Abstract
Recently, the critical roles played by genetic variants of TREM2 (Triggering Receptor Expressed on Myeloid cells 2) in Alzheimer’s disease have been aggressively highlighted. However, few studies have focused on the deleterious roles of Nasu-Hakola disease (NHD) associated TREM2 variants. In order to get insights into the contributions made by these variants to neurodegeneration, we investigated the influences of four NHD associated TREM2 mutations (Y38C, W50C, T66M, and V126G) on loss-of-function, and followed this with in silico prediction and conventional molecular dynamics simulation. NHD mutations were predicted to be highly deleterious by eight different in silico bioinformatics tools and found to induce conformational changes by molecular dynamics simulation. As compared with the wild-type, the four variants produced substantial differences in the collective motions of loop regions, which not only promoted structural remodeling in the CDR2 (complementarity-determining region 2) loop but also in the CDR1 loop, by changing inter- and intra-loop hydrogen bonding networks. In addition, structural studies in a free energy landscape analysis showed that Y38, T66, and V126 are crucial for maintaining the structural features of CDR1 and CDR2 loops, and that mutations in these positions produced steric clashes and loss of ligand binding. These results showed the presence of mutations in the TREM2 ectodomain induced flexibility and caused structural alterations. Dynamical scenarios, as provided by the present study, may be critical to our understanding of the roles of these TREM2 mutations in neurodegenerative diseases.
Similar content being viewed by others
Introduction
The accumulation of amyloid-β (Aβ) in brain parenchyma is the main hallmark of Alzheimer’s disease (AD), which results in a slow progressive decline of cognitive function by causing the formation of intracellular neurofibrillary tangles, synapse loss, and cell death1,2. Although the precise mechanisms and molecular determinants of this neurodegenerative disease are incomplete, recent whole-genome sequencing studies have demonstrated altered genetic loci including those in TREM2 (Triggering Receptor Expressed on Myeloid cells 2) are associated with a markedly higher risk of progression to AD3,4.
TREM2 is a V-type immunoglobulin (Ig) domain-containing transmembrane protein that is expressed in osteoclasts, microglia, alveolar macrophages, and other mononuclear phagocytes4. The activation of TREM2 is initiated by anionic lipids, such as bacterial lipopolysaccharide (LPS) and phospholipids, and several putative ligands like apolipoprotein J (ApoJ) and apolipoprotein E (ApoE) have been reported to bind to TREM25,6,7. Upon ligand binding, TREM2 recruits protein tyrosine kinase SYK through an adapter protein known as DNAX-activating protein of 12 kDa (DAP12), which interacts with the transmembrane region of TREM28,9,10. This binding is followed by a cascade of signaling events, which include the activations of MAPK and PI3K, and regulates the phagocytosis of cellular debris and the inflammatory responses of microglia10,11,12. During early and mid-term AD, TREM2 plays a protective role13, and its overexpression is associated with clearing soluble and insoluble Aβ42 aggregates from brain14,15. Furthermore, TREM2 has also been reported to suppress the accumulation and diffusion of Aβ by modulating microglial activation around amyloid plaques16,17,18,19.
During the past six years, cell biological research studies have suggested links between several TREM2 variants and failure of Aβ clearance, and several rare variants have been shown to be associated with AD progression (Table S1)20. Mutations, such as, R47H and R62H variants have been shown to present significant risks of AD, and, as a result, are considered AD-associated variants21. Interestingly, at the molecular level, genetic variations in TREM2 have also been linked with frontotemporal dementia (FTD) and Nasu-Hakola disease (NHD), the latter of which is characterized by demyelination, early-onset dementia, and bone cyst lipoma and known to be associated with Y38C, W50C, T66M, and V126G mutations in the ectodomains of TREM222,23,24,25.
In this regard, Yeh et al. showed that TREM2 variants, including AD-associated (R47H, R62H, and D87N) and NHD linked mutations (Y38C and T66M) reduce binding between TREM2 and its ligands. Moreover, loss of ligand binding was found to be more severe for NHD associated mutations than AD-associated mutations7. The present study provides in silico insights of the magnitudes of the damaging effects of TREM2 variants, particularly of NHD associated mutations, and provides classical molecular dynamics simulation-based descriptions of the structural dynamic behavior of TREM2 protein in the wild and mutated states.
Results and Discussions
Assessment of pathogenicity of TREM2 variants
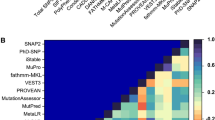
Human TREM2 is composed of 230 amino acids and a polypeptide chain that consists of three distinct regions, namely, an N-terminal mature ectodomain (ECD, residues 19–174), a membrane-spanning region (residues 175–195), and a C-terminal cytosolic tail (residues 196–230). The other amino acids, especially residues 1–18 act, as a signaling peptide in the TREM2 signaling cascade. As shown in Fig. 1, the tertiary structure of the TREM2 ECD domain is mainly composed of nine β-strands (βA - βF), which include three major complementarity-determining regions (dubbed CDR loops), that is, CDR1 (residues Pro37 to Arg47), CDR2 (residues Thr66 to Arg76), and CDR3 (His144 to Glu117). Like the other members of the Ig superfamily, ligands bind to TREM2 ECD near apical CDR loops. Previous studies have shown that CDR2 maintains a stable conformation in normal conditions by maintaining H-bonding using the CDR1 loop, which is necessary for ligand interactions26. However, genetic variations result in the destabilizations of these loops, and thus, by impairing ligand binding may have deleterious effects. According to X-ray diffraction analysis, the H-bonding network between CDR loops appears to be lost in the R47H variant and result in conformational remodeling of the CDR2 loop. In the present study, we used in silico deleterious prediction analysis to re-rank the risk associations of known disease-associated TREM2 variants. Eight in silico state-of-the-art-tools were utilized to predict deleteriousness: SIFT (= 0), PolyPhen‐2 (>0.9), PROVEAN (<−2.5), I‐Mutant 3.0 (<−0.5), FATHMM (<−3.0 or >3.0), MutPred (>0.75), CADD (>20), and Condel (>0.8), where parenthesis show the cutoffs used. Of these tools, I-Mutant 3 predicted the highest number of deleterious variants, though all predictions substantially concurred (Fig. S1). In fact, in silico predictions of any two tools were found to be significantly associated for most combinations (P < 0.0001 by the Student’s t‐test). Analysis showed the NHD W50C mutation was the most deleterious by all tools, whereas other NHD variants, including Y38C, T66M, and V126G, were classified as significantly deleterious (P = 0.0001) by seven of the eight computational tools. Interestingly, these results are consistent with those of a previous experimental study, in which the Y38C and T66M mutations (located in the CDR1 and CDR2 regions, respectively) were found to be involved in loss of ligand binding. Since NHD variants were found to be more deleterious by in silico analysis and experimental findings, we systematically analyzed how these mutations contribute to the pathological behavior of TREM2.
