Abstract
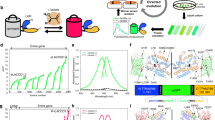
As a key glycolytic metabolite, lactate has a central role in diverse physiological and pathological processes. However, comprehensive multiscale analysis of lactate metabolic dynamics in vitro and in vivo has remained an unsolved problem until now owing to the lack of a high-performance tool. We recently developed a series of genetically encoded fluorescent sensors for lactate, named FiLa, which illuminate lactate metabolism in cells, subcellular organelles, animals, and human serum and urine. In this protocol, we first describe the FiLa sensor-based strategies for real-time subcellular bioenergetic flux analysis by profiling the lactate metabolic response to different nutritional and pharmacological conditions, which provides a systematic-level view of cellular metabolic function at the subcellular scale for the first time. We also report detailed procedures for imaging lactate dynamics in live mice through a cell microcapsule system or recombinant adeno-associated virus and for the rapid and simple assay of lactate in human body fluids. This comprehensive multiscale metabolic analysis strategy may also be applied to other metabolite biosensors using various analytic platforms, further expanding its usability. The protocol is suited for users with expertise in biochemistry, molecular biology and cell biology. Typically, the preparation of FiLa-expressing cells or mice takes 2 days to 4 weeks, and live-cell and in vivo imaging can be performed within 1–2 hours. For the FiLa-based assay of body fluids, the whole measuring procedure generally takes ~1 min for one sample in a manual assay or ~3 min for 96 samples in an automatic microplate assay.
Key points
-
This protocol describes the use of FiLa biosensors, fluorescence-based sensors for the analysis of lactate metabolism. FiLa biosensors can be used in both in vitro assays and in vivo assays, and under different nutritional and pharmacological conditions.
-
Unlike traditional methods of lactate analysis, FiLa biosensors allow subcellular analysis of lactate in real time and show a larger fluorescence ratio response than existing biosensors.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The data used to generate the example results presented in Table 2 are available in the supporting primary research paper39. All other data supporting the findings of this study are available for research purposes from the authors upon reasonable request. Source data are provided with this paper.
References
Harjes, U. Metabolism: more lactate, please. Nat. Rev. Cancer 17, 707 (2017).
Vander Heiden, M. G., Cantley, L. C. & Thompson, C. B. Understanding the Warburg effect: the metabolic requirements of cell proliferation. Science 324, 1029–1033 (2009).
Rabinowitz, J. D. & Enerbäck, S. Lactate: the ugly duckling of energy metabolism. Nat. Metab. 2, 566–571 (2020).
Martinez-Reyes, I. & Chandel, N. S. Waste not, want not: lactate oxidation fuels the TCA cycle. Cell Metab. 26, 803–804 (2017).
Hui, S. et al. Glucose feeds the TCA cycle via circulating lactate. Nature 551, 115–118 (2017).
Zhang, W. et al. Lactate is a natural suppressor of RLR signaling by targeting MAVS. Cell 178, 176–189 (2019).
Daw, C. C. et al. Lactate elicits ER–mitochondrial Mg2+ dynamics to integrate cellular metabolism. Cell 183, 474–489 (2020).
Zhang, D. et al. Metabolic regulation of gene expression by histone lactylation. Nature 574, 575–580 (2019).
Brooks, G. A. The science and translation of lactate shuttle theory. Cell Metab. 27, 757–785 (2018).
Boussouar, F. & Benahmed, M. Lactate and energy metabolism in male germ cells. Trends Endocrinol. Metab. 15, 345–350 (2004).
Oginuma, M. et al. A gradient of glycolytic activity coordinates FGF and Wnt signaling during elongation of the body axis in amniote embryos. Dev. Cell 40, 342–353 (2017).
Du, J. et al. A small-molecule cocktail promotes mammalian cardiomyocyte proliferation and heart regeneration. Cell Stem Cell 29, 545–558 (2022).
Velentzas, P. D. et al. The proton-coupled monocarboxylate transporter hermes is necessary for autophagy during cell death. Dev. Cell 47, 281–293 (2018).
Jia, M. et al. ULK1-mediated metabolic reprogramming regulates Vps34 lipid kinase activity by its lactylation. Sci. Adv. 9, eadg4993 (2023).
