Abstract
Bacteria occupy all major ecosystems and maintain an intensive relationship to the eukaryotes, developing together into complex biomes (i.e., phycosphere and rhizosphere). Interactions between eukaryotes and bacteria range from cooperative to competitive, with the associated microorganisms affecting their host`s development, growth and health. Since the advent of non-culture dependent analytical techniques such as metagenome sequencing, consortia have been described at the phylogenetic level but rarely functionally. Multifaceted analysis of the microbial consortium of the ancient phytoplankton Botryococcus as an attractive model food web revealed that its all abundant bacterial members belong to a niche of biotin auxotrophs, essentially depending on the microalga. In addition, hydrocarbonoclastic bacteria without vitamin auxotrophies seem adversely to affect the algal cell morphology. Synthetic rearrangement of a minimal community consisting of an alga, a mutualistic and a parasitic bacteria underpins the model of a eukaryote that maintains its own mutualistic microbial community to control its surrounding biosphere. This model of coexistence, potentially useful for defense against invaders by a eukaryotic host could represent ecologically relevant interactions that cross species boundaries. Metabolic and system reconstruction is an opportunity to unravel the relationships within the consortia and provide a blueprint for the construction of mutually beneficial synthetic ecosystems.
Similar content being viewed by others
Introduction
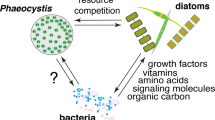
Bacteria are omnipresent and affect likely every eukaryotic system through their presence1. Microalgae are the dominant primary producers and the base of the food web within aquatic ecosystems and sustain in the natural environment in close relationship with multiple associated microorganisms2,3. Thus, the interactions between phytoplankton and bacteria represent a fundamental ecological relationship within aquatic environments2, by controlling nutrient cycling and biomass production at the food web base. The nature of the exchange of micro- and macronutrients, diverse metabolites including complex products such as polysaccharides, and infochemicals defines the relationship of the symbiotic partners4,5,6, which span mutualism, commensalism and parasitism3. Within a parasitic association, the bacteria are known to adversely affect the algae7, while in commensalism the partners can survive independently, with the commensal feeding on algae-provided substrates, but without harming the host3. Mutualistic interactions include i.a. the obligate relationships between vitamin-synthesizing bacteria and vitamin-auxotrophic phytoplankton species8. Many microalgae cannot synthesize several growth-essential vitamins (like B12 and B1)8,9,10 and obtain them through a symbiotic relationship with bacteria in exchange for organic carbon11. Recent observations suggest that not only certain microalgae require vitamins, but many bacteria also have to compete with other organisms for the B-vitamins12,13.
However, in microbial environments, where the distinction between host and symbionts is less clear, the identification of the associated partners and the nature of their relationship is challenging. Looking for a suitable model food web system to elucidate the complexity of eukaryote-prokaryote interactions, we focused on the microalga-bacteria consortia with an organic carbon-rich phycosphere. The planktonic chlorophyta Botryococcus braunii excretes long-chain hydrocarbons and a variety of exo-polysaccharides14 which allows this single celled alga to form large agglomerates1. Both organic carbon-based products are exposed to the surrounding ecosystem and enable the alga to sustain in the natural environment (phycosphere) in close relationship with multiple associated microorganisms2,3. Botryococcus exudes up to 46% of their photosynthetically fixed carbon into the phycosphere15, similarly to processes, however with lesser extent, observed for seedlings as well as adult plants within the rhizosphere in the form of poorly characterized rhizodeposits16,17,18. Additionally, it is generally considered that a progenitor of Botryococcus impacted the development of today's oil reserves and coal shale deposits19. Thus, the microbiome associated with Botryococcus braunii presents a unique opportunity to dissect the roles of the eukaryote and prokaryotes within the consortium and test how the eukaryote responds to perturbations in its environmental context.
The interactions between B. braunii and the associated bacteria are not fully understood, but are known to influence the microalgal growth performance and product formation capacity14,20,21,22. The grouping of interdependent organisms living and functionally interacting with each other in the same habitat is also called biocoenosis, a term coined by Karl Möbius in 187723. To understand the complex nature of the unique habitat of a microalgal phycosphere and to unravel the food web between alga and bacteria, we investigated the consortia accompanying the hydrocarbon/carbohydrate-producing green microalga Botryococcus braunii.
Results
Metagenomic survey of the Botryococcus braunii consortia
To determine the inhabitants of the B. braunii ecosystem, we profiled four different Botryococcus communities (races A and B) using high-throughput 16S rDNA gene amplicon and read-based metagenome sequencing approaches (Figures S1, S2, S3; Fig. 1a). Assembly and functional annotation of the metagenomes together with single strain isolation and sequencing reconstructed the capabilities of the most abundant consortial microorganisms and were used to design and test synthetic alga-bacteria ecosystems. The analysis of the 16S rDNA amplicons and a read-based metagenome approach showed that each of the B. braunii communities represents a comparable conserved consortium of a single algal and at least 25 different bacterial species as well as traces of Archaea and viruses (Figure S2). In total, 16S rDNA amplicon sequencing identified 33 bacterial taxa, 22 of which were present in both race A and B consortia (Figure S3, Supplementary Discussion, Chapter 1). The relative abundance of plastid 16S rDNA sequences accounted up to 49 and 59% of all detected amplicon reads in race A and B samples, respectively. Combined metagenome de novo assembly and differential coverage and tetra-nucleotide signature based binning resulted in ten high quality and ten fragmentary bacterial metagenome assembled genomes (MAGs). Three additional genomes were closed by single strain sequencing (Fig. 1; Tables S1 and S2; Supplementary Discussion, Chapter 1). The bacterial community varied quantitatively, but not qualitatively between growth phases and strains (Figure S3). B. braunii consortia are inhabited by the bacterial phyla (Proteobactreria, Bacteroidetes, Acidobacteria and Actinobacteria), comparable to those occupying the plant rhizosphere24, albeit in different relative abundances.
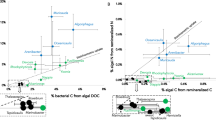
Taxonomy of the B. braunii bacterial community and selected functional categories encoded in reconstructed MAGs and complete genomes. (a) Normalized read-based comparison and taxonomic assignment of different B. braunii metagenome datasets. The circle size represents the amount of classified reads for the respective taxonomic group (minimum 50 reads per taxonomic group). Abbreviations: P, phylum; G, genus; O, order. (b) Abundance of selected functional categories encoded in the high-quality draft and complete bacterial genomes. Numbers in parentheses represent the maximal total number of genes within each pathway. Percentage: per cent of total number of genes per pathway identified within each genome. All annotated genes within the respective pathways are presented on the EMGB platform (https://emgb.cebitec.uni-bielefeld.de/Bbraunii-bacterial-consortium/). The pathways statistics are summarized in the Table S5.