Changes in the conformational stabilities of TREM2 variants
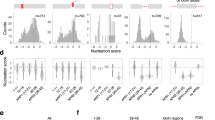
Over past decades, molecular simulation has provided means of characterizing in detail the structural configurations of macromolecules in various environments, as determined by their functions and interactions with other molecular species27,28. In this context, molecular dynamics simulations of wild-type and Y38C, W50C, T66M, and V126G variants were conducted for 100 ns to access their structural dynamic and stability characteristics. The conformational stabilities of wild and variant types during simulations were analyzed by calculating RMSD values for the backbones of all proteins from starting structures (Fig. 2). RMSD analysis revealed that the wild-type and Y38C variant achieved equilibrium after 5 ns and that equilibrium states were maintained until the end of simulations, whereas the three other variants achieved equilibrium after 20 ns. Furthermore, wild-type and Y38C structures showed similar deviations from their starting structures at 100 ns, which resulted in backbone RMSD values of ~1.4 to 1.7 Å during simulations. However, W50C, T66M, and V126G structures showed significantly different deviations from wild-type and Y38C structures (range ~1.9 to 3 Å). After the relaxation period, the magnitudes of fluctuations of the Y38C, W50C, and V126G variants were stably maintained29. However, the RMSD of the T66M variant reduced at 80 ns but then remained stable from 84 ns to the end of the simulation, indicating a major conformational change occurred. A similar pattern was also observed for the W50C variant. Overall, RMSD analysis suggested that the trajectories produced provided an appropriate basis for further analysis.
Root-mean-square deviation (RMSD) of Cα atoms for wild-type and TREM2 variants at 100 ns. Here, the dark green line represents the wild-type, and the blue, red, orange and violet lines represent Y38C, W50C, T66M, and V126G variant RMSDs. In addition, the bottom panel shows density plots for the wild-type and variants and illustrates the distributions of sampled conformations during simulation.
Conformational in changes in variants as compared with the wild-type were evaluated using radius of gyration (Rg) (Fig. 3), which describes overall dimensions of protein structures in terms of compactness (higher Rg values represent lower compactness)30,31. As shown in Fig. 3a and c, the overall dimensions of Y38C and T66M were similar to those of the wild-type, whereas V126G (Fig. 3d) showed increased compactness from 50 ns to the end of the simulation. On the other hand, W50C (Fig. 3c) had a higher Rg than the wild-type. SASA provides a measure of solvent accessibility, and SASA profiles were similar to Rg profiles for variants and the wild-type. The wild-type and Y38C, W50C, and T66M variants showed similar deviations from their initial structures (Fig. 4a–c), but V126G showed an increase in SASA after 50 ns (Fig. 4d), which indicated bloating of its structure as compared with the wild-type32. Although RMSD analysis showed conformational alterations in T66M and V126G variants versus the wild-type, only the V126G variant showed conformational alterations by Rg and SASA. T66M showed a decrease in RMSD before 80 ns, and Rg and SASA values also reduced at this time point. Summarizing, Rg, SASA, and RMSD analyses (Table S1) showed that the Y38C, W50C, T66M, and V126G variants modulated protein dimensions, which in turn suggested misfolding and changes in protein-protein interactions33.
Changes in total solvent accessible surface areas (SASA) of the Y38C (a), W50C (b), T66M (c), and V126G (d) variants as compared with wild-type TREM2. Here, the dark green line represents the wild-type and blue, red, orange, and violet lines represent Y38C, W50C, T66M, and V126G variants, respectively.
Figure 5 shows RMSF values for protein backbones and depicts local changes in protein structure caused by mutations. Local residue fluctuations in proteins underlie biological functions, as the many functional sites in proteins are uniquely coupled with structural fluctuations34,35,36. As represented in Fig. 5a, TREM2 contains ten loops, including three CDR loops, which all fluctuated throughout simulations. Degrees of fluctuation were greater in variant structures than in the wild type. Interestingly, Y38C and T66M variants showed high levels of fluctuation in CDR1 and CDR2 regions, respectively. The V126G variant showed increased local flexibility, especially in the regions including residues 50 to 60 and 85 to 95 and in the CDR1 loop and N-terminal tail. On the other hand, no significant changes were observed for the W50C variant as compared with the wild-type with the exception of residues 86 to 93, which had a lower RMSF value, suggesting conformational changes in this region. For more insight, SASA values were calculated per residue for the wild-type and variants. Variant structures showed an increase in residual SASA values, especially W50C and T66M in CDR1 and CDR2 regions (Fig. 5b). SASA deviations were also observed for Y38C and V126G variants in the CDR1 and CDR2 loop regions. SASA per residue values reflect structural conformational changes at protein surfaces; lower values mean residues are buried inside globular folded conformations37,38,39. This analysis together and the RMSF study suggest that the Y38C, W50C, T66M, and V126G increase solvent exposure, modulate protein-protein interactions, and possibly increase aggregation propensity40,41,42.
Investigation of the local structural effects of mutations. Root mean square fluctuations of Cα atoms for wild-type TREM2 and the four variants (a). SASA values were computed on a per residue basis for the wild-type and TREM2 variants (b). In all plots, residues index is level with colored bar, representing the structural propensity. In each graph, the dark green line represents the wild-type, while blue, red, orange, and violet lines represent the Y38C, W50C, T66M, and V126G variants, respectively.
Effects of mutations on protein dynamics
In different functional states, proteins undergo conformational transitions due to global domain motions facilitated by the collective motions of backbone atoms43. Generally, the residues located in the regions with secondary structures move in a concerted manner44. For additional insight into the dynamics of wild-type TREM2 and its variants, dynamic cross-correlation maps (DCCM) and principal component analyses were performed on the MD trajectories of different systems. DCCM addresses the concerted motions of protein residues and highlights strongly correlated motions between specific residues in red (Fig. 6), and conversely, highlights highly anti-correlated motions in blue. Analysis revealed that as compared with the wild-type, the Y38C, W50C, T66M, and V126G mutations induced completely different motions. Y38C (Fig. 6b) produced significant anti-correlative motion between residues 17–27 (CDR1 loop, residues 37 to 47) and residues 45 to 56 (CDR2 loop, residues 65 to 76), and 65 to 74 (residues, 85 to 94). The T66M variant (Fig. 6d) slightly reduced the degrees of correlated and anti-correlated motions observed in the wild-type, although no significant correlation has been seen in the DCCM analysis. The W50C variant exhibited correlative motions not observed in Y38C, T66M, or V126G variants, and showed strong anti-correlative motions between residues 5 to 11 (residues, 25 to 31) and residues 38 to 46 (residues, 58 to 66), 56 to 60 (residues, 76 to 80), 62 to 67 (residues, 82 to 87), 73 to 76 (residues, 93 to 96), respectively (Fig. 6c). Furthermore, significant positive correlations were observed between region 40 to 46 (residues, 60 to 66) and 63 to 69 (residues, 83 to 89), the latter of which showed anti-correlative motions with residues in CDR2 loops 50–56 (residues, 70 to 76) and the region including residues from 77 to 85 (residues, 97 to 105). For the V126G variant (Fig. 6e), results showed mutation increased correlative intra-residue movement, specifically between the loop residues, 34 to 40 (residues, 54 to 60) and the c-terminal portion 80 to 105 (residues, 100 to 125). A negative correlation was also observed in the CDR1 and CDR2 loop regions, suggesting the mutation had a deleterious effect on TREM2 functions.
Calculated dynamic cross-correlation matrix of Cα atoms around their mean positions for 100 ns molecular dynamics simulations. Extents of correlated motions and anti-correlated motions are color-coded from red to blue, which represent positive and negative correlations, respectively. Wild-type (a), Y38C (b), W50C (c), T66M (d), and V126G (e).
To further support the changes on the protein dynamics, principal component analysis (PCA) was conducted based on Cα atoms (Fig. S2). PCA analysis, which reduces the complexity of collective motions is usually performed to obtain insight of the dynamics and mechanical properties of simulated systems28. Furthermore, the dynamics of two different proteins are best compared by characterizing their phase space behaviors, which are directly associated with protein stability and function45,46. We described the overall combined motions of Cα atoms of protein structures using eigenvectors of covariance matrix, which is correlated with the coincident eigenvalues. Trace values for the wild-type and the Y38C, W50C, T66M, and V126G variants were found to be 46.707 Å2, 47.487 Å2, 58.94 Å2, 49.822 Å2, and 99.309 Å2, respectively, which suggested that mutations induced collective flexibility. V126G had a higher trace value than the wild-type or Y38C, W50C, or T66M variants, although its Rg value reduced after 50 ns of simulation. This observation suggests V126G exhibits significantly greater protein local flexibility rather than overall flexibility, as can be seen in the RMSF plot (Fig. 5), which shows greater flexibility in residues 50 to 60 and 85 to 95. To investigate further, three PCAs were considered for further analysis. Resultantly, simulated systems of the wild-type and Y38C, W50C, T66M, and V126G variants showed variances of 19.81%, 33.43%, 24.50%, 20.94%, and 29.67%, respectively, for the first three PCAs. The motions of different proteins can be visualized using projections of trajectories in phase spaces of the first two principal components (PC1, PC2), whereas variants show different patterns of conformational spaces, indicating significant changes in protein conformations. Using these projections, Y38C showed a uniform conformational distribution in a larger phase space, indicating greater flexibility, than the wild-type (Fig. S2b). The T66M variant also showed conformational transitions during simulation that indicated less flexibility over time (Fig. S2d). On the other hand, the W50C and V126G variants exhibited higher periodic jumps, indicating greater fluctuations than the wild-type and the occupation of more phase space (Fig. S2c and e).
The variations in atomic movement were presented by all first PCAs (Fig. 7), and PC1 exhibited the most dynamic motion. In the figure, the displacement of atoms is denoted by wide tubes, whereas narrow tubes mark regions that remained rigid during the simulation. Structural deviations in the wild-type and in variants were obvious for first PCAs (Fig. 7), in which the contributions of residues to first principal components are represented in the plot rendered in the bottom panel (Fig. 7f), which describes residual mobilities during the simulation. As shown by the figure, Y38C induced flexibility in several loops of TREM2, including CDR1, CDR2, and CDR3 (Fig. 7b), whereas T66M induced significant flexibility in CDR2 (Fig. 7d).
Representation of atomic displacements of PCA1 as determined by simulations of wild-type TREM2 (a), and of the Y38C (b), W50C (c), T66M (d), and V126G (e) variants, showing significant movement differences. The green circles indicate regions of greater flexibility. Residue-wise loadings for PC1, where the dark green line represents the wild-type and blue, red, orange and violet lines represent the Y38C, W50C, T66M and V126G variants (f).
Conformational changes in the wild-type structure were observed in the loop region, including residues 47 to 60, and this region also exhibited significant changes in the W50C variant (Fig. 7c and e). The W50C variant also exhibited more residual mobility in the CDR1 loop than the wild-type. Interestingly, V126G induced the highest residual mobility in the region from 47 to 60, which is consistent with RMSF and DCCM results and supports mutation-induced local conformational changes.
Mutation-induced conformational changes
Although MD simulations well explained conformational changes in protein structures, we also investigated mutation-driven changes in secondary structures. RMSF, DCCM and PCA analysis showed Y38C and T66M variants exhibited considerable flexibility in CDR regions that might alter structural orientations in these regions. Therefore, variations in secondary structural elements for wild-type and variant proteins during simulation were characterized using Schrödinger 2017–1 simulation interaction diagrams (LLC, New York, NY, USA)47. During simulation, the appearance of secondary structural elements determines protein structural flexibility, for example, conformations such as α-helices and β-sheets are naturally more rigid than coil or turn conformations28. Total percentages of secondary structural elements in total simulation time are shown in Figs. S3,S4 to S8. Interestingly, variants increased total secondary structures as compared with the wild-type. Moreover, the CDR1 (residue index 17–27) and CDR2 (residue index 46–57) regions gained a short α-helix conformation in variant structures, though the percentage of helix formation was greater in Y38C (Table S2). During simulation, Y38C showed higher α-helix propensity in the CDR1 loop from 20 to 40 ns, although α-helix formation was observed to persist over the entire simulation (Fig. S5). On the other hand, W50C and V126G variants increased the total secondary structure elements in the TREM2, and increases in α-helix conformation in the CDR2 region was greater in W50C and V126G than in Y38C and T66M. The W50C variant only showed α-helix formation in CDR1 loop, but its percentage was less than that of Y38C and T66M. Furthermore, the W50C variant demonstrated significant conformational changes in the region residue 48 to 64 (mainly in β-strands, βc and βc′) within the simulation time 68 to 78 ns (Fig. S6). Notably, the residues of βc and βc′ strands are located near the W50C mutation site, where formation and reduction of β-strand propensity were observed, as a result, the RMSD pattern was seen to change in this time point. In addition, T66M had low RMSD values at simulation times <80 ns. Interestingly, during this period, T66M showed aggressive α-helix formation in the CDR2 loop, which supported the dramatic behavior of T66M during the simulation (Fig. S7). Previous studies on AD associated variants have shown that substitution of R47 by H47 facilitates CDR2 loop remodeling to a short α-helix, which contributes to loss of ligand binding26. In summary, our results suggest that the variants examined facilitate conformation remodeling of CDR2 loop and of CDR1 regions.
Mechanistic insights into conformational remodeling
Intra-residue H-bonding influences the secondary structures of biological macromolecules48. Thus, H-bond occupancy analyses within CDR1 and CDR2 loops of the wild-type and variants types were undertaken as both loops undergo conformational remodeling (Fig. S9). As shown in Fig. S9, mutations induced diverse intra and inter loop H-bond interactions. Furthermore, these loops were identified as flexible regions by RMSF and PCA studies, indicating that increasing flexibility by mutations render residues liable to different interaction networks. According to the RMSF study, the substitution of tyrosine with cystine at the 38th position induced high flexibility next to D39 and facilitated H-bond interactions within the CDR1 loop, especially with M41 and H43 (Fig. S9a). H-bond occupancy between D39 and H43 in Y38C variant was maintained at 12.34% during simulation, whereas it was 3.91% in the wild-type. Furthermore, D39 maintained 6.6% H-bond occupancy with M41, but this interaction was absent in the wild-type. This observation indicates that maintaining intra-loop H-bonds between D39 and M41 or H43 may facilitate short helix formation of the CDR1 loop in Y38C. In the case of the CDR2 loop, the Y38C variant showed less H-bonding between S73 and W70, L72, and S75 than in the wild-type. In addition, Y38C exhibited two new H-bonds between T66 and S73 and between S73 and L69, which were lacking in the wild-type structure (Fig. S9b).
