Abstract
Selection of the pre-mRNA branch site (BS) by the U2 small nuclear ribonucleoprotein (snRNP) is crucial to prespliceosome (A complex) assembly. The RNA helicase PRP5 proofreads BS selection but the underlying mechanism remains unclear. Here we report the atomic structures of two sequential complexes leading to prespliceosome assembly: human 17S U2 snRNP and a cross-exon pre-A complex. PRP5 is anchored on 17S U2 snRNP mainly through occupation of the RNA path of SF3B1 by an acidic loop of PRP5; the helicase domain of PRP5 associates with U2 snRNA; the BS-interacting stem-loop (BSL) of U2 snRNA is shielded by TAT-SF1, unable to engage the BS. In the pre-A complex, an initial U2–BS duplex is formed; the translocated helicase domain of PRP5 stays with U2 snRNA and the acidic loop still occupies the RNA path. The pre-A conformation is specifically stabilized by the splicing factors SF1, DNAJC8 and SF3A2. Cancer-derived mutations in SF3B1 damage its association with PRP5, compromising BS proofreading. Together, these findings reveal key insights into prespliceosome assembly and BS selection or proofreading by PRP5.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The atomic coordinates for human 17S U2 snRNP and the pre-A complex have been deposited in the Protein Data Bank (PDB) under accession codes PDB 7EVO and PDB 7VPX, respectively. The EM maps of 17S U2 snRNP core region, PRP5-α1, PRP5-α2, TAT-SF1 and BSL have been deposited in the EMDB with accession codes EMD-31334, EMD-31335, EMD-31336, EMD-31337 and EMD-31338, respectively. The EM map of the pre-A complex core region, DNAJC8 region and U1 snRNP region have been deposited in EMDB with accession codes EMD-32074, EMD-32075 and EMD-32076, respectively. The accession number for the RNA sequencing data reported in this paper is from the Gene Expression Omnibus (GEO GSE185713). Source data are provided with this paper.
References
Wahl, M. C., Will, C. L. & Luhrmann, R. The spliceosome: design principles of a dynamic RNP machine. Cell 136, 701–718 (2009).
Yan, C., Wan, R. & Shi, Y. Molecular mechanisms of pre-mRNA splicing through structural biology of the spliceosome. Cold Spring Harb. Perspect. Biol. 11, a032409 (2019).
Das, R., Zhou, Z. L. & Reed, R. Functional association of U2 snRNP with the ATP-independent spliceosomal complex E. Mol. Cell 5, 779–787 (2000).
Ruby, S. W., Chang, T. H. & Abelson, J. Four yeast spliceosomal proteins (PRP5, PRP9, PRP11, and PRP21) interact to promote U2 snRNP binding to pre-mRNA. Genes Dev. 7, 1909–1925 (1993).
Xu, Y. Z. et al. Prp5 bridges U1 and U2 snRNPs and enables stable U2 snRNP association with intron RNA. EMBO J. 23, 376–385 (2004).
Liang, W. W. & Cheng, S. C. A novel mechanism for Prp5 function in prespliceosome formation and proofreading the branch site sequence. Gene Dev. 29, 81–93 (2015).
Wang, E. T. et al. Alternative isoform regulation in human tissue transcriptomes. Nature 456, 470–476 (2008).
Chen, M. & Manley, J. L. Mechanisms of alternative splicing regulation: insights from molecular and genomics approaches. Nat. Rev. Mol. Cell Biol. 10, 741–754 (2009).
Lee, Y. & Rio, D. C. Mechanisms and regulation of alternative pre-mRNA splicing. Annu. Rev. Biochem. 84, 291–323 (2015).
Bonnal, S. C., Lopez-Oreja, I. & Valcarcel, J. Roles and mechanisms of alternative splicing in cancer—implications for care. Nat. Rev. Clin. Oncol. 17, 457–474 (2020).
