Abstract
During peri-implantation development in mice, X chromosome inactivation (XCI) status changes dynamically. Here, we examined the expression of Xist and its antisense partner, Tsix, via whole-mount 3D RNA-FISH using strand-specific probes and evaluated XCI status. The results indicate that Xist expression disappears completely by embryonic day (E) 4.5 without Tsix activation in the ICM and that Xist re-expression occurs at E4.75 in some cells, suggesting that random XCI is already initiated in these cells. Intriguingly, epiblast cells exhibiting biallelic Xist expression were observed frequently (~15%) at E5.25 and E5.5. Immunostaining analysis of epigenetic modifications suggests that global change in epigenomic status occurs concomitantly with the transition from imprinted to random XCI. However, global upregulation of H3K27me3 modifications initiated earlier than other modifications, occurring specifically in ICM during progression of Xist erasure. Although both Xist expression and imprinted XCI are thought to be stable in the primitive endoderm/visceral endoderm and trophectoderm/extraembryonic ectoderm lineages, transient loss of Xist clouds was noted only in a subset of extraembryonic ectodermal cells, suggesting distinct features of Xist regulation among the three different embryonic tissue layers. These results will serve as a basis for future functional studies of XCI regulation in vivo.
Similar content being viewed by others
Introduction
In female mammals, one of the two X chromosomes is inactivated for gene dosage compensation between XX females and XY males1. This phenomenon, termed X chromosome inactivation (XCI), is regulated by several factors, such as the noncoding RNA Xist and its antisense sequence Tsix. Xist is exclusively expressed from the inactive X (Xi) and accumulates on it, leading to a chromosome-wide inactivation of gene expression2,3,4,5. Tsix, with its repressive effect on Xist expression, is expressed normally from the active X and is silenced on Xi6,7. Imprinted XCI occurs in preimplantation-stage embryos and Xist is essential for its initiation8. During this process, the paternal X (Xp) is preferentially selected as Xi according to a maternal imprint causing Xist repression on maternal allele9,10,11. The maternal imprint is now thought to be H3K27me3 modifications laid onto maternal X during oogenesis12. The imprinted XCI is then erased in the embryonic lineage, and XCI is resumed later as random XCI, in which Xi is chosen randomly. The erasure of imprinted XCI initiates in the inner cell mass (ICM) of early blastocysts. This is accompanied by the loss of Xist RNA accumulation, EED/EZH2 association and histone H3 lysine 27 trimethylation (H3K27me3) modifications from the Xp, and derepression of genes that are subjected to inactivation on the Xp13,14,15,16. During this erasure process, epigenetic memories for imprinted XCI are thought to be erased and both X chromosomes become epigenetically equivalent. Random XCI takes place after this imprinted XCI erasure. Although these event sequences have been described17, the precise timing of XCI erasure and initiation of random XCI during the development of peri-implantation embryos in vivo is not understood fully.
The reasons for studying the precise kinetics of XCI during embryonic development are at least twofold. First, basic information on the dynamics of XCI will provide clues that will contribute to understanding the regulatory mechanisms that operate in vivo. Traditionally, the in vitro differentiation system of embryonic stem (ES) cells has been used extensively in studying XCI. Despite its great experimental advantages, the in vitro ES cell system cannot cover all aspects of the XCI dynamics that occur in vivo. For example, imprinted XCI does not exist in the ES cell system. To reevaluate the usefulness of the ES cell system and to understand the XCI phenomenon in general, studies of embryos developing in vivo are essential, as they add complementary knowledge to the data accumulated from in vitro studies. Second, changes in XCI status are likely to be coupled with epigenomic or nuclear reorganization in developing peri-implantation mouse embryos. ICM and mouse ES cells (mESCs) represent a ground state (naïve state) of pluripotency, whereas epiblasts of postimplantation-stage embryos or epiblast stem cells (EpiSCs) correspond to a “primed” pluripotent state18. XCI is one of the key features of EpiSCs. In contrast with female mESCs, where the two X chromosomes are both active, a random XCI operates in female EpiSCs. It is becoming increasingly clear that there are significant differences in epigenetic status or an “epigenetic barrier” between the naïve and primed states of pluripotent stem cells19,20,21, and that the imprinted XCI–random XCI conversion that takes place in peri-implantation mammalian embryos might be a reflection of epigenomic reorganizations that are not restricted to X chromosomes. Therefore, we believe that the XCI status could be a useful indicator of large-scale epigenomic reprogramming events that have remained unexplored to date.
Xist RNA clouds or coatings (i.e., the accumulation of Xist RNA over the entire Xi) are one of the indicators of whether cells are in XCI state22,23,24, and the accumulation of Xist RNA is lost during imprinted XCI erasure13,14,16. As Xist RNA is essential for the establishment of XCI, re-expression of Xist is thought to be the sign of random XCI commencement. However, the repression of Xist itself might not represent an active state of the X chromosome, because it is known that the expression/repression status of Xist does not necessarily coincide with the expression status of other X-linked genes. For example, it has been reported that Xist expression is dispensable for X inactivation in mouse embryonic fibroblasts (MEFs)25 or in developing primordial germ cells (PGCs)26. Moreover, it has been demonstrated that several X-linked genes located on the imprinted Xi are reactivated even in the presence of Xist coatings15,16. Therefore, it is necessary to examine both Xist repression and activation of X-linked gene(s) to judge whether XCI reversal occurs or not. Among the numerous X-linked genes, we have paid particular attention to Tsix, an antisense RNA partner of Xist that plays a repressive role in Xist expression7,27,28. It is known that continued expression of an inducible Tsix allele causes stable repression of Xist in imprinted XCI during preimplantation development28. However, whether Tsix has an active role in induction of Xist repression during imprinted XCI erasure in vivo is not fully understood. Furthermore, in female ES cells, Tsix is expressed at high levels from both alleles, and it is proposed that this biallelic Tsix expression induces changes in histone modifications along the Xist locus that ensure the epigenetic equivalency of both X chromosomes in undifferentiated pluripotent cells29, implying that the cells that have undergone X reactivation will show biallelic expression of Tsix both in vitro and in vivo. Therefore, we decided to track the kinetics of both Xist and Tsix expressions to know whether Tsix reactivation trigger the initiation of Xist repression and make a careful estimate of XCI status during XCI reversal and random XCI establishment. In addition, as Tsix reactivation would not necessarily reflect the overall state of the X chromosome, expression statuses of other X-linked genes (Lamp2 and Pgk1) were also examined to confirm progression of X reactivation at the developmental stage when biallelic expression of Tsix became predominant. To perform a quantitative expression analysis in peri-implantation-stage embryos, we used whole-mount RNA fluorescence in situ hybridization (FISH), which had been successfully utilized for study of X reactivation in PGCs26. RNA-FISH is better suited to assessing the on-and-off state of transcription because it can detect nascent RNA and not the steady-state level of RNA. Furthermore, the whole-mount technique, which does not destroy embryonic structures, enables the unambiguous detection of RNA expression in different embryonic lineages, such as the trophectoderm/extraembryonic ectoderm (TE/ExE), primitive endoderm/visceral endoderm (PE/VE), and ICM/epiblast lineages.
