Abstract
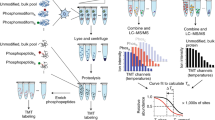
Proteins regulate biological processes by changing their structure or abundance to accomplish a specific function. In response to a perturbation, protein structure may be altered by various molecular events, such as post-translational modifications, protein–protein interactions, aggregation, allostery or binding to other molecules. The ability to probe these structural changes in thousands of proteins simultaneously in cells or tissues can provide valuable information about the functional state of biological processes and pathways. Here, we present an updated protocol for LiP-MS, a proteomics technique combining limited proteolysis with mass spectrometry, to detect protein structural alterations in complex backgrounds and on a proteome-wide scale. In LiP-MS, proteins undergo a brief proteolysis in native conditions followed by complete digestion in denaturing conditions, to generate structurally informative proteolytic fragments that are analyzed by mass spectrometry. We describe advances in the throughput and robustness of the LiP-MS workflow and implementation of data-independent acquisition–based mass spectrometry, which together achieve high reproducibility and sensitivity, even on large sample sizes. We introduce MSstatsLiP, an R package dedicated to the analysis of LiP-MS data for the identification of structurally altered peptides and differentially abundant proteins. The experimental procedures take 3 d, mass spectrometric measurement time and data processing depend on sample number and statistical analysis typically requires ~1 d. These improvements expand the adaptability of LiP-MS and enable wide use in functional proteomics and translational applications.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout









Similar content being viewed by others
Data availability
The mass spectrometry proteomics data presented in Fig. 8 and Extended Data Fig. 7 and 9 have been deposited to the ProteomeXchange Consortium via the PRIDE partner repository27 with the dataset identifier PXD031616 and PXD031627 (http://www.proteomexchange.org/). The mass spectrometry proteomics data presented in Fig. 6, Fig. 7 and Extended Data Fig. 8 were generated as part of ref. 4. The mass spectrometry data presented in Fig. 9 were generated as part of ref. 5. The corresponding spectral libraries, Spectronaut Reports and statistical source data presented in Figs. 6–9 and Extended Data Figs. 7 and 9 are available at https://doi.org/10.5281/zenodo.5749994.
Code availability
The R Markdown notebooks for data analysis are available at https://doi.org/10.5281/zenodo.5749994. The MSstatsLiP R-package can be installed from Bioconductor (https://www.bioconductor.org/packages/release/bioc/html/MSstats.html).
Change history
18 January 2023
A Correction to this paper has been published: https://doi.org/10.1038/s41596-023-00808-9
References
Feng, Y. et al. Global analysis of protein structural changes in complex proteomes. Nat. Biotechnol. 32, 1036–1044 (2014).
Leuenberger, P. et al. Cell-wide analysis of protein thermal unfolding reveals determinants of thermostability. Science 355, eaai7825 (2017).
Piazza, I. et al. A map of protein-metabolite interactions reveals principles of chemical communication. Cell 172, 358–372.e23 (2018).
Piazza, I. et al. A machine learning-based chemoproteomic approach to identify drug targets and binding sites in complex proteomes. Nat. Commun. 11, 4200 (2020).
Cappelletti, V. et al. Dynamic 3D proteomes reveal protein functional alterations at high resolution in situ. Cell 184, 545–559.e22 (2021).
Schopper, S. et al. Measuring protein structural changes on a proteome-wide scale using limited proteolysis-coupled mass spectrometry. Nat. Protoc. 12, 2391–2410 (2017).
Mackmull, M.-T. et al. Global, in situ analysis of the structural proteome in individuals with Parkinson’s disease to identify a new class of biomarker. Nat. Struct. Mol. Biol. 29, 978–989 (2022).
Leitner, A. et al. Probing native protein structures by chemical cross-linking, mass spectrometry, and bioinformatics. Mol. Cell. Proteom. 9, 1634–1649 (2010).
Rappsilber, J. The beginning of a beautiful friendship: cross-linking/mass spectrometry and modelling of proteins and multi-protein complexes. J. Struct. Biol. 173, 530–540 (2011).
Sinz, A. Chemical cross-linking and mass spectrometry to map three-dimensional protein structures and protein-protein interactions. Mass Spectrom. Rev. 25, 663–682 (2006).
Kiselar, J. G. & Chance, M. R. Future directions of structural mass spectrometry using hydroxyl radical footprinting. J. Mass Spectrom. 45, 1373–1382 (2010).