Smith, C. W. J. & Valcarcel, J. Alternative pre-mRNA splicing: the logic of combinatorial control. Trends Biochem. Sci. 25, 381–388 (2000).
O’Day, C. L., Dalbadie-McFarland, G. & Abelson, J. The Saccharomyces cerevisiae Prp5 protein has RNA-dependent ATPase activity with specificity for U2 small nuclear RNA. J. Biol. Chem. 271, 33261–33267 (1996).
Abu Dayyeh, B. K., Quan, T. K., Castro, M. & Ruby, S. W. Probing interactions between the U2 small nuclear ribonucleoprotein and the DEAD-box protein, Prp5. J. Biol. Chem. 277, 20221–20233 (2002).
Perriman, R., Barta, I., Voeltz, G. K., Abelson, J. & Ares, M. Jr. ATP requirement for Prp5p function is determined by Cus2p and the structure of U2 small nuclear RNA. Proc. Natl Acad. Sci. USA 100, 13857–13862 (2003).
Yang, F. et al. Mechanisms of the RNA helicases DDX42 and DDX46 in human U2 snRNP assembly. Nat. Commun. 14, 897 (2023).
Zhang, Z. W. Molecular architecture of the human 17S U2 snRNP. Nature 583, 310–313 (2020).
Perriman, R. & Ares, M. Invariant U2 snRNA nucleotides form a stem loop to recognize the intron early in splicing. Mol. Cell 38, 416–427 (2010).
Xu, Y. Z. & Query, C. C. Competition between the ATPase prp5 and branch region-U2 snRNA pairing modulates the fidelity of spliceosome assembly. Mol. Cell 28, 838–849 (2007).
Tang, Q. et al. SF3B1/Hsh155 HEAT motif mutations affect interaction with the spliceosomal ATPase Prp5, resulting in altered branch site selectivity in pre-mRNA splicing. Gene Dev. 30, 2710–2723 (2016).
Carrocci, T. J., Zoerner, D. M., Paulson, J. C. & Hoskins, A. A. SF3b1 mutations associated with myelodysplastic syndromes alter the fidelity of branchsite selection in yeast. Nucleic Acids Res. 45, 4837–4852 (2017).
Yoshida, K. & Ogawa, S. Splicing factor mutations and cancer. WIREs RNA 5, 445–459 (2014).
Bonnal, S., Vigevani, L. & Valcarcel, J. The spliceosome as a target of novel antitumour drugs. Nat. Rev. Drug Discov. 11, 847–859 (2012).
Golas, M. M., Sander, B., Will, C. L., Luhrmann, R. & Stark, H. Molecular architecture of the multiprotein splicing factor SF3b. Science 300, 980–984 (2003).
Cretu, C. et al. Molecular architecture of SF3b and structural consequences of Its cancer-related mutations. Mol. Cell 64, 307–319 (2016).
Yan, C. Y., Wan, R. X., Bai, R., Huang, G. X. Y. & Shi, Y. G. Structure of a yeast activated spliceosome at 3.5 Å resolution. Science 353, 904–911 (2016).
Will, C. L. et al. Characterization of novel SF3b and 17S U2 snRNP proteins, including a human Prp5p homologue and an SF3b DEAD-box protein. EMBO J. 21, 4978–4988 (2002).
Kastner, B. et al. GraFix: sample preparation for single-particle electron cryomicroscopy. Nat. Methods 5, 53–55 (2008).
Zhang, X. et al. An atomic structure of the human spliceosome. Cell 169, 918–929 (2017).
Kaida, D. et al. Spliceostatin A targets SF3b and inhibits both splicing and nuclear retention of pre-mRNA. Nat. Chem. Biol. 3, 576–583 (2007).
Berget, S. M. Exon recognition in vertebrate splicing. J. Biol. Chem. 270, 2411–2414 (1995).