The results presented here demonstrate Xist/Tsix dynamics during peri-implantation development at an unprecedented resolution, implying the period of imprinted XCI erasure and the timing of random XCI commencement in the ICM/epiblast lineage in vivo. Moreover, we found differences in Xist/Tsix dynamics and changes in epigenetic status among the three different lineages of embryos. The information regarding Xist/Tsix dynamics in vivo obtained in this study will serve as a basis for future functional studies on XCI regulation.
Results
Kinetic changes in Xist/Tsix status during development of the embryonic lineage
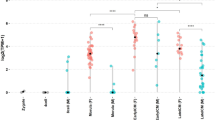
Xist and Tsix expression in ICM/epiblast, PE/VE and TE/ExE lineages were examined by using strand-specific RNA-FISH (see Methods for details of the strand-specific probe preparation) combined with immunofluorescence against those cell lineage markers during a period between E3.5 and E5.5 at intervals of 6 h (Figs 1, 2, Table 1 and Supplementary Fig. S1a). Embryonic lineage cells were marked by an antibody against the pluripotent cell marker POU5F1 (OCT3/4) for E3.5 blastocysts and for embryos sampled between E4.75 and E5.5. As POU5F1 expression was observed in both the primitive endoderm and the epiblast at stages from E3.75 to E4.5, anti-NANOG was used instead to stain embryonic lineage cells (Supplementary Fig. S1a). Images were acquired on a confocal microscope, and z-stack images were used to analyze Xist/Tsix expression patterns in each cell of the whole embryo (Fig. 1b–i; see Supplementary Video S1). We used this three-dimensional (3D) whole-mount RNA-FISH technique26 to determine the kinetic changes in Xist/Tsix expression in almost all the cells that constituted entire embryos at E3.5–5.5, and the analysis was also performed at E6.5 (Table 1, 4,127 nuclei in 107 embryos in total). Xist-positive cells were classified into four categories: cells with one Xist cloud per nucleus, cells with single pinpoint or dispersed signals, cells with two Xist signals, or cells with no Xist signal. Tsix-positive cells were categorized into cells with two, one, or no signals. To understand the relationships between the expression patterns of Xist and Tsix in the same individual cells, a matrix illustrating proportional data (%) for each category of Xist- or Tsix-positive cells was constructed (Fig. 1a).
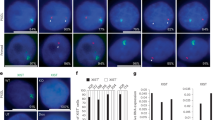
Xist/Tsix expression kinetics in the embryonic cell lineage. (a) The matrices represent the Xist and Tsix expression patterns of embryonic-lineage cells at each developmental stage. The rows represent Tsix expression patterns (no signal, one, or two signals) and the columns represent Xist expression patterns (one cloud, single pinpoint or dispersed signal, no signal, or two signals). The numbers displayed at the intersections of rows and columns indicate the percentage of cells with the Tsix and Xist expression pattern indicated by the row and the column, respectively. (b–g) Representative images of cells showing different patterns of Xist (green)/Tsix (magenta) expression. The blue and white colors indicate lineage-marker expression and nuclear DNA staining, respectively. (b) One Xist cloud and a single Tsix signal; (c) no Xist and a single Tsix signal; (d) no Xist and two Tsix signals; (e) one Xist pinpoint signal and two Tsix signals; (f) one Xist dispersed signal and a single Tsix spot; and (g) two Xist clouds and a single Tsix signal. (h,i) Examples of single optical sections of E4.75 (h) and E5.5 (i) embryos. The cells enclosed by a dotted circle correspond to the cells shown in (d,g), respectively. (j) The expression patterns of two ordinary X-linked genes, Lamp2 (77 nuclei in 3 embryos) and Pgk1 (57 nuclei in 2 embryos), and Tsix in the embryonic cell lineage at E4.75.
Xist/Tsix expression kinetics in extraembryonic lineages. (a,b) The matrices represent Xist and Tsix expression patterns in PE/VE lineage cells (a) and TE/ExE lineage cells (b) in each developmental stage, respectively. (c,d) Representative images of Xist (green)/Tsix (magenta) RNA-FISH combined with GATA6 (for PE/VE lineage (c)) and CDX2 (for TE/ExE lineage (d)) immunofluorescence with nuclear DNA staining (white). (e) Image of an E4.5 embryo. Cell 1 exhibited a Xist cloud, whereas cell 2 was negative for the Xist signal. (f,g) Serial sections of cells 1 (f) and 2 (g) respectively.