Previous studies have shown that conformational changes of the CDR2 loop are facilitated by π-π stacking interactions between the imidazole moieties of H47 and H67, which cause H67 to swing by ~180 °26. When we investigated the inter-loop interaction between CDR1 and CDR2 we found that Y38C increased H-bonding between H47 and H67 (Fig. S9c), which indicated conformational changes in the CDR1 loop.
Similarly, the T66M variant showed significant local fluctuations in S65 and H67 residues as compared with the wild-type; H67 maintained H-bonding with L72 by >20%. Interestingly, mutation at T66 increased H-bonding capacity with R47 versus the wild-type. However, unusual H-bonding between R47 and S65 was observed after replacing M66 in T66M, which possibly contributed to conformational changes of CDR1. Nevertheless, the interaction between M66 and R47 dominated, and as a result, the degree of helix formation in the CDR1 loop was observed to decrease subsequently during the simulation (Fig. S9c).
The W50C mutation reduced intra-loop H-bonding in the CDR1 loop versus the wild-type, and K42 maintained H-bonding only with the D39 residue with an occupancy of 39.41%. Furthermore, the W50C mutation changed the H-bonding pattern in the CDR loop, where it increased the interaction between T66 and S73 and between H67 and L72. Additionally, the interaction between T66 and R47 was also greater in W50C, in which R47 also interacted with the H67 residue with an occupancy of 8.22%. On the other hand, V126G mutation reduced H-bonding between R47 and T66 and induced interaction between R47 and N68, which might facilitate conformational CDR1 loop changes. Besides, conformational changes in CDR2 loop could be induced by numerous H-bonding interactions between H67 and M41, H43, and T66.
In order to validate our H-bonding findings, we performed additional free energy landscape (FEL) analysis to select the most stable conformer from the simulation ensemble. FEL results are rendered in Fig. 8, where different colors represent structures with different energies. Lower energy conformations are colored blue and are usually more stable than other conformations (red) generated by simulation49. The thermodynamic stabilities of proteins are represented by depths of energy minima, and protein kinetic stabilities are indicated by the height of barriers between energy minima.
Free energy contour maps derived from radius of gyration and RMSD values, where the dark blue color area indicates lower energy. Each graph accompanied by a corresponding 3-D structure of lowest energy and a residue interaction network. Nodes in networks represent residues, whereas edges represent interactions. Edge colors signifying indicate interaction types, i.e., green = hydrogen bond, orange = van der Walls contact, cyan = salt bridge, blue = pi-pi stacking, and pink = disulfide bridge.
The depths of energy minima may also suggest conformational changes, where the width of an energy minimum associated with the number of conformational ensemble within the energy well50. As shown in Fig. 8, the region of conformational space corresponding to the basin changed in the wild-type and in variants, which indicated that the overall conformational stability TREM2 is affected by mutations. In order to understand the disordered state, the conformation from the favorable energy minima has been visualized. We also analyzed the intra-residue H-bonding networks using residue interaction network analysis. The lowest minima in the wild-type trajectory was found to have average Rg and RMSD values of 1.33 and 1.32 nm, respectively, which exhibit no conformational changes in the CDR1 or CDR2 loops.
In detailed structural view, T66 was found to interact with R47 by forming two H-bonds. A short helix was found in the CDR2 loop of the most stable conformer of variant trajectories. In Y38C, M41 maintained a H-bond with D39, and R47 H-bonded with H67 and L72, while in T66M, R47 maintained a π-π stacking interaction with H67. Intra and inter-loop H-bonding analysis showed a similar residual interaction in W50C, but when whole trajectory systems were considered, the interaction between D39 and H43 was maintained in the residual interaction network (Fig. S10c). On the other hand, the V126G variant showed unusual H-bonding between residues 66 and 67 and between R47 and S65. Although no conformational change in the CDR1 loop was visualized, interaction between D39 and M43 was also seen within the CDR1 loop. These observations were consistent with the residual interaction networks generated from total MD trajectories (Fig. S10).
Computational analyses in mutational landscapes, particularly MD simulations, have proven to be crucial tools in terms of deciphering the structural basis of protein aggregation and misfolding at the molecular level51. MD simulation provides a complete understanding of the phenotypic expressions of mutations by generating detailed information on conformational and structural consequences at a reasonable quality and adequate time scale52. Several recent studies have investigated the correlation between experimental studies and MDS analysis, and have suggested MDS studies are helpful for uncovering underlying mutation-associated disease mechanisms, especially in the context of neurodegeneration40,53,54,55,56,57. It has been suggested in previous MD simulation studies that R47H is responsible for loop distortions in TREM258 and to induce flexibility, particularly in CDR2 loops59. Furthermore, an X-ray crystallography study on the R47H mutation concluded that it mediated CDR2 remodeling by causing short helix formation in the loop region that ultimately abolished the ligand interaction and caused loss of ligand binding26.
Consistent with the mechanisms deemed responsible for loss of function by AD-associated variants, as described in studies mentioned above26, NHD mutations in the present study also showed significant structural alterations during molecular dynamics simulations, especially in the loop regions. It has been demonstrated mutations involving the substitution of an amino acid with different properties can disrupt molecular function by disturbing domain organization28,60. Likewise, in the present study, substituents at the 38 and 66 positions of TREM2 were found to induce effects that differed from those of native residues, and consequently to cause steric clashes between junctions of the CDR1 and CDR2 loops. Other W50C and V126G mutations had similar effects, and V126G was found to induce steric clash in CDR1. As was revealed by H-bond analysis, all of these mutations altered H-bonding patterns in CDR loops in the most native conformational states and in overall MD ensembles. Sudom et al. reported residues S65, T66, H67, N68, and H67 were critical for maintaining conformation stability of the CDR2 loop by forming a H-bonded network with residue R47 in the CDR1 loop26. Similarly, NHD mutations caused also deleterious effects similar to those caused by the AD-associated R47H mutation, which indicates this H-bonding network is important for maintaining the stabilities of CDR1 and CDR2 loops. Kober et al.61 suggested that NHD mutations might cause conformational changes of buried TREM2 residues, promote misfolding, and inconsistently impact TREM2 surface expression and aggregation. In particular, our simulation-based study showed that the Y38C, W50C, T66M, and V126G mutations cause CDR aggregations by changing secondary structural preferences, and hence, contribute to NHD-associated loss of function.
