Abstract
We report that long double-stranded DNA confined to quasi-1D nanochannels undergoes superdiffusive motion under the action of the enzyme T4 DNA ligase in the presence of necessary co-factors. Inside the confined environment of the nanochannel, double-stranded DNA molecules stretch out due to self-avoiding interactions. In absence of a catalytically active enzyme, we see classical diffusion of the center of mass. However, cooperative interactions of proteins with the DNA can lead to directed motion of DNA molecules inside the nanochannel. Here we show directed motion in this configuration for three different proteins (T4 DNA ligase, MutS, E. coli DNA ligase) in the presence of their energetic co-factors (ATP, NAD+).
Similar content being viewed by others
Introduction
The statistical mechanics of DNA confined to long and narrow nanofluidic channels have been explored extensively, and an increasingly complete picture is emerging1,2,3,4,5,6. For the purpose of this paper, we only consider quasi-onedimensional channels with a cross-section on the order of 100 × 100 nm2, and tens to hundreds of microns long. In these nanochannels, DNA is stretched out to yield a constant stored contour length of DNA per unit distance along the channel axis. The mapping of genetic landmarks on nanochannel-stretched DNA has become sufficiently mature to meaningfully contribute to genome science at large7,8. Epigenetic profiling is emergent9,10,11,12.
The stretching technique has been extended from DNA to DNA-protein complexes, and the analysis of compaction or extension of nucleic acids by DNA-binding proteins has been developed into an analytic technique with considerable power13,14,15,16,17. The majority of studies of DNA-protein interactions are strictly in equilibrium, and even studies that are not in equilibrium are performed in a regime where reversibility can reasonably be assumed18.
However, this assumption of reversibility does not hold for protein-DNA interactions where enzymatic activity coupled with modification of covalent chemical bonds is a key feature. For instance, the activity of endonucleases is profoundly irreversible, and has been monitored within nanochannels19,20. It appears though that endonucleases are not a good model for a general DNA-modifying enzyme because the energy source is the DNA itself. In contrast, wide classes of DNA-modifying enzymes such as helicases, isotopomerases and ligases utilize adenosine triphosphate (ATP) or a similar energetic compound to establish the preferred direction of the reaction cascade21.
In this publication, we present a detailed test of one such enzyme, T4 DNA ligase, and an exploratory overview of others. We find that the catalytic activity is linked to a superdiffusive displacement of the DNA molecule within the nanochannel that is consistent with a constant mean velocity for each molecule. We proceed to propose a speculative model for the superdiffusive motion. While the motion superficially resembles that of a protein filament transported by a set of surface-tethered proteins22,23,24, we argue that a motor-like action in that narrow sense is not present here. Instead, we propose that the transport effect is due to self-maintaining concentration polarization of ions within the channel.
Results
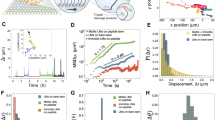
Experiments were performed with fluorescently stained λ-DNA in Tris-Borate (TB) buffer. The buffer was supplemented with T4 DNA ligase, ATP, MgCl2 and EDTA if required. The sample was introduced into large microchannels (1 × 100 μm2 cross-section, 1 cm long) of the fused silica device (Fig. 1A), and then driven into the nanochannels (100 × 100 nm2 cross-section, 200 μm long) using pressure. The pressure was appropriately low such that the majority of molecules were introduced in a linear configuration25, and any folded molecules were discarded.
(A) Schematic of the device. (B,C) Kymographs showing the fluorescent intensity along nanochannel axis as a function of time for bare λ-DNA (B) and λ-DNA with T4 DNA ligase in a catalytically active buffer (C), respectively. (D) Center of mass of the molecules in (B) (red, diamonds) and (C) (blue, circles) as function of time.