During the growth phase, the Botryococcus phycosphere is dominated by the genera of Devosia, Dyadobacter, Dokdonella, Acidobacteria, Brevundimonas, Mesorhizobium and Hydrogenophaga which account for over 80% of the consortial bacteria (Table S1; Figure S4). The re-processing of the B. braunii Guadeloupe strain metagenome data25 and direct comparison of the different datasets revealed the coinciding presence of the individual bacterial genera, although massively varying in the abundance pattern in dependence of cultivation conditions (Figure S5).The same taxa are also present i.a. in the active microbiome associated roots and rhizosphere of oilseed rape26, as well as in the lettuce27 and global citrus rhizosphere24, strongly suggesting an overall mutual dependence or preferred coexistence.
To identify genes that enable the bacterial community members to contribute to and to benefit from the food web of the consortium, we performed a KEGG-based quantitative functional assignment of the annotated high-quality MAGs and the sequenced genomes of isolated associates (for details see Methods; Supplementary Discussion, Chapter 2).
We originally expected a uniform distribution of abilities and a high degree of interconnectedness between all members of the consortium. Surprisingly, the consortium divides into two functionally distinct groups of bacteria: The dominant consortia members such as Devosia (DevG), Dyadobacter (DyaG) and Brevundimonas (Bre1G, B.Bb-A) do not oxyfunctionalize hydrocarbons28, but their genomes are rich in genes involved in the degradation/assimilation of carbohydrates (Fig. 1b). On the contrary, the bacterial isolates Mycobacterium and Pimelobacter (M.Bb-A and P.Bb-B), which are of low abundance within the consortia (Figure S3), contain a large portfolio of genes encoding monooxygenases/hydroxylases for hydrocarbon degradation28 and hold the potential for a pronounced catabolic capacity towards lipids, fatty acids and xenobiotics (Fig. 1b). Besides, Mycobacterium and Pimelobacter genomes were the only genomes among the examined community members to carry non-heme membrane-associated monooxygenase alkB genes (Table S6)28. To test these observed genetic predisposition of the consortia, we assessed algal capacities for growth and hydrocarbon production in co-cultivation of the axenic B. braunii with one representative of each group.
Effect of the bacterial community on the B. braunii host
One representative of the abundant as well as less abundant community members (Brevundimonas sp. Bb-A and Mycobacterium sp. Bb-A, respectively) were isolated in axenic culture and used for the co-cultivation experiments. Synthetic algae-bacteria consortia were formed with an axenic hydrocarbon- and carbohydrate producing B. braunii race A strain SAG30.81 under strict phototrophic conditions (e.g. without the supplementation of vitamins or organic carbon). Axenic Botryococcus braunii shows a continuous increase in cell biomass and hydrocarbon content, yielding 1.36 ± 0.1 g L−1 and 0.5 ± 0.1 g L−1, respectively (Fig. 2a, b; Figure S6). Co-cultivation of the Brevundimonas sp. Bb-A promotes the growth of the alga and increases the biomass by 60.5 ± 16.9% after 15 days of cultivation compared to axenic control (Fig. 2b). While co-cultivation with the Mycobacterium sp. Bb-A results in reduced growth of the consortia by 12.6 ± 4.1% at the end of cultivation with a maximal reduction of 30.6 ± 4.8% after 9 days (Fig. 2a).
Physiological effect of the individual bacterial isolates on the microalgae during co-cultivation. (a–d) Determination of algal growth and product formation performance of axenic and xenic B. braunii cultures, supplemented with bacterial isolates. Shown are growth analysis of B. braunii in the presence of (a) Mycobacterium sp. Bb-A and (b) Brevundimonas sp. Bb-A. The inserts within the graph show the maximal observed growth difference in comparison to the control. (c) Comparison of the hydrocarbon levels observed in samples cultivated with bacterial isolates in relation to the axenic B. braunii culture (values set to 100%) in the course of the cultivation; (d) Morphological characteristic of algal cells during the cultivation (pictures were taken after 9 days of cultivation) with the bacterial isolates (red arrows indicate the hydrocarbons produced by the microalga). Graphs imbedded in the pictures show the progress of the bacterial growth (cell number on log10 scale). The error bars represent standard error of mean values of three biological and three technical replicates (SE; n = 9). Asterisks represent p values as determined via Student’s t-test (*= < 0.05, **= < 0.01, ***= < 0.001).
Axenic B. braunii accumulates hydrocarbons up to 38% of its dry weight (DW) in a bacteria-free environment (Figure S6a), (the value was set to 100% for easier comparison in Fig. 2c). In contrast, co-cultivation with Mycobacterium limits hydrocarbon accumulation down to 4.4 ± 0.2% compared to the axenic control (Fig. 2c) and alters colony morphology (Fig. 2d). Loss of the hydrocarbon matrix reduces massively the colony size (Fig. 2d) and likely increases susceptibility to grazers29. The dramatic decrease in hydrocarbon content and disturbed colony formation cannot be observed in cultures supplemented with Brevundimonas or in the xenic Botryococcus braunii algae-bacteria communities (Fig. 2c, d; Figures S1 and S6), pointing to a balanced equilibrium between the alga and its microbiome in natural environments.
Since Botryococcus readily releases huge amounts of organic carbon into the extracellular milieu15, creating a phycosphere that naturally attracts many microorganisms, including those with potential damaging effects, its evolutionary fitness will depend on efficient management of the “bacterial zoo”. We hypothesized that there are strong control mechanisms that allow the algae host to attract and control symbiotic / beneficial bacteria. Therefore, we examined the genetic portfolio of the bacterial community for essential cofactors and interdependencies.
The core community of Botryococcus phycosphere belongs to a niche of biotin auxotrophs
At first glance, the Botryococcus alga does not appear to be obligatory dependent on a bacterial community, since axenic strains can be fully photoautotrophically cultured without vitamin supplements (Fig. 2 a, b) and contains the vitamin B12-independent form of methionine synthase (metE)30. (Meta)Genome reconstruction revealed that the B. braunii bacterial core community is auxotrophic for various B-vitamins (Fig. 3a), making them dependent on an exogenous supply of vitamins for survival. The complete thiamin synthesis pathway (vitamin B1, essential for citrate cycle and pentose phosphate pathway12) could not be reconstructed for four assembled genomes. Five genera are auxotrophic for cobalamin (vitamin B12). All abundant genera within the Botryococcus consortia (including the growth-promoting Brevundimonas, (Fig. 2b)) lack the complete gene portfolio for biotin (vitamin B7) synthesis (Fig. 3a), but contained the complete biosynthesis pathway of fatty acids for which biotin is essential12. In contrast, low abundant consortial members such as Mycobacterium are B-vitamin prototrophs (Fig. 3a).
