Abstract
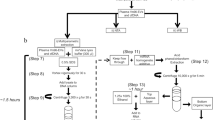
Extracellular vesicles (EVs) are increasingly being recognized as important vehicles for intercellular communication and as promising sources for biomarker discovery. Because the state of protein post-translational modifications (PTMs) such as phosphorylation and glycosylation can be a key determinant of cellular physiology, comprehensive characterization of protein PTMs in EVs can be particularly valuable for early-stage diagnostics and monitoring of disease status. However, the analysis of PTMs in EVs has been complicated by limited amounts of purified EVs, low-abundance PTM proteins, and interference from proteins and metabolites in biofluids. Recently, we developed an approach to isolate phosphoproteins and glycoproteins in EVs from small volumes of human plasma that enabled us to identify nearly 10,000 unique phosphopeptides and 1,500 unique N-glycopeptides. The approach demonstrated the feasibility of using these data to identify potential markers to differentiate disease from healthy states. Here we present an updated workflow to sequentially isolate phosphopeptides and N-glycopeptides, enabling multiple PTM analyses of the same clinical samples. In this updated workflow, we have improved the reproducibility and efficiency of EV isolation, protein extraction, and phosphopeptide/N-glycopeptide enrichment to achieve sensitive analyses of low-abundance PTMs in EVs isolated from 1 mL of plasma. The modularity of the workflow also allows for the characterization of phospho- or glycopeptides only and enables additional analysis of total proteomes and other PTMs of interest. After blood collection, the protocol takes 2 d, including EV isolation, PTM/peptide enrichment, mass spectrometry analysis, and data quantification.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The raw MS data from this study have been deposited into the ProteomeXchange Consortium via the PRIDE Archive with the identifier PXD013893 (https://www.ebi.ac.uk/pride/archive/projects/PXD013893).
References
Harel, M., Oren-Giladi, P., Kaidar-Person, O., Shaked, Y. & Geiger, T. Proteomics of microparticles with SILAC Quantification (PROMIS-Quan): a novel proteomic method for plasma biomarker quantification. Mol. Cell Proteom. 14, 1127–1136 (2015).
Milane, L., Singh, A., Mattheolabakis, G., Suresh, M. & Amiji, M. M. Exosome mediated communication within the tumor microenvironment. J. Control Release 219, 278-294 (2015).
Cocucci, E. & Meldolesi, J. Ectosomes and exosomes: shedding the confusion between extracellular vesicles. Trends Cell Biol. 25, 364–372 (2015).
An, T. et al. Exosomes serve as tumour markers for personalized diagnostics owing to their important role in cancer metastasis. J. Extracell. Vesicles 4, 27522 (2015).
Dobrowolski, R. & De Robertis, E. M. Endocytic control of growth factor signalling: multivesicular bodies as signalling organelles. Nat. Rev. Mol. Cell Biol. 13, 53–60 (2011).
Melo, S. A. et al. Glypican-1 identifies cancer exosomes and detects early pancreatic cancer. Nature 523, 177–182 (2015).
Gonzales, P. A. et al. Large-scale proteomics and phosphoproteomics of urinary exosomes. J. Am. Soc. Nephrol. 20, 363–379 (2009).
Boukouris, S. & Mathivanan, S. Exosomes in bodily fluids are a highly stable resource of disease biomarkers. Proteom. Clin. Appl. 9, 358–367 (2015).
Wu, A. Y., Ueda, K. & Lai, C. P. Proteomic analysis of extracellular vesicles for cancer diagnostics. Proteomics 19, e1800162 (2019).
Xu, R., Greening, D. W., Zhu, H. J., Takahashi, N. & Simpson, R. J. Extracellular vesicle isolation and characterization: toward clinical application. J. Clin. Invest. 126, 1152–1162 (2016).
Bandu, R., Oh, J. W. & Kim, K. P. Mass spectrometry-based proteome profiling of extracellular vesicles and their roles in cancer biology. Exp. Mol. Med. 51, 30 (2019).
Emmanouilidi, A., Paladin, D., Greening, D. W. & Falasca, M. Oncogenic and non-malignant pancreatic exosome cargo reveal distinct expression of oncogenic and prognostic factors involved in tumor invasion and metastasis. Proteomics 19, e1800158 (2019).
Hurwitz, S. N. & Meckes, D. G., Jr. Extracellular vesicle integrins distinguish unique cancers. Proteomes 7, 7020014 (2019).