Sharma, S., Kohlstaedt, L. A., Damianov, A., Rio, D. C. & Black, D. L. Polypyrimidine tract binding protein controls the transition from exon definition to an intron defined spliceosome. Nat. Struct. Mol. Biol. 15, 183–191 (2008).
Cretu, C. et al. Structural basis of intron selection by U2 snRNP in the presence of covalent inhibitors. Nat. Commun. 12, 4491 (2021).
Schneider, M. et al. Exon definition complexes contain the tri-snRNP and can be directly converted into B-like precatalytic splicing complexes. Mol. Cell 38, 223–235 (2010).
Tholen, J., Razew, M., Weis, F. & Galej, W. P. Structural basis of branch site recognition by the human spliceosome. Science 375, 50–57 (2022).
Shao, W., Kim, H. S., Cao, Y., Xu, Y. Z. & Query, C. C. A U1–U2 snRNP interaction network during intron definition. Mol. Cell. Biol. 32, 470–478 (2012).
Zhang, X. et al. Structure of the human activated spliceosome in three conformational states. Cell Res 28, 307–322 (2018).
Zhan, X., Yan, C., Zhang, X., Lei, J. & Shi, Y. Structures of the human pre-catalytic spliceosome and its precursor spliceosome. Cell Res 28, 1129–1140 (2018).
Rauhut, R. et al. Molecular architecture of the Saccharomyces cerevisiae activated spliceosome. Science 353, 1399–1405 (2016).
Cretu, C. et al. Structural basis of splicing modulation by antitumor macrolide compounds. Mol. Cell 70, 265 (2018).
Finci, L. I. et al. The cryo-EM structure of the SF3b spliceosome complex bound to a splicing modulator reveals a pre-mRNA substrate competitive mechanism of action. Genes Dev. 32, 309–320 (2018).
Pomeranz Krummel, D. A., Oubridge, C., Leung, A. K., Li, J. & Nagai, K. Crystal structure of human spliceosomal U1 snRNP at 5.5 Å resolution. Nature 458, 475–480 (2009).
Nesic, D. & Kramer, A. Domains in human splicing factors SF3a60 and SF3a66 required for binding to SF3a120, assembly of the 17S U2 snRNP, and prespliceosome formation. Mol. Cell. Biol. 21, 6406–6417 (2001).
Selenko, P. et al. Structural basis for the molecular recognition between human splicing factors U2AF(65) and SF1/mBBP. Mol. Cell 11, 965–976 (2003).
Abovich, N. & Rosbash, M. Cross-intron bridging interactions in the yeast commitment complex are conserved in mammals. Cell 89, 403–412 (1997).
Crisci, A. et al. Mammalian splicing factor SF1 interacts with SURP domains of U2 snRNP-associated proteins. Nucleic Acids Res. 43, 10456–10473 (2015).
Liu, Z. H. et al. Structural basis for recognition of the intron branch site RNA by splicing factor 1. Science 294, 1098–1102 (2001).
Jacewicz, A., Chico, L., Smith, P., Schwer, B. & Shuman, S. Structural basis for recognition of intron branchpoint RNA by yeast Msl5 and selective effects of interfacial mutations on splicing of yeast pre-mRNAs. RNA 21, 401–414 (2015).
Kao, C. Y., Cao, E. C., Wai, H. L. & Cheng, S. C. Evidence for complex dynamics during U2 snRNP selection of the intron branchpoint. Nucleic Acids Res. 49, 9965–9977 (2021).
Gozani, O., Feld, R. & Reed, R. Evidence that sequence-independent binding of highly conserved U2 snRNP proteins upstream of the branch site is required for assembly of spliceosomal complex A. Gene Dev. 10, 233–243 (1996).
Ma, C. T. et al. Ordered multi-site phosphorylation of the splicing factor ASF/SF2 by SRPK1. J. Mol. Biol. 376, 55–68 (2008).