Essential cofactors (inter-)dependencies of the bacterial core community and isolates of B. braunii. (a) Heatmap of the reconstructed de novo biosynthesis of B-vitamins encoded in the high-quality draft and complete bacterial genomes. Numbers in parentheses represent the maximal total number of genes within each pathway. Percentage: per cent of total number of genes per pathway identified within each genome. All annotated genes within the respective pathways are presented on the EMGB platform (https://emgb.cebitec.uni-bielefeld.de/Bbraunii-bacterial-consortium/). The pathways statistics are summarized in the Table S5 and S8. (b) Growth assay to confirm B-vitamin auxotrophy and prototrophy in axenic Brevundimonas sp. Bb-A (blue) and Mycobacterium sp. Bb-A (orange) strains, respectively. (c) Biotin concentration levels detected in cell-free supernatants from the cultures of axenic B. braunii as well as in the presence of Mycobacterium sp. Bb-A (orange) and Brevundimonas sp. Bb-A (blue). The error bars represent standard error of mean values of three biological and three technical replicates (SE; n = 9).
Extended experimental analysis confirmed the biotin (vitamin B7) auxotrophy in Brevundimonas sp. Bb-A and vitamin B prototrophy in Mycobacterium sp. Bb-A as deduced from the assembled genomes (Fig. 3a, b). Botryococcus braunii secretes and accumulates biotin into the surrounding environment (Fig. 3c). The biotin secretion appears to be induced by the presence of bacteria, in the cultures with the B7-auxotrophic Brevundimonas as well as in the presence of the biotin-prototrophic Mycobacterium sp. Bb-A, although to a lesser extent (cf. Figure 3c at timepoint 3 days). The subsequent decrease in biotin amount during the progress of the cultivation further hints at the active vitamin absorption from the culture supernatant in both co-cultivation approaches in comparison to the axenic control (Fig. 3c), especially during bacterial proliferation, with the biotin levels increasing again when bacterial growth stagnates (Fig. 2d, bacterial growth determination for M.Bb-A and B.Bb-A).
Collectively, in a healthy (natural) community, non-hydrocarbon degrading bacterial genera which are auxotrophic for one or more vitamins appear to dominate the Botryococcus phycosphere, while fully prototrophic hydrocarbonoclastic bacteria are present but not abundant. We hypothesized that B. braunii cultivates its own auxotrophic “bacterial zoo” to combat hydrocarbonoclastic bacteria. To test the hypothesis, synthetic consortia of alga–”good bacteria”– “bad bacteria” were formed and analyzed.
Bacterial biotin-auxotrophic symbionts defend their eukaryotic host
Based on the abovementioned findings, we have hypothesized that the alga maintains its balanced, ‘healthy' microbiome by recruiting beneficial, vitamin-auxotrophic microorganisms, inter alia, with the provision of photosynthetically fixed carbon and biotin, and thus counteracts pathogen assault. To test this hypothesis, one representative of each group, the vitamin-auxotrophic Brevundimonas and the prototroph Mycobacterium were co-cultivated with the axenic Botryococcus alga under photoautrophic conditions with three different inoculum ratios (10:90, 50:50 or 90:10 per cent based on cfu) (Fig. 4a).
Bacterial symbionts fend their eukaryotic host from hydrocarbonoclastic invaders. Physiology of a synthetic alga-bacteria-bacteria community, consisting of Brevundimonas sp. Bb-A (blue) and Mycobacterium sp. Bb-A (orange) in co-culture with B. braunii (green) under photoautrophic conditions with three different inoculum ratios (10:90, 50:50 or 90:10 per cent based on cfu (colony-forming units)). Shown are (a) the progression of bacterial proliferation (cfu) and (b) detected relative hydrocarbon levels (normalized to the axenic control) during the co-cultivation. The error bars represent standard error of mean values of three biological and three technical replicates (SE; n = 9).
Independently from the initial amount of inoculum, biotin-auxotrophic Brevundimonas sp. Bb-A proliferated strongly within this synthetic community and reached in each case a comparable cfu-density of up to 2.1*108 cells mL−1. Mycobacterium did not proliferate in the presence of Brevundimonas (Fig. 4a), but was clearly capable of growth in the presence of the microalga alone (Fig. 2d). The propagation of Mycobacterium sp. Bb-A, limited in each approach to a maximum cfu of up to 1.7*106 cells mL−1 at the end of the cultivation, is therefore likely to be attributable to the bioactive compounds with bacteriostatic properties produced by Brevundimonas. Indeed, all reconstructed genomes encode metabolic gene cluster for antibiotic compounds (especially against Gram-positive bacteria) and for bacteriocins (Fig. 1b, Table S9) with significant antimicrobial potency32. The protective effect of Brevundimonas appears to be further evident from the hydrocarbon content of cultures, which increases with decreasing proportions of Mycobacterium (Fig. 4b). Experimental analyses (Figs. 2, 3, 4) thus support key conjectures derived from metagenomic assemblies (Figs. 1 and 3a).
Discussion
Microalgae exist in the natural environment as part of a complex microbial consortium, where the different species may exert considerable influence on each other for the exploitation of unique biological functions and/or for the metabolite exchange2. Bacterial communities within the Botryococcus phycosphere appear to be highly conserved and illustrate relatively low species biodiversity (Figures S2a and S3), with most of the detected genera occurring in all tested consortia, albeit differentially abundant (Fig. 2; Figure S5;25). The evaluation further emphasizes dynamic consortia33 that vary considerably depending on physiological conditions, especially in artificial systems (e.g. supplementation of vitamins and/or antibiotics25). Under strict photoautotrophic conditions, most abundant genera identified within the Botryococcus communities (such as Devosia, Dyadobacter, Brevundimonas etc.; Fig. 1a) also occur in other biomes such as phycosphere and/or rhizosphere24,26,27,34, strongly indicating preferential coexistence of certain bacteria and eukaryotes.
The fact that Botryococcus readily excretes huge amounts of their photosynthetically fixed organic carbon into its phycosphere15 (Figure S1) naturally attracts many microorganisms, that can be potentially harmful, so the evolutionary fitness of the algae host depends heavily on the efficient management of the "bacterial zoo", which requires strong control mechanisms that facilitate the recruitment and control of beneficial bacterial partners.
Based on the genetically encoded features, the bacterial associates of B. braunii consortia can be apportioned into two functionally distinct categories of (i) B-vitamin-auxotrophic carbohydrate-degrader (controllable), and (ii) prototrophic hydrocarbonoclastic species (uncontrollable) (Fig. 5).