Figure 1B shows a kymograph of the intensity along a nanochannel axis after a single λ-DNA molecule was introduced and the pressure was removed. No T4 DNA ligase, Mg2+, or ATP was present. We observe the commonly reported intensity and length fluctuations linked to internal fluctuations, and an apparently random displacement of the center of mass of the molecule. Figure 1C shows the outcome of the same experimental procedure, but with T4 DNA ligase, Mg2+, and ATP present. DNA appears somewhat less extended - an effect previously attributed to the presence of ATP26 (see Supp. Mat. C for length analysis). While this molecule undergoes the same internal and length fluctuations as the molecule in Fig. 1B, its center of mass mass appears to drift into one direction.
We determined the center of mass and extension for each video frame by fitting the brightness along the molecule backbone to the convolution of a Gaussian and a boxcar function1. In Fig. 1D, we compare the center of mass position for the two molecules presented in Fig. 1B and Fig. 1C. This analysis was performed for all molecules of the set, and we find in general that DNA with catalytically active T4 DNA ligase (Fig. 1C) undergoes directed motion. The motion appears locally piecewise linear.
The natural question arising is whether the motion under protein action can be described by a directed translocation, a classical diffusion with increased diffusion coefficient (when compared to DNA without any cofactors), or an entirely different process. To answer this question, we have prepared mean square displacement (MSD) graphs for both DNA without T4 DNA ligase or co-factors and with T4 DNA ligase and co-factors over a set of 20 molecules for each condition (Fig. 2). If the motion were governed by classical diffusion, we would anticipate that the MSD grows linearly with time, i.e. \(\langle {\rm{\Delta }}{x}^{2}({\rm{\Delta }}{t})\rangle \propto {\rm{\Delta }}{{t}}^{{\alpha }}\) with α = 1. If it grows as a power law with \(\alpha > 1\) we call the motion “superdiffusive”, and if α = 2 we have a the special case of directed translocation in a strict sense with a well-defined mean speed.
Average mean-square displacement curves calculated from the center of mass position for 20 molecules for each condition. Blue circles correspond to data DNA with T4 DNA ligase in the presence of its cofactors, while red diamonds indicate bare DNA. Error bars are the standard deviation between molecules of the experimental set, and are depicted only for select data points to illustrate the trend. The continuous and dashed curves correspond to the fits for the conditions as described in the text.
The graph for protein-free DNA in Fig. 2 shows a well defined single-power-law relationship \(\langle {\rm{\Delta }}{x}^{2}({\rm{\Delta }}t)\rangle =\) \(\mathrm{(0.21}\pm \mathrm{0.01)}\mu {m}^{2}{({\rm{\Delta }}t\frac{1}{s})}^{1.04\pm 0.01}\) (error is the 95% confidence interval of the fit). The curve is evidently consistent with classical diffusion, except for a mildly subdiffusive regime at very small times that likely is caused by fitting errors. The MSD curve for DNA with catalytically active T4 DNA ligase clearly shows superdiffusive behavior. We find a good fit to the data by using a double-power law of the form \(\langle {\rm{\Delta }}{x}^{2}({\rm{\Delta }}t)\rangle =\mathrm{(0.23}\pm \mathrm{0.01)}\mu {m}^{2}\) \({({\rm{\Delta }}t\frac{1}{s})}^{0.77\pm 0.01}+\mathrm{(0.45}\pm \mathrm{0.01)}\mu {m}^{2}{({\rm{\Delta }}t\frac{1}{s})}^{1.96\pm 0.01}\), indicating translocation at constant speed of 0.7 μm/s for long times, and a mildly subdiffusive regime at very short times. For times between 1 s and 15 s, the motion is very well described by directed translocation in a single direction.
For finding a working hypothesis for the translocation of DNA in presence of catalytically active T4 DNA ligase, we first evaluated whether the relatively complex buffer system could be a source of the motion (Table 1 and Supp. Mat. D.i.). We found that all of the buffer components alone, as well as combinations of components, did not induce a superdiffusive motion of DNA within the nanochannel. We then evaluated whether the presence of the protein itself could be a source of translocation. We tested T4 DNA ligase without any co-factors, with only ATP (as in a prior publication18), with only Mg2+ and EDTA, and with the full buffer system as described above. Only the full system supported translocation. We further note that using human genomic DNA also gave rise to the directed translocation, indicating that the motion is not substrate-specific.