Bae, S., Brumbaugh, J. & Bonavida, B. Exosomes derived from cancerous and non-cancerous cells regulate the anti-tumor response in the tumor microenvironment. Genes Cancer 9, 87–100 (2018).
Ghosh, A. et al. Rapid isolation of extracellular vesicles from cell culture and biological fluids using a synthetic peptide with specific affinity for heat shock proteins. PLoS One 9, e110443 (2014).
Lobb, R. J. et al. Optimized exosome isolation protocol for cell culture supernatant and human plasma. J. Extracell. Vesicles 4, 27031 (2015).
Gallart-Palau, X. et al. Extracellular vesicles are rapidly purified from human plasma by potein organic solvent precipitation (PROSPR). Sci. Rep. 5, 14664 (2015).
Moreno-Gonzalo, O., Villarroya-Beltri, C. & Sanchez-Madrid, F. Post-translational modifications of exosomal proteins. Front. Immunol. 5, 383 (2014).
Zhang, Y., Wu, X. & Tao, W. A. Characterization and applications of extracellular vesicle proteome with post-translational modifications. Trends Analyt. Chem. 107, 21–30 (2018).
Gerlach, J. Q. & Griffin, M. D. Getting to know the extracellular vesicle glycome. Mol. Biosyst. 12, 1071–1081 (2016).
Oeyen, E. et al. Bladder cancer diagnosis and follow-up: the current status and possible role of extracellular vesicles. Int. J. Mol. Sci. 20, 821 (2019).
Mann, M. & Jensen, O. N. Proteomic analysis of post-translational modifications. Nat. Biotechnol. 21, 255–261 (2003).
Aebersold, R. & Goodlett, D. R. Mass spectrometry in proteomics. Chem. Rev. 101, 269–295 (2001).
Kettenbach, A. N., Rush, J. & Gerber, S. A. Absolute quantification of protein and post-translational modification abundance with stable isotope-labeled synthetic peptides. Nat. Protoc. 6, 175–186 (2011).
Chen, I. H. et al. Phosphoproteins in extracellular vesicles as candidate markers for breast cancer. Pro. c. Natl Acad. Sci. USA 114, 3175–3180 (2017).
Chen, I. H. et al. Analytical pipeline for discovery and verification of glycoproteins from plasma-derived extracellular vesicles as breast cancer biomarkers. Anal. Chem. 90, 6307–6313 (2018).
Jaros, J. A. et al. Clinical use of phosphorylated proteins in blood serum analysed by immobilised metal ion affinity chromatography and mass spectrometry. J. Proteom. 76(Spec No.), 36–42 (2012).
Hu, L. et al. Profiling of endogenous serum phosphorylated peptides by titanium (IV) immobilized mesoporous silica particles enrichment and MALDI-TOFMS detection. Anal. Chem. 81, 94–104 (2009).
Sokolova, V. et al. Characterisation of exosomes derived from human cells by nanoparticle tracking analysis and scanning electron microscopy. Colloids Surf. B Biointerfaces 87, 146–150 (2011).
Palmisano, G. et al. Characterization of membrane-shed microvesicles from cytokine-stimulated beta-cells using proteomics strategies. Mol. Cell. Proteom. 11, 230–243 (2012).
Royo, F. et al. Different EV enrichment methods suitable for clinical settings yield different subpopulations of urinary extracellular vesicles from human samples. J. Extracell. Vesicles 5, 29497 (2016).
Cvjetkovic, A., Lotvall, J. & Lasser, C. The influence of rotor type and centrifugation time on the yield and purity of extracellular vesicles. J. Extracell. Vesicles 3, 1–11 (2014).
Enderle, D. et al. Characterization of RNA from exosomes and other extracellular vesicles isolated by a novel spin column-based method. PLoS One 10, e0136133 (2015).
Niu, Z. et al. Polymer-based precipitation preserves biological activities of extracellular vesicles from an endometrial cell line. PLoS One 12, e0186534 (2017).
Yu, L. L. et al. A comparison of traditional and novel methods for the separation of exosomes from human samples. Biomed. Re. s. Int. 2018, 3634563 (2018).
Tauro, B. J. et al. Two distinct populations of exosomes are released from LIM1863 colon carcinoma cell-derived organoids. Mol. Cell. Proteom. 12, 587–598 (2013).