Conclusion
The effects of genetic variants on TREM2 loss of function is crucial to understanding its involvement in late-onset Alzheimer’s disease (AD). Using molecular dynamics simulation, this study presents novel findings on the deleterious roles of NHD variants, which were found to promote structural alterations in the ectodomain of TREM2. The Y38C, W50C, T66M, and V126G mutations examined all increased flexibility and altered hydrogen bonding patterns, and Y38C, T66M, and V126G induced structural remodeling in the CDR1 and CDR2 loops by inducing steric clashes. This study supports previous findings and provides additional insight of the mechanism responsible for loss of ligand binding, which is critical to our understanding of the role of TREM2 in neurodegenerative diseases.
Methods
Prediction of pathogenicity from in silico tools
The deleterious effects of selected variants were characterized using several in silico bioinformatics tools and information retrieved from the NCBI dbSNP62. Eight tools were utilized, namely, Sorting Intolerant From Tolerant (SIFT)63, Polymorphism Phenotyping v2 (PolyPhen-2)64, Functional Analysis through Hidden Markov Models (FATHMM)65, PROVEAN66, Mutpred67, I‐Mutant 3.068, Combined Annotation Dependent Depletion (CADD)69, and Condel 2.070. SIFT using sequence homology approaches to predict deleterious effects, whereas PolyPhen-2 uses the position‐specific independent counts (PSIC) scoring method and the Bayesian classifier. Pathogenicity prediction by FATHMM is typically accomplished using hidden Markov models (HMMs), and I-Mutant 3.0 uses a support vector machine algorithm to calculate the stabilities of variant protein structure based on free energy changes. Mutpred predicts gain and loss of 14 different structural and functional properties of proteins caused by mutations. CADD considers protein level scores derived by SIFT and Polyphen analysis and a broad range of data, including functional genomics data, to predict the pathogenicities of nonsynonymous mutations. On the other hand, Condel uses a consensus approach to predict SNP impact by integrating Log R Pfam E-value (LogRE)71, MAPP[8,19,72, Mutation Assessor18,73, Polyphen2 (PPH2)64, and SIFT63.
Preparation of the simulation system
The three-dimensional crystal structure of the ectodomain of TREM2 (PDB id of 5ELI61) was retrieved from the protein databank (http://www.rcsb.org/pdb)74. The structure of wild-type TREM2 was initially prepared by adding bond orders, hydrogens, and charges and refined by removing water molecules and optimizing it at neutral pH. The structure was further fixed by correcting for asparagine amide groups, some thiol and hydroxyl groups, protonation states of glutamic acids, aspartic acids, and histidines. In order to adjust heavy atom Root Mean Square Deviation (RMSD), minimization was applied by using Optimized Potentials for Liquid Simulation (OLPS3) force field down to 0.30 Å. Y38C, W50C, T66M, and V126G variant structures were constructed by computational mutagenesis using the Mutate Residues script from Schrödinger suite 2017-1 (LLC, New York, NY, USA)75.
Additional short molecular dynamics refinement simulation was performed to better resolve the structures (wild-type TREM2 and those of the Y38C, W50C, T66M, and V126G variants). Thus, molecular dynamics simulation was conducted by applying YAMBER3 force field76 for 500 ps at pH 7.4 and 298 K at a solvent density of 0.997 g/cc. The simulation was run using YASARA software by a default md_refine macro, and the final structures were selected based on the lowest free energy energies.
Molecular dynamics simulation
Molecular dynamics simulation was used to study the changes in the dynamic behaviors of protein caused by mutations using the Desmond module of Schrödinger suite 2017-1 (LLC, New York, NY, USA)77,78. Here, OLPS3 all-atom force field was utilized to visualize molecular behavior79,80,81. Structures were prepared for wild-type TREM2 and the Y38C, W50C, T66M, and V126G variants solvated in the presence of explicit solvent in a triclinic periodic boundary box. Each system was submerged to a Monte-Carlo equilibrated TIP3P solvation model extending to ~10 Å in each direction, which is known to provide best experimental outputs82. Additional counter ions were added to the water model to neutralization the system. Physiological condition and ionic strength of the solvent system were maintained by maintaining a default salt (NaCl) concentration of 0.15 M in the simulation box. The default relaxation protocol83,84 was used to relax the 8-stage system. Followed by Brownian dynamics, the second stage simulation was started at 10 K in an NVT ensemble for 12 ps with restraints on solute heavy atoms. Third stage simulation was done for 12 ps in the same ensemble at 10 K with restraints on solute heavy atoms. The fourth stage was begun by running 12 ps simulation under the NPT ensemble at 1 bar pressure while maintaining solute heavy atom restraints. In the fifth stage, protein cavity was solvated using the solvate pocket script. Stage 6 involved 12 ps simulation in an NPT ensemble with restraints on solute heavy atoms, and stage 7 involved a 24 ps simulation with no restraints on solute heavy atoms (both stages were performed at 300 K at 1 bar. Finally, a 100 ns MD simulation was performed for each system with no constraints applied, and resulting trajectories were subjected to further analysis. Throughout the simulation, RESPA85 integrator (a motion integration package) was employed with 2 fs as inner time step with the M-SHAKE86 algorithm to constrain all covalent bonds connecting hydrogen atoms. The Particle Mesh Ewald method was used to calculate long-range electrostatic interactions and 9.0 Å was used for short-range electrostatic contacts; uniform density approximation was select for the cutoff of long-range van der Waals (VDW) interactions. Conditions during simulations were maintained by Nose–Hoover thermostats87 at 300 K and 1 atmosphere using the Martyna–Tobias–Klein method88.
Resultant trajectories were used to evaluate protein conformational changes and stabilities using Root Mean Square Deviation (RMSD), Root Mean Square Fluctuation (RMSF), and SSE (Secondary Structure Elements) in the Simulation Interactions Diagram panel of Schrödinger 2017-1 (LLC, New York, NY, USA). Radius of gyration (Rg), Solvent Accessible Surface Areas (SASA), and H-bond occupancies were calculated using VMD software89. Dynamic cross-correlation maps were generated from trajectories to explain time-correlated protein motions, which were assembled using Bio3D90 software by R programming. Time-correlated information between protein atoms i and j (cij) was represented as a matrix in DCCM, which was obtained using the following expression:
Displacements between current and average positions of atom i and j are represented by d, and mean times overall trajectories are represented by the angled brackets. Calculated values in DCCM ranged between −1 and +1, which denoted negative and positive correlations, respectively. Principle Component Analysis (PCA) was further utilized to describe the collective motions of TREM2 variants44. PCA eigenvectors were calculated by superimposing atomic coordinates on reference structures without translational or rotational movements. Eigenvectors represented mean square displacements (MSD) of atoms and were associated with eigenvalues. Mathematical details have been previously described in detail91,92.
Free energy landscape (FEL)
The free energy landscape technique is used to map all possible conformational changes of macromolecules using energy levels derived from the spatial dispositions of interacting molecules93,94. For FEL analysis, Gibb’s free energy is calculated as a function of protein enthalpy and entropy and also identifies conformational states related to protein structure-function correlations. In the present study, the energy bases of conformational diversity in various TREM2 structures were investigated using the following equation:
where kB is Boltzmann’s constant, T is the temperature (which was set at 300 K), Ni is the population in bin i and Nmax is the population in the most populated bin. An artificial barrier scale is set to the bin with no population with the lowest provability. Color-code modes were used to display different energy levels.
Residue interaction network
To analyze the Residue Interaction Network (RIN), the most stable 3-D coordinates of the wild-type and of variants were transferred to the RING server95, which provides intra-residue interactions in the exhaustive network view. In the network model, protein residues are represented by nodes and interaction modes as edges. Results from the RING server were then processed using Cytoscape to construct interactive RIN 3.2.196 using the plug-in RINalyzer. Types of interactions are described by dashed or dotted edges, and salt bridge, H-bonding, and van der Waal interactions. In addition, residual interaction networks were constructed by considering the whole trajectories of 100 ns molecular dynamics simulations. For that 1000 representative structures with an interval of 100 ps from each simulation system were visualized and analyzed using a combination of UCSF Chimera97, structureViz98, and Cytoscape99.
References
Selkoe, D. J. & Hardy, J. The amyloid hypothesis of Alzheimer’s disease at 25 years. EMBO molecular medicine 8, 595–608 (2016).
Jack, C. R. Jr. et al. Tracking pathophysiological processes in Alzheimer’s disease: an updated hypothetical model of dynamic biomarkers. The Lancet Neurology 12, 207–216 (2013).
Guerreiro, R. et al. TREM2 variants in Alzheimer’s disease. New England Journal of Medicine 368, 117–127 (2013).
Jonsson, T. et al. Variant of TREM2 associated with the risk of Alzheimer’s disease. New England Journal of Medicine 368, 107–116 (2013).
Atagi, Y. et al. Apolipoprotein E is a ligand for triggering receptor expressed on myeloid cells 2 (TREM2). Journal of Biological Chemistry 290, 26043–26050 (2015).
Bailey, C. C., DeVaux, L. B. & Farzan, M. The triggering receptor expressed on myeloid cells 2 binds apolipoprotein E. Journal of Biological Chemistry, jbc. M115. 677286 (2015).
Yeh, F. L., Wang, Y., Tom, I., Gonzalez, L. C. & Sheng, M. TREM2 Binds to Apolipoproteins, Including APOE and CLU/APOJ, and Thereby Facilitates Uptake of Amyloid-Beta by Microglia. Neuron 91, 328–340, https://doi.org/10.1016/j.neuron.2016.06.015 (2016).
Daws, M. R. et al. Pattern recognition by TREM-2: binding of anionic ligands. The Journal of Immunology 171, 594–599 (2003).
Cannon, J. P., O’Driscoll, M. & Litman, G. W. Specific lipid recognition is a general feature of CD300 and TREM molecules. Immunogenetics 64, 39–47 (2012).
Poliani, P. L. et al. TREM2 sustains microglial expansion during aging and response to demyelination. The Journal of clinical investigation 125, 2161–2170 (2015).
Peng, Q. et al. TREM2-and DAP12-dependent activation of PI3K requires DAP10 and is inhibited by SHIP1. Science signaling 3, ra38–ra38 (2010).
Zhao, Y. et al. TREM2 Is a Receptor for beta-Amyloid that Mediates Microglial Function. Neuron 97, 1023–1031.e1027, https://doi.org/10.1016/j.neuron.2018.01.031 (2018).
Cheng, J. et al. TREMs in Alzheimer’s disease: Genetic and clinical investigations. Clinica Chimica Acta 463, 88–95 (2016).
Jiang, T. et al. Upregulation of TREM2 ameliorates neuropathology and rescues spatial cognitive impairment in a transgenic mouse model of Alzheimer’s disease. Neuropsychopharmacology 39, 2949 (2014).
Jiang, T. et al. TREM2 overexpression has no improvement on neuropathology and cognitive impairment in aging APPswe/PS1dE9 mice. Molecular neurobiology 54, 855–865 (2017).
Wang, Y. et al. TREM2 lipid sensing sustains the microglial response in an Alzheimer’s disease model. Cell 160, 1061–1071 (2015).
Wang, Y. et al. TREM2-mediated early microglial response limits diffusion and toxicity of amyloid plaques. Journal of Experimental Medicine, jem. 20151948 (2016).
Yuan, P. et al. TREM2 haplodeficiency in mice and humans impairs the microglia barrier function leading to decreased amyloid compaction and severe axonal dystrophy. Neuron 90, 724–739 (2016).
Keren-Shaul, H. et al. A unique microglia type associated with restricting development of Alzheimer’s disease. Cell 169, 1276–1290. e1217 (2017).
Thelen, M. et al. Investigation of the role of rare TREM2 variants in frontotemporal dementia subtypes. Neurobiology of Aging 35, 2657.e2613–2657.e2619, https://doi.org/10.1016/j.neurobiolaging.2014.06.018 (2014).
Jin, S. C. et al. Coding variants in TREM2 increase risk for Alzheimer’s disease. Human molecular genetics 23, 5838–5846 (2014).
Guerreiro, R. et al. A novel compound heterozygous mutation in TREM2 found in a Turkish frontotemporal dementia-like family. Neurobiology of aging 34, 2890. e2891–2890. e2895 (2013).
Guerreiro, R. J. et al. Using exome sequencing to reveal mutations in TREM2 presenting as a frontotemporal dementia–like syndrome without bone involvement. JAMA neurology 70, 78–84 (2013).
Le Ber, I. et al. Homozygous TREM2 mutation in a family with atypical frontotemporal dementia. Neurobiology of aging 35, 2419. e2423–2419. e2425 (2014).
Dardiotis, E. et al. A novel mutation in TREM2 gene causing Nasu-Hakola disease and review of the literature. Neurobiology of Aging 53, 194.e113–194.e122, https://doi.org/10.1016/j.neurobiolaging.2017.01.015 (2017).
Sudom, A. et al. Molecular basis for the loss-of-function effects of the Alzheimer’s disease–associated R47H variant of the immune receptor TREM2. Journal of Biological Chemistry 293, 12634–12646 (2018).
Elber, R. Perspective: Computer simulations of long time dynamics. The Journal of chemical physics 144, 060901 (2016).
Hosen, S. Z., Dash, R., Junaid, M., Mitra, S. & Absar, N. Identification and structural characterization of deleterious non-synonymous single nucleotide polymorphisms in the human SKP2 gene. Computational biology and chemistry 79, 127–136 (2019).
Schreiner, W., Karch, R., Knapp, B. & Ilieva, N. Relaxation estimation of RMSD in molecular dynamics immunosimulations. Computational and Mathematical Methods in Medicine 2012 (2012).
Priya Doss, C. G., Chakraborty, C., Chen, L. & Zhu, H. Integrating in silico prediction methods, molecular docking, and molecular dynamics simulation to predict the impact of ALK missense mutations in structural perspective. BioMed research international 2014 (2014).
Tanwar, H. et al. Chapter Five-A Computational Approach to Identify the Biophysical and Structural Aspects of Methylenetetrahydrofolate Reductase (MTHFR) Mutations (A222V, E429A, and R594Q) Leading to Schizophrenia. Advances in Protein Chemistry and Structural Biology 108, 105–125 (2017).
Eisenberg, D. & McLachlan, A. D. Solvation energy in protein folding and binding. Nature 319, 199 (1986).