Lee, D. C. et al. A lactate-induced response to hypoxia. Cell 161, 595–609 (2015).
Torrini, C. et al. Lactate is an epigenetic metabolite that drives survival in model systems of glioblastoma. Mol. Cell 82, 3061–3076 (2022).
Scheiman, J. et al. Meta-omics analysis of elite athletes identifies a performance-enhancing microbe that functions via lactate metabolism. Nat. Med. 25, 1104–1109 (2019).
Zeni, A. I., Hoffman, M. D. & Clifford, P. S. Energy expenditure with indoor exercise machines. J. Am. Med. Assoc. 275, 1424–1427 (1996).
Marin, E. et al. Human tolerogenic dendritic cells regulate immune responses through lactate synthesis. Cell Metab. 30, 1075–1090 (2019).
Xu, K. et al. Glycolysis fuels phosphoinositide 3-kinase signaling to bolster T cell immunity. Science 371, 405–410 (2021).
Suzuki, A. et al. Astrocyte–neuron lactate transport is required for long-term memory formation. Cell 144, 810–823 (2011).
Magistretti, P. J. & Allaman, I. Lactate in the brain: from metabolic end-product to signalling molecule. Nat. Rev. Neurosci. 19, 235–249 (2018).
Zeng, X. et al. Gut bacterial nutrient preferences quantified in vivo. Cell 185, 3441–3456 (2022).
Iatsenko, I., Boquete, J. P. & Lemaitre, B. Microbiota-derived lactate activates production of reactive oxygen species by the intestinal NADPH oxidase Nox and shortens Drosophila lifespan. Immunity 49, 929–942 (2018).
Dou, X. et al. PDK4-dependent hypercatabolism and lactate production of senescent cells promotes cancer malignancy. Nat. Metab. 5, 1887–1010 (2023).
Lund, J., Clemmensen, C. & Schwartz, T. W. Outrunning obesity with Lac-Phe? Cell Metab. 34, 1085–1087 (2022).
Li, V. L. et al. An exercise-inducible metabolite that suppresses feeding and obesity. Nature 606, 785–790 (2022).
Lin, Y. et al. Lactate is a key mediator that links obesity to insulin resistance via modulating cytokine production from adipose tissue. Diabetes 71, 637–652 (2022).
Watson, M. J. et al. Metabolic support of tumour-infiltrating regulatory T cells by lactic acid. Nature 591, 645–651 (2021).
Martinez-Reyes, I. & Chandel, N. S. Cancer metabolism: looking forward. Nat. Rev. Cancer 21, 669–680 (2021).
Wang, Y. et al. Saturation of the mitochondrial NADH shuttles drives aerobic glycolysis in proliferating cells. Mol. Cell 82, 3270–3283 (2022).
Gomez, H. & Kellum, J. A. Lactate in sepsis. J. Am. Med. Assoc. 313, 194–195 (2015).
Immke, D. C. & McCleskey, E. W. Lactate enhances the acid-sensing Na+ channel on ischemia-sensing neurons. Nat. Neurosci. 4, 869–870 (2001).
Zhang, J. et al. Endothelial lactate controls muscle regeneration from ischemia by inducing M2-like macrophage polarization. Cell Metab. 31, 1136–1153 (2020).
Cluntun, A. A. et al. The pyruvate–lactate axis modulates cardiac hypertrophy and heart failure. Cell Metab. 33, 629–648 (2021).
Pan, R. Y. et al. Positive feedback regulation of microglial glucose metabolism by histone H4 lysine 12 lactylation in Alzheimer’s disease. Cell Metab. 34, 634–648 (2022).
Glancy, B. et al. Mitochondrial lactate metabolism: history and implications for exercise and disease. J. Physiol. 599, 863–888 (2021).
Wan, N. et al. Cyclic immonium ion of lactyllysine reveals widespread lactylation in the human proteome. Nat. Methods 19, 854–864 (2022).
Li, X. et al. Ultrasensitive sensors reveal the spatiotemporal landscape of lactate metabolism in physiology and disease. Cell Metab. 35, 200–211 (2023).