Wu, X., Li, L., Iliuk, A. & Tao, W. A. Highly efficient phosphoproteome capture and analysis from urinary extracellular vesicles. J. Proteome Res. 17, 3308–3316 (2018).
Ramirez, M. I. et al. Technical challenges of working with extracellular vesicles. Nanoscale 10, 881–906 (2018).
Contreras-Naranjo, J. C., Wu, H. J. & Ugaz, V. M. Microfluidics for exosome isolation and analysis: enabling liquid biopsy for personalized medicine. Lab Chip 17, 3558–3577 (2017).
Rosa-Fernandes, L., Rocha, V. B., Carregari, V. C., Urbani, A. & Palmisano, G. A perspective on extracellular vesicles proteomics. Front Chem. 5, 102 (2017).
Wisniewski, J. R., Zougman, A., Nagaraj, N. & Mann, M. Universal sample preparation method for proteome analysis. Nat. Methods 6, 359–362 (2009).
Heath, N. et al. Rapid isolation and enrichment of extracellular vesicle preparations using anion exchange chromatography. Sci. Rep. 8, 5730 (2018).
Feist, P. & Hummon, A. B. Proteomic challenges: sample preparation techniques for microgram-quantity protein analysis from biological samples. Int. J. Mol. Sci. 16, 3537–3563 (2015).
Gundry, R. L. et al. Preparation of proteins and peptides for mass spectrometry analysis in a bottom-up proteomics workflow. Curr. Protoc. Mol. Biol. Unit 10.25, supplement 88 (2009).
Bereman, M. S., Egertson, J. D. & MacCoss, M. J. Comparison between procedures using SDS for shotgun proteomic analyses of complex samples. Proteomics 11, 2931–2935 (2011).
Hsu, C. C. et al. Universal plant phosphoproteomics workflow and its application to tomato signaling in response to cold stress. Mol. Cell. Proteom. 17, 2068–2080 (2018).
Bayramoglu, G., Celikbicak, O., Arica, M. Y. & Salih, B. Trypsin Immobilized on Magnetic Beads via Click Chemistry: Fast Proteolysis of Proteins in a Microbioreactor for MALDI-ToF-MS Peptide Analysis. Ind. Eng. Chem. Res. 53, 4554–4564 (2014).
Sielaff, M. et al. Evaluation of FASP, SP3, and iST protocols for proteomic sample preparation in the low microgram range. J. Proteome Res. 16, 4060–4072 (2017).
Ludwig, K. R., Schroll, M. M. & Hummon, A. B. Comparison of in-solution, FASP, and S-trap based digestion methods for nottom-up proteomic studies. J. Proteome Res. 17, 2480–2490 (2018).
Swaney, D. L. & Villen, J. Enrichment of phosphopeptides via immobilized metal affinity chromatography. Cold Spring Harb. Protoc. 2016, 088005 (2016).
Fila, J. & Honys, D. Enrichment techniques employed in phosphoproteomics. Amino Acids 43, 1025–1047 (2012).
Yue, X., Schunter, A. & Hummon, A. B. Comparing multistep immobilized metal affinity chromatography and multistep TiO2 methods for phosphopeptide enrichment. Anal. Chem. 87, 8837–8844 (2015).
Montoya, A., Beltran, L., Casado, P., Rodriguez-Prados, J. C. & Cutillas, P. R. Characterization of a TiO(2) enrichment method for label-free quantitative phosphoproteomics. Methods 54, 370–378 (2011).
Huang, J. et al. Highly efficient release of glycopeptides from hydrazide beads by hydroxylamine assisted PNGase F deglycosylation for N-glycoproteome analysis. Anal. Chem. 87, 10199–10204 (2015).
Zhang, H., Li, X. J., Martin, D. B. & Aebersold, R. Identification and quantification of N-linked glycoproteins using hydrazide chemistry, stable isotope labeling and mass spectrometry. Nat. Biotechnol. 21, 660–666 (2003).
Cao, W., Huang, J., Jiang, B., Gao, X. & Yang, P. Highly selective enrichment of glycopeptides based on zwitterionically functionalized soluble nanopolymers. Sci. Rep. 6, 29776 (2016).
Zhu, R., Zacharias, L., Wooding, K. M., Peng, W. & Mechref, Y. Glycoprotein enrichment analytical techniques: advantages and disadvantages. Methods Enzymol. 585, 397–429 (2017).