It would seem that DNA is transported in a fashion superficially similar to a system in which motor proteins are coupled to the surface of a channel with sub-persistence length width in which a filament is transported on its surface24. A preliminary, and very unlikely hypothesis thus would require T4 DNA ligase to have the action of a motor protein. While T4 DNA ligase is a widely-used “workhorse” enzyme, it is not as well-studied as T7 DNA ligase27. Its known functions include blunt-ended ligation, ligation coupled to hybridization, lyase activity, nick-sealing, gapped-ligase, and single-stranded ligation28,29,30,31,32. The influence of adenylation of T4 DNA ligase (through ATP) and the impact of Mg2+ on binding and catalytic activity have been studied33,34,35. However, under no circumstance has a motor-like search mechanism coupled to ATP hydrolysis been even suggested. We thus decided to test whether two other DNA-binding proteins with known catalytic activity coupled to an energetic compound could cause the directed motion.
E. coli DNA ligase is an interesting alternative to T4 DNA ligase as it utilizes NAD+ instead of ATP for adenylation36. It also likely has fewer functions than T4 DNA ligase and thus presents a “cleaner” system. Using λ-DNA and the manufacturer-supplied Tris-based buffer supplemented with 2 mM ATP, we find that E. coli DNA ligase does catalyze translocation in presence of NAD+ with a close-to linear displacement versus time relationship but about half the translocation speed observed using T4 DNA ligase (Table 1 and Supp. Mat. D.iii.). In a buffer that contained ATP instead of NAD+, double-power law curve was also required to fit the MSD versus time graph, but even the faster of the two components merely had α = 1.03 indicating classical diffusion and not directed translocation (Table 1 and Supp. Mat. D.iii.). Thus, the translocation appears distinctively linked to the specific interaction of an energetic compound and protein, and not some unspecific interaction of ATP with protein and DNA.
MutS is an extensively characterized protein that is part of the mismatch recognition and repair pathway with an ATPase domain37,38. We repeated the experiment with the same buffer set and substrate as described for T4 DNA ligase, and observed the characteristic directed displacement of DNA due to MutS with a drift velocity similar to the achieved by T4 DNA ligase (Table 1 and Supp. Mat. D.ii.). Using MutS in a buffer containing Mg2+ and EDTA but no ATP did show classical diffusion. It is established that MutS cannot utilize γ-ATP efficiently. We repeated the experiments with γ-ATP instead of ATP, and found strongly retarded directional motion with a dislocation in 10 s more than an order of magnitude lower than with ATP. Since MutS certainly does not contain a motor protein domain, we strongly believe that the translocation of DNA in presence of T4 DNA ligase is also not due to motor protein action.
Discussion
Since we have established that the motion is unlikely due to a distinctive “motor action”, we must propose a mechanism that only employs the interactions of liquid and DNA, mediated by the ions in solution.
We note that the symmetry along the channel axis must be broken, either through composition of the solution, the DNA, or the protein distribution along the channel or the DNA. Furthermore, we have to identify a mechanism that propels the DNA while maintaining the asymmetry along the channel axis. In our case, possible candidates for the asymmetry is the liquid flow during insertion of the DNA molecule into the nanochannel in the form of a concentration polarization of ions, or and stochastic asymmetry in protein concentration of density of nicks along the DNA substrate. Such concentration polarization is well established at nanopore entrances39, and anticipated around DNA molecules40. In particular, ATP, AMP, and PPi will establish a non-homogeneous concentration profile. Once a concentration polarization is established, we follow observation by Pallaci et al. that DNA drifts into the direction of the higher ionic strength41. For colloidal particles the effect is well established together with a well-accepted model42,43, and theoretical description for DNA drift been given by Hsu44. A possible complication here is that of strong confinement of DNA, but we note that our experiment is similar to nanopore experiments in that aspect45. In the supplementary materials (section H) we summarize our experiments in which we placed nanochannel-stretched DNA into an ionic strength gradient with stationary liquid, and observed that DNA indeed drifts towards the high ionic strength nanochannel entrance. Thus, we propose that an imbalance of ionic strength between the termini of the DNA is responsible for the apparent drift.