Alga-bacteria interactions within the Botryococcus braunii phycosphere. The depiction summarizes the observed generic potential encoded in the (meta)genomes of bacterial associates (Figs. 1 and 3), along with experimental validation within the minimal synthetic community (Figs. 2 and 4). The color code indicates the functional grouping of the community members: supportive B-vitamin-auxotrophic (blue) and hydrocarbonoclastic prototrophic (red). The potential for degradation of hydrocarbons and carbohydrates of the bacterial associates is illustrated via yellow and striped pacmans, respectively. The genetic predisposition to the synthesis of bacteriocins and antibiotics encoded by the secondary metabolite biosynthesis gene clusters, is indicated by the halos and triangles, respectively. Abbreviation: HC, Hydrocarbons; CH, carbohydrates; B7, Biotin; N, nitrogen; P, phosphorus.
The first category includes the most abundant bacterial members of the Botryococcus consortia that do not assimilate hydrocarbons and display B-vitamin-auxotrophy, strongly depending on exogenous supply for their survival. Although, auxotrophic for various B-vitamins and non-viable on its own, the consortia’s most abundant species Devosia (DevG, up to 30% abundancy of all bacteria, Table S3) is the only representative of the first category to contain the complete pathway for vitamin B12 synthesis (Fig. 3a). In view of the fact that the bacterial abundance determines their functional relevance for the community35,36, this consortia-dominant species (DevG) is likely to provide both microalga and bacteria with the essential co-factor8, thus supporting the recent proposition that the B12 supply is realized only by a few certain bacteria within the alga-bacteria consortia34. The release of vitamins and vitamin precursors by bacteria is a well known mediator of algae-bacteria interactions9,33, where the transfer of vitamins like cobalamin and thiamin8,37 supports algal growth and can be considered as ‘bacterial farming’ of algae as providers of organic resources. However, our observations show that B. braunii does not essentially depend on microbial vitamin supply. Further, according to the (meta)genome reconstruction results, the Botryococcus bacterial consortia cultivated under strictly phototrophic conditions is dominated by vitamin-auxotrophs (Fig. 3a). Especially the biotin-auxotrophic bacteria are prevalent within the Botryococcus consortia, so it is doubtful that the large required supply of biotin is of bacterial origin. Because of its predominance within the community (Figure S2) and the observed active biotin secretion (Fig. 3c), the microalgae tend to be the main source for the consortial biotin supply. The excretion of some B-vitamins has been observed in microalgae and plants38,39, however the increasingly induced vitamin secretion by the presence of bacteria, as well as their simultaneous active uptake, remained so far unrecognized. This observation supports the recent proposition that many microorganisms although capable of vitamin synthesis themselves if needed, likely prefer to take up the compounds when available in the environment, because synthesis requires usually more metabolic energy than transport31. However, we cannot rule out that the lower level of induction of biotin secretion observed for the biotin-prototrophic Mycobacterium sp. Bb-A (Fig. 3C), may result from a possible suppression of biotin formation in B. braunii by the bacterium or the recognition of the bacterium as a non-symbiont by the algae. Nevertheless, since some of the missing cofactors may only be complemented by bacteria (e.g. vitamin B12), strong bacteria-bacteria relations within the consortia occur in addition to the algae-bacteria interactions, signifying a deep degree of interconnectedness within the Botryococcus phycosphere.
Completely autonomous species, such as the isolate Mycobacterium sp. Bb-A are rare among the consortia and assigned to the second category (uncontrollable). These phylotypes are known to be hydrocarbonoclastic microorganisms40,41 targeting mainly the organic carbon source and therefore pose a potential risk to the microalga, strongly implying a parasitic relationship. And although these phylotypes are able to provide the Botryococcus alga with vitamin B12, nutrients and probiotic compounds, their activity results in a less complex colony structure due to the degradation of the hydrocarbon matrix (Fig. 2). The observed change of the colony morphology might affect the survival of the typically colonized microalgae in the natural environment, i.e. by higher susceptibility to grazers29. The malfunction of colony formation could thus be attributed to imbalanced and dysbiotic host-microbiome interaction similar to imbalanced interactions previously observed42,43, so that prototrophic, mainly carbon-targeting bacterial species may essentially act as pathogens.
In this context, it seems reasonable to assume that specific bacteria that require essential co-factors are deliberately promoted and controlled by Botryococcus braunii in exchange for protection as well as nutrients and probiotic metabolites. The observed growth-promoting effect of Brevundimonas (Fig. 2b), which depends on exogenous biotin supply for survival (Fig. 3), is thus based on the close interaction with the microalga. This mutualistic alga-bacteria relationship seems efficiently to prevent uncontrolled overgrowing of hydrocarbonoclastic competitors, as demonstrated by the experiments with the minimal synthetic consortia (Fig. 4).
Thus, the interactions between the microalga and bacteria within the Botryococcus braunii consortia depends on a complex interplay between all members, with the microalga as the “conductor” in this play (Fig. 5). However, the elucidation of the microalga-bacteria dynamics as well as of the communication signals exchanged between the participating parties requires more precise kinetic analyses and will be subject for further research. In this context, the metagenomic analyses of the Botryococcus consortium might represent a path towards mechanistic resolution of ecological interactions at the microscopic scale.
Materials and methods
Strains, cultivation conditions and growth determination parameters
Liquid, xenic cultures of Botryococcus braunii race A (CCALA778, isolated in 1997 in Serra da Estrela, Baragem da Erva da Fome, Portugal) and race B (AC761, isolated in 1983 from Paquemar, Martinique, France) as well as the axenic B. braunii race A strain (SAG30.81, Peru, Dpto.Cuzco, Laguna Huaypo) were used for co-cultivation studies. The cultures were phototrophically cultivated at room temperature (~ 22 °C) under 16:8 light–dark illumination of 350 ± 400 μmol photons m−2 s−1 white light in modified Chu-13 media (without the addition of vitamin and organic carbon source) as described before15. Carbon supply and agitation were achieved by bubbling the cultures with moisture pre-saturated carbon dioxide-enriched air (5%, v/v) with a gas flow rate of 0.05 vvm. Cultures growth was monitored by measurements of biomass organic dry weight (DW) by drying 10 mL culture per sample (in three biological and technical replicates) following the protocol described before44. Additionally, microalgae and bacteria cells were regularly checked by optical microscopy.
The bacterial isolates were obtained by plating highly diluted xenic B. braunii cultures (CCALA778 and AC761) on LB agar plates supplemented with vitamins (0.1 nM cobalamin, 3.26 nM thiamine and 0.1 nM biotin, Sigma Aldrich). After 1–2 weeks single cell colonies were picked, separately plated and analyzed via 16S rDNA Sanger-sequencing by using the universal primer pair 27F (5′-AGAGTTTGATCCTGGCTCAG-3′) and 1385R (5′-CGGTGTGTRCAAGGCCC-3′). Two isolates, deriving from algal race A and one from race B, were classified based on 16S rDNA similarity as Mycobacterium, Brevundimonas and Pimelobacter species, respectively. The purity of the axenic alga and bacterial strains was regularly verified by polymerase chain reaction (PCR) amplification and Sanger-sequencing of the 16S rRNA gene using a universal primer pair 27F and 1385R. All primers were purchased by Sigma Aldrich.