In E3.5 blastocysts, more than 80% of ICM cells possessed one Xist cloud and one pinpoint Tsix signal per nucleus (Fig. 1b), indicating that most ICM cells were still subjected to imprinted XCI at this stage. From E3.75 onward, Xist cloud signals began to disappear and cells showing biallelic expression of Tsix began to appear (Fig. 1c,d).
At E4.5, all epiblast cells were Xist negative. A large proportion of the cells (58%) showed a single Tsix signal, whereas 11% and 31% of cells exhibited no or two Tsix signals, respectively. This indicates that the erasure of Xist precedes the derepression of Tsix from the silent allele and that Tsix activation might not be a prerequisite for the disappearance of Xist. The numbers of cells showing Tsix biallelic expression increased after E4.5, whereas the numbers of Tsix single-positive cells decreased. Most cells (>53%) exhibited biallelic expression of Tsix and no expression of Xist (Fig. 1h) at E4.75 and E5.0, and the numbers of cells showing Tsix biallelism then decreased from 72% at E5.0 to 29% at E5.25. These results suggest that reactivation of the silent Tsix allele occurred increasingly after E4.5 and plateaued at E4.75–5.0. At E4.75, the proportions of the cells showing biallelic expression of Lamp2 and Pgk1 were almost the same as that of the cells exhibiting Tsix biallelic expression (Figs 1j and S2). The result indicates that reactivation of “ordinary” X-linked genes, in addition to Tsix, was also progressing at this stage. A small number of cells in the epiblast had already begun to show weak Xist signals at E4.75 and the proportion of such cells increased thereafter, indicating the random XCI process had already begun in those cells. These Xist weakly positive cells were not clustered or positioned in any specific region of the epiblast. These results suggest that switching from the XCI reversal to random XCI had already been initiated in some nuclei at E4.75–5.0 before all the epiblast cells had completed XCI reversal. The numbers of Xist-positive cells increased further so that at E5.0, 26% of the epiblast cells showed weak Xist expression as a pinpoint signal (Fig. 1e), whereas more than half of the cells (55%) exhibited Xist signals at E5.25. Although most of the cells showed dispersed Xist signals (Fig. 1f), cells displaying Xist clouds began to appear from this stage onward, suggesting that the expression level of Xist increased gradually between E4.75 and E5.25. During this period, the proportion of cells with two Tsix signals and one Xist signal decreased, whereas the numbers of cells with one Tsix and one Xist signal increased, implying that elevated expression of Xist seems to cause Tsix silencing. At E5.5, a large proportion (>50%) of the cells showed one Xist cloud. Xist /Tsix expression patterns in embryonic lineage at E6.5 were also examined and the result showed that almost all of the cells displayed a single Xist cloud with or without one Tsix signal and cells with biallelic Tsix expression were absent (Fig. 1a and Table 1). This suggests that Xi had already been chosen in almost all the epiblast cells by E6.5.
Intriguingly, a significant number of epiblast cells exhibited biallelic expression of Xist at E5.25 (13.9%) and E5.5 (9.3%) (Fig. 1a,g,i). Cells with two Xist clouds were also detected, and the frequency of such cells increased at E5.5 (Figs 1g,i and S3). Combined analysis of RNA and DNA-FISH can directly address whether those cells were normal diploid or tri-/higher-ploid cells, but such combined analysis is not compatible with our whole-mount protocol (see Discussion). Instead, we reasoned that potential triploid cells exhibit two Xist clouds and one Tsix signal on the three different positions and looked for such cases. In this analysis, we hardly observed the cells in which one or more Tsix signals were localized distantly from two Xist clouds. In fact, there was only one cell showing potential triploidy or higher ploidy in the epiblast cells at E5.25 (1 cell in 136 cells observed), as well as one such cell at E5.5 (1 in 258 cells). In both cases, the cell exhibited two Tsix signals and one Xist signal localized distantly from Tsix signals. Therefore, although the possibility of aneuploidy cannot be ruled out completely, we believe that it is highly likely that those cells exhibiting the two Xist signals observed in this study were normal diploid cells.
Xist expression appears to be stable in the PE/VE lineage during peri-implantation development
Most of the cells (88–98%) in the PE/VE lineage marked by GATA6 expression (Supplementary Fig. S1a) exhibited one Xist cloud per nucleus throughout the developmental period examined here (Fig. 2a,c). This result suggests that the imprinted repression of maternal Xist and paternal Xist expression are basically maintained in this lineage from E3.5 to E5.5. However, pinpoint signals for Tsix were lost specifically at E4.0 in nearly 50% of the cells. Loss of Tsix expression of this kind has never been observed in ExE cells. At E4.0 and E4.25, some cells (12.1% at E4.0 and 4.4% at E4.25) exhibited weak or no signals for Xist. As differentiation of the PE and epiblasts could not be delineated clearly by the lineage markers NANOG and GATA6 at this stage (Supplementary Fig. S1b), PE/VE cells with weaker Xist expression might represent precursor cells of the epiblast lineage, in which Xist will disappear.
Transient loss of Xist clouds in a subset of ExE cells around the implantation stage
The TE or ExE cells were marked by CDX2 expression (Supplementary Fig. S1a and Supplementary Video S2). At all stages examined, most of the TE/ExE cells (70–95%) possessed one Xist cloud and a single Tsix spot (Fig. 2b,d–f), suggesting that the imprinted XCI persisted throughout the development of this lineage. However, careful examination of the expression of Xist revealed that cells with no Xist signals were present and that the number of these cells increased transiently, reaching a peak at around E4.75 (21% of all the ExE cells showed no Xist expression; Fig. 2b,e,g). These Xist-negative cells were not clustered or positioned in a specific region of ExE tissues. Most of these Xist-negative cells had either a single or no Tsix signal, and very few cells showed two Tsix signals. This suggests that the transient loss of Xist expression occurs independently of the status of Tsix expression from the Xi.