The imbalance in ionic strength has to be maintained by the enzymatic reaction of T4 DNA ligase. The obvious overall process that changes the ionic strength is the consumption of ATP to form AMP and free pyrophosphate (PPi) that is coupled to nick-sealing34,35. We propose here that under the prevailing conditions (low DNA concentration, incubation of T4 DNA ligase in ATP and Mg2+), the majority of T4 DNA ligase is in an adenylated state. Our experiment proceeds in a fluorescence microscope where DNA nicking is present at all times46, and thus a strong background rate of nick-sealing events should be anticipated. Thus, when the adenylated enzyme encounters a nick site, we propose that that AMP is released, thereby locally raising the ionic strength. The analogue leaving groups to PPi and AMP for the E. coli DNA ligase are β-nicotinamide mononucleotide (β-NMN) and AMP, while the leaving groups for MutS would be ADP and phosphate. This release of reaction products and linked increase of ionic strength would be occurring on the end of DNA that encounters the higher fraction of ligases poised for activity. DNA then would drift towards that terminus, enhancing the fraction of ligases ready for catalytic activity, and thus sustaining the reaction and the drift. The self-consistency of the model was numerically verified (Supp. Mat. I.), and the drift dynamics as well as the ionic profiles around DNA are are show Fig. 3. Note that removal of catalytic activity without any other modification of the model lead to complete ceasing of the drift.
In the description of the data in Fig. 1 we pointed out that the motion appeared piecewise linear in presence of T4 DNA ligase and co-factors. We analyzed this by fitting a the minimal set of piecewise linear functions to the position versus time graphs. The number of fitted segments to a kymograph was chosen such that the average squared error between fit and data became smaller than the expected error of a linear fit to purely diffusing DNA (examples and statistics shown in Supp. Mat. E). We obtain segments on average 12 seconds in duration, about the same time as covered by Fig. 2.
The displacement probability density for a set of 20 DNA molecules presence of catalytically active T4 DNA ligase as function of drift velocity (slope in position versus time plot) is shown in Fig. 4. The histogram is dominated by two maxima at 0.7 ± 0.1 μm/s in the forward/backward direction with a distribution half-width of 0.22 ± 0.07 μm/s. The coincidence of the maximum likely drift speeds in Fig. 4 (forward/backward peak location) and Fig. 2 (drift speed of quadratic component) strengthens our initial interpretation that the motion can be viewed as translocation with constant speed at times up to roughly 10 s. At times longer than that, Fig. 2 has insufficient statistical weight. The long-term behavior is thus best characterized by the transitions between segments the underlie Fig. 4. We categorized segments into forward, stationary, backward by setting thresholds at ±0.2 μm/s, and found that no molecule contained both forward and backward motion. We found that the majority of transitions (32/55) between a moving state and a temporary stall, and the rest contained changes of speed while keeping direction (23/55). Note that an equal number of molecules in Fig. 4 were introduced from both the left and the right. If the entrance direction was taken into account, we observed that the majority of molecules travelled into the direction of introduction.
The robustness of the drift direction raises the question whether it is imprinted to the pair or molecule and channel in some fashion. We probed this by applying ultra-low pressure (<0.1 psi) to push back the stretched DNA against its direction of travel before releasing it. We found that molecules continue to travel in the direction that was initially observed (Supp. Mat. F).