Zhong, X. Y., Ding, J. H., Adams, J. A., Ghosh, G. & Fu, X. D. Regulation of SR protein phosphorylation and alternative splicing by modulating kinetic interactions of SRPK1 with molecular chaperones. Gene Dev. 23, 482–495 (2009).
Yoshida, K. et al. Frequent pathway mutations of splicing machinery in myelodysplasia. Blood 118, 212–212 (2011).
Liu, Z. Q. et al. Mutations in the RNA splicing factor SF3B1 promote tumorigenesis through MYC stabilization. Cancer Discov. 10, 806–821 (2020).
DeBoever, C. et al. Transcriptome sequencing reveals potential mechanism of cryptic 3′ splice site selection in SF3B1-mutated cancers. PLoS Comput. Biol. 11, e1004105 (2015).
Darman, R. B. et al. Cancer-associated SF3B1 hotspot mutations induce cryptic 3′ splice site selection through use of a different branch point. Cell Rep. 13, 1033–1045 (2015).
Alsafadi, S. et al. Cancer-associated SF3B1 mutations affect alternative splicing by promoting alternative branchpoint usage. Nat. Commun. 7, 10615 (2016).
Yin, S. Y. et al. A murine model of chronic lymphocytic leukemia based on B cell-restricted expression of Sf3b1 mutation and Atm deletion. Cancer Cell 35, 283 (2019).
Brooks, A. N. et al. Conservation of an RNA regulatory map between Drosophila and mammals. Genome Res. 21, 193–202 (2011).
Buonamici, S. et al. Abstract 2040: Mutations in SF3B1 lead to aberrant splicing through cryptic 3′ splice site selection and impair hematopoietic cell differentiation. Cancer Res 75, https://doi.org/10.1158/1538-7445.Am2015-2040 (2015).
Zhang, Z. et al. Structural insights into how Prp5 proofreads the pre-mRNA branch site. Nature 596, 296–300 (2021).
Zeng, Y. et al. Profiling lariat intermediates reveals genetic determinants of early and late co-transcriptional splicing. Mol. Cell 82, 4681–4699 e4688 (2022).
Buonamici, S. et al. H3B-8800, an orally bioavailable modulator of the SF3b complex, shows efficacy in spliceosome-mutant myeloid malignancies. Blood 128, 966 (2016).
Seiler, M. et al. H3B-8800, an orally available small-molecule splicing modulator, induces lethality in spliceosome-mutant cancers. Nat. Med. 24, 497–504 (2018).
Steensma, D. P. et al. Results of a clinical trial of H3B-8800, a splicing modulator, in patients with myelodysplastic syndromes (MDS), acute myeloid leukemia (AML) or chronic myelomonocytic leukemia (CMML). Blood 134, 673 (2019).
Ghosh, A. K., Mishevich, J. L. & Jurica, M. S. Spliceostatins and derivatives: chemical syntheses and biological properties of potent splicing inhibitors. J. Nat. Prod. 84, 1681–1706 (2021).
Bai, R. et al. Mechanism of spliceosome remodeling by the ATPase/helicase Prp2 and its coactivator Spp2. Science 371, 141 (2021).
Zhan, X., Yan, C., Zhang, X., Lei, J. & Shi, Y. Structure of a human catalytic step I spliceosome. Science 359, 537–545 (2018).
Chiara, M. D. et al. Identification of proteins that interact with exon sequences, splice sites, and the branchpoint sequence during each stage of spliceosome assembly. Mol. Cell. Biol. 16, 3317–3326 (1996).
Jurica, M. S., Licklider, L. J., Gygi, S. P., Grigorieff, N. & Moore, M. J. Purification and characterization of native spliceosomes suitable for three-dimensional structural analysis. RNA 8, 426–439 (2002).
Dignam, J. D., Lebovitz, R. M. & Roeder, R. G. Accurate transcription initiation by RNA polymerase II in a soluble extract from isolated mammalian nuclei. Nucleic Acids Res. 11, 1475–1489 (1983).