Epigenetic dynamics during peri-implantation development
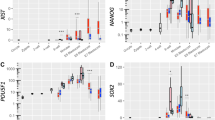
To examine global epigenomic changes during peri-implantation development, we performed immunofluorescence analysis on modifications of DNA, i.e., 5-methylcytosine (5 mC) and 5-hydroxymethylcytosine (5 hmC) (Fig. 3a) as well as histone modifications such as di- and trimethylation of histone H3 lysine9 (H3k9me2 and me3) (Fig. 3b) and trimethylation of histone H3 lysine 27 (H3K27me3) (Fig. 3c).
Epigenetic dynamics during peri-implantation development. (a–c) Immunofluorescence analysis of epigenetic modifications in peri-implantation embryos using antibodies against 5 mC, 5 hmC (a) H3K9me2, H3K9me3 (b) and H3K27me3. (c) The green and magenta colors indicate immunofluorescence signals of 5mC and each other modification, respectively and white dotted circles indicate the location of the ICM/epiblast lineage cells. 5 mC and 5 hmC staining were performed in the same embryos. The images of H3K9me2, H3K9me3, and H3K27me3 are overlapped with nuclear staining (white). Magnified views of epiblast and ExE cells at E5.25–5.5 (a,b) and E4.5, 4.75, and E5.25–5.5 (c) are shown on the right side (a,b) and bottom of each image (c) respectively. (d) The images of DAPI staining at E4.5 and 5.5. Magnified views of 1: ICM/epiblast, 2: PE/VE and 3: TE/ExE cells at each developmental stage are shown. The yellow and white scale bars indicate 25 and 5 μm, respectively.
As shown in Fig. 3a,b, the levels of 5 mC, 5 hmC, H3K9me2 and H3K9me3 in ICM/epiblasts were comparable with or somewhat lower than those in ExE at E4.5 and E4.75, whereas the signals appeared to be more prominent in epiblasts relative to ExE at E5.25–5.5. The results suggest that changes in the four epigenetic modifications showed a similar trend. Namely, global levels of the modifications increase after E4.75.
In contrast to the four modifications mentioned above, the H3K27me3 modification behaves differently (Fig. 3c). At E3.5, H3K27me3 staining signals were detected as a single, heavy deposit probably corresponding to Xi over a relatively weak background in each ICM and TE cell. However, at E4.5, H3K27me3 modifications were globally upregulated specifically in ICM and this global upregulation was continuously observed at E4.75 and E5.25–5.5. Therefore, upregulation of global H3K27me3 level in epiblast initiated earlier compared to the increase in other four modifications. Interestingly, the H3K27me3 staining was seen as particulate structures distinct from chromocenters as shown in Fig. 3c.
In addition, we noted that the heterochromatin organization, revealed by DAPI staining, changed dramatically in different cell lineages at different developmental stages (Fig. 3d). The number and morphologies of DAPI-dense chromocenters were roughly similar to each other between ICM, PE, and TE cell lineages at E4.5. However, at E5.5, each cell in different lineages appeared to develop unique heterochromatin organization. In epiblasts, chromocenters were distributed over the entire nucleus, whereas VE lineage cells were characterized by smaller chromocenters distributed along the nuclear membrane and ExE lineage cells showed relatively few and large chromocenters. These observations suggest that distinct epigenetic status and chromatin organizations characteristic to these three lineages are established in a relatively short period during peri-implantation development.
Discussion
We traced Xist/Tsix expression during peri-implantation embryonic development and attempt to address assessment of XCI dynamics in vivo using whole-mount 3D RNA-FISH (Fig. 4). The results showed that Xist erasure began at E3.75 and was completed by E4.5, whereas the cells that exhibited biallelic expression of Tsix and no expression of Xist became dominant at the following stage between E4.75 and E5.0. The expression patterns of two X-linked genes, Pgk1 and Lamp2, were also examined at E4.75. Proportions of cells showing biallelic expression of these genes were very similar to that of cells exhibiting Tsix biallelic expression, indicating that X reactivation certainly occurred for X-linked genes other than Tsix. Borensztein et al. clearly demonstrated that the timing of reactivation varied among individual X-linked genes and classified X-linked genes into ‘early reactivated (reactivated from E3.5)’, ‘late reactivated (from E4.0)’ and ‘very late reactivated’. The ‘very late reactivated’ genes were not reactivated even at E4.016. Pgk1 gene is classified as late reactivated gene, and not fully reactivated at E4.0. Here, we demonstrated that Pgk1, one of the late reactivated genes, was activated in about half of the embryonic lineage cells at E4.75. Tsix was activated in only ~15% of the ICM cells at E4.0–E4.25 and ~50% at E4.75, suggesting that Tsix is categorized into the ‘late reactivated’ genes or possibly later than “late reactivated” genes. Combined, reactivation of Pgk1, Tsix and Lamp2 likely occurs at the late-to-very late stage of XCI reversal. The cells exhibiting two Tsix without Xist signals may not always mean X-reactivated state. Nevertheless, as all of X-linked genes including Tsix should be biallelically expressed in the cells subjected to XCI reversal, biallelic expression of Tsix without Xist is, at least, necessary condition for X-reactivated state. Therefore, based on the Xist/Tsix expression pattern as well as results of two other X-linked genes, we think that the proportion of X-reactivated cells in the embryonic lineage appears to peak at E4.75–5.0. We found that, already at E4.75, a few epiblast cells exhibited weak Xist expression while the same cells continued to show Tsix biallelism. Such cells are considered to have initiated random XCI. These cells showing ‘early’ Xist expression were not clustered or positioned spatially in specific regions of the embryos. The numbers of these cells further increased until E5.0, and then Tsix biallelic expression declined at later stages.
