Abstract
Kidney mesenchyme (KM) and nephron progenitors (NPs) depend on WNT activity, and their culture in vitro requires extensive repertoire of recombinant proteins and chemicals. Here we established a robust, simple culture of mouse KM using a combination of 3D Matrigel and growth media supplemented with Fibroblast Growth Factor 2 (FGF2) and Src inhibitor PP2. This allows dissociated KM to spontaneously self-organize into spheres. To reassess the requirement of WNT activity in KM self-organization and NPs maintenance, cells were cultured with short pulse of high-dose GSK3β inhibitor BIO, on a constant low-dose or without BIO. Robust proliferation at 48 hours and differentiation at 1 week were observed in cultures with high BIO pulse. Importantly, dissociated KM cultured without BIO, similarly to that exposed to constant low dose of BIO, maintained NPs up to one week and spontaneously differentiated into nephron tubules at 3 weeks of culture. Our results show that KM is maintained and induced to differentiate in a simple culture system. They also imply that GSK3β/WNT-independent pathways contribute to the maintenance and induction of mouse KM. The robust and easy 3D culture enables further characterization of NPs, and may facilitate disease modeling when applied to human cells.
Similar content being viewed by others
Introduction
The mammalian kidney develops from the intermediate mesoderm through reciprocal interactions of the ureteric bud (UB) and kidney mesenchyme (KM) known also as metanephric mesenchyme1. Nephrons, the functional units of the kidney that filter the waste products from the blood, form by orchestrated differentiation of the nephron progenitor cells (NPs) located in close proximity to UB tips. Mesenchymal NPs are first induced to aggregate and form renal vesicle, which then fully epithelializes via pretubular aggregate, comma- and S-shaped bodies1. It has been demonstrated that WNT pathway is a key paracrine signaling for maintenance of nephron progenitors and their timely differentiation into nephron epithelium2,3. Wnt11 expressed by UB tips facilitates UB branching via positively reinforcing glial cell line derived growth factor (GDNF)/rearranged in transformation (RET) signaling, and participates in NPs maintenance3,4. Wnt9b, expressed by the UB trunk and to lesser extent in tips, maintains and induces SIX2-expressing NPs for differentiation5,6,7. In the differentiating nephron structures Wnt4 acting cell-autonomously is needed for completion of mesenchyme-to- epithelium transition (MET)8. Both canonical pathway involving Glycogen Synthase Kinase 3 Beta (GSK3β) inhibition and leading to β-catenin stabilization, as well as non-canonical pathway mediate WNT functions in self-renewal and nephrogenesis7,9.
Organotypic cultures, such as the transfilter system and explants of the whole kidney anlagen, have profoundly deepened our knowledge of the developmental mechanisms of kidney development10. However, it has been shown that when the KM is cultured in isolation, it stops proliferating, does not differentiate and decays in a few days10,11. First attempts towards the long-term KM in vitro culture were undertaken by dissociating and reaggregating whole embryonic kidneys where UB cells were present12. A variety of factors required for the maintenance, proliferation, and differentiation of the KM has been discovered13,14,15,16. Recently, the maintenance and propagation of purified nephrogenic progenitor cells was achieved17,18. Although efficient in maintaining the purified SIX2+ NPs, these cultures exclude stromal niche cells. In addition, the maintenance protocols are tedious, depend on complex procedures and require multiple synthetic agents and growth factors.
The majority of renal cell cultures rely on 2D monolayers19,20,21,22 that do not accurately model the 3D architecture of the tissue. The 3D architecture and cell-cell contacts are essential for propagation SIX2+ NPs in vitro both in isolation and when pluripotent stem cells are differentiated into kidney organoids17,23.
Until recently, the induction of an isolated kidney mesenchyme for differentiation was based on its recombination with the UB or heterologous inducing tissues such as embryonic spinal cord21,24,25. The GSK3β inhibitor 6-bromoindirubin-3-oxime (BIO) has been shown to induce differentiation of isolated rat and mouse mesenchyme via the canonical WNT signaling pathway9. Chemical induction with GSK3β inhibitors has been utilized not only for kidney explants but also for generating organoids from hiPSC that show nephron marker patterns typical for that of nephrogenesis in vivo26.
Here we describe a robust, simple culture of mouse KM using a combination of 3D Matrigel and growth media supplemented with FGF2 and PP2. We show that this method facilitates dissociated KM to self-organize into spheres where also renal stromal cells are present. We further demonstrate that NPs and stromal niche propagate for a week, and thereafter NPs undergo spontaneous MET to form nephron tubules without additional WNT activation. Our results imply that GSK3β/WNT-independent pathways complement regulation of self-renewal and differentiation decisions in mouse KM.