Lei, J. L. & Frank, J. Automated acquisition of cryo-electron micrographs for single particle reconstruction on an FEI Tecnai electron microscope. J. Struct. Biol. 150, 69–80 (2005).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Zhang, K. Gctf: real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Chen, S. X. et al. High-resolution noise substitution to measure overfitting and validate resolution in 3D structure determination by single particle electron cryomicroscopy. Ultramicroscopy 135, 24–35 (2013).
Rosenthal, P. B. & Henderson, R. Optimal determination of particle orientation, absolute hand, and contrast loss in single-particle electron cryomicroscopy. J. Mol. Biol. 333, 721–745 (2003).
Swint-Kruse, L. & Brown, C. S. Resmap: automated representation of macromolecular interfaces as two-dimensional networks. Bioinformatics 21, 3327–3328 (2005).
Steiner, R. A. & Murshudov, G. N. Flat model bulk solvent correction in the program REFMAC. Acta Crystallogr. 56, S301 (2000).
Nicholls, R. & Murshudov, G. ProSMART—procrustes structural matching alignment and restraints tool. Acta Crystallogr. 67, C745–C745 (2011).
Amunts, A. et al. Structure of the yeast mitochondrial large ribosomal subunit. Science 343, 1485–1489 (2014).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Harrow, J. et al. GENCODE: the reference human genome annotation for The ENCODE Project. Genome Res. 22, 1760–1074 (2012).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Acknowledgements
We thank the Cryo-EM Facility, the Computing Center, the Crystallography Facility and the Mass Spectrometry & Metabolomics Core Facility of Westlake University for technical support. We thank N. A. Larsen from H3 Biomedicine for providing SSA. This work was supported by funds from the National Natural Science Foundation of China (31930059 to Y.S.), the China Postdoctoral Science Foundation (2020M671806 to X. Zhang; 2021M692888 to X. Zhan), the National Postdoctoral Program for Innovative Talents of China (BX20200305 to X. Zhang; BX2021268 to X. Zhan) and Start-up funds from Westlake University (to Y.S.).
Author information
Authors and Affiliations
Contributions
Y.S. conceived the project. X. Zhang, T.B. and F.Y. designed and performed the experiments. X. Zhang prepared cryo-EM samples and collected the EM data. X. Zhan, X. Zhang, Y.L. and Z.X. processed the EM data, calculated the EM maps, and built the atomic models. F.Y., R.F., P.L. and Q.Z. performed RNA-seq data analysis. Y.S. and X. Zhang wrote the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Structural & Molecular Biology thanks Jeffrey Pleiss and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Sara Osman, in collaboration with the Nature Structural & Molecular Biology team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Purification and cryo-EM reconstruction of human 17S U2 snRNP.
a, Only U2 snRNA is detectable in the purified human 17S U2 snRNP as shown in this denaturing PAGE gel. b, Analysis of the purified human 17S U2 snRNP on a silver-stained SDS-PAGE gel. The protein components were identified by mass spectrometry. c, A representative cryo-EM micrograph (upper panel) and representative 2D class averages (lower panel) of the human 17S U2 snRNP sample. The experiments were repeated independently for more than three times with similar results. d, A flow chart diagram of cryo-EM data processing for human 17S U2 snRNP. Please refer to Methods for details. e, The final reconstructions for the masked core (red), PRP5-α1 (blue), PRP5-α2 (yellow), TAT-SF1(magenta) and BSL (cyan) display average resolutions of 2.5 Å, 2.9 Å, 2.9 Å, 3.8 Å and 2.8 Å, respectively, on the basis of the FSC value of 0.143. f, Two overall views of the EM density map. The local resolutions are color-coded for different regions of U2 snRNP. g, Angular distribution of the particles used for the reconstruction. h, The Fourier-shell correlation (FSC) curves for cross-validation between the model and the cryo-EM map of human 17S U2 snRNP.