This then suggests that the temporary stall events are due to short times during which the DNA is retarded by temporary adhesion to the channel wall. Tentative indication of this mechanism can be found in Table 1, where all conditions that contain T4 DNA ligase do not show a strict classical diffusion in the short-term limit (or overall), but are instead mildly subdiffusive with α < 1 as would be characteristic in a system retarded by binding traps. We note that the addition of a high concentration of bovine serum albumin (BSA, 0.6 μg/ml) suppressed the drift motion (Table 1). Typically BSA is used to reduce adhesion of proteins to channel sidewalls. Thus it appears feasible that at least part of the adenylated protein that is poised to release AMP is adsorbed onto the wall, but can currently not point to its specific role.
To conclude, we report that DNA translocates through nanofluidic channels in presence of a number of catalytically active enzymes that hydrolyze an energetic compound. We propose a kinetic model that depends on an asymmetry of ionic strengths between the termini of the molecule within the channel. We further propose that the ionic strength imbalance is maintained by the enzymatic reaction. The mechanism is specific to nanochannel confined DNA since confinement causes the macromolecule to sample a spatial zone and prevents rapid dissipation of the ionic disturbance. We propose that this mechanism may be a sensitive assay for hydrolysis of energetic compounds by a wide class of enzymes that utilize DNA as a substrate.
Methods
Biological Materials
λ-DNA (Roche) was stained with YOYO-1 (Invitrogen) at a ratio of 1 dye per 20 base pairs for 48 hours at 4 °C in 0.5× TBE buffer (pH 8). The contour length of unstained λ-DNA (48.5 kbp) is around 16 μm, which increases somewhat after staining47. T4 DNA ligase (Roche) had a final concentration of 40 units/ml in a 0.5× TBE buffer (pH 8) that was modified as follows. In order to obtain catalytically active ligase, 1 mM ATP (Roche) and 5 mM MgCl2 were added as T4 DNA ligase cofactors to catalytically active solutions. This solution was incubated for 1 hour at 16 °C to allow full equilibrization of Mg2+, ATP, and T4 DNA ligase. After this incubation the free Mg2+ concentration was lowered by adding 2 mM EDTA. This is necessary since YOYO-1 is not a stable intercalator at a free Mg2+ concentration of 5 mM. Stained DNA was added to the protein yield a final concentration of 5 μg/mL followed by stabilization for 2 hours at 25 °C. Tests using alternative DNA (human genomic DNA from Roche) and an alternative enzyme provider (New England Biolabs) were conducted to exclude possible contamination. In a subset of experiments, a high concentration (0.6 mg/ml) bovine serum albumin (BSA) was added to prevent sticking to the nanochannel walls.
Devices and observation
All experiments used mixed micro- and nanofluidic devices made from fused silica, which were prepared by methods discussed elsewhere48. Nanofluidic channels with a 100 × 100 nm2 cross-section and a length of 200 μm were placed between microchannels (see Supp. Mat. A.). Molecules were observed using an inverted fluorescence microscope with a 100×, 1.3 N.A. oil-immersion microscope objective under illumination from a metal halide lamp, and observation by EM-CCD camera (Andor) with a frame duration of 90 ms.
Each DNA molecule is driven from microchannel to nanochannel by a static pressure difference of 35 psi applied over the full length of the nanochannel. After the DNA molecule has entered the nanochannel, the pressure gradient is removed and the dynamics of molecules are observed.
Data availability statement
The datasets generated during and analysed during the current study are available from the corresponding author on reasonable request.
References
Tegenfeldt, J. O. et al. The dynamics of genomic-length DNA molecules in 100-nm channels. Proceedings of the National Academy of Sciences of the United States of America 101, 10979–83, https://doi.org/10.1073/pnas.0403849101 (2004).
Reisner, W. et al. Statics and Dynamics of Single DNA Molecules Confined in Nanochannels. Physical Review Letters 94, 196101, https://doi.org/10.1103/PhysRevLett.94.196101 (2005).
Odijk, T. Scaling theory of DNA confined in nanochannels and nanoslits. Physical Review E 77, 060901, https://doi.org/10.1103/PhysRevE.77.060901 (2008).