Results
Self-organization and formation of metanephric niche
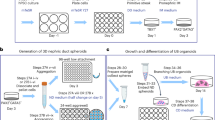
To characterize NPs self-organization in vitro, KM from E11.5 mouse embryos were microsurgically dissected from the UB and dissociated into single cells (Fig. 1A–C). Previous studies have shown that FGF family proteins are essential for the KM maintenance27,28,29. Therefore, basic culture medium (BM) was supplemented with 50 ng/ml of FGF2. We then plated 30,000 dissociated KM cells on top of 3D Matrigel per well of 96-plate in 200 µl either BM or BM + FGF2 (Fig. 1D, E respectively). Time-lapse imaging revealed that single cell suspensions self-organized into spheroids within 17 hours of culture (Supplemental video 1). To reassess the role of WNT in the self-organization and propagation of the NPs, the dissociated KM cell culture was pulsed with 10 µM of BIO for 1 hour (BIO pulse), thereafter the medium was replaced with fresh BM + FGF2 without BIO (Fig. 1F, Supplemental video 2). Bright field imaging after 72 hours of culture showed that dissociated KM cultured in BM-only did not form spheres and decayed with time (Fig. 2A), whereas in the BM + FGF2 and BM + FGF2 + BIO conditions dissociated KM self-organized into cell clusters comparably (Fig. 2B,C).
Schematic representation of the 3D culture method of dissociated kidney mesenchyme. (A) E11.5 embryonic mouse kidneys are isolated and subjected to collagenase IV treatment after which the KM is separated from the epithelial ureteric bud (UB) by manual microdissection. (B) Dissociation of KM into single cells assisted by TrypLE enzymatic digestion, resulting in single cell suspension (dissociated KM). (C) After pelleting the cells with centrifugation, the dissociated KM is resuspended into Basic Media (BM). Dissociated KM cells are then plated on top of the 3D Matrigel submerged in either (D) BM, (E) BM + 50 ng/ml FGF2 or (F) BM + 50 ng/ml FGF2 and pulsed with 10 µM BIO for 1 hour. In some experiments BIO pulse is replaced by constant BIO application at 50 nM concentration.
Self-organization, survival and maintenance of dissociated kidney mesenchyme in 3D culture. (A–C) Representative bright field images of dissociated KM after 72 hours of culture (n = 4). (A) Dissociated KM spheres cultured in basic media (BM) do not form spheres and decay in three days (B) Dissociated KM cultured in BM + FGF2 self-organizes into spheres. (C) Dissociated KM cultured in BM + FGF2 with pulse of 10 µM BIO 1 hour. FGF2 promotes dissociated KM cells to form aggregates resembling pretubular aggregates of an intact kidney. Further culture facilitates aggregates to form spheres and enhances dissociated KM maintenance. Dissociated KM cultured in FGF2 + 10 µM BIO 1 hour pulse self-organized to large, dense spheres. (D–F) Flow cytometric graphs of cell survival analyzed by cytotoxicity Sytox assay after 72 hours of culture. (D) Dead cell fraction (gray) dominates in dissociated KM cultured with BM. (E) Viable cell fraction increases drastically in dissociated KM grown with addition of FGF2. (F) BM + FGF2 + 10 µM BIO 1 hour pulse additionally boosts cell survival in dissociated KM spheres. (G) Quantification of flow cytometry data. Sytox assay’s values are inverted to present viability in Y axis. Dissociated KM survival increase from 20.1% ± 8.8% in BM culture to 57.4% ± 20.9% when cultured in the presence of FGF2, and reached 79.8% ± 14.4% in dissociated KM cultured in BM + FGF2 + BIO pulse. Asterisks indicate statistically significant difference (p < 0.05, paired sample t-test, n = 4, error bars - standard deviation). (H) Representative bright field images of dissociated KM cultures in BM + FGF2 (upper panel) and in BM + FGF2 + 50 nM constant BIO concentration. Dissociated KM cells form spheres and survive up to 3 weeks without major morphological differences between these two conditions. Scale bars in bright field images = 100 µm.
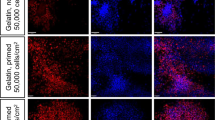
To better understand the composition and self-organization of KM spheres we examined the cell identities by immunofluorescence at 72 hours of culture and compared these to E11.5 whole kidney reaggregation cultured in classical Trowell-type culture12. Cross-sections of the KM spheres were immunostained with different cell type markers to identify whether both stromal and nephrogenic lineages are present. This revealed that the spheres were composed of MEIS1/2+ stroma, SIX2+ and PAX2 + NPs progenitors (Fig. 3A–F). The BM + FGF2 cultured KM spheres comprised of MEIS1/2 labeled cells (Fig. 3A, C) neighboring SIX2+ progenitors (Fig. 3B,C), whereas in spheres cultured in BM + FGF2 + BIO pulse PAX2+ NPs formed pretubular aggregate-like condensates, which were surrounded by MEIS1/2 positive stroma cells. Quantification of immunofluorescence images demonstrated, that KM spheres cultured in BM + FGF2 for 72 hours contained 48.56% ± 6.34% of SIX2 + cells, whereas BIO pulse significantly reduced the SIX2 + progenitors (4.31% ± 2.13%, p < 0.05, n = 6, Supplemental Fig. 1). Analysis of niche self-organization in the whole kidney dissociation and reaggregation culture revealed that E11.5 reaggregated kidneys were decreased in size at 72 hours (Supplemental Fig. 2) and were negative for NP marker SIX2 and nephron epithelial markers CDH1 and lotus tetragonolobus lectin (LTL) (Supplemental Fig. 3E–H). These immunostainings revealed that in our dissociated KM culture method, stroma and NPs spontaneously self-organize into stroma-NP niche with a capacity to recapitulate structures such as pretubular aggregates.