Extended Data Fig. 2 Purification and cryo-EM reconstruction of human pre-A complex.
a, Analysis of RNA components on 8% urea-PAGE gel after glycerol gradient centrifugation. The fractions that contain pre-mRNA, U1 and U2 snRNAs are indicated by a red rectangular box. Fractions 11–15, thought to contain the pre-A complex, were pooled for cryo-EM analysis. b, Analysis of the purified pre-A complex on a silver-stained SDS-PAGE gel. The protein components were identified by mass spectrometry. c, A representative cryo-EM micrograph of the pre-A complex sample. d, Representative 2D class averages. Scale bar: 44 nm. The experiments were repeated independently for more than three times with similar results. e, A flow chart diagram of cryo-EM data processing for human pre-A complex. Please refer to Methods for details. f, The final reconstruction for the masked U2 region displays an average resolution of 3.0 Å on the basis of the FSC value of 0.143. g, Two overall views of the EM density map. The local resolutions are color-coded for different regions of U2 snRNP in the pre-A complex. h, Angular distribution of the particles used for the reconstruction of U2 snRNP region of human pre-A complex. i, The Fourier-shell correlation (FSC) curves for cross-validation between the model and the cryo-EM map for the human pre-A complex.
Extended Data Fig. 3 The EM density map of human 17S U2 snRNP.
a, The EM density map for the small molecule E7107 and its surrounding structural elements. Two related views are shown. b, The EM density map for the RRM domain of the splicing factor TAT-SF1 (left panel). The EM density maps for two representative α-helices are shown (middle and right panels). c, The EM density map for the Linker domain of TAT-SF1. d, The EM density map for SF3A2. The separator helix is indicated by a black dashed box. e, The EM density map for the α1 helix of PRP5 and its interacting elements from SF3B1. Two related views are shown. f, The EM density map for the acidic loop of PRP5 and its interacting elements from SF3B1. Two related views are shown. g, The EM density map for the α2 helix of PRP5 and its interacting elements from SF3B1. Two related views are shown.
Extended Data Fig. 4 The EM density map for different regions of the human pre-A complex.
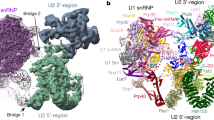
a, The EM density map for the initial U2/BS duplex in the pre-A complex. b, The EM density map for the zinc finger (ZnF) of SF3A2. c, The EM density map for the structurally resolved region of DNAJC8. Two related views are shown. d, Docking of SF1 into the EM density map. e, Docking of U1 snRNP into the EM density map. f, Superimposition of the initial U2/BS duplex from pre-A complex and the three-way junction of U2 snRNA from human 17S U2 snRNP (grey). g, The low-pass filtered density map for the 5′-end sequences of U2 snRNA can be traced to the helicase domain of PRP5.
Extended Data Fig. 5 PRP5 knockdown in HEK293F cells induces altered splicing site selection.
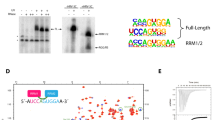
a, Analysis of PRP5 mRNA level (top panel) and protein expression level (bottom panel) in control (NC) and siRNA-transfected cells (mean ± s.d., n = 3 independent replicates). b, The volcano plot of log2 fold change versus log10(Pvalue) of gene expression level in siRNA-transfected cells and WT cells. Red and blue dots indicate up-regulated and down-regulated genes, respectively (Pvalue < 0.001). c, The volcano plot of log2 fold change of ∆PSI versus log10(Pvalue) of all splicing changes. Red dots indicate significantly altered splicing events (|∆PSI| > 10 and Pvalue < 0.05). d, Distribution of novel (orange) and known (grey) aberrant splicing events in PRP5 knockdown cells. Intron retention and alternative 3’SS are the most frequently occurring events. See also Extended Data Table 4. e, Validation of representative aberrantly spliced genes in control (n = 3) and PRP5 knockdown (n = 5) cells through quantitative RT-PCR assays. f, A schematic diagram of normal and aberrant splicing of ANKHD1, a representative gene that is sensitive to PRP5 knockdown. Positions of the cryptic (red) and canonical (green) 3’SS are indicated. P values were calculated by a two-sided two-test without adjustment.