Muralidhar, A., Tree, D. R., Wang, Y. & Dorfman, K. D. Interplay between chain stiffness and excluded volume of semiflexible polymers confined in nanochannels. Journal of Chemical Physics 140, 084905, https://doi.org/10.1063/1.4865965 (2014).
Gupta, D. et al. Experimental Evidence of Weak Excluded Volume Effects for Nanochannel ConfinedDNA. ACS Macro Letters 4, 759–763, https://doi.org/10.1021/acsmacrolett.5b00340 (2015).
Iarko, V. et al. Extension of nanoconfined DNA: Quantitative comparison between experiment and theory. Physical Review E 92, 1–8, https://doi.org/10.1103/PhysRevE.92.062701 (2015).
Mak, A. C. Y. et al. Genome-Wide Structural Variation Detection by Genome Mapping on Nanochannel Arrays. Genetics 202, 351–62, https://doi.org/10.1534/genetics.115.183483 (2016).
Mostovoy, Y. et al. A hybrid approach for de novo human genome sequence assembly and phasing. Nature Methods. https://doi.org/10.1038/nmeth.3865 (2016).
Cipriany, B. R. et al. Single molecule epigenetic analysis in a nanofluidic channel. Analytical Chemistry 82, 2480–7, https://doi.org/10.1021/ac9028642 (2010).
Lim, S. F. et al. DNA methylation profiling in nanochannels. Biomicrofluidics 5, 034106, https://doi.org/10.1063/1.3613671 (2011).
Matsuoka, T. et al. Nanoscale squeezing in elastomeric nanochannels for single chromatin linearization. Nano Letters 12, 6480–4, https://doi.org/10.1021/nl304063f (2012).
Lim, S. F., Karpusenko, A., Blumers, A. L., Streng, D. E. & Riehn, R. Chromatin modification mapping in nanochannels. Biomicrofluidics 7, 64105, https://doi.org/10.1063/1.4833257 (2013).
Streng, D. E., Lim, S. F., Pan, J., Karpusenka, A. & Riehn, R. Stretching chromatin through confinement. Lab on a Chip 9, 2772, https://doi.org/10.1039/b909217j (2009).
Zhang, C. et al. Effect of H-NS on the elongation and compaction of single DNA molecules in a nanospace. Soft Matter 9, 9593, https://doi.org/10.1039/c3sm51214b (2013).
Frykholm, K. et al. Probing Physical Properties of a DNA-Protein Complex Using Nanofluidic Channels. Small 10, 884–7, https://doi.org/10.1002/smll.201302028 (2014).
Jiang, K. et al. Effects of Hfq on the conformation and compaction of DNA. Nucleic Acids Research 43, 4332–4341, https://doi.org/10.1093/nar/gkv268 (2015).
Frykholm, K. et al. DNA compaction by the bacteriophage protein Cox studied on the single DNA molecule level using nanofluidic channels. Nucleic Acids Research gkw352 https://doi.org/10.1093/nar/gkw352 (2016).
Roushan, M. et al. Probing transient protein-mediated DNA linkages using nanoconfinement. Biomicrofluidics 8, 034113, https://doi.org/10.1063/1.4882775 (2014).
Riehn, R. et al. Restriction mapping in nanofluidic devices. Proceedings Of The National Academy Of Sciences Of The United States Of America 102, 10012–10016, https://doi.org/10.1073/pnas.0503809102 (2005).
Persson, F. et al. Lipid-based passivation in nanofluidics. Nano Letters 12, 2260–5, https://doi.org/10.1021/nl204535h (2012).
Alberts, B. et al. Molecular Biology of the Cell (Garland Science, New York, 2014).
Nicolau, D. V., Suzuki, H., Mashiko, S., Taguchi, T. & Yoshikawa, S. Actin motion on microlithographically functionalized myosin surfaces and tracks. Biophysical Journal 77, 1126–1134, https://doi.org/10.1016/S0006-3495(99)76963-8 (1999).