A model of Xist/Tsix expression dynamics during the development of peri-implantation mouse embryos. (1) Imprinted XCI; the Xp is inactivated and a Xist cloud is observed on it, whereas Tsix shows a single spot on the maternal X (blue region). (2) Cessation of Xist expression commences at E3.75, and the Xist cloud disappears by E4.5 (light pink region). (3) The silenced Tsix gene is derepressed on the Xp in the embryonic lineage. Most cells show biallelic Tsix expression between E4.75 and E5.0 (deep pink region). Cells undergoing X-reactivation are included in this region. (4) A single Xist spot, rather than a cloud, appears in a small percentage of cells showing Tsix biallelic expression (green region). (5) A single Xist spot or Xist cloud is observed in cells showing monoallelic Tsix expression. Generally, Xist and Tsix are expressed in a mutually exclusive manner on X chromosomes. However, a significant number of cells exhibit double Xist signals. These ‘double Xist’ cells appear transiently in E5.25–5.5. (6) Random XCI is fully established by E6.5.
At E6.5, almost all of the cells displayed a single Xist cloud and cells with biallelic Tsix expression were absent. Using the same RNA-FISH technique, it was demonstrated that there was no epiblast cell exhibiting biallelic expression of a X-linked gene, Mecp2, at E6.530. These results suggest that random XCI is established by E6.5. Previous cytogenetic studies based on Kanda staining31 or DNA replication timing32 demonstrated that random XCI is established by E5.5 or around E6.5, respectively. Those results were generally consistent with ours based on Xist/Tsix expression. The slight differences in the timing found between studies might have been caused by differences in the genetic background of the strains used or could be a reflection of the use of different indices representing different stages of the random XCI process.
We also found that X-reactivation seemed to progress in a cell division-independent manner. Although NANOG-positive ICM cells did not proliferate actively from E3.75 to E4.5 (Table 1), imprinted XCI reversal can progress at these stages. Conversely, after E4.75, when the epiblast cells resume Xist expression to establish random XCI, cell proliferation was accelerated dramatically (Table 1). Such a XCI reversal event is also observed during PGC development and is accompanied by genome-wide epigenetic reprogramming26,33,34. Here, we demonstrated that a large-scale change in epigenetic modifications and in chromatin organizations also occurred between E3.5 and E5.5, in particular during E4.75 and E5.25–5.5 when Xist accumulation on one of the two X chromosomes is progressively established. Interestingly, H3K27me3 upregulation in the ICM/epiblast lineage, in contrast to all the other epigenetic modifications examined, was observed between E3.5 and E4.5. This developmental period exactly matches the period of Xist erasure. It has been reported that H3K27me3 is required for Tsix-independent Xist repression in naïve ES cells35. In addition, Inoue et al. demonstrated that H3K27me3 is required for maternal Xist silencing in imprinted XCI during pre-implantation development12. Therefore, it is possible that paternal Xist repression observed in ICM/epiblasts may be caused by H3K27me3 modifications laid onto paternal Xist regulatory regions after E3.5. These results indicate that the major de- and reprogramming events of the epigenomic status occur at around the peri-implantation stage, and thus the conversion of XCI regulation from imprinted to the random process seems to be one of those major epigenomic events.
Here we showed that Xist erasure occurred without any activation of Tsix from the Xi, suggesting that Tsix expression is not involved in initiating the erasure of the Xist RNA. Forced induction of Tsix expression caused Xist repression in cis, resulting in Xi reactivation in a lineage- and stage-specific manner28. In contrast with this ‘gain-of-function’ type of experiment, Maclary et al. showed that XCI reversal in the ICM could occur in the absence of functional Tsix, suggesting that Tsix is dispensable for X chromosome reactivation at this stage36. Moreover, Payer et al. observed a loss of H3K27me3 enrichment on the Xi of epiblast cells in Tsix-null mutant embryos, although the loss was significantly delayed in the mutants compared with normal embryos37. The results of these two studies based on Tsix null mutants were consistent with our findings. Thus, the onset of the loss of inactive markers of the Xi can occur in the absence of Tsix expression. Pasque et al. demonstrated that Tsix expression is dispensable for XCI reversal during the reprogramming of MEFs to induced pluripotent stem cells (iPSCs)38. Therefore, is Tsix not essential for X chromosome reactivation at all? Considering that the X reactivation process can be divided into at least two phases—the initiation of release from the inactive state and the establishment of epigenetic equivalency between the two X chromosomes—Tsix is probably not involved in the former phase, but is likely to be involved in the latter, thus assuring the epigenetic parity of both X chromosomes, as suggested by Navarro et al.29.