Extended Data Fig. 6 The open and closed conformations of SF3B1.
a, The open conformation of SF3B1. SF3B1 exists in an open conformation in both 17S U2 snRNP and the pre-A complex. Shown here is an overlay of the SF3B1 structures from these two complexes. b, The closed conformation of SF3B1. Pre-mRNA engagement by U2 snRNP results in a closed conformation of SF3B1. Shown here is the structure of SF3B1 from the human Bact complex. c, Structural alignment using HEAT repeats 16 through 20 of SF3B1 reveals major conformational differences between the open and closed states of SF3B1. Two perpendicular views are shown. d, Structural alignment using HEAT repeats 4 through 9 of SF3B1 reveals major conformational differences between the open and closed states of SF3B1.
Extended Data Fig. 7 Structural comparison between the yeast intron-defined pre-A complex and the human exon-defined pre-A complex.
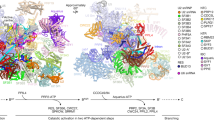
a, Structure of the pre-A complex from S. cerevisiae. The yeast complex was assembled on an intron-definition pre-mRNA. A schematic diagram of the pre-mRNA and its recognition by the pre-A complex is shown above. b, Structure of the human pre-A complex. The human complex was assembled on an exon-definition pre-mRNA. A schematic diagram of the pre-mRNA and its recognition by the pre-A complex is shown above.
Extended Data Fig. 8 The relative orientations of U1 and U2 snRNPs in yeast and human spliceosomes.
a, Molecular organizations of yeast U1 (pink) and U2 (green and light blue) snRNPs in the pre-A, A and pre-B complexes. Overall, the U1 snRNP moves away from U2 snRNP to allow the recruitment of tri-snRNP. b, In human, the cross-exon pre-A complex is expected to be converted to cross-intron complex for splicing reaction to proceed; this step may involve interaction of U2 snRNP with another U1 snRNP that bound an upstream 5′SS.
Extended Data Fig. 9 Recognition of the splicing inhibitors E7107 and SSA by the Hinged pocket.
a, A close-up view on the EM density of E7107 in human 17S U2 snRNP. b, A close-up view on the interface between E7107 and the Hinged pocket. The Hinged pocket is exactly where the BPA binds in the assembled spliceosomes. c, A close-up view on the EM density of SSA in the human pre-A complex. SSA is covalently linked to Cys26 of PHF5A. d, A close-up view on the interface between SSA and the Hinged pocket.
Supplementary information
Supplementary Information
Supplementary Tables 1–3.
Supplementary Table 4
List of aberrant splicing events associated with PRP5 knockdown in HEK293F cells.
Source data
Source Data Fig. 6
Unprocessed western blots.
Source Data Extended Data Fig. 1
Unprocessed gels.
Source Data Extended Data Fig. 2
Unprocessed gels.
Source Data Extended Data Fig. 5
Unprocessed western blots.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, X., Zhan, X., Bian, T. et al. Structural insights into branch site proofreading by human spliceosome. Nat Struct Mol Biol 31, 835–845 (2024). https://doi.org/10.1038/s41594-023-01188-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-023-01188-0
This article is cited by
-
Decoding branch points and unlocking splicing secrets
Nature Structural & Molecular Biology (2024)
-
Structural insights into the cross-exon to cross-intron spliceosome switch
Nature (2024)
-
Structural insights into human exon-defined spliceosome prior to activation
Cell Research (2024)
-
Molecular basis for the activation of human spliceosome
Nature Communications (2024)