Anwar, M. A. & Choi, S. Structure-activity relationship in TLR4 mutations: atomistic molecular dynamics simulations and residue interaction network analysis. Scientific reports 7, 43807 (2017).
Eaton, W. A., Henry, E. R. & Hofrichter, J. Application of linear free energy relations to protein conformational changes: the quaternary structural change of hemoglobin. Proceedings of the National Academy of Sciences 88, 4472–4475 (1991).
Ikeguchi, M., Ueno, J., Sato, M. & Kidera, A. Protein structural change upon ligand binding: linear response theory. Physical review letters 94, 078102 (2005).
Levy, Y. & Onuchic, J. N. Mechanisms of protein assembly: lessons from minimalist models. Accounts of chemical research 39, 135–142 (2006).
Lee, Y. et al. Dissecting the critical factors for thermodynamic stability of modular proteins using molecular modeling approach. Plos One 9, e98243 (2014).
Benson, N. C. & Daggett, V. A comparison of multiscale methods for the analysis of molecular dynamics simulations. The Journal of Physical Chemistry B 116, 8722–8731 (2012).
Agrahari, A. & Doss, C. G. P. Impact of I30T and I30M substitution in MPZ gene associated with Dejerine–Sottas syndrome type B (DSSB): A molecular modeling and dynamics. Journal of theoretical biology 382, 23–33 (2015).
Padhi, A. K., Jayaram, B. & Gomes, J. Prediction of functional loss of human angiogenin mutants associated with ALS by molecular dynamics simulations. Scientific reports 3, 1225 (2013).
Chan, P., Curtis, R. A. & Warwicker, J. Soluble expression of proteins correlates with a lack of positively-charged surface. Scientific reports 3, 3333 (2013).
Barz, B., Turner, B. S., Bansil, R. & Urbanc, B. Folding of pig gastric mucin non-glycosylated domains: a discrete molecular dynamics study. Journal of biological physics 38, 681–703 (2012).
Gaieb, Z. & Morikis, D. Detection of Side Chain Rearrangements Mediating the Motions of Transmembrane Helices in Molecular Dynamics Simulations of G Protein-Coupled Receptors. Computational and structural biotechnology journal 15, 131–137 (2017).
Ichiye, T. & Karplus, M. Collective motions in proteins: a covariance analysis of atomic fluctuations in molecular dynamics and normal mode simulations. Proteins: Structure, Function, and Bioinformatics 11, 205–217 (1991).
Amadei, A., Linssen, A. B. & Berendsen, H. J. Essential dynamics of proteins. Proteins: Structure, Function, and Bioinformatics 17, 412–425 (1993).
Agrahari, A. K., George, P. D. C., Siva, R., Magesh, R. & Zayed, H. Molecular insights of the G2019S substitution in LRRK2 kinase domain associated with Parkinson’s disease: A molecular dynamics simulation approach. Journal of theoretical biology (2019).
Kabsch, W. & Sander, C. Dictionary of protein secondary structure: pattern recognition of hydrogen−bonded and geometrical features. Biopolymers 22, 2577–2637 (1983).
Bai, Y. & Englander, S. W. Hydrogen bond strength and beta-sheet propensities: the role of a side chain blocking effect. Proteins 18, 262–266, https://doi.org/10.1002/prot.340180307 (1994).
Guan, S. et al. Exploration of binding and inhibition mechanism of a small molecule inhibitor of influenza virus H1N1 hemagglutinin by molecular dynamics simulation. Scientific reports 7, 3786 (2017).
Gershenson, A., Gierasch, L. M., Pastore, A. & Radford, S. E. Energy landscapes of functional proteins are inherently risky. Nature chemical biology 10, 884 (2014).
Redler, R. L. et al. Computational approaches to understanding protein aggregation in neurodegeneration. J. Mol. Cell. Biol. 6, 104–115, https://doi.org/10.1093/jmcb/mju007 (2014).
Hollingsworth, S. A. & Dror, R. O. Molecular Dynamics Simulation for All. Neuron 99, 1129–1143, https://doi.org/10.1016/j.neuron.2018.08.011 (2018).
Kumar Ghosh, D., Nanaji Shrikondawar, A. & Ranjan, A. Local structural unfolding at the edge-strands of beta sheets is the molecular basis for instability and aggregation of G85R and G93A mutants of superoxide dismutase 1. Journal of Biomolecular Structure and Dynamics, 1–13 (2019).
Kumar, V., Pandey, P., Idrees, D., Prakash, A. & Lynn, A. M. Delineating the effect of mutations on the conformational dynamics of N-terminal domain of TDP-43. Biophysical chemistry 250, 106174, https://doi.org/10.1016/j.bpc.2019.106174 (2019).
Padhi, A. K. & Hazra, S. Insights into the role of d-amino acid oxidase mutations in amyotrophic lateral sclerosis. Journal of cellular biochemistry 120, 2180–2197 (2019).
Padhi, A. K., Kumar, H., Vasaikar, S. V., Jayaram, B. & Gomes, J. Mechanisms of loss of functions of human angiogenin variants implicated in amyotrophic lateral sclerosis. Plos One 7, e32479, https://doi.org/10.1371/journal.pone.0032479 (2012).
Padhi, A. K., Narain, P. & Gomes, J. Rare Angiogenin and Ribonuclease 4 variants associated with amyotrophic lateral sclerosis exhibit loss-of-function: a comprehensive in silico study. Metabolic brain disease, 1–17 (2019).
Abduljaleel, Z. et al. Evidence of trem2 variant associated with triple risk of Alzheimer’s disease. Plos One 9, e92648 (2014).
Menzies, G. E., Sims, R. & Williams, J. Molecular Dynamics simulations of Alzheimer’s variants, R47H and R62H, in TREM2 provide evidence for structural alterations behind functional changes. BioRxiv, 536540 (2019).
Arifuzzaman, M. et al. In silico analysis of nonsynonymous single-nucleotide polymorphisms (nsSNPs) of the SMPX gene. Annals of human genetics (2019).
Kober, D. L. et al. Neurodegenerative disease mutations in TREM2 reveal a functional surface and distinct loss-of-function mechanisms. eLife 5, e20391, https://doi.org/10.7554/eLife.20391 (2016).
Sherry, S. T. et al. dbSNP: the NCBI database of genetic variation. Nucleic Acids Res. 29, 308–311, https://doi.org/10.1093/nar/29.1.308 (2001).
Ng, P. C. & Henikoff, S. J. N. A. R. SIFT: Predicting amino acid changes that affect protein function. 31, 3812–3814 (2003).
Adzhubei, I. A. et al. A method and server for predicting damaging missense mutations. 7, 248 (2010).
Shihab, H. A. et al. Predicting the functional, molecular, and phenotypic consequences of amino acid substitutions using hidden Markov models. 34, 57–65 (2013).
Choi, Y., Sims, G. E., Murphy, S., Miller, J. R. & Chan, A. P. J. P. O. Predicting the functional effect of amino acid substitutions and indels. 7, e46688 (2012).