Zhang, C. et al. Evaluation of different N-glycopeptide enrichment methods for N-glycosylation sites mapping in mouse brain. J. Proteome Res. 15, 2960–2968 (2016).
Nilsson, J. et al. Enrichment of glycopeptides for glycan structure and attachment site identification. Nat. Methods 6, 809–811 (2009).
Yang, W. et al. Comparison of enrichment methods for intact N- and O-linked glycopeptides using strong anion exchange and hydrophilic interaction liquid chromatography. Anal. Chem. 89, 11193–11197 (2017).
Liebler, D. C. & Zimmerman, L. J. Targeted quantitation of proteins by mass spectrometry. Biochemistry 52, 3797–3806 (2013).
Osinalde, N., Aloria, K., Omaetxebarria, M. J. & Kratchmarova, I. Targeted mass spectrometry: An emerging powerful approach to unblock the bottleneck in phosphoproteomics. J. Chromatogr. B 1055-1056, 29–38 (2017).
Thery, C. et al. Minimal information for studies of extracellular vesicles 2018 (MISEV2018): a position statement of the International Society for Extracellular Vesicles and update of the MISEV2014 guidelines. J. Extracell. Vesicles 7, 1535750 (2018).
Shao, H. et al. New technologies for analysis of extracellular vesicles. Chem. Rev. 118, 1917–1950 (2018).
Szatanek, R. et al. The methods of choice for extracellular vesicles (EVs) characterization. Int. J. Mol. Sci. 18, 1153 (2017).
Lotvall, J. et al. Minimal experimental requirements for definition of extracellular vesicles and their functions: a position statement from the International Society for Extracellular Vesicles. J. Extracell. Vesicles 3, 26913 (2014).
Masuda, T., Saito, N., Tomita, M. & Ishihama, Y. Unbiased quantitation of Escherichia coli membrane proteome using phase transfer surfactants. Mol. Cell. Proteom. 8, 2770–2777 (2009).
Rappsilber, J., Mann, M. & Ishihama, Y. Protocol for micro-purification, enrichment, pre-fractionation and storage of peptides for proteomics using StageTips. Nat. Protoc. 2, 1896–1906 (2007).
Humphrey, S. J., Karayel, O., James, D. E. & Mann, M. High-throughput and high-sensitivity phosphoproteomics with the EasyPhos platform. Nat. Protoc. 13, 1897–1916 (2018).
Mertins, P. et al. Reproducible workflow for multiplexed deep-scale proteome and phosphoproteome analysis of tumor tissues by liquid chromatography-mass spectrometry. Nat. Protoc. 13, 1632–1661 (2018).
Hernandez-Valladares, M. et al. Reliable FASP-based procedures for optimal quantitative proteomic and phosphoproteomic analysis on samples from acute myeloid leukemia patients. Biol. Proced. Online 18, 13 (2016).
Witwer, K.W. et al. Standardization of sample collection, isolation and analysis methods in extracellular vesicle research. J. Extracell. Vesicles 2, 20360 (2013).
Nakai, W. et al. A novel affinity-based method for the isolation of highly purified extracellular vesicles. Sci. Rep. 6, 33935 (2016).
Kowal, J. et al. Proteomic comparison defines novel markers to characterize heterogeneous populations of extracellular vesicle subtypes. Proc. Natl Acad. Sci. USA 113, E968–977 (2016).
Coumans, F. A. et al. Reproducible extracellular vesicle size and concentration determination with tunable resistive pulse sensing. J. Extracell. Vesicles 3, 25922 (2014).
Yuana, Y. et al. Cryo-electron microscopy of extracellular vesicles in fresh plasma. J. Extracell. Vesicles 2, 21494 (2013).
Iliuk, A. B., Martin, V. A., Alicie, B. M., Geahlen, R. L. & Tao, W. A. In-depth analyses of kinase-dependent tyrosine phosphoproteomes based on metal ion-functionalized soluble nanopolymers. Mol. Cell. Proteom. 9, 2162–2172 (2010).
Ye, J. et al. Optimized IMAC-IMAC protocol for phosphopeptide recovery from complex biological samples. J. Proteome Res. 9, 3561–3573 (2010).
Larsen, M. R., Thingholm, T. E., Jensen, O. N., Roepstorff, P. & Jorgensen, T. J. Highly selective enrichment of phosphorylated peptides from peptide mixtures using titanium dioxide microcolumns. Mol. Cell. Proteom. 4, 873–886 (2005).