Kraut, J. A. & Madias, N. E. Lactic acidosis. N. Engl. J. Med. 371, 2309–2319 (2014).
van der Windt, G. J. W., Chang, C. H. & Pearce, E. L. Measuring bioenergetics in T cells using a seahorse extracellular flux analyzer. Curr. Prot. Immunol. 113, 16B.11–16B.14 (2016).
Choe, M. & Titov, D. V. Genetically encoded tools for measuring and manipulating metabolism. Nat. Chem. Biol. 18, 451–460 (2022).
Zou, Y. et al. Analysis of redox landscapes and dynamics in living cells and in vivo using genetically encoded fluorescent sensors. Nat. Protoc. 13, 2362–2386 (2018).
Zhang, Z., Cheng, X., Zhao, Y. & Yang, Y. Lighting up live-cell and in vivo central carbon metabolism with genetically encoded fluorescent sensors. Annu. Rev. Anal. Chem. 13, 293–314 (2020).
Zhao, Y. et al. In vivo monitoring of cellular energy metabolism using SoNar, a highly responsive sensor for NAD+/NADH redox state. Nat. Protoc. 11, 1345–1359 (2016).
Zhao, Y. & Yang, Y. Profiling metabolic states with genetically encoded fluorescent biosensors for NADH. Curr. Opin. Biotechnol. 31, 86–92 (2015).
Zhao, Y. et al. Genetically encoded fluorescent sensors for intracellular NADH detection. Cell Metab. 14, 555–566 (2011).
Zhao, Y. et al. SoNar, a highly responsive NAD+/NADH sensor, allows high-throughput metabolic screening of anti-tumor agents. Cell Metab. 21, 777–789 (2015).
Tao, R. et al. Genetically encoded fluorescent sensors reveal dynamic regulation of NADPH metabolism. Nat. Methods 14, 720–728 (2017).
Zou, Y. et al. Illuminating NAD+ metabolism in live cells and in vivo using a genetically encoded fluorescent sensor. Dev. Cell 53, 240–252 (2020).
San Martin, A. et al. A genetically encoded FRET lactate sensor and its use to detect the Warburg effect in single cancer cells. PloS ONE 8, e57712 (2013).
Harada, K. et al. Green fluorescent protein-based lactate and pyruvate indicators suitable for biochemical assays and live cell imaging. Sci. Rep. 10, 19562 (2020).
Nasu, Y. et al. A genetically encoded fluorescent biosensor for extracellular l-lactate. Nat. Commun. 12, 7058 (2021).
Bekdash, R. et al. GEM-IL: a highly responsive fluorescent lactate indicator. Cell Rep. Methods 1, 100092 (2021).
Koveal, D. et al. A high-throughput multiparameter screen for accelerated development and optimization of soluble genetically encoded fluorescent biosensors. Nat. Commun. 13, 2919 (2022).
Aburto, C. et al. Single-fluorophore indicator to explore cellular and subcellular lactate dynamics. ACS Sens. 7, 3278–3286 (2022).
Aguilera, L. et al. Dual role of LldR in regulation of the lldPRD operon, involved in l-lactate metabolism in Escherichia coli. J. Bacteriol. 190, 2997–3005 (2008).
Li, Z. & Ai, H. W. Illuminating lactate in cells, mice, and patient samples. Cell Metab. 35, 5–7 (2023).
Wishart, D. S. et al. HMDB 5.0: the human metabolome database for 2022. Nucleic Acids Res. 50, D622–d631 (2022).
Chen, W. W., Freinkman, E., Wang, T., Birsoy, K. & Sabatini, D. M. Absolute quantification of matrix metabolites reveals the dynamics of mitochondrial metabolism. Cell 166, 1324–1337.e1311 (2016).
Zhao, Y. et al. An expanded palette of genetically encoded Ca2+ indicators. Science 333, 1888–1891 (2011).
Wiederkehr, A. & Demaurex, N. Illuminating redox biology using NADH- and NADPH-specific sensors. Nat. Methods 14, 671–672 (2017).
Gu, W. et al. Glycolytic metabolism plays a functional role in regulating human pluripotent stem cell state. Cell Stem Cell 19, 476–490 (2016).
Hernández, G. et al. Effect of a resuscitation strategy targeting peripheral perfusion status vs serum lactate levels on 28-day mortality among patients with septic shock: the ANDROMEDA-SHOCK randomized clinical trial. J. Am. Med. Assoc. 321, 654–664 (2019).