For experiments testing the requirement of B-vitamins, mono-cultures of Mycobacterium sp. Bb-A and Brevundimonas sp. Bb-A were pre-washed three times in minimal medium45 and then starved of vitamins for two days prior to the start of the experiment. Then, bacterial cells were diluted to 0.2 (oD600) with minimal medium supplemented with 1gL−1 glucose and B-vitamins (at the following concentrations: 0.1 nM (each) of cobalamin, riboflavin, biotin and pantothenate as well as 3.26 nM thiamine, Sigma Aldrich) and grown for seven days.
For the co-cultivation experiments, 500 mL axenic algal cultures (~ 0.25 g L−1) were inoculated with the isolates (Mycobacterium sp. Bb-A or Brevundimonas sp. Bb-A, with ~ 2 × 106 cells mL−1 for single isolate cultivation and with ~ 2 × 106–2 × 107 cells mL−1 (in different ratios 10:90, 50:50 and 90:10 (cfu/cfu)) for co-cultivation with both bacterial strains) and cultured under photoautotrophic conditions. The bacterial cell density was determined by manual cell counting using hemocytometer or by determination of the colony-forming units (cfu) as a proxy for viable population density using the replica plating method46.
The quantitative determination of the biotin concentration in cell-free supernatants of the co-cultures and the axenic control was accomplished by using microbiological microtiter plate test (VitaFast, r-biopharm, Darmstadt, Germany) according to manufacturer instructions.
Hydrocarbon extraction and analysis
Hydrocarbon extraction was performed according to the protocol published previously15. The dry extracted hydrocarbons were resuspended in 500 µL of n-hexane, containing an internal standard n-hexatriacontane (C36H74, Sigma Aldrich) and analysed via GC–MS and GC-FID as described before15,47.
DNA sample collection and preparation
The samples from xenic B. braunii race A and B cultures (three biological replicates) were taken in the linear and stationary growth phases (after 9 and 24 days). Total community genomic DNA was extracted as previously described48. The quality of the DNA was assessed by gel electrophoresis and the quantity was estimated using the Quant-iT PicoGreen dsDNA Assay Kit (Invitrogen) and a Tecan Infinite 200 Microplate Reader (Tecan Deutschland GmbH).
High-throughput 16S rDNA amplicon sequencing
To get insight into the community composition of B. braunii Race A and B, high-throughput 16S rDNA amplicon sequencing was performed as described recently by Maus et al49. The primer pair Pro341F (5′-CCTACGGGGNBGCASCAG-3′) and Pro805R (5′-GACTACNVGGGTATCTAATCC-3′) was used to amplify the hypervariable regions V3 and V4 of diverse bacterial and archaeal 16S rRNAs50. In addition, the primers also cover the 16S rDNA gene of algal chloroplasts and eukaryotic mitochondrial genomes. Furthermore, in a two PCR steps based approach, multiplex identifier (MID) tags and Illumina-specific sequencing adaptors were added to each amplicon. Only amplicons featuring a size of ~ 460 bp were purified using AMPureXP magnetic beads (Beckman Coulter GmbH, Brea, California, USA). Resulting amplicons were qualitatively and quantitatively analyzed using the Agilent 2100 Bioanalyzer system (Agilent Inc., Santa Clara, California, USA) and pooled in equimolar amounts for paired-end sequencing on the Illumina MiSeq system (Illumina, San Diego, California USA). This sequencing approach provided ~ 150,000 reads per sample. An in-house pipeline as described previously51 was used for adapter and primer trimming of all samples. For amplicon processing, a further in house amplicon processing pipeline including FLASH52, USEARCH53 (v8.1), UPARSE54 and the RDP55 classifier (v2.9) was applied as described recently56,57. In summary, unmerged sequences resulting by FLASH 52 (default settings + −M 300) were directly filtered out. Furthermore, sequences with > 1Ns (ambiguous bases) and expected errors > 0.5 were also discarded. Resulting data was further processed and operational taxonomic units (OTUs) were clustered by applying USEARCH53 (v8.1). The resulting OTUs were taxonomically classified using the RDP55 classifier (v2.9) in 16S modus (Threshold > 0.8) and compared to the nt database by means of BLASTN58. In the last step, raw sequence reads were mapped back onto the OTU sequences in order to get quantitative assignments.
(Meta)Genome sequencing, assembly, binning and annotation
For each of the four samples (B. braunii race A and B from linear (T1) and stationary (T2) growth phase), the genomic DNA from three replicates was pooled. To obtain the metagenome sequence, four whole-genome-shotgun PCR-free sequencing libraries (Nextera DNA Sample Prep Kit; Illumina, Munich, Germany) were generated based on the manufacturer’s protocol representing different time points of the cultivation and different algae communities. The libraries were sequenced in paired-end mode in a MiSeq run (2 × 300 bp) resulting in 11,963,558 and 11,567,552 reads for race A T1 and T2; 6,939,996 and 12,031,204 reads for race B T1 and T2, respectively (total 12.75 Gb). The de novo metagenome assembly was performed using the Ray Meta assembler59 (v2.3.2) using a k-mer size of 41 and default settings. For contigs with length larger than 1 kb (n = 127,914), the total assembly size was 338.5 Mb, the N50 = 2884 and the largest contig 1.59 Mb. All processed raw sequence reads were aligned to the assembled metagenome contigs by means of Bowtie 260 (v2.2.4). By using SAMtools61 (v1.6), the SAM files were converted to BAM files. These files were sorted and read mapping statistics were calculated. The following portions of reads could be assembled to the draft metagenomes: 70.1% and 77.5% for race A (T1 and T2); 51.0% and 62.9% for race B (T1 and T2). For binning of the metagenome assembled genomes (MAGs), the tool MetaBAT (v0.21.3)62 was used with default settings. Completeness and contamination of the resulting MAGs were tested with BUSCO63,64 (v3.0.), using the Bacteria and Metazoa data set (Table S1).