Tyanova, S., Temu, T. & Cox, J. The MaxQuant computational platform for mass spectrometry-based shotgun proteomics. Nat. Protoc. 11, 2301–2319 (2016).
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26, 1367–1372 (2008).
Tyanova, S. et al. The Perseus computational platform for comprehensive analysis of (prote)omics data. Nat. Methods 13, 731–740 (2016).
Tyanova, S. & Cox, J. Perseus: a bioinformatics platform for integrative analysis of proteomics data in cancer tesearch. Methods Mol. Biol. 1711, 133–148 (2018).
Rauniyar, N. Parallel reaction monitoring: a targeted experiment performed using high resolution and high mass accuracy mass spectrometry. Int. J. Mol. Sci. 16, 28566–28581 (2015).
Cordwell, S. J. & White, M. Y. Targeted proteomics for determining phosphorylation site-specific associations in cardiovascular disease. Circulation 126, 1803–1807 (2012).
Zauber, H., Kirchner, M. & Selbach, M. Picky: a simple online PRM and SRM method designer for targeted proteomics. Nat. Methods 15, 156–157 (2018).
Acknowledgements
This project was funded by NIH grants 5R01GM088317 and 1R01GM111788 and NSF grant 1506752.
Author information
Authors and Affiliations
Contributions
H.A.A., A.B.I. and W.A.T. designed the studies. H.A.A., A.B.I. and I.-H.C. developed methods. H.A.A. performed the experiments and analyzed the data. H.A.A. and W.A.T. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
A.B.I and W.A.T. are the co-founders of Tymora Analytical Operations, which commercialized a polyMAC kit used in parts of the described protocol.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
Chen, I.-H. et al. Proc. Natl. Acad. Sci. USA 114, 3175–3180 (2017): https://www.pnas.org/content/114/12/3175
Chen, I.-H. et al. Anal. Chem. 90, 6307–6313 (2018): https://pubs.acs.org/doi/10.1021/acs.analchem.8b01090
Wu X., Li, L., Iliuk, A. & Tao, W. A. J. Proteome Res. 17, 3308–3316 (2018): https://pubs.acs.org/doi/10.1021/acs.jproteome.8b00459
Integrated supplementary information
Supplementary Figure 1 Venn diagrams showing the overlap between the EV database from Vesiclepedia and EV data from control and breast cancer.
The Venn diagrams show a 77% overlap between both, the EV database and control EV proteins, and the EV database and EV breast cancer proteins. Overlap of the proteome from the control and the breast cancer EV data is also shown.
Supplementary Figure 2 Quantitation results from MaxQuant and Perseus showing Pearson correlations from proteome analysis across each condition and replicate.
Scatterplots and Pearson correlation coefficients depicting the log2-transformed intensities from proteome analysis of control and breast cancer samples, each in triplicate.
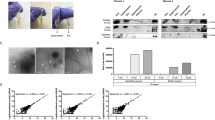
Supplementary Figure 3 Scatterplots representing the targeted proteomic (PRM) analysis.
Raw intensities from five individual control samples corresponding to five phosphopeptides from five EV proteins were plotted. See Table S1 for details.
Supplementary information
Supplementary Information
Supplementary Figs. 1–3 and Supplementary Tables 1 and 2.
Rights and permissions
About this article
Cite this article
Andaluz Aguilar, H., Iliuk, A.B., Chen, IH. et al. Sequential phosphoproteomics and N-glycoproteomics of plasma-derived extracellular vesicles. Nat Protoc 15, 161–180 (2020). https://doi.org/10.1038/s41596-019-0260-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-019-0260-5
This article is cited by
-
Extracellular vesicle-based liquid biopsy biomarkers and their application in precision immuno-oncology
Biomarker Research (2023)
-
Advanced extracellular vesicle bioinformatic nanomaterials: from enrichment, decoding to clinical diagnostics
Journal of Nanobiotechnology (2023)
-
Comprehensive profiling of extracellular vesicles in uveitis and scleritis enables biomarker discovery and mechanism exploration
Journal of Translational Medicine (2023)
-
Recent advances of small extracellular vesicle biomarkers in breast cancer diagnosis and prognosis
Molecular Cancer (2023)
-
Proteomic strategies for characterizing ubiquitin-like modifications
Nature Reviews Methods Primers (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.