Intriguingly, here we showed that the biallelic expression of Xist, including two Xist clouds, could take place at the onset of random XCI during normal mouse embryonic development. In general, to examine the ploidy of those cells, a combined analysis of RNA- and DNA-FISH can be used. For the combined analysis, samples are fixed and anchored on a glass slide so that DNA-FISH images are superimposed to the recorded RNA-FISH signals. However, as the embryos examined in our whole-mount protocol were not anchored, we could not perform a combined analysis of RNA- and DNA-FISH. Alternatively, we searched potential triploidy or higher ploidy cells exhibiting one or more Tsix signals localized distantly from two Xist clouds. However, we rarely found such cases. Therefore, we think that the cells with two Xist clouds were likely to be normal diploid cells. This conclusion is also supported by in vitro studies performed by other groups: biallelic expression of Xist was detected in a subset of differentiating diploid ES cells39,40,41,42; live-cell imaging of the fluorescently tagged nascent Xi in vitro showed survival of some cells with two Xi signals41; and two Xist signals could be also detected in diploid ICM cells allowed to differentiate in vitro40. In addition, ectopic expression of Xist was also observed in epiblast cells of male embryos at the same stage (Supplementary Fig. S4) although frequency of detection was much lower in male (1.2 cells per embryo at E5.5 (n = 5)) than in female (6 cells per embryo at E5.5 (n = 4)). This trend is also true for differentiating ES cells40,42. Therefore, it is possible that such biallelic expression of Xist in female, and also ectopic Xist expression in male, can occur in the normal diploid cells during peri-implantation development in mice. Similar results have been reported recently by Sousa et al.42. At E4.75 when re-expression of Xist began to be observed in the ICM/epiblast cells of female embryos, the cells exhibiting biallelic expression of Xist already appeared. In addition, the percentage of the cells showing asymmetric expression of Xist, e.g. one pinpoint (or dispersed) and one cloud, was lower than that of the cells showing two Xist pinpoints (or dispersed) signals or two clouds (Supplementary Fig. S3). Therefore, it appears that the Xist biallelism is caused by simultaneous Xist activation from both alleles rather than an occasional up-regulation of the second Xist allele in the cells that have already initiated Xist expression from one allele. It was unexpected and intriguing observation, because Xist RNA accumulation on the future Xi is supposed to occur after determining the number of X chromosomes and election of the future Xi and Xa43. Since the present study only provides snap shots of initiation process of random XCI, longitudinal observations using different techniques will be needed to delineate how exactly random XCI initiates or to determine the fate of cells with Xist biallelic expression. Biallelic Xist activation can be found in a high proportion of cells during early development in human and rabbit which have no imprinted XCI mechanism44. Although these phenomena are superficially similar to the one in mice, Xist biallelism in mice may have different biological significance. Proportion of the epiblast cells exhibiting Xist biallelic expression in post-implantation epiblast of mice is low compared to those in human pre-implantation embryo (at most 15% in mice vs >80% in human) and cells showing Xist biallelic expression were only transiently detected (from E5.25 and E5.5), whereas Xist biallelism is observed in relatively long period of human preimplantation development, implying that the transient Xist biallelic expression may be harmful for mouse epiblast cells, thereby causing XCI on both X chromosomes. On the other hand, Xist accumulation does not induce chromosome-wide XCI in human preimplantation embryos44. Interestingly, as described above, ectopic expression of Xist in mice was observed at almost the same stage in both female and male epiblast cells. It is thus possible to speculate that such ectopic Xist expression in mice might be a reflection of large-scale changes in epigenomic states and/or in nuclear architecture that occur in both male and female embryos at a similar developmental stage corresponding to the beginning of random XCI establishment in female embryos.
Our in vivo analysis revealed that Xist/Tsix expression dynamics were significantly different among the three different embryonic lineages studied: ICM/epiblast, TE/ExE, and PE/VE. Interestingly, Xist expression was lost transiently in some ExE cells, from E4.0 to E5.0, reaching a peak of 21% at E4.75. One possible explanation for the loss of Xist clouds is that those cells were in early G1 phase because Xist clouds may not be observed in early G1 phase depending on pretreatment (permeabilization) of the cells prior to RNA-FISH45. As another explanation, we speculate that loss of Xist clouds is possibly linked to the change in regulatory mechanism of imprinted XCI in the ExE lineage. Imprinted Xp inactivation is thought to continue in ExE cells. However, the imprinted XCI can be reversed in a subset of female ExE cells that carry a paternally derived Xist mutation, suggesting that the primary imprint for imprinted XCI, which is likely to be H3K27me3 modification in maternal Xist locus12, might be erased in ExE cells46. Oikawa et al. also demonstrated that the primary imprint mark must be erased in the TE/ExE cells of E4.5 embryos by nuclear transfer experiments10. These results suggest that imprinted XCI is somehow maintained during ExE development, although its regulatory mechanism might be converted to a different one. The timing of the transient loss of Xist in ExE cells, shown in this study, roughly matches the transition period of the mechanism for imprinted XCI regulation in ExE cells suggested by Oikawa et al.10. Therefore, we speculate that the transient loss of Xist observed in the ExE might be involved in the conversion of the imprinted XCI regulatory mechanism in the ExE lineage.
Methods
Mice
The mouse strain C57BL/6J was used throughout the experiments. As an exception, embryos with mixed genetic background, BDF1xC57BL/6J, were used for Lamp2 and Pgk1 RNA-FISH experiments, because only very faint or no FISH signals were detected around E4.75 when we used embryos of C57BL/6J strain. E3.5–5.5 embryos were recovered at intervals of 6 h from naturally mated female mice. Noon was set as E0.5 on days when vaginal plugs were detected. We collected embryos between E3.5 and E4.0 by flushing the uteri. In many studies, E4.5 stage embryos were obtained by culturing preimplantation blastocyst or by flushing out the uteri. However, we experienced that the number of embryos collected by flushing out around E4.5 stage is substantially smaller than those obtained at E3.5, indicating that embryos firmly attached (=implanted) to the uterus cannot be recovered by flushing out, and it is likely that only embryos of relatively early stage (not yet firmly attached) were collected. Therefore, embryos after E4.25 were collected by manual dissection. The collected embryos were used immediately for the subsequent analysis without embryo culture. All animal experiments were performed in accordance with the guidelines for the experimental use and care of laboratory animals of the RIKEN Tsukuba Institute with prior approval from the Institutional Animal Experiment Committee of the RIKEN Tsukuba Institute.
Strand-specific probe synthesis for whole-mount 3D RNA-FISH
Strand-specific DNA probes for detecting the Xist, Tsix, Lamp2 and Pgk1 RNAs were prepared according to our published protocol47. For details, see the Supplementary Methods.
Expression analysis of Xist/Tsix via whole-mount 3D RNA-FISH combined with immunofluorescence
Whole embryos were dissected then treated with 0.1% Triton X-100 in phosphate-buffered saline (PBS) for 10 s on ice for permeabilization. After fixation with 4% paraformaldehyde in PBS with 0.1% Triton X-100 for 10 min at room temperature, RNA-FISH and immunofluorescence analyses were performed. Details of the procedure, including the antibodies used, are provided in the Supplementary Methods.