Maleknia, S. D. & Downard, K. M. Advances in radical probe mass spectrometry for protein footprinting in chemical biology applications. Chem. Soc. Rev. 43, 3244–3258 (2014).
Espino, J. A. & Jones, L. M. Illuminating biological interactions with in vivo protein footprinting. Anal. Chem. 91, 6577–6584 (2019).
Rinas, A., Espino, J. A. & Jones, L. M. An efficient quantitation strategy for hydroxyl radical-mediated protein footprinting using Proteome Discoverer. Anal. Bioanal. Chem. 408, 3021–3031 (2016).
Liu, F., Rijkers, D. T. S., Post, H. & Heck, A. J. R. Proteome-wide profiling of protein assemblies by cross-linking mass spectrometry. Nat. Methods 12, 1179–1184 (2015).
Savitski, M. M. et al. Tracking cancer drugs in living cells by thermal profiling of the proteome. Science 346, 1255784 (2014).
Tan, C. S. H. et al. Thermal proximity coaggregation for system-wide profiling of protein complex dynamics in cells. Science 359, 1170–1177 (2018).
Potel, C. M. et al. Impact of phosphorylation on thermal stability of proteins. Nat. Methods 18, 757–759 (2021).
Mateus, A. et al. Thermal proteome profiling for interrogating protein interactions. Mol. Syst. Biol. 16, e9232 (2020).
Ludwig, C. et al. Data‐independent acquisition‐based SWATH‐ MS for quantitative proteomics: a tutorial. Mol. Syst. Biol. 14, e8126 (2018).
Clough, T., Thaminy, S., Ragg, S., Aebersold, R. & Vitek, O. Statistical protein quantification and significance analysis in label-free LC-MS experiments with complex designs. BMC Bioinforma. 13, 1–17 (2012).
Tsai, T. & Vitek, O. Introduction to MSstatsPTM. Available at https://rdrr.io/bioc/MSstatsPTM/f/vignettes/MSstatsPTM.Rmd (2020).
Moulder, R., Goo, Y. A. & Goodlett, D. R. Label-free quantitation for clinical proteomics. Methods Mol. Biol. 1410, 65–76 (2016).
Richards, A. L. et al. One-hour proteome analysis in yeast. Nat. Protoc. 10, 701–714 (2015).
Feng, Y. & Picotti, P. Selected reaction monitoring to measure proteins of interest in complex samples: a practical guide. Methods Mol. Biol. 1394, 43–56 (2016).
Rappsilber, J., Mann, M. & Ishihama, Y. Protocol for micro-purification, enrichment, pre-fractionation and storage of peptides for proteomics using StageTips. Nat. Protoc. 2, 1896–1906 (2007).
Perez-Riverol, Y. et al. The PRIDE database and related tools and resources in 2019: improving support for quantification data. Nucleic Acids Res. 47, D442–D450 (2019).
Murtagh, F. & Legendre, P. Ward’s hierarchical agglomerative clustering method: which algorithms implement Ward’s criterion?. J. Classif. 31, 274–295 (2014).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Kohler, D. et al. Bioconductor - MSstatsPTM. Available at https://www.bioconductor.org/packages/release/bioc/html/MSstatsPTM.html (2021).
Choi, M. et al. MSstats: an R package for statistical analysis of quantitative mass spectrometry-based proteomic experiments. Bioinformatics 30, 2524–2526 (2014).
Ueda, K. et al. Molecular cloning of cDNA encoding an unrecognized component of amyloid in Alzheimer disease. Proc. Natl Acad. Sci. USA 90, 11282–11286 (1993).
Acknowledgements
We thank the Picotti laboratory for continuous input on the protocol optimization and the R package. I.P. and L.M. were supported by long-term EMBO postdoctoral fellowships (ALTF 846-2014 and ALTF 538-2016). This project received funding from the European Research Council (grant agreement no. 866004) and through the EPIC-XS Consortium (grant agreement no. 823839) both under the European Union’s Horizon 2020 research and innovation program. It was supported by a Personalized Health and Related Technologies (PHRT) grant (PHRT-506), a Sinergia grant from the Swiss National Science Foundation (SNSF grant CRSII5_177195) and the National Center of Competence in Research AntiResist funded by the SNSF (grant number 51NF40_180541).