Dissociated kidney mesenchyme self-organizes into nephron progenitor-stromal niche, with the capacity to form pretubular aggregates. Representative immunofluorescence image of dissociated KM cultured for 72 hours in BM + FGF2 (A–C) and BM + FGF2 + BIO pulse (D–F). (A) Stromal lineage marker MEIS1/2 strongly stains most of the KM spheres cells. (B) Immunostaining for nephron progenitor marker SIX2 shows abundant NPs in KM spheres. (C) Merged image of separate channels for MEIS1/2 and SIX2 co-immunostaining, counterstained with nuclear stain DAPI shows spontaneous self-organization into spheres of both stroma and NPs resembling the niche of intact kidney. (D) MEIS1/2 immunostaining of BM + FGF2 + BIO pulse cultured spheres labels stromal cells (E) PAX2+ stains pretubular aggregate-resembling cell clusters. (F) Merged image of separate channels for MEIS1/2 and PAX2 co-immunostaining, counterstained with nuclear stain DAPI reveals that PAX2+ pretubular aggregates are wrapped inside of MEIS1/2+ stromal cells. Scale bars = 50 µm, F scale bar = 10 µm. Dotted line (D–F) marks pretubular aggregate border with stroma cells, n = 3.
FGF2 promotes maintenance of kidney mesenchyme spheres while BIO induces their expansion
To assess the survival of KM spheres in different conditions we re-dissociated and incubated the spheres with Sytox, a live cell impermeant agent allowing analysis of live cell proportions by flow cytometry (Fig. 2D–G). This revealed, in accordance with previous findings, that dissociated KM cultured in BM + FGF2 showed almost three times higher cell survival than those cultured in BM only (57.4% ± 20.9% and 20.1% ± 8.8% respectively, n = 4, Fig. 2D, E). A four-fold increase in cell survival was observed when dissociated KM cells were cultured with BM + FGF2 + BIO pulse (79.8% ± 14.4%, n = 4, Fig. 2B).
We then characterized proliferation in KM spheres for up to three weeks by EdU incorporation assay and Click-it fluorescent labeling of cells in S-phase of cell cycle (Supplemental Fig. 4). At 48 hours of culture, proliferating cell ratio of the KM spheres cultured in BM + FGF2 was 0.20 ± 0.05 while the same ratio was 0.38 ± 0.06 in spheres supplemented with BIO pulse (Supplemental Fig. 4A, B). This difference in proliferating cell ratios was abolished after one week of culture and remained similar also at three weeks of culture (Supplemental Fig. 4B). The deceleration of proliferation in KM spheres cultured with BIO pulse coincided with a rapid switch from progenitor to differentiation state in response to strong BIO pulse, as evident from immunofluorescence staining of one week cultured KM spheres (Supplemental Fig. 5).
Kidney mesenchyme spheres WNT-independently undergo tubulogenesis
Several studies have reported that constant basic WNT activity, induced by continual presence of low dose of BIO or CHIR (another potent WNT-activator) facilitates NPs maintenance17,30. Since strong BIO pulse failed to maintain NP state and proliferation in long-term, we set to investigate the effect of low, constant BIO-dose on our 3D dissociated KM cultures. For this, the BIO concentration was titrated to identify appropriate concentration. After one week of culture, 50 nM BIO concentration maintained SIX2+ progenitors without much of the positivity for HPA (Supplemental Fig. 6) indicating maintenance of the progenitor status. No differences were observed by bright field microscopy between dissociated KM cultured in BM + FGF2 and BM + FGF2 + constant BIO in self-organization or KM sphere size (see Fig. 2H, one week time point). This concentration of BIO was chosen for following studies addressing progenitor status (SIX2+) and tubulogenesis as a readout of differentiation (lectin Helix pomatia agglutinin (HPA), a basolateral marker of nephron epithelium).
We next analyzed the long-term maintenance of SIX2+ NPs in KM spheres cultured in BM + FGF2 and BM + FGF2 + constant BIO. Whole mount immunofluorescence staining showed that at one week of culture, SIX2+ NPs were maintained in both BM + FGF2 and BM + FGF2 + constant BIO cultured KM spheres (Fig. 4A, left panel). Quantification of the whole mount immunofluorescence images, using custom image analysis pipelines on the modular workflow system ANIMA31, showed no significant difference in ratios of SIX2+ and Pax2+-to-total cell counts between BM + FGF2 and BM + FGF2 + constant BIO (SIX2+ 15.32% ± 2.52%, 18.70% ± 3.78% n = 3, p = 0.25, and PAX2+ 10.32% ± 4.81%, 16.68% ± 6.28%, n = 3, p = 0.26, respectively, Fig. 4A’). Importantly, these data demonstrate a significantly longer maintenance of cultured NP cells than previously reported in such a simple culture system22. To assess the nephrogenic potential of NP cells cultured for one week in BM + FGF2 and BM + FGF2 + constant BIO we next exposed them to WNT activation, which revealed equal epithelization capacity comparable to that seen with naïve, freshly isolated MM (Supplemental Fig. 8). To test the long-term survival of NP cells in KM spheres we next analyzed SIX2 expression in spheres cultured for three weeks. Very few clusters of SIX2+ NP cells were detected in spheres cultured for three weeks regardless whether they were supplemented with FGF2 alone or together with constant BIO (4.46% ± 2.56% and 4.80% ± 2.56%, respectively, n = 3, p = 0.93, Fig. 4B, B’, and Supplemental Fig. 10). However PAX2 + -to-total cell counts between BM + FGF2 and BM + FGF2 + constant BIO were much higher and with later culture condition causing significantly higher PAX2 expression (31.25 ± 15.33%, and 48.73% ± 17.50% respectively, n = 4, p < 0.05). At this stage, the sphere cultures appeared also not suitable for recombination experiments, which failed due to poor quality of the cells. Of note, epithelial nephron tubules were detected in KM spheres cultured in both conditions for three weeks indicating loss of NP cells by spontaneous differentiation (Supplementary Figs 9, 10).