Analysis of Xist/Tsix expression patterns
Xist and Tsix expression patterns in individual cells from each lineage were examined as follows. The whole embryo was subjected to Xist/Tsix RNA-FISH followed by immunostaining with a marker that was specific to a particular lineage: POU5F1 or NANOG for ICM/embryonic ectoderm, GATA6 for PE/VE, and CDX2 for TE/ExE. Images were acquired using confocal microscopy. Optical sections of the confocal images were examined manually, and Xist/Tsix expression patterns in each cell of the embryos were determined by considering the overlaps of cellular images between the z-stack images. Xist expression patterns in each cell were classified into four categories: (1) cells with one Xist cloud per nucleus; (2) cells with single pinpoint or dispersed signals; (3) cells with two Xist signals; or (4) cells with no Xist signal. Tsix-positive cells were categorized into cells with two, one, or no signals. The combinations of Xist and Tsix expression patterns in each cell were determined, and the proportion of each combination was calculated and shown as a matrix. We determined Xist and Tsix expression patterns and their combinations in almost all the cells that constituted a whole embryo at a particular stage. We performed these analyses for nine different embryonic stages and for three different cell lineages of each stage. We repeated these experiments at least three times for each embryonic stage. At E6.5, more than 250 embryonic ectoderm cells in total were randomly chosen from two embryos (115 and 139 cells, respectively) and their Xist and Tsix expression patterns were examined. Confocal image data used in this study have been deposited to Systems Science of Biological Dynamics (SSBD) database (http://ssbd.qbic.riken.jp) and will be open to the public.
Immunofluorescence analysis of epigenetic modifications
Whole embryos were dissected, treated with 0.1% Triton X-100 in PBS for 10 s on ice for permeabilization, and fixed with 4% paraformaldehyde in PBS with 0.1% Triton X-100 for 10 min at room temperature followed by immunofluorescence analysis. For details, including antibody information, see the Supplementary Methods.
References
Lyon, M. F. Gene action in the X-chromosome of the mouse (Mus musculus L.). Nature 190, 372–373 (1961).
Borsani, G. et al. Characterization of a murine gene expressed from the inactive X chromosome. Nature 351, 325–329 (1991).
Brockdorff, N. et al. Conservation of position and exclusive expression of mouse Xist from the inactive X chromosome. Nature 351, 329–331 (1991).
Penny, G. D., Kay, G. F., Sheardown, S. A., Rastan, S. & Brockdorff, N. Requirement for Xist in X chromosome inactivation. Nature 379, 131–137 (1996).
Marahrens, Y., Panning, B., Dausman, J., Strauss, W. & Jaenisch, R. Xist-deficient mice are defective in dosage compensation but not spermatogenesis. Genes Dev. 11, 156–166 (1997).
Lee, J. T., Davidow, L. S. & Warshawsky, D. Tsix, a gene antisense to Xist at the X-inactivation centre. Nat. Genet. 21, 400–404 (1999).
Stavropoulos, N., Lu, N. & Lee, J. T. A functional role for Tsix transcription in blocking Xist RNA accumulation but not in X-chromosome choice. Proceedings of the National Academy of Sciences of the United States of America 98, 10232–10237 (2001).
Borensztein, M. et al. Xist-dependent imprinted X inactivation and the early developmental consequences of its failure. Nat. Struct. Mol. Biol. 24, 226–233 (2017).
Tada, T. et al. Imprint switching for non-random X-chromosome inactivation during mouse oocyte growth. Development 127, 3101–3105 (2000).
Oikawa, M. et al. Understanding the X chromosome inactivation cycle in mice: a comprehensive view provided by nuclear transfer. Epigenetics 9, 204–211 (2014).
Okamoto, I. et al. Evidence for de novo imprinted X-chromosome inactivation independent of meiotic inactivation in mice. Nature 438, 369–373 (2005).
Inoue, A., Jiang, L., Lu, F. & Zhang, Y. Genomic imprinting of Xist by maternal H3K27me3. Genes Dev. 31, 1927–1932 (2017).
Mak, W. et al. Reactivation of the paternal X chromosome in early mouse embryos. Science 303, 666–669 (2004).
Okamoto, I., Otte, A. P., Allis, C. D., Reinberg, D. & Heard, E. Epigenetic dynamics of imprinted X inactivation during early mouse development. Science 303, 644–649 (2004).
Williams, L. H., Kalantry, S., Starmer, J. & Magnuson, T. Transcription precedes loss of Xist coating and depletion of H3K27me3 during X-chromosome reprogramming in the mouse inner cell mass. Development 138, 2049–2057 (2011).
Borensztein, M. et al. Contribution of epigenetic landscapes and transcription factors to X-chromosome reactivation in the inner cell mass. Nature communications 8, 1297, https://doi.org/10.1038/s41467-017-01415-5 (2017).
Mohammed, H. et al. Single-Cell Landscape of Transcriptional Heterogeneity and Cell Fate Decisions during Mouse Early Gastrulation. Cell reports 20, 1215–1228 (2017).
Nichols, J. & Smith, A. Naive and primed pluripotent states. Cell stem cell 4, 487–492 (2009).
Bao, S. et al. Epigenetic reversion of post-implantation epiblast to pluripotent embryonic stem cells. Nature 461, 1292–1295 (2009).
Hiratani, I. et al. Genome-wide dynamics of replication timing revealed by in vitro models of mouse embryogenesis. Genome Res. 20, 155–169 (2010).
Smith, Z. D. et al. A unique regulatory phase of DNA methylation in the early mammalian embryo. Nature 484, 339–344 (2012).
Brockdorff, N. et al. The product of the mouse Xist gene is a 15 kb inactive X-specific transcript containing no conserved ORF and located in the nucleus. Cell 71, 515–526 (1992).
Brown, C. J. et al. The human XIST gene: analysis of a 17 kb inactive X-specific RNA that contains conserved repeats and is highly localized within the nucleus. Cell 71, 527–542 (1992).