Author information
Authors and Affiliations
Contributions
L.M., V.C. and N.d.S. wrote the paper. L.M., V.C. and I.P. optimized the original version of the protocol. M.P., P.S., C.D., F.E., L.K., N.B. and L.R. contributed to protocol optimization. L.M., V.C., D.K., T.-H.T. and F.S. performed bioinformatic analysis of the data. D.K. and T.-H.T. optimized algorithms and tools for the R package MSstatsLiP. O.V. supervised the development of the R package. P.P. supervised the optimization of the protocol.
Corresponding authors
Ethics declarations
Competing interests
N.B. and L.R. are employees of Biognosys AG (Zurich, Switzerland). P.P. is a scientific advisor for the company Biognosys AG (Zurich, Switzerland) and an inventor of a patent licensed by Biognosys AG that covers the LiP-MS method used in this protocol. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Protocols thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Related Links
Key references using this protocol
Piazza, I. et al. Nat. Commun. 11, 4200 (2020): https://doi.org/10.1038/s41467-020-18071-x
Cappelletti, V. et al. Cell 184, 545–559.e22 (2021): https://doi.org/10.1016/j.cell.2020.12.021
This protocol is an update to Nat. Protoc. 12, 2391–2410 (2017): https://doi.org/10.1038/nprot.2017.100.
Extended data
Extended Data Fig. 1 Analysis of cleavage preference of PK.
a and b, WebLogo diagram of the cleavage site of PK and the neighboring five amino acids in a mammalian cell lysate (a) and yeast cell lysate (b). Amino acids are plotted according to the frequency of occurrence at the cleavage side and colored on the basis of their physicochemical properties. c, WebLogo diagram of the cleavage site of a specific protease (trypsin) in a mammalian cell extract. d, Boxplots showing the distribution of secondary structure elements in half-tryptic peptides in different organisms. Box shows the quartiles of the dataset, median values are represented by a vertical line in the center of the box, bars extend to the rest of the distribution and dots represent outliers. AA, amino acid.
Extended Data Fig. 2 Improvement of reproducibility and robustness of a LiP-MS experiment.
a, Coefficient of variation of a LiP experiment performed in a thermocycler as described in this protocol (new) and performed in a boiling pot as described in Schopper et al.6 (old). The median value is displayed below the plot. b, Coefficient of variation of a LiP experiment with DOC removal through centrifugation in individual reaction tubes as described in Schopper et al.6 (old) and with DOC removal through filtration (new). The median value is displayed below the plot.
Extended Data Fig. 3 Comparison of PK cleavage pattern at different sample concentrations.
Principal component (PC) analysis of MS1 features. The colors indicate the different samples amounts in 50 μl.
Extended Data Fig. 4 Influence of E:S ratio and incubation time.
a, Summary characterization of peptides in LiP-MS experiments performed as described in Fig. 2a. Left pie chart: Number of peptides identified across the different incubation times. Right pie chart: Number of peptides identified across the different E:S ratios. b and c, Pairwise comparisons of peptide identifications and intensity changes for each incubation time (1–60 min) versus the shortest incubation time (30 s). b, Fraction of peptides identified only in the higher digestion condition (red), only in the lower digestion condition (blue) or shared between both conditions (white). c, Fraction of peptides with significantly changed intensities (|log2(fold change)| > 2, q-value < 0.05); increase indicated in blue, decrease indicated in red and no change indicated in gray. d and e, Distribution of peptide length (number of amino acids) in groups defined in b and c. Boxplot of peptide length in amino acids, colored by groups. d, Peptides identified only in the higher digestion condition (red), only in the lower digestion condition (blue) or shared between both conditions (green). e, Peptides with increased intensity (blue), decreased intensity (red) or no change (gray). A significant difference in mean peptide length is indicated by asterisks (P < 0.05, Wilcoxon test).
Extended Data Fig. 5 Experimental characteristics in different organisms.
Quantification in three different species: E. coli, S. cerevisiae and human (Homo sapiens) cell lines, both in LiP samples (light blue) and TrP samples (dark blue). a, HT content (percentage). b, Median CV (percentage). c, Number of peptide identifications. d, Number of protein identifications.
Extended Data Fig. 6 Comparison of LiP peptide intensities in different proteinaceous backgrounds.
Correlation of peptide intensities of monomeric α-synuclein spiked into 1.8 mg/ml BSA, whole-cell lysate of HeLa cells, SH-SY5Y cells and yeast S. cerevisiae cells. Pearson correlation coefficient of log2-transformed fragment ion intensities visualized as colored squares (upper triangle) with values ranging from 0.983 (gray) to 0.968 (white). Scatter plot of log2-transformed fragment ion intensity correlation (lower triangle) and data distribution density plot (diagonal).