For sequencing of the isolated species, 4 µg of purified chromosomal DNA48 was used to construct PCR-free sequencing library and sequenced applying the paired protocol on an Illumina MiSeq system, yielding ~ 1.71 Mio Reads (517 Mb in 3 Scaffolds, 58 Contigs) for Mycobacterium sp. Bb-A and ~ 1.70 Mio Reads (503 Mb in 6 Scaffolds, 65 Contigs) for Pimelobacter sp. Bb-B. Obtained sequences were de novo assembled using the GS de novo Assembler software (v2.8, Roche). An in silico gap closure approach was performed as described previously65. MinION sequencing library with genomic DNA from Brevundimonas sp. Bb-A was prepared using the Nanopore Rapid DNA Sequencing kit (SQK-RAD04) according to the manufacturer's instructions with the following changes: The entry DNA amount was increased to 800 ng and an AMPure XP bead cleanup was carried out after transposon fragmentation. Sequencing was performed on an Oxford Nanopore MinION Mk1b sequencer using a R9.5 flow cell, which was prepared according to the manufacturer's instructions. MinKNOW (v1.13.1) was used to control the run using the 48 h sequencing run protocol; base calling was performed offline using albacore (v2.3.1). The assembly was performed using canu v1.766, resulting in a single, circular contig (170 Mb, 1 kb to 17.5 kb). This contig was then polished with Illumina short read data from the metagenome data set using pilon67, run for sixteen iterative cycles. bwa-mem68 was used for read mapping in the first eight iterations and bowtie2 v2.3.260 in the second set of eight iterations. Annotation of the draft chromosomes was performed within the PROKKA69 (v1.11) pipeline and visualized using the GenDB 2.0 system70.
Gene prediction, taxonomic assignment and functional characterization
Gene prediction on the assembled contigs of the MAGs was performed with Prodigal71 (v2.6.1) using the metagenome mode (“-p meta”). All genes were annotated and analyzed with the in-house EMGB annotation system using the databases NCBI NR58, Pfam72 and KEGG73. Based on DIAMOND74 (v0.8.36) hits against the NCBI-NR58 database, all genes were subject to taxonomic assignment using MEGAN75. Relationships to metabolic pathways were assigned based on DIAMOND74 hits against KEGG73. For Pfam72 annotations, pfamscan was used with default parameters. The genomes were manually inspected using an E-value cutoff of 1 × 10–10 for several specific sequences listed in Additional file 2, Table S6. The profiling of the carbohydrate-active enzymes (CAZy) encoded by the B. braunii community members was accomplished using dbCAN276 metaserver (Table S7). The identification, annotation and analysis of the secondary metabolite biosynthesis gene clusters within the MAGs and complete genomes were accomplished via the antiSMASH77 (v4.0) web server with default settings (Table S9).
Phylogenetic analyses
To assign and phylogenetically classify the MAGs, a core genome tree was created by applying EDGAR78 (v2.0). For calculation of a core-genome-based phylogenetic tree, the core genes of all MAGs and the three isolates Mycobacterium sp. Bb-A, Brevundimonas sp. Bb-A and Pimelobacter sp. Bb-B, corresponding to selected reference sequences were considered. The implemented version of FASTtree79 (using default settings) in EDGAR78 (v2.0) was used to create a phylogenetic Maximum-Likelihood tree based on 53 core genes (see Table S4) of all genomes. Additionally, the MAGs were taxonomically classified on species level by calculation of average nucleic and amino acid identities (ANI, AAI) by applying EDGAR78 (v2.0). For phylogenetic analysis of the 16S rDNA amplicon sequences, MEGA80 (v7.0.26) was used. Phylogenetic tree was constructed by means of Maximum-Likelihood Tree algorithm using Tamura 3-parameter model. The robustness of the inferred trees was evaluated by bootstrap (1000 replications).
Data availability
The high-throughput 16S rDNA amplicon sequencing datasets obtained during the present work are available at the EMBL-EBI database under BioProjectID PRJEB21978. The chloroplastic 16S rRNA genes sequencing data of B. braunii SAG 30.81 are available under the GenBank accession number MN956340. The metagenome datasets of Botryococcus braunii consortia as well as the resulting MAGs have been deposited under the BioProjectIDs PRJEB26344 and PRJEB26345, respectively. The genomes of the isolates Brevundimonas sp. Bb-A, Mycobacterium sp. BbA or Pimelobacter sp. Bb-B are accessible under BioProjectIDs PRJNA528993, PRJEB28031 und PRJEB28032, respectively. In addition, the communities most abundant ten high-quality draft as well as isolate genomes are also accessible via EMGB platform (https://emgb.cebitec.uni-bielefeld.de/Bbraunii-bacterial-consortium/).
References
Grosberg, R. K. & Strathmann, R. R. The evolution of multicellularity: a minor major transition?. Annu. Rev. Ecol. Evol. Syst. 38, 621–654 (2007).
Seymour, J. R., Amin, S. A., Raina, J. B. & Stocker, R. Zooming in on the phycosphere: the ecological interface for phytoplankton-bacteria relationships. Nat. Microbiol. 2, 1–12 (2017).
Ramanan, R., Kim, B.-H., Cho, D.-H., Oh, H.-M. & Kim, H.-S. Algae–bacteria interactions: evolution, ecology and emerging applications. Biotechnol. Adv. 34, 14–29 (2016).
Amin, S. A. et al. Interaction and signalling between a cosmopolitan phytoplankton and associated bacteria. Nature 522, 98–101 (2015).
Cole, J. J. Interactions between bacteria and algae in aquatic ecosystems. Annu. Rev. Ecol. Syst. 13, 291–314 (1982).
Zengler, K. & Zaramela, L. S. The social network of microorganisms: how auxotrophies shape complex communities. Nat. Rev. Microbiol. 16, 383–390 (2018).
Wang, X. et al. Lysis of a red-tide causing alga, Alexandrium tamarense, caused by bacteria from its phycosphere. Biol. Control 52, 123–130 (2010).
Croft, M. T., Lawrence, A. D., Raux-Deery, E., Warren, M. J. & Smith, A. G. Algae acquire vitamin B12 through a symbiotic relationship with bacteria. Nature 438, 90–93 (2005).
Croft, M. T., Warren, M. J. & Smith, A. G. Algae need their vitamins. Eukaryot. Cell 5, 1175–1183 (2006).
Tang, Y. Z., Koch, F. & Gobler, C. J. Most harmful algal bloom species are vitamin B1 and B12 auxotrophs. Proc. Natl. Acad. Sci. 107, 20756–20761 (2010).
Grant, M. A. A., Kazamia, E., Cicuta, P. & Smith, A. G. Direct exchange of vitamin B12 is demonstrated by modelling the growth dynamics of algal-bacterial cocultures. ISME J. 8, 1418–1427 (2014).
Sañudo-Wilhelmy, S. A., Gómez-Consarnau, L., Suffridge, C. & Webb, E. A. The role of B vitamins in marine biogeochemistry. Ann. Rev. Mar. Sci. 6, 339–367 (2014).
Gómez-Consarnau, L. et al. Mosaic patterns of B-vitamin synthesis and utilization in a natural marine microbial community. Environ. Microbiol. 20, 2809–2823 (2018).
Banerjee, A., Sharma, R., Chisti, Y. & Banerjee, U. C. Botryococcus braunii: a renewable source of hydrocarbons and other chemicals. Crit. Rev. Biotechnol. 22, 245–279 (2002).