Nephron progenitors cultured without WNT activation spontaneously differentiate into tubules. (A) At one week, abundant SIX2+ NPs self-organize similarly into clusters regardless whether cultured in BM + FGF2 or BM + FGF2 + constant BIO (left panel). PAX2, a marker for progenitors and early nephron precursors, labels clustered cells (asterisks) and lumen forming tubular cells (arrowheads) in dissociated KM spheres cultured in BM + FGF2, whereas PAX2 labeling predominates in lumen forming tubular cells (arrowheads) when dissociated KM is cultured in BM + FGF2 + constant BIO (right panel). (A’) Quantification of staining at one week of KM cultured in BM + FGF2 or BM + FGF2 + constant BIO shows similar amounts of SIX2+ and PAX2+ progenitors (n = 3, p = 0.26). (B) At three weeks of culture, most of SIX2+ progenitor cells (left panel) have differentiated into PAX2+ complex tubular structures (right panel) in both conditions. (B’) Quantification of staining at three weeks of KM cultured in BM + FGF2 or BM + FGF2 + constant BIO shows small amount of SIX2+ progenitors 4.80% ± 3.13% and 4.46% ± 3.12% (n = 3, p = 0.93), respectively. In contrast, PAX2+ progenitors occupied 31.25% ± 15.33% and 48,73% ± 17,50% (n = 4, p < 0.05), of KM spheres respectively. C) 3D reconstruction rendered from PAX2 immunofluorescent stacks. Upper panel shows folded epithelial aggregates in BM + FGF2 and BM + FGF2 + constant BIO cultured KM spheres at one week time point. Lower panel shows complexed tubular network in both BM + FGF2 and BM + FGF2 + constant BIO KM spheres at three weeks of culture. Scale bar: A and B = 50 µm, C = 30 µm, perspective angle in C = 45°, representative images of 3 independent experiments.
We finally characterized nephrogenesis in the KM spheres cultured for one and three weeks. Staining with PAX2, a protein upregulated in nephron precursors undergoing epithelialization32, revealed PAX2+ cell aggregates, reminiscent of pretubular aggregates and lumen containing nephron tubules in spheres cultured with BM + FGF2 for one week (Fig. 4A, right panel). In BM + FGF2 + constant BIO conditions PAX2+ cells were predominantly found in lumen forming tubules (Fig. 4A, right panel arrowheads). This suggests that the MET was initiated, and to some extent also completed at one week of KM spheres culture. At three weeks, however, we observed a robust PAX2 signal with varying intensities in KM spheres cultured in BM + FGF2 (Fig. 4B, left panel). To ascertain the presence of the tubules, we performed 3D reconstruction of PAX2 immunofluorescence within KM spheres cultured for one and three weeks. At one week of culture, we again confirmed the presence of PAX2+ in the early cell aggregate of spheres cultured in BM + FGF2 and larger aggregates in spheres cultured in BM + FGF2 + constant BIO (Fig. 4C, upper panel). At three weeks a network of PAX2 labelled complex tubular structures were observed in both BM + FGF2 and BM + FGF2 + constant BIO cultured KM spheres (Fig. 4C). We have further confirmed that these tubules are CDH1 positive (Supplemental Fig. 10), however lack of segment specific markers such as LTL in CHD1 + tubules suggests that the nephron epithelium is fairly immature. Taken together, the panel of immunofluorescence staining and their 3D reconstructions indicate that KM spheres spontaneously differentiate into tubular epithelium even without activation of WNT signaling.
Discussion
The organization of an in vivo NP niche involves presence of all three tissue layers contributing to normal kidney organogenesis, namely KM itself, stroma and UB epithelium. In vitro, the self-organization and propagation of kidney nephron progenitors require activation of a complex mixture of different cascades aiming to mimic signals normally secreted by UB and stroma17,30. Simple systems, which allow manipulation of the cells, have utilized dissociation and re-aggregation together with the UB-derived inductive signals such as GSK3 inhibition, UB itself or spinal cord expressing several Wnts1,12,20,22,33. Until recently, in vitro expansion of nephron progenitors was not possible and current protocols now in use are not only expensive but also tedious17,34.
Various in vivo and in vitro studies have indicated that FGF signaling is indispensable for survival and maintenance of metanephric mesenchyme18,22,27,28,29. Our 3D cell culture model confirms this. Contrary to previous studies, here we present a simple 3D culture method to aggregate and propagate dissociated KM into spheres resembling NP niche and maintaining nephrogenic potential up to one week. This new method utilizes the 3D Matrigel in combination with FGF2 and PP2 to generate conditions allowing self-organization into nephron progenitors and stromal cells where both lineages interact with each other and support their maintenance. As indicated by previous studies, a clear lineage separation of the nephron and stromal progenitor is established early in renal differentiation35,36,37. Stromal compartment in cooperation with ureteric epithelium is also shown to be crucial for NP self-renewal and differentiation38,39,40. In the light of a growing interest towards generating kidney organoids from human cells41,42, our culture method presents more simple approach by providing an opportunity to investigate the cell aggregation43, self-organization, cross-talk between stroma and NPs in the conditions where signals from ureteric epithelium or other inductive tissues are not needed.
Likely, the most interesting finding with our newly developed dissociated KM culture is the observation that NPs within KM spheroids, cultured without exogenous WNT-activation, undergo spontaneous epithelization. This exciting finding underlines the robustness of the nephron induction and tubulogenesis programs, implying that similarly to rat kidneys44, signaling pathways other than just GSK3β/WNT-activation supplement regulation of self-renewal and differentiation decisions in mouse KM. Indeed, Schmidt-Ott and colleagues previously identified conserved UB tip expressed gene signature in mouse and rat kidneys, and demonstrated that e.g. secreted cytokine receptor, cytokine receptor-like factor 1 (CLF-1) in complex with its ligand induces tubulogenesis in isolated mouse MM45. Moreover, several studies also support the involvement of FGF induced signaling in differentiation of nephrons43,46,47. Thus, the robust and easy 3D culture method presented here provides a tool to study niche-NPs interaction and can be utilized to generate new knowledge about NPs characteristics and nephron differentiation.
Material and Methods
Tissue culture
Mouse embryonic kidneys were microdissected from E11.5 NMRI embryos as described48. The kidneys were dissected in PBS, and subsequently treated with collagenase type IV for 15 min at 37 °C. Thereafter, the kidney mesenchyme (KM) was separated from the ureteric epithelium (UB) by microdissection in the DMEM supplemented with 10% fetal calf serum. The KM was incubated with TrypLE Express and dispersed into single cells in Basic Media (BM) (DMEM/F12, 10% fetal calf serum (FCS), and 10 µM PP2 (to reduce neuronal outgrowths and augment sphere aggregation) or BM with 50 ng/ml FGF29. The cell suspension was seeded on normal growth factor 3D Matrigel (Basement Membrane Matrix from EHS mouse tumor, BD Biosciences), at the density of 3 × 104 cells/well, 50 µl/well to Lab-Tek Chamber Slide system 16 chamber glass slide (LabTek/ThermoFisher Scientific) or to a flat bottom 96 well plate (Costar/Corning Inc.). For BIO ((2′Z,3′E)-6-Bromoindirubin-3′-oxime; Calbiochem) pulse experiments, BIO was added at a final concentration of 10 µM during cell plating and removed after 1 hour by replacing medium with 200 µl of fresh BM + FGF2 without BIO. For low dose experiments, BIO was added to 50 nM final concentration and kept constant during the culture. All media were replaced with fresh after three days.
Embryonic kidney dissociation and reaggregation
Embryonic kidneys were dissociated and reaggregated as described in (Unbekandt and Davies, 2010)12 with following modifications: E11.5 embryonic kidneys were isolated as described48, the isolated kidneys were dissociated into single cells by pipetting after incubation of 5 min in TrypLE at 37 °C. The digestion was blocked by washing in 10 parts of DMEM supplemented in 10% FCS. Cell were counted and at least 80,000 cells were used for reaggregation by centrifugation at 500 × g for 3 min. Pellets were transferred on nucleopore carbon filter and cultured on Trowell-type system at air liquid interface in DMEM supplemented with 10% FCS.
Tissue recombination
After five days of culture, the KM spheres were recovered from Matrigel and transferred on top of a piece of E11.5 mouse spinal cord (Supplementary Fig. 7). The recombinant tissue was cultured in Trowell type tissue culture for 72 hours, followed by fixation with 4% PFA for 1 hour at room temperature and subjected to immunofluorescence staining.
Immunofluorescence
The KM spheres were fixed for 20 min at RT with 4% PFA (unless otherwise stated), washed with PBS, pelleted in 1.5 ml low protein binding tubes, embedded in Richard-Allan Scientific HistoGel (for paraffin sections) or OCT embedding compound (for cryosections), and subsequently cut at 5 µm section in microtome or cryostat. The sections were blocked with secondary antibody host specific 10% serum for 1 hour at RT. Samples were incubated with primary antibodies overnight at +4 °C, followed washes in PBS and 1 hour incubation with secondary antibodies. The samples were then counterstained with 10 µg/ml of Hoechst or DAPI, and mounted for imaging.
Whole mount immunofluorescence
The KM spheres were fixed for 30 min at RT with 4% PFA, washed with PBS, pelleted in 1.5 ml low protein binding tubes and pellets were transferred on microscopic slide. The pellets were permeabilized for 20 min with PBS Triton x-100 0.5%, blocked for 1 hour with secondary antibody host specific 10% serum at RT. Samples were rinsed with PBS and incubated with primary antibodies at +4 °C o.n., followed by wash and incubation with secondary antibody o.n. at +4 °C. After wash samples were counterstained with DAPI mounted and imaged.
Proliferation assay
At the indicated time points the proliferating cells were labeled with 10 µM of 5-ethynyl-2′-deoxyuridine (EdU) for 2 hours and detected by Click-IT EdU Imaging Kit (AlexaFluor-594 conjugated, Life Technologies) according to the standard protocol. Subsequently, the samples were co-immunostained as described above.
Flow cytometric analysis of cell viability
After 72 hours in culture the KM spheres were harvested from Matrigel, transferred to low protein binding tubes, and collected by centrifugation and subsequently dissociated into single cells by TrypLE Express. Cells were stained with Sytox Red Dead Cell Stain (ThermoScientific) according manufacturer’s instruction, and analyzed with Accuri Flow Cytometer (BD Biosciences) at the HiLIFE Biomedicum Flow Cytometry core facility.
Imaging and image analysis
The samples were imaged by Zeiss AxioImager Z1 or Leica SP8 confocal microscope. Bright field images were taken by Nikon Eclipse Ti-E. The proliferation and protein expression ratios (labeled cells to-total cell count) were quantified using custom developed pipelines on the modular workflow system ANIMA31. In short, images were pre-processed, segmented for cell nuclei by Graphcut algorithm using Farsight library49, and mapped for mask, which was used to measure intensity of label channel. The intensity values and their standard deviations were clustered by k-mean algorithm to determine negative staining threshold and signal intensity classes. KM spheres 3D structure was reconstructed using Imaris software (Bitplane) from convoluted (Huygens Professional software, SVI) immunofluorescence and DAPI counterstained-stained confocal microscopy images. Imaging, image deconvolution and 3D rendering has been performed at HiLIFE Biomedicum imaging core facility.
Data analysis and statistical methods
For all figures the independent experiments (different mouse litters with no less than 8 embryos) were performed at least three times. The proliferation ratio was calculated from at least three independent experiments, each of which contained pooled KM sample from at least eight embryos. To determine the proliferation ratio and cell survival percentage no less than 10,000 cell counts were used. For SIX2 and PAX2-to-total cell count positive cell quantification, images form at least 3 independent experiments were used and calculated as described in image analysis section above. The independent two-sample t-test (significance p < 0.05) analysis was performed on the means of the ratios (calculated with OriginPro statistical analysis software).
Ethical approval
All methods were carried out in accordance with relevant guidelines and regulations. All animal husbandry and experimental procedures were approved by EU legislation and Finnish Animal Care and Use Committee.
Significance statement
The kidney mesenchyme (KM), also known as metanephric mesenchyme, contains both nephron progenitors (NPs) and stromal cells. In vitro maintenance and propagation of dissociated KM and uninduced NPs is not trivial, and current culture protocols rely on complex procedures. This study demonstrates that a combination of three dimensional (3D) cell culture and Fibroblast Growth Factor 2 (FGF2) expands NPs that self-organize into spheres resembling the niche. They also form epithelial tubes without inductive tissues or exogenous source for WNT activation. This robust and easy 3D culture method provides a tool to study niche-NPs interaction and can be utilized to generate new knowledge about NPs characteristics.
Data Availability
The datasets and quantification pipelines generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Saxén, L. & Sariola, H. Early organogenesis of the kidney. Pediatr Nephrol 1, 385–392 (1987).
Krause, M., Rak-Raszewska, A., Pietila, I., Quaggin, S. E. & Vainio, S. Signaling during Kidney Development. Cells 4, 112–132, https://doi.org/10.3390/cells4020112 (2015).
O’Brien, L. L. et al. Wnt11 directs nephron progenitor polarity and motile behavior ultimately determining nephron endowment. eLife 7, https://doi.org/10.7554/eLife.40392 (2018).
Costantini, F. Genetic controls and cellular behaviors in branching morphogenesis of the renal collecting system. Wiley interdisciplinary reviews. Developmental biology 1, 693–713, https://doi.org/10.1002/wdev.52 (2012).
Carroll, T. J., Park, J.-S., Hayashi, S., Majumdar, A. & McMahon, A. P. Wnt9b plays a central role in the regulation of mesenchymal to epithelial transitions underlying organogenesis of the mammalian urogenital system. Dev Cell 9, 283–292, https://doi.org/10.1016/j.devcel.2005.05.016 (2005).
Kobayashi, A. et al. Six2 defines and regulates a multipotent self-renewing nephron progenitor population throughout mammalian kidney development. Cell Stem Cell 3, 169–181, https://doi.org/10.1016/j.stem.2008.05.020 (2008).
Karner, C. M. et al. Canonical Wnt9b signaling balances progenitor cell expansion and differentiation during kidney development. Development 138, 1247–1257, https://doi.org/10.1242/dev.057646 (2011).
Stark, K., Vainio, S., Vassileva, G. & McMahon, A. P. Epithelial transformation of metanephric mesenchyme in the developing kidney regulated by Wnt-4. Nature 372, 679–683, https://doi.org/10.1038/372679a0 (1994).
Kuure, S., Popsueva, A., Jakobson, M., Sainio, K. & Sariola, H. Glycogen synthase kinase-3 inactivation and stabilization of beta-catenin induce nephron differentiation in isolated mouse and rat kidney mesenchymes. J Am Soc Nephrol 18, 1130–1139, https://doi.org/10.1681/ASN.2006111206 (2007).
Grobstein, C. Inductive epitheliomesenchymal interaction in cultured organ rudiments of the mouse. Science 118, 52–55 (1953).
Yang, J. et al. An epithelial precursor is regulated by the ureteric bud and by the renal stroma. Dev Biol 246, 296–310, https://doi.org/10.1006/dbio.2002.0646 (2002).
Unbekandt, M. & Davies, J. A. Dissociation of embryonic kidneys followed by reaggregation allows the formation of renal tissues. Kidney Int 77, 407–416, https://doi.org/10.1038/ki.2009.482 (2010).
Lanza, R. P. et al. Generation of histocompatible tissues using nuclear transplantation. Nature biotechnology 20, 689–696, https://doi.org/10.1038/nbt703 (2002).
Dressler, G. R. The cellular basis of kidney development. Annu Rev Cell Dev Biol 22, 509–529, https://doi.org/10.1146/annurev.cellbio.22.010305.104340 (2006).
Costantini, F. & Kopan, R. Patterning a Complex Organ: Branching Morphogenesis and Nephron Segmentation in Kidney Development. Developmental Cell 18, 698–712, https://doi.org/10.1016/j.devcel.2010.04.008 (2010).
Kurtzeborn, K., Cebrian, C. & Kuure, S. Regulation of Renal Differentiation by Trophic Factors. Front Physiol 9, 1588, https://doi.org/10.3389/fphys.2018.01588 (2018).
Li, Z. et al. 3D Culture Supports Long-Term Expansion of Mouse and Human Nephrogenic Progenitors. Cell Stem Cell 19, 516–529, https://doi.org/10.1016/j.stem.2016.07.016 (2016).
Tanigawa, S., Taguchi, A., Sharma, N., Perantoni, A. O. & Nishinakamura, R. Selective In Vitro Propagation of Nephron Progenitors Derived from Embryos and Pluripotent Stem Cells. Cell Rep, https://doi.org/10.1016/j.celrep.2016.03.076 (2016).
Bryant, D. M. et al. A molecular network for de novo generation of the apical surface and lumen. Nat Cell Biol 12, 1035–1045, https://doi.org/10.1038/ncb2106 (2010).
Rosines, E. et al. Constructing kidney-like tissues from cells based on programs for organ development: toward a method of in vitro tissue engineering of the kidney. Tissue Eng Part A 16, 2441–2455, https://doi.org/10.1089/ten.TEA.2009.0548 (2010).
Morizane, R. et al. Kidney specific protein-positive cells derived from embryonic stem cells reproduce tubular structures in vitro and differentiate into renal tubular cells. PLoS One 8, e64843, https://doi.org/10.1371/journal.pone.0064843 (2014).
Junttila, S. et al. Functional genetic targeting of embryonic kidney progenitor cells ex vivo. J Am Soc Nephrol 26, 1126–1137, https://doi.org/10.1681/ASN.2013060584 (2015).
Morizane, R. et al. Nephron organoids derived from human pluripotent stem cells model kidney development and injury. Nat Biotechnol 33, 1193–1200, https://doi.org/10.1038/nbt.3392 (2015).
Steer, D. L., Bush, K. T., Meyer, T. N., Schwesinger, C. & Nigam, S. K. A strategy for in vitro propagation of rat nephrons. Kidney Int 62, 1958–1965, https://doi.org/10.1046/j.1523-1755.2002.00694.x (2002).
Morizane, R., Monkawa, T. & Itoh, H. Differentiation of murine embryonic stem and induced pluripotent stem cells to renal lineage in vitro. Biochemical and Biophysical Research Communications 390, 1334–1339, https://doi.org/10.1016/j.bbrc.2009.10.148 (2009).
Morizane, R., Miyoshi, T. & Bonventre, J. V. Concise Review: Kidney Generation with Human Pluripotent Stem Cells. Stem Cells 35, 2209–2217, https://doi.org/10.1002/stem.2699 (2017).
Barasch, J. et al. Ureteric bud cells secrete multiple factors, including bFGF, which rescue renal progenitors from apoptosis. The American Journal of Physiology 273, F757–767 (1997).
Barak, H. et al. FGF9 and FGF20 Maintain the Stemness of Nephron Progenitors in Mice and Man. Developmental Cell 22, 1191–1207, https://doi.org/10.1016/j.devcel.2012.04.018 (2012).
Gallegos, T. F., Kamei, C. N., Rohly, M. & Drummond, I. A. Fibroblast growth factor signaling mediates progenitor cell aggregation and nephron regeneration in the adult zebrafish kidney. Dev Biol, https://doi.org/10.1016/j.ydbio.2019.06.011 (2019).
Brown, A. C., Muthukrishnan, S. D. & Oxburgh, L. A synthetic niche for nephron progenitor cells. Dev Cell 34, 229–241, https://doi.org/10.1016/j.devcel.2015.06.021 (2015).
Rantanen, V., Valori, M. & Hautaniemi, S. Anima: modular workflow system for comprehensive image data analysis. Frontiers in bioengineering and biotechnology 2, 25, https://doi.org/10.3389/fbioe.2014.00025 (2014).
