Abstract
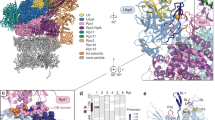
Proteasome inhibitors are widely used as therapeutics and research tools, and typically target one of the three active sites, each present twice in the proteasome complex. An endogeneous proteasome inhibitor, PI31, was identified 30 years ago, but its inhibitory mechanism has remained unclear. Here, we identify the mechanism of Saccharomyces cerevisiae PI31, also known as Fub1. Using cryo-electron microscopy (cryo-EM), we show that the conserved carboxy-terminal domain of Fub1 is present inside the proteasome’s barrel-shaped core particle (CP), where it simultaneously interacts with all six active sites. Targeted mutations of Fub1 disrupt proteasome inhibition at one active site, while leaving the other sites unaffected. Fub1 itself evades degradation through distinct mechanisms at each active site. The gate that allows substrates to access the CP is constitutively closed, and Fub1 is enriched in mutant CPs with an abnormally open gate, suggesting that Fub1 may function to neutralize aberrant proteasomes, thereby ensuring the fidelity of proteasome-mediated protein degradation.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
Cryo-EM maps and atomic model coordinates have been deposited in the Electron Microscopy Data Bank and the Protein Data Bank, respectively, as α3∆ 20S (EMD-25847, PDB 7TEJ) and α3∆ 20S + Fub1 complex (EMD-25848, PDB 7TEO). Additional structures referenced here include 3MG0, 5CZ4 and 7LS5. Source data are provided with this paper.
References
Bard, J. A. M. et al. Structure and function of the 26S proteasome. Annu. Rev. Biochem. 87, 697–724 (2018).
Rousseau, A. & Bertolotti, A. Regulation of proteasome assembly and activity in health and disease. Nat. Rev. Mol. Cell Biol. 19, 697–712 (2018).
Tomko, R. J.Jr. & Hochstrasser, M. Molecular architecture and assembly of the eukaryotic proteasome. Annu. Rev. Biochem. 82, 415–445 (2013).
Hanna, J. et al. Deubiquitinating enzyme Ubp6 functions noncatalytically to delay proteasomal degradation. Cell 127, 99–111 (2006).
Crosas, B. et al. Ubiquitin chains are remodeled at the proteasome by opposing ubiquitin ligