/10.1038/ni.3324 (2016).
Phan, A. T., Goldrath, A. W. & Glass, C. K. Metabolic and epigenetic coordination of T cell and macrophage immunity. Immunity 46, 714–729. https://doi.org/10.1016/j.immuni.2017.04.016 (2017).
Shi, C. & Pamer, E. G. Monocyte recruitment during infection and inflammation. Nat. Rev. Immunol. 11, 762–774. https://doi.org/10.1038/nri3070 (2011).
Mege, J. L., Mehraj, V. & Capo, C. Macrophage polarization and bacterial infections. Curr. Opin. Infect. Dis. 24, 230–234. https://doi.org/10.1097/QCO.0b013e328344b73e (2011).
Alfano, M., Graziano, F., Genovese, L. & Poli, G. Macrophage polarization at the crossroad between HIV-1 infection and cancer development. Arterioscler. Thromb. Vasc. Biol. 33, 1145–1152. https://doi.org/10.1161/ATVBAHA.112.300171 (2013).
Mantovani, A. et al. The chemokine system in diverse forms of macrophage activation and polarization. Trends Immunol. 25, 677–686. https://doi.org/10.1016/j.it.2004.09.015 (2004).
Noy, R. & Pollard, J. W. Tumor-associated macrophages: From mechanisms to therapy. Immunity 41, 49–61. https://doi.org/10.1016/j.immuni.2014.06.010 (2014).
Chittezhath, M. et al. Molecular profiling reveals a tumor-promoting phenotype of monocytes and macrophages in human cancer progression. Immunity 41, 815–829. https://doi.org/10.1016/j.immuni.2014.09.014 (2014).
Colegio, O. R. et al. Functional polarization of tumour-associated macrophages by tumour-derived lactic acid. Nature 513, 559–563. https://doi.org/10.1038/nature13490 (2014).
Barkal, A. A. et al. Engagement of MHC class I by the inhibitory receptor LILRB1 suppresses macrophages and is a target of cancer immunotherapy. Nat. Immunol. 19, 76–84. https://doi.org/10.1038/s41590-017-0004-z (2018).
Jinushi, M. & Komohara, Y. Tumor-associated macrophages as an emerging target against tumors: Creating a new path from bench to bedside. Biochim. Biophys. Acta 123–130, 2015. https://doi.org/10.1016/j.bbcan.2015.01.002 (1855).
Ries, C. H. et al. Targeting tumor-associated macrophages with anti-CSF-1R antibody reveals a strategy for cancer therapy. Cancer Cell 25, 846–859. https://doi.org/10.1016/j.ccr.2014.05.016 (2014).
Chen, D. et al. Chloroquine modulates antitumor immune response by resetting tumor-associated macrophages toward M1 phenotype. Nat. Commun. 9, 873. https://doi.org/10.1038/s41467-018-03225-9 (2018).
Mori, K., Hiroi, M., Shimada, J. & Ohmori, Y. Infiltration of m2 tumor-associated macrophages in oral squamous cell carcinoma correlates with tumor malignancy. Cancers (Basel) 3, 3726–3739. https://doi.org/10.3390/cancers3043726 (2011).
Akdis, M. et al. Interleukins, from 1 to 37, and interferon-gamma: Receptors, functions, and roles in diseases. J. Allergy Clin. Immunol. 127(701–721), e701-770. https://doi.org/10.1016/j.jaci.2010.11.050 (2011).
Zhang, J. H., Zhang, M., Wang, Y. N. & Zhang, X. Y. Correlation between IL-4 and IL-13 gene polymorphisms and asthma in Uygur children in Xinjiang. Exp. Ther. Med. 17, 1374–1382. https://doi.org/10.3892/etm.2018.7096 (2019).
Tabata, Y. & Khurana, H. G. K. IL-13 receptor isoforms: Breaking through the complexity. Curr. Allergy Asthma Rep. 7, 338–345. https://doi.org/10.1007/s11882-007-0051-x (2007).
Fichtner-Feigl, S., Strober, W., Kawakami, K., Puri, R. K. & Kitani, A. IL-13 signaling through the IL-13alpha2 receptor is involved in induction of TGF-beta1 production and fibrosis. Nat. Med. 12, 99–106. https://doi.org/10.1038/nm1332 (2006).
Fujisawa, T., Joshi, B. H. & Puri, R. K. IL-13 regulates cancer invasion and metastasis through IL-13Ralpha2 via ERK/AP-1 pathway in mouse model of human ovarian cancer. Int. J. Cancer 131, 344–356. https://doi.org/10.1002/ijc.26366 (2012).
Kramer, O. H., Mahboobi, S. & Sellmer, A. Drugging the HDAC6-HSP90 interplay in malignant cells. Trends Pharmacol. Sci. 35, 501–509. https://doi.org/10.1016/j.tips.2014.08.001 (2014).
Narita, T., Weinert, B. T. & Choudhary, C. Functions and mechanisms of non-histone protein acetylation. Nat. Rev. Mol. Cell Biol 20, 156–174. https://doi.org/10.1038/s41580-018-0081-3 (2019).
Zhang, M. et al. HDAC6 regulates DNA damage response via deacetylating MLH1. J. Biol. Chem. 294, 5813–5826. https://doi.org/10.1074/jbc.RA118.006374 (2019).
Lee, Y. S. et al. The cytoplasmic deacetylase HDAC6 is required for efficient oncogenic tumorigenesis. Cancer Res. 68, 7561–7569. https://doi.org/10.1158/0008-5472.CAN-08-0188 (2008).
Zhang, M. et al. HDAC6 deacetylates and ubiquitinates MSH2 to maintain proper levels of MutSalpha. Mol. Cell 55, 31–46. https://doi.org/10.1016/j.molcel.2014.04.028 (2014).
Yang, M. H. et al. HDAC6 and SIRT2 regulate the acetylation state and oncogenic activity of mutant K-RAS. Mol. Cancer Res 11, 1072–1077. https://doi.org/10.1158/1541-7786.MCR-13-0040-T (2013).
Li, Y., Zhang, X., Polakiewicz, R. D., Yao, T. P. & Comb, M. J. HDAC6 is required for epidermal growth factor-induced beta-catenin nuclear localization. J. Biol. Chem. 283, 12686–12690. https://doi.org/10.1074/jbc.C700185200 (2008).
Li, Y., Shin, D. & Kwon, S. H. Histone deacetylase 6 plays a role as a distinct regulator of diverse cellular processes. FEBS J. 280, 775–793. https://doi.org/10.1111/febs.12079 (2013).
Bergman, J. A. et al. Selective histone deacetylase 6 inhibitors bearing substituted urea linkers inhibit melanoma cell growth. J. Med. Chem. 55, 9891–9899. https://doi.org/10.1021/jm301098e (2012).
Nakashima, H. et al. Histone deacetylase 6 inhibition enhances oncolytic viral replication in glioma. J. Clin. Invest. 125, 4269–4280. https://doi.org/10.1172/JCI80713 (2015).
Vieson, M. D. et al. Treatment with a selective histone deacetylase 6 inhibitor decreases lupus nephritis in NZB/W mice. Histol. Histopathol. 32, 1317–1332. https://doi.org/10.14670/HH-11-885 (2017).
Iliopoulos, D. et al. Fragile genes as biomarkers: Epigenetic control of WWOX and FHIT in lung, breast and bladder cancer. Oncogene 24, 1625–1633. https://doi.org/10.1038/sj.onc.1208398 (2005).
Zhang, M. Z. et al. IL-4/IL-13-mediated polarization of renal macrophages/dendritic cells to an M2a phenotype is essential for recovery from acute kidney injury. Kidney Int. 91, 375–386. https://doi.org/10.1016/j.kint.2016.08.020 (2017).
Laino, A. S. et al. HDAC6 selective inhibition of melanoma patient T-cells augments anti-tumor characteristics. J. Immunother. Cancer 7, 33. https://doi.org/10.1186/s40425-019-0517-0 (2019).
Li, C., Shintani, S., Terakado, N., Nakashiro, K. & Hamakawa, H. Infiltration of tumor-associated macrophages in human oral squamous cell carcinoma. Oncol. Rep. 9, 1219–1223 (2002).
El-Rouby, D. H. Association of macrophages with angiogenesis in oral verrucous and squamous cell carcinomas. J. Oral. Pathol. Med. 39, 559–564. https://doi.org/10.1111/j.1600-0714.2010.00879.x (2010).
Yan, B. et al. HDAC6 deacetylase activity is critical for lipopolysaccharide-induced activation of macrophages. PLoS ONE 9, e110718. https://doi.org/10.1371/journal.pone.0110718 (2014).
Wang, G. et al. HDAC inhibition prevents white matter injury by modulating microglia/macrophage polarization through the GSK3beta/PTEN/Akt axis. Proc. Natl. Acad. Sci. U S A 112, 2853–2858. https://doi.org/10.1073/pnas.1501441112 (2015).
Guha, M. & Mackman, N. LPS induction of gene expression in human monocytes. Cell Signal 13, 85–94. https://doi.org/10.1016/s0898-6568(00)00149-2 (2001).
Gwinn, M. R. & Vallyathan, V. Respiratory burst: Role in signal transduction in alveolar macrophages. J. Toxicol. Environ. Health B Crit. Rev. 9, 27–39. https://doi.org/10.1080/15287390500196081 (2006).
Chuang, J. Y. et al. Syk/JNK/AP-1 signaling pathway mediates interleukin-6-promoted cell migration in oral squamous cell carcinoma. Int. J. Mol. Sci. 15, 545–559. https://doi.org/10.3390/ijms15010545 (2014).
Youn, G. S., Lee, K. W., Choi, S. Y. & Park, J. Overexpression of HDAC6 induces pro-inflammatory responses by regulating ROS-MAPK-NF-kappaB/AP-1 signaling pathways in macrophages. Free Radic. Biol. Med. 97, 14–23. https://doi.org/10.1016/j.freeradbiomed.2016.05.014 (2016).
Aziz, S. et al. Salivary immunosuppressive cytokines IL-10 and IL-13 are significantly elevated in oral squamous cell carcinoma patients. Cancer Invest. 33, 318–328. https://doi.org/10.3109/07357907.2015.1041642 (2015).
Acknowledgements
We would like to thank the all patients who cooperated to realize this research and provided their tissue samples as well as clinical characteristics for research use. This research was funded by Ministry of Science and Technology of Taiwan (grant numbers MOST 110-2314-B-650-004), E-Da Hospital/E-Da Cancer Hospital (grant numbers EDCHT109016, EDCHT110015 and EDCHS111005) and by ZBKAFGH RESEARCH PROJECT (KAFGH-ZY-A-109022).
Author information
Authors and Affiliations
Contributions
Conceptualization, T.-H.H. and C.-C.T.; methodology, S.-Y.H.; validation, C.-W.W.; resources, H.-P.T.; writing—original draft preparation, T.-H.H. and C.-W.W.; writing—review and editing, T.-H.H.; supervision, T.-H.H.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Tseng, CC., Huang, SY., Tsai, HP. et al. HDAC6 is a prognostic biomarker that mediates IL-13 expression to regulate macrophage polarization through AP-1 in oral squamous cell carcinoma. Sci Rep 12, 10513 (2022). https://doi.org/10.1038/s41598-022-14052-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-14052-w
This article is cited by
-
Targeting histone deacetylases in head and neck squamous cell carcinoma: molecular mechanisms and therapeutic targets
Journal of Translational Medicine (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.