Hess, H., Clemmens, J., Qin, D., Howard, J. & Vogel, V. Light-Controlled Molecular Shuttles Made from Motor Proteins Carrying Cargo on Engineered Surfaces. Nano Letters 1, 235–239, https://doi.org/10.1021/nl015521e (2001).
van den Heuvel, M. G. L. Molecular Sorting by Electrical Steering of Microtubules in Kinesin-Coated Channels. Science 312, 910–914, https://doi.org/10.1126/science.1124258 (2006).
Chen, P. et al. Probing single DNA molecule transport using fabricated nanopores. Nano Letters 4, 2293–2298, https://doi.org/10.1021/nl048654j (2004).
Roushan, M., Azad, Z., Lim, S. F., Wang, H. & Riehn, R. Interference of ATP with the fluorescent probes YOYO-1 and YOYO-3 modifies the mechanical properties of intercalator-stained DNA confined in nanochannels. Microchimica Acta 182, 1561–1565, https://doi.org/10.1007/s00604-015-1495-7 (2015).
Subramanya, H. S., Doherty, A. J., Ashford, S. R. & Wigley, D. B. Crystal structure of an atp-dependent dna ligase from bacteriophage t7. Cell 85, 607–615, https://doi.org/10.1016/S0092-8674(00)81260-X (1996).
Tomkinson, A. E., Vijayakumar, S., Pascal, J. M. & Ellenberger, T. DNA Ligases: Structure, Reaction Mechanism, and Function. Chemical Reviews 106, 687–699, https://doi.org/10.1021/cr040498d (2006).
Du, Q., Vologodskaia, M., Kuhn, H., Frank-Kamenetskii, M. & Vologodskii, A. Gapped DNA and cyclization of short DNA fragments. Biophysical Journal 88, 4137–45, https://doi.org/10.1529/biophysj.104.055657 (2005).
Kuhn, H. & Frank-Kamenetskii, M. D. Template-independent ligation of single-stranded DNA by T4 DNA ligase. FEBS Journal 272, 5991–6000, https://doi.org/10.1111/j.1742-4658.2005.04954.x (2005).
Nilsson, S. V. & Magnusson, G. Sealing of gaps in duplex DNA by T4 DNA ligase. Nucleic Acids Research 10, 1425–37, https://doi.org/10.1093/nar/10.5.1425 (1982).
Bogenhagen, D. F. & Pinz, K. G. The Action of DNA Ligase at Abasic Sites in DNA. Journal of Biological Chemistry 273, 7888–7893, https://doi.org/10.1074/jbc.273.14.7888 (1998).
Rossi, R., Montecucco, A. & Ciarrocchi, G. Functional characterization of the T4 DNA ligase: a new insight into the mechanism of action. Nucleic Acids Research 25, 2106–2113, https://doi.org/10.1093/nar/25.11.2106 (1997).
Cherepanov, A. V. & De Vries, S. Kinetics and thermodynamics of nick sealing by T4 DNA ligase. European Journal of Biochemistry 270, 4315–4325, https://doi.org/10.1046/j.1432-1033.2003.03824.x (2003).
Lohman, G. J. S., Chen, L. & Evans, T. C. Kinetic Characterization of Single Strand Break Ligation in Duplex DNA by T4 DNA Ligase. Journal of Biological Chemistry 286, 44187–44196, https://doi.org/10.1074/jbc.M111.284992 (2011).
Zimmerman, S. B., Little, J. W., Oshinsky, C. K. & Gellert, M. Enzymatic joining of DNA strands: a novel reaction of diphosphopyridine nucleotide. Proceedings of the National Academy of Sciences of the United States of America 57, 1841–1848, https://doi.org/10.1073/pnas.57.6.1841 (1967).