Pejaver, V. et al. MutPred2: inferring the molecular and phenotypic impact of amino acid variants. 134981 (2017).
Capriotti, E., Fariselli, P. & Casadio, R. I-Mutant2. 0: predicting stability changes upon mutation from the protein sequence or structure. 33, W306–W310 (2005).
Kircher, M. et al. A general framework for estimating the relative pathogenicity of human genetic variants. 46, 310 (2014).
González-Pérez, A. & López-Bigas, N. J. T. A. J. O. H. G. Improving the assessment of the outcome of nonsynonymous SNVs with a consensus deleteriousness score. Condel. 88, 440–449 (2011).
Clifford, R. J., Edmonson, M. N., Nguyen, C. & Buetow, K. H. Large-scale analysis of non-synonymous coding region single nucleotide polymorphisms. Bioinformatics 20, 1006–1014, https://doi.org/10.1093/bioinformatics/bth029 (2004).
Binkley, J. et al. ProPhylER: a curated online resource for protein function and structure based on evolutionary constraint analyses. Genome Res 20, 142–154, https://doi.org/10.1101/gr.097121.109 (2010).
Reva, B., Antipin, Y. & Sander, C. Determinants of protein function revealed by combinatorial entropy optimization. Genome Biol 8, R232–R232, https://doi.org/10.1186/gb-2007-8-11-r232 (2007).
Zardecki, C. et al. (ACS Publications, 2016).
Raju, D. et al. Molecular Insight and Binding Pattern Analysis of Shikonin as a Potential VEGFR-2 Inhibitor. Current Enzyme Inhibition 13, 1–10, https://doi.org/10.2174/1573408013666161227162452 (2017).
Krieger, E., Darden, T., Nabuurs, S. B., Finkelstein, A. & Vriend, G. Making optimal use of empirical energy functions: force‐field parameterization in crystal space. Proteins: Structure, Function, and Bioinformatics 57, 678–683 (2004).
Tools, M. (Version, 2011).
Fazil, M. H. U. T. et al. Comparative structural analysis of two proteins belonging to quorum sensing system in Vibrio cholerae. Journal of Biomolecular Structure and Dynamics 30, 574–584 (2012).
Jorgensen, W. L., Chandrasekhar, J., Madura, J. D., Impey, R. W. & Klein, M. L. Comparison of simple potential functions for simulating liquid water. The Journal of chemical physics 79, 926–935 (1983).
Bowers, K. J. et al. In Proceedings of the 2006 ACM/IEEE conference on Supercomputing. 84 (ACM).
Shaw, D. E. A fast, scalable method for the parallel evaluation of distance‐limited pairwise particle interactions. Journal of computational chemistry 26, 1318–1328 (2005).
Shaik, M. S., Liem, S. Y. & Popelier, P. L. Properties of liquid water from a systematic refinement of a high-rank multipolar electrostatic potential. The Journal of chemical physics 132, 174504 (2010).
Mitra, S. & Dash, R. Structural dynamics and quantum mechanical aspects of shikonin derivatives as CREBBP bromodomain inhibitors. Journal of Molecular Graphics and Modelling 83, 42–52 (2018).
Arifuzzaman, M. et al. A Computational workflow for the identification of the potent inhibitor of type II secretion system traffic ATPase of Pseudomonas aeruginosa. Computational biology and chemistry 76, 191–201 (2018).
Humphreys, D. D., Friesner, R. A. & Berne, B. J. A multiple-time-step molecular dynamics algorithm for macromolecules. The Journal of Physical Chemistry 98, 6885–6892 (1994).
Kräutler, V., Van Gunsteren, W. F. & Hünenberger, P. H. A fast SHAKE algorithm to solve distance constraint equations for small molecules in molecular dynamics simulations. Journal of computational chemistry 22, 501–508 (2001).
Hoover, W. G. Canonical dynamics: Equilibrium phase-space distributions. Physical review A 31, 1695 (1985).
Martyna, G. J., Tobias, D. J. & Klein, M. L. Constant pressure molecular dynamics algorithms. The Journal of Chemical Physics 101, 4177–4189 (1994).
Humphrey, W., Dalke, A. & Schulten, K. VMD: visual molecular dynamics. Journal of molecular graphics 14, 33–38 (1996).
Grant, B. J., Rodrigues, A. P., ElSawy, K. M., McCammon, J. A. & Caves, L. S. J. B. Bio3d: an R package for the comparative analysis of protein structures. 22, 2695–2696 (2006).
Shlens, J. A tutorial on principal component analysis. arXiv preprint arXiv:1404.1100 (2014).
Salmas, R. E., Yurtsever, M. & Durdagi, S. Investigation of inhibition mechanism of chemokine receptor CCR5 by micro-second molecular dynamics simulations. Scientific reports 5 (2015).
Frauenfelder, H., Sligar, S. G. & Wolynes, P. G. The energy landscapes and motions of proteins. Science 254, 1598–1603 (1991).
Lange, O. F. & Grubmüller, H. Generalized correlation for biomolecular dynamics. Proteins: Structure, Function, and Bioinformatics 62, 1053–1061 (2006).
Martin, A. J. et al. RING: networking interacting residues, evolutionary information and energetics in protein structures. Bioinformatics 27, 2003–2005 (2011).
Baryshnikova, A. Exploratory analysis of biological networks through visualization, clustering, and functional annotation in cytoscape. Cold Spring Harbor Protocols 2016, pdb. prot077644 (2016).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. Journal of computational chemistry 25, 1605–1612 (2004).
Morris, J. H., Huang, C. C., Babbitt, P. C. & Ferrin, T. E. structureViz: linking Cytoscape and UCSF Chimera. Bioinformatics 23, 2345–2347 (2007).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Acknowledgements
The study was supported by the Basic Science Research Program through the Korean National Research Foundation (#NRF-2018R1A2B6002232 to ISM).
Author information
Authors and Affiliations
Contributions
I.S.M. conceived the study and designed the experimental work. R.D. and H.J.C. conducted the experiments. I.S.M., R.D. and H.J.C. analyzed the data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Dash, R., Choi, H.J. & Moon, I.S. Mechanistic insights into the deleterious roles of Nasu-Hakola disease associated TREM2 variants. Sci Rep 10, 3663 (2020). https://doi.org/10.1038/s41598-020-60561-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-60561-x
This article is cited by
-
Computational saturation mutagenesis to explore the effect of pathogenic mutations on extra-cellular domains of TREM2 associated with Alzheimer’s and Nasu-Hakola disease
Journal of Molecular Modeling (2023)
-
Defects in lysosomal function and lipid metabolism in human microglia harboring a TREM2 loss of function mutation
Acta Neuropathologica (2023)
-
Dynamic insights into the effects of nonsynonymous polymorphisms (nsSNPs) on loss of TREM2 function
Scientific Reports (2022)
-
In silico chemical profiling and identification of neuromodulators from Curcuma amada targeting acetylcholinesterase
Network Modeling Analysis in Health Informatics and Bioinformatics (2021)
-
N-acetyl-D-glucosamine kinase binds dynein light chain roadblock 1 and promotes protein aggregate clearance
Cell Death & Disease (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.