Clemson, C. M., McNeil, J. A., Willard, H. F. & Lawrence, J. B. XIST RNA paints the inactive X chromosome at interphase: evidence for a novel RNA involved in nuclear/chromosome structure. J. Cell Biol. 132, 259–275 (1996).
Csankovszki, G., Panning, B., Bates, B., Pehrson, J. R. & Jaenisch, R. Conditional deletion of Xist disrupts histone macroH2A localization but not maintenance of X inactivation. Nat. Genet. 22, 323–324 (1999).
Sugimoto, M. & Abe, K. X chromosome reactivation initiates in nascent primordial germ cells in mice. PLoS Genet. 3, e116, https://doi.org/10.1371/journal.pgen.0030116 (2007).
Sado, T., Hoki, Y. & Sasaki, H. Tsix silences Xist through modification of chromatin structure. Dev. Cell 9, 159–165 (2005).
Ohhata, T., Senner, C. E., Hemberger, M. & Wutz, A. Lineage-specific function of the noncoding Tsix RNA for Xist repression and Xi reactivation in mice. Genes Dev. 25, 1702–1715 (2011).
Navarro, P., Pichard, S., Ciaudo, C., Avner, P. & Rougeulle, C. Tsix transcription across the Xist gene alters chromatin conformation without affecting Xist transcription: implications for X-chromosome inactivation. Genes Dev. 19, 1474–1484 (2005).
Hosoi, Y. et al. Female mice lacking Ftx lncRNA exhibit impaired X-chromosome inactivation and a microphthalmia-like phenotype. Nature communications 9, 3829, https://doi.org/10.1038/s41467-018-06327-6 (2018).
Rastan, S. Timing of X-chromosome inactivation in postimplantation mouse embryos. J Embryol Exp Morphol 71, 11–24 (1982).
Takagi, N., Sugawara, O. & Sasaki, M. Regional and temporal changes in the pattern of X-chromosome replication during the early post-implantation development of the female mouse. Chromosoma 85, 275–286 (1982).
Seki, Y. et al. Extensive and orderly reprogramming of genome-wide chromatin modifications associated with specification and early development of germ cells in mice. Dev. Biol. 278, 440–458 (2005).
Seki, Y. et al. Cellular dynamics associated with the genome-wide epigenetic reprogramming in migrating primordial germ cells in mice. Development 134, 2627–2638 (2007).
Ohhata, T. et al. Histone H3 Lysine 36 Trimethylation Is Established over the Xist Promoter by Antisense Tsix Transcription and Contributes to Repressing Xist Expression. Mol. Cell. Biol. 35, 3909–3920 (2015).
Maclary, E. et al. Differentiation-dependent requirement of Tsix long non-coding RNA in imprinted X-chromosome inactivation. Nature communications 5, 4209, https://doi.org/10.1038/ncomms5209 (2014).
Payer, B. et al. Tsix RNA and the germline factor, PRDM14, link X reactivation and stem cell reprogramming. Mol. Cell 52, 805–818 (2013).
Pasque, V. et al. X chromosome reactivation dynamics reveal stages of reprogramming to pluripotency. Cell 159, 1681–1697 (2014).
Augui, S. et al. Sensing X chromosome pairs before X inactivation via a novel X-pairing region of the Xic. Science 318, 1632–1636 (2007).
Monkhorst, K., Jonkers, I., Rentmeester, E., Grosveld, F. & Gribnau, J. X. inactivation counting and choice is a stochastic process: evidence for involvement of an X-linked activator. Cell 132, 410–421 (2008).
Guyochin, A. et al. Live Cell Imaging of the Nascent Inactive X Chromosome during the Early Differentiation Process of Naive ES Cells towards Epiblast Stem Cells. Plos one 9, e116109, https://doi.org/10.1371/journal.pone.0116109 (2014).
Sousa, E. J. et al. Exit from Naive Pluripotency Induces a Transient X Chromosome Inactivation-like State in Males. Cell stem cell 22, 919–928 (2018).
Avner, P. & Heard, E. X-chromosome inactivation: counting, choice and initiation. Nat. Rev. Genet. 2, 59–67 (2001).
Okamoto, I. et al. Eutherian mammals use diverse strategies to initiate X-chromosome inactivation during development. Nature 472, 370–374 (2011).
Jonkers, I. et al. Xist RNA is confined to the nuclear territory of the silenced X chromosome throughout the cell cycle. Mol. Cell. Biol. 28, 5583–5594 (2008).
Mugford, J. W., Yee, D. & Magnuson, T. Failure of extra-embryonic progenitor maintenance in the absence of dosage compensation. Development 139, 2130–2138 (2012).
Shiura, H., Sakata, Y., Abe, K. & Sado, T. RNA-FISH and Immunofluorescence of Mouse Preimplantation and Postimplantation Embryos. Methods in molecular biology 1861, 161–176 (2018).
Acknowledgements
We are grateful to Dr. T. Sado for critical reading of this manuscript. We thank Dr. S. Suzuki for the mouse work. We especially acknowledge all the members of the K.A. laboratory for their advice and suggestions. This work was supported by RIKEN Special Postdoctoral Researchers Program to H.S. and in part by a grant to K.A. from the Ministry of Education, Culture, Sports, Science and Technology of Japan.
Author information
Authors and Affiliations
Contributions
H.S. and K.A. conceived the project and wrote the manuscript. H.S. performed all the experiments.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Shiura, H., Abe, K. Xist/Tsix expression dynamics during mouse peri-implantation development revealed by whole-mount 3D RNA-FISH. Sci Rep 9, 3637 (2019). https://doi.org/10.1038/s41598-019-38807-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-38807-0
This article is cited by
-
GATA transcription factors drive initial Xist upregulation after fertilization through direct activation of long-range enhancers
Nature Cell Biology (2023)
-
Preventing erosion of X-chromosome inactivation in human embryonic stem cells
Nature Communications (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.