Extended Data Fig. 7 Commonly observed errors or issues.
a, CV (ratio of the standard deviation to the mean) of peptide intensity for biological replicates in samples with varying viscosity. Mixing efficiency is visualized by the addition of red color to the samples. In solutions with high viscosity, poor mixing can lead to non-uniform distribution of PK. Poor distribution of PK results in differential extents of cleavage and thus high variability (median CV > 0.25). Proper mixing can counteract this effect. b, High CV (median CV > 0.25) of peptide intensity for biological replicates in three samples. Evaporation can occur during PK inactivation as a result of opening lids. This leads to a global increase in variability in all samples of an experiment. c, Distribution of HT peptides (gray) and fully tryptic peptides (black) in two samples. Insufficient inactivation of PK can lead to variations in the proportions of HT peptides as reflected by the inconsistent trypticity content among replicates in sample 2. In this case, replicate 1 of sample 2 cannot be used for the statistical analysis. d, PCA of fragment ion intensities in four samples. Spike-in of single proteins at high concentrations into a complex background can display poor separation of experimental groups when using log2-transformed data (left panel). Using non-transformed data emphasizes the separation effect of the spike-in protein (right panel). e, Differential analysis of structurally altered peptides, visualized in a volcano plot. Each point represents a peptide. Peptides passing the significance cutoff (|log2(FC)| > 1, q-value < 0.05) are colored by trypticity (blue: fully tryptic; red: HT). Impaired PK activity in one of the conditions results in predominantly unidirectional fold changes of peptide classes (upregulation of almost all fully tryptic peptides and downregulation of mostly semi-tryptic peptides).
Extended Data Fig. 8 Multiple dose analysis of yeast cells treated with rapamycin at nine different concentrations.
a, Volcano plots of altered LiP peptide intensities generated with MSstatsLiP. Significantly altered peptides (|log2FC| > 1, q-value < 0.01) are indicated in black. Significantly changing peptides mapping to the direct target of rapamycin (FPR1) are colored in orange. b, Bar plot showing in how many conditions a significant FPR1 peptide was found. Only two LiP peptides were found in six conditions: LGLSNEDFFHK mapping to Sac1p and TGDLVTIHYTGTLENGQK mapping to Fpr1p, the know target of rapamycin. c, The Fpr1p LiP peptide found altered in six conditions is mapped onto the 3D protein structure complexed with rapamycin obtained by homology modeling (PDB: 2DG3). Rapamycin bound to target protein is indicated in violet. Orange indicates the TGDLVTIHYTGTLENGQK LiP peptide.
Extended Data Fig. 9 Benchmarking abundance correction approaches for LiP-MS.
a, Schematic of the experimental setup. Spike-in of intact proteins. The M and F conformation of α-synuclein (aSyn) are spiked into yeast lysate at two different concentrations (5 and 20 pmol/µg lysate). b, Schematic of the experimental setup. Spike-in of peptides. Peptides obtained from the digestion of M and F aSyn are spiked into a yeast lysate digest at the same concentrations as in a. c, Differential analysis of LiP peptides in the comparison M1 versus M2 (i.e., two different concentrations of monomer) described in b. Each dot represents an individual peptide, and green indicates peptides of aSyn. Dark green, peptides passing the significance cutoff (|log2(FC)| > 2, q-value < 0.05); light green, non-significant peptides. d, Structural barcodes visualizing fully tryptic peptides mapped along the sequence of aSyn in the M1 versus M2 comparison described in c and corrected for protein abundance as indicated. Significant peptides are colored in dark green, and non-significant peptides are colored in light green. Gray regions indicate no identified fully tryptic peptide matching this region. The position of the NAC region is indicated as a line below the barcode.
Supplementary information
Supplementary Information
Supplementary Discussion and Tables 1 and 2
Supplementary Data 1
FASTA file containing a list of known contaminants
Source data
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Malinovska, L., Cappelletti, V., Kohler, D. et al. Proteome-wide structural changes measured with limited proteolysis-mass spectrometry: an advanced protocol for high-throughput applications. Nat Protoc 18, 659–682 (2023). https://doi.org/10.1038/s41596-022-00771-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-022-00771-x
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.