Blifernez-Klassen, O. et al. Metabolic survey of Botryococcus braunii: Impact of the physiological state on product formation. PLoS ONE 13, e0198976 (2018).
Marschner, P. Marschner’s Mineral Nutrition of Higher Plants 3rd edn. (Elsevier Ltd., Amsterdam, 2012).https://doi.org/10.1016/C2009-0-63043-9.
Bisseling, T., Dangl, J. L. & Schulze-Lefert, P. Next-generation communication. Science 324(80), 691 (2009).
Nguyen, C. Rhizodeposition of organic C by plant: mechanisms and controls. In Sustainable Agriculture (eds Lichtfouse, E. et al.) 97–123 (Springer, Dordrecht, 2009). https://doi.org/10.1007/978-90-481-2666-8_9.
McKirdy, D. M., Cox, R. E., Volkman, J. K. & Howell, V. J. Botryococcane in a new class of Australian non-marine crude oils. Nature 320, 57–59 (1986).
Fuentes, J. L. et al. Impact of microalgae-bacteria interactions on the production of algal biomass and associated compounds. Mar. Drugs 14, 100 (2016).
Tanabe, Y. et al. A novel alphaproteobacterial ectosymbiont promotes the growth of the hydrocarbon-rich green alga Botryococcus braunii. Sci. Rep. 5, 10467 (2015).
Chirac, C., Casadevall, E., Largeau, C. & Metzger, P. Bacterial influence upon growth and hydrocarbon production of the green alga Botryococcus braunii. J. Phycol. 21, 380–387 (1985).
Jax, K. Ecological units: definitions and application. Q. Rev. Biol. 81, 237–258 (2006).
Xu, J. et al. The structure and function of the global citrus rhizosphere microbiome. Nat. Commun. 9, 4894 (2018).
Sambles, C. et al. Metagenomic analysis of the complex microbial consortium associated with cultures of the oil-rich alga Botryococcus braunii. Microbiologyopen 6, e00482 (2017).
Gkarmiri, K. et al. Identifying the active microbiome associated with roots and rhizosphere soil of oilseed rape. Appl. Environ. Microbiol. 83, e01938-e2017 (2017).
Schreiter, S. et al. Effect of the soil type on the microbiome in the rhizosphere of field-grown lettuce. Front. Microbiol. 5, 144 (2014).
Rojo, F. Enzymes for aerobic degradation of alkanes. Handb. Hydrocarb. Lipid Microbiol. 2, 781–797 (2010).
Pančić, M. & Kiørboe, T. Phytoplankton defence mechanisms: traits and trade-offs. Biol. Rev. 93, 1269–1303 (2018).
Molnár, I. et al. Bio-crude transcriptomics: gene discovery and metabolic network reconstruction for the biosynthesis of the terpenome of the hydrocarbon oil-producing green alga, Botryococcus braunii race B (Showa). BMC Genom. 13, 576 (2012).
Jaehme, M. & Slotboom, D. J. Diversity of membrane transport proteins for vitamins in bacteria and archaea. Biochim. Biophys. Acta Gen. Subj. 1850, 565–576 (2015).
Cotter, P. D., Ross, R. P. & Hill, C. Bacteriocins: a viable alternative to antibiotics?. Nat. Rev. Microbiol. 11, 95–105 (2013).
Cirri, E. & Pohnert, G. Algae−bacteria interactions that balance the planktonic microbiome. New Phytol. 223, 100–106 (2019).
Krohn-Molt, I. et al. Insights into Microalga and bacteria interactions of selected phycosphere biofilms using metagenomic, transcriptomic, and proteomic approaches. Front. Microbiol. 8, 1941 (2017).
Reed, H. & Martiny, J. Testing the functional significance of microbial composition in natural communities. FEMS Microbiol. Ecol. 62, 161–170 (2007).
Rivett, D. W. & Bell, T. Abundance determines the functional role of bacterial phylotypes in complex communities. Nat. Microbiol. 3, 767–772 (2018).
Paerl, R. W. et al. Use of plankton-derived vitamin B1 precursors, especially thiazole-related precursor, by key marine picoeukaryotic phytoplankton. ISME J. 11, 753 (2017).
Aaronson, S. et al. The cell content and secretion of water-soluble vitamins by several freshwater algae. Arch. Microbiol. 112, 57–59 (1977).
Higa, A., Miyamoto, E., Rahman, L. & Kitamura, Y. Root tip-dependent, active riboflavin secretion by Hyoscyamus albus hairy roots under iron deficiency. Plant Physiol. Biochem. 46, 452–460 (2008).
Thompson, H., Angelova, A., Bowler, B., Jones, M. & Gutierrez, T. Enhanced crude oil biodegradative potential of natural phytoplankton-associated hydrocarbonoclastic bacteria. Environ. Microbiol. 19, 2843–2861 (2017).
Dashti, N. et al. Most hydrocarbonoclastic bacteria in the total environment are diazotrophic, which highlights their value in the bioremediation of hydrocarbon contaminants. Microbes Environ. 30, 70–75 (2015).
Gilbert, J. A. et al. Microbiome-wide association studies link dynamic microbial consortia to disease. Nature 535, 94 (2016).
Durán, P. et al. Microbial interkingdom interactions in roots promote Arabidopsis survival. Cell 175, 973–983 (2018).
Astals, S., Nolla-Ardèvol, V. & Mata-Alvarez, J. Anaerobic co-digestion of pig manure and crude glycerol at mesophilic conditions: Biogas and digestate. Bioresour. Technol. 110, 63–70 (2012).
Schlegel, H. G., Kaltwasser, H. & Gottschalk, G. Ein Submersverfahren zur Kultur wasserstoffoxydierender Bakterien: Wachstumsphysiologische Untersuchungen. Arch. Mikrobiol. 38, 209–222 (1961).
Jett, B. D., Hatter, K. L., Huycke, M. M. & Gilmore, M. S. Simplified agar plate method for quantifying viable bacteria. Biotechniques 23, 648–650 (1997).
Schwarzhans, J. P. et al. Dependency of the fatty acid composition of Euglena gracilis on growth phase and culture conditions. J. Appl. Phycol. 27, 1389–1399 (2014).
Zhou, J., Bruns, M. A. & Tiedje, J. M. DNA recovery from soils of diverse composition. Appl. Environ. Microbiol. 62, 316–322 (1996).
Maus, I. et al. Biphasic study to characterize agricultural biogas plants by high-throughput 16S rRNA gene amplicon sequencing and microscopic analysis. J. Microbiol. Biotechnol. 27, 321–334 (2017).
Takahashi, S., Tomita, J., Nishioka, K., Hisada, T. & Nishijima, M. Development of a prokaryotic universal primer for simultaneous analysis of bacteria and archaea using next-generation sequencing. PLoS ONE 9, e105592 (2014).