Larsen, T. Fluorometric determination of d-lactate in biological fluids. Anal. Biochem. 539, 152–157 (2017).
Belousov, V. V. et al. Genetically encoded fluorescent indicator for intracellular hydrogen peroxide. Nat. Methods 3, 281–286 (2006).
Berg, J., Hung, Y. P. & Yellen, G. A genetically encoded fluorescent reporter of ATP:ADP ratio. Nat. Methods 6, 161–166 (2009).
Tantama, M., Hung, Y. P. & Yellen, G. Imaging intracellular pH in live cells with a genetically encoded red fluorescent protein sensor. J. Am. Chem. Soc. 133, 10034–10037 (2011).
Xue, S. et al. A synthetic-biology-inspired therapeutic strategy for targeting and treating hepatogenous diabetes. Mol. Ther. 25, 443–455 (2017).
Zhou, Y. et al. A small and highly sensitive red/far-red optogenetic switch for applications in mammals. Nat. Biotechnol. 40, 262–272 (2022).
Wu, Z., Asokan, A. & Samulski, R. J. Adeno-associated virus serotypes: vector toolkit for human gene therapy. Mol. Ther. 14, 316–327 (2006).
Fripont, S., Marneffe, C., Marino, M., Rincon, M. Y. & Holt, M. G. Production, purification, and quality control for adeno-associated virus-based vectors. J. Vis. Exp. https://doi.org/10.3791/58960 (2019).
Gutscher, M. et al. Real-time imaging of the intracellular glutathione redox potential. Nat. Methods 5, 553–559 (2008).
Acknowledgements
This research is supported by National Key Research and Development Program of China (2019YFA0904800 to Y. Zhao and 2021YFA0804900 to A.W.), National Natural Science Foundation of China (32150030, 32030065, 32121005 and 92049304 to Y. Zhao; 91857202, 21937004 and 32150028 to Y.Y.; 82030039 to Z.J; 32000920 to A.W.; 32201230 to Y. Zou), the Shanghai Science and Technology Commission (20JC1412000), the Shanghai Frontiers Science Center of Optogenetic Techniques for Cell Metabolism (Y. Zhao), the Research Unit of New Techniques for Live-cell Metabolic Imaging (Chinese Academy of Medical Sciences, 2019-I2M-5-013 to Y. Zhao), the innovative research team of high-level local universities in Shanghai, the State Key Laboratory of Bioreactor Engineering, the Fundamental Research Funds for the Central Universities, US National Institutes of Health (HL155107, HL155096 and HL166137 to J.L.) and the American Heart Association (AHA2020CV-19 and AHA957729 to J.L.).
Author information
Authors and Affiliations
Contributions
Y. Zhao, Y.Y., T. Li, A.W. and Y. Zou conceived and designed the live-cell and live-mice imaging experiments. Y. Zhao and Ru Wang designed the body fluid analysis experiment. A.W., Y. Zou, T. Li, S.L., X.Z., L.Z. and Ruwen Wang performed experiments. Y.X., X.L., Z.Z., T. Liu, Z.J. and J.L. gave technical support and conceptual advice. Y. Zhao, Y.Y., R.W., A.W., Y. Zou, T. Li and J.L. analyzed the data and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Protocols thanks Jonathan Rocheleau, Masayuki Yazawa and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
Li, X. et al. Cell Metab. 35, 200–211 (2023): https://doi.org/10.1016/j.cmet.2022.10.002
Jia, M. et al. Sci. Adv. 9, eadg4993 (2023): https://doi.org/10.1126/sciadv.adg4993
Dou, X. et al. Nat. Metab. 5, 1887–1910 (2023): https://doi.org/10.1038/s42255-023-00912-w
Supplementary information
Supplementary Information
Supplementary Figs. 1–8.
Supplementary Data
Source data for Supplementary Figs. 1–7.
Source data
Source Data Figs. 1–7
Statistical source data for Figs. 1–7.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wang, A., Zou, Y., Liu, S. et al. Comprehensive multiscale analysis of lactate metabolic dynamics in vitro and in vivo using highly responsive biosensors. Nat Protoc 19, 1311–1347 (2024). https://doi.org/10.1038/s41596-023-00948-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-023-00948-y
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.