Dressler, G. R., Deutsch, U., Chowdhury, K., Nornes, H. O. & Gruss, P. Pax2, a new murine paired-box-containing gene and its expression in the developing excretory system. Development 109, 787–795 (1990).
Ganeva, V., Unbekandt, M. & Davies, J. A. An improved kidney dissociation and reaggregation culture system results in nephrons arranged organotypically around a single collecting duct system. Organogenesis 7, 83–87, https://doi.org/10.4161/org.7.2.14881 (2011).
Takasato, M., Er, P. X., Chiu, H. S. & Little, M. H. Generation of kidney organoids from human pluripotent stem cells. Nat Protoc 11, 1681–1692, https://doi.org/10.1038/nprot.2016.098 (2016).
Levinson, R. S. et al. Foxd1-dependent signals control cellularity in the renal capsule, a structure required for normal renal development. Development 132, 529–539, https://doi.org/10.1242/dev.01604 (2005).
Humphreys, B. D. et al. Fate tracing reveals the pericyte and not epithelial origin of myofibroblasts in kidney fibrosis. The American journal of pathology 176, 85–97, https://doi.org/10.2353/ajpath.2010.090517 (2010).
Kobayashi, A. et al. Identification of a multipotent self-renewing stromal progenitor population during mammalian kidney organogenesis. Stem Cell Reports 3, 650–662, https://doi.org/10.1016/j.stemcr.2014.08.008 (2014).
Das, A. et al. Stromal-epithelial crosstalk regulates kidney progenitor cell differentiation. Nat Cell Biol 15, 1035–1044, https://doi.org/10.1038/ncb2828 (2013).
Hatini, V., Huh, S. O., Herzlinger, D., Soares, V. C. & Lai, E. Essential role of stromal mesenchyme in kidney morphogenesis revealed by targeted disruption of Winged Helix transcription factor BF-2. Genes &. development 10, 1467–1478 (1996).
Hum, S., Rymer, C., Schaefer, C., Bushnell, D. & Sims-Lucas, S. Ablation of the renal stroma defines its critical role in nephron progenitor and vasculature patterning. PLoS One 9, e88400, https://doi.org/10.1371/journal.pone.0088400 (2014).
Takasato, M. et al. Kidney organoids from human iPS cells contain multiple lineages and model human nephrogenesis. Nature 526, 564–568, https://doi.org/10.1038/nature15695 (2015).
Tan, Z., Shan, J., Rak-Raszewska, A. & Vainio, S. J. Embryonic Stem Cells Derived Kidney Organoids as Faithful Models to Target Programmed Nephrogenesis. Scientific reports 8, 16618, https://doi.org/10.1038/s41598-018-34995-3 (2018).
Ihermann-Hella, A. et al. Dynamic MAPK/ERK Activity Sustains Nephron Progenitors through Niche Regulation and Primes Precursors for Differentiation. Stem Cell Reports 11, 912–928, https://doi.org/10.1016/j.stemcr.2018.08.012 (2018).
Barasch, J. et al. Mesenchymal to epithelial conversion in rat metanephros is induced by LIF. Cell 99, 377–386 (1999).
Schmidt-Ott, K. M. et al. Novel regulators of kidney development from the tips of the ureteric bud. J Am Soc Nephrol 16, 1993–2002, https://doi.org/10.1681/asn.2004121127 (2005).
Grieshammer, U. et al. FGF8 is required for cell survival at distinct stages of nephrogenesis and for regulation of gene expression in nascent nephrons. Development 132, 3847–3857, https://doi.org/10.1242/dev.01944 (2005).
Perantoni, A. O. et al. Inactivation of FGF8 in early mesoderm reveals an essential role in kidney development. Development 132, 3859–3871, https://doi.org/10.1242/dev.01945 (2005).
Sainio, K. In The Kidney (eds Woolf, A. S. & Bard, J. B. L.) 327–342 (Academic Press, 2003).
Luisi, J., Narayanaswamy, A., Galbreath, Z. & Roysam, B. The FARSIGHT trace editor: an open source tool for 3-D inspection and efficient pattern analysis aided editing of automated neuronal reconstructions. Neuroinformatics 9, 305–315, https://doi.org/10.1007/s12021-011-9115-0 (2011).
Acknowledgements
The authors acknowledge, Agnes Viherä, Solja Eurola (Department of Biochemistry and Developmental Biology, Faculty of Medicine, University of Helsinki) for excellent technical support. The authors thank Marc Baumann, MD PhD (Department of Biochemistry and Developmental Biology, Faculty of Medicine, University of Helsinki) for critical comments on the manuscript. All authors agreed on publication. Disclosures: None.
Author information
Authors and Affiliations
Contributions
A.D., S.K., K.S. and H.S. conceived and designed the study; A.D., K.S., M.L., S.K. and H.S. drafted and revised the manuscript; Y.G. and A.D. designed organoid embedding methodology. V.R. and A.D. designed image analysis methods; A.D. performed experiments and data analysis; H.S. gave the final approval the manuscript for publication.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Dapkunas, A., Rantanen, V., Gui, Y. et al. Simple 3D culture of dissociated kidney mesenchyme mimics nephron progenitor niche and facilitates nephrogenesis Wnt-independently. Sci Rep 9, 13433 (2019). https://doi.org/10.1038/s41598-019-49526-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-49526-x
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.