Wibberg, D. et al. Genome analysis of the sugar beet pathogen Rhizoctonia solani AG2-2IIIB revealed high numbers in secreted proteins and cell wall degrading enzymes. BMC Genom. 17, 245 (2016).
Magoč, T. & Salzberg, S. L. FLASH: Fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27, 2957–2963 (2011).
Edgar, R. C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26, 2460–2461 (2010).
Edgar, R. C. UPARSE: highly accurate OTU sequences from microbial amplicon reads. Nat. Methods 10, 996–998 (2013).
Wang, Q., Garrity, G. M., Tiedje, J. M. & Cole, J. R. Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl. Environ. Microbiol. 73, 5261–5267 (2007).
Liebe, S. et al. Taxonomic analysis of the microbial community in stored sugar beets using high-throughput sequencing of different marker genes. FEMS Microbiol. Ecol. 92, 1–12 (2016).
Klassen, V. et al. Highly efficient methane generation from untreated microalgae biomass. Biotechnol. Biofuels 10, 186 (2017).
O’Leary, N. A. et al. Reference sequence (RefSeq) database at NCBI: Current status, taxonomic expansion, and functional annotation. Nucleic Acids Res. 44, D733–D745 (2016).
Boisvert, S., Raymond, F., Godzaridis, E., Laviolette, F. & Corbeil, J. Ray Meta: scalable de novo metagenome assembly and profiling. Genome Biol. 13, R122 (2012).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Kang, D. D., Froula, J., Egan, R. & Wang, Z. MetaBAT, an efficient tool for accurately reconstructing single genomes from complex microbial communities. PeerJ 3, e1165 (2015).
Simão, F. A., Waterhouse, R. M., Ioannidis, P., Kriventseva, E. V. & Zdobnov, E. M. BUSCO: Assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 31, 3210–3212 (2015).
Waterhouse, R. M. et al. BUSCO applications from quality assessments to gene prediction and phylogenomics. Mol. Biol. Evol. 35, 543–548 (2018).
Wibberg, D. et al. Complete genome sequencing of Agrobacterium sp. H13–3, the former Rhizobium lupini H13–3, reveals a tripartite genome consisting of a circular and a linear chromosome and an accessory plasmid but lacking a tumor-inducing Ti-plasmid. J. Biotechnol. 155, 50–62 (2011).
Koren, S. et al. Canu: Scalable and accurate long-read assembly via adaptive κ-mer weighting and repeat separation. Genome Res. 27, 722–736 (2017).
Walker, B. J. et al. Pilon: An integrated tool for comprehensive microbial variant detection and genome assembly improvement. PLoS ONE 9, e112963 (2014).
Li, H. Toward better understanding of artifacts in variant calling from high-coverage samples. Bioinformatics 30, 2843–2851 (2014).
Seemann, T. Prokka: rapid prokaryotic genome annotation. Bioinformatics 30, 2068–2069 (2014).
Meyer, F. et al. GenDB: an open source genome annotation system for prokaryote genomes. Nucleic Acids Res. 31, 2187–2195 (2003).
Hyatt, D. et al. Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinform. 11, 119 (2010).
Finn, R. D. et al. Pfam: the protein families database. Nucleic Acids Res. 42, D222–D230 (2014).
Kanehisa, M., Furumichi, M., Tanabe, M., Sato, Y. & Morishima, K. KEGG: New perspectives on genomes, pathways, diseases and drugs. Nucleic Acids Res. 45, D353–D361 (2017).
Buchfink, B., Xie, C. & Huson, D. H. Fast and sensitive protein alignment using DIAMOND. Nat. Methods 12, 59–60 (2015).
Huson, D. H. et al. MEGAN community edition: interactive exploration and analysis of large-scale microbiome sequencing data. PLoS Comput. Biol. 12, e1004957 (2016).
Zhang, H. et al. dbCAN2: a meta server for automated carbohydrate-active enzyme annotation. Nucleic Acids Res. 46, W95–W101 (2018).
Blin, K. et al. antiSMASH 4.0: improvements in chemistry prediction and gene cluster boundary identification. Nucleic Acids Res. 45, W36–W41 (2017).
Blom, J. et al. EDGAR 2.0: an enhanced software platform for comparative gene content analyses. Nucleic Acids Res. 44, W22–W28 (2016).
Price, M. N., Dehal, P. S. & Arkin, A. P. FastTree 2: approximately maximum-likelihood trees for large alignments. PLoS ONE 5, e9490 (2010).
Kumar, S., Stecher, G. & Tamura, K. MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 33, 1870–1874 (2016).
Acknowledgements
We are thankful to Joao Gouveia and Douwe van der Veen (Wageningen University, The Netherlands) for providing Botryococcus braunii race A and B (CCALA778 and AC761) stock cultures. Furthermore, we are grateful to the Center for Biotechnology (CeBiTec) at Bielefeld University for access to the Technology Platforms. This work was supported by the European Union Seventh Framework Programme (FP7/2007-2013) under grant agreement 311956 (relating to project “SPLASH – Sustainable PoLymers from Algae Sugars and Hydrocarbons”). The bioinformatics support by the BMBF-funded project “Bielefeld-Gießen Center for Microbial Bioinformatics” – BiGi (Grant 031A533) within the German Network for Bioinformatics Infrastructure (de. NBI) is gratefully acknowledged.
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
O.B.K., V.K, and O.K. conceived this study. O.B.K., A.S., A.B., J.K. and O.K. supervised experiments and analyses. O.B.K., V.K., E.C. and S.C. accomplished all physiological studies. O.B.K., D.W., A.S., C.H., C.R., A.B., V.K., A.A.D., A.W., J.K., O.R., J.B. and A.G. performed the sequencing and the bioinformatic analyses. O.B.K., V.K., A.B. and O.K. wrote the original draft manuscript with contributions from all authors. O.B.K., V.K., A.B and O.K. reviewed and edited the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Blifernez-Klassen, O., Klassen, V., Wibberg, D. et al. Phytoplankton consortia as a blueprint for mutually beneficial eukaryote-bacteria ecosystems based on the biocoenosis of Botryococcus consortia. Sci Rep 11, 1726 (2021). https://doi.org/10.1038/s41598-021-81082-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-81082-1
This article is cited by
-
Characterization of a bloom-associated alphaproteobacterial lineage, ‘Candidatus Phycosocius’: insights into freshwater algal-bacterial interactions
ISME Communications (2023)
-
The overlooked role of a biotin precursor for marine bacteria - desthiobiotin as an escape route for biotin auxotrophy
The ISME Journal (2022)
-
Large-scale screening of natural genetic resource in the hydrocarbon-producing microalga Botrycoccus braunii identified novel fast-growing strains
Scientific Reports (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.