Kunkel, T. A. & Erie, D. A. Eukaryotic Mismatch Repair in Relation to DNA Replication. Annual Review of Genetics 49, 291–313, https://doi.org/10.1146/annurev-genet-112414-054722 (2015).
Junop, M. S., Obmolova, G., Rausch, K., Hsieh, P. & Yang, W. Composite Active Site of an ABC ATPase. Molecular Cell 7, 1–12, https://doi.org/10.1016/S1097-2765(01)00149-6 (2001).
Sparreboom, W., Van Den Berg, A. & Eijkel, J. C. Principles and applications of nanofluidic transport. Nature Nanotechnology 4, 713–720, https://doi.org/10.1038/nnano.2009.332 (2009).
Das, S., Dubsky, P., van den Berg, A. & Eijkel, J. Concentration Polarization in Translocation of DNA through Nanopores and Nanochannels. Physical Review Letters 108, 138101, https://doi.org/10.1103/PhysRevLett.108.138101 (2012).
Palacci, J., Abécassis, B., Cottin-Bizonne, C., Ybert, C. & Bocquet, L. Colloidal motility and pattern formation under rectified diffusiophoresis. Physical Review Letters 104, 1–4, https://doi.org/10.1103/PhysRevLett.104.138302 (2010).
Ebel, J. P., Anderson, J. L. & Prieve, D. C. Diffusiophoresis of latex particles in electrolyte gradients. Langmuir 4, 396–406, https://doi.org/10.1021/la00080a024 (1988).
Anderson, J. L. Colloid transport by interfacial forces. Annual Review of Fluid Mechanics 21, 61–99 (1989).
Hsu, J. P., Liu, K. L. & Tseng, S. Diffusiophoresis of polyelectrolytes in nanodevices: Importance of boundary. Journal of Physical Chemistry C 117, 9469–9476, https://doi.org/10.1021/jp312702r (2013).
Wanunu, M., Morrison, W., Rabin, Y., Grosberg, A. Y. & Meller, A. Electrostatic focusing of unlabelled DNA into nanoscale pores using a salt gradient. Nature Nanotechnology 5, 160–5, https://doi.org/10.1038/nnano.2009.379 (2010).
Åkerman, B. & Tuite, E. Single- and double-strand photocleavage of DNA by YO, YOYO and TOTO. Nucleic Acids Research 24, 1080–90, https://doi.org/10.1093/nar/24.6.1080 (1996).
Reuter, M. & Dryden, D. T. F. The kinetics of YOYO-1 intercalation into single molecules of double-stranded DNA. Biochemical and Biophysical Research Communications 403, 225–9, https://doi.org/10.1016/j.bbrc.2010.11.015 (2010).
Riehn, R. et al. Nanochannels for Genomic DNA Analysis: The Long and Short of It. In Liu, R. H. & Lee, A. P. (eds) Integrated Biochips for DNA Analysis, 1 edn, chap. 12, 151–186 (Landes Bioscience, Austin, Texas, 2007).
Acknowledgements
We acknowledge support from the National Institutes of Health (GM107559). This work was performed in part at the NCSU Nanofabrication Facility (NNF), a member of the North Carolina Research Triangle Nanotechnology Network (RTNN), and GA Tech Institute for Electronics and Nanotechnology, which are supported by the National Science Foundation (Grant ECCS-1542015 and ECCS-1626153). ZA acknowledges support through a Fulbright scholarship (9005-SP).
Author information
Authors and Affiliations
Contributions
R.R., M.R., K.R.W. and S.F.L. conceived the experiments, M.R., Z.A., R.R., P.D.R. and K.R.W. conducted the experiments, M.R., Z.A. and G.I.L. analyzed the results. S.M. performed numerical simulations. M.R., R.R. and G.I.L. wrote the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Roushan, M., Azad, Z., Movahed, S. et al. Motor-like DNA motion due to an ATP-hydrolyzing protein under nanoconfinement. Sci Rep 8, 10036 (2018). https://doi.org/10.1038/s41598-018-28278-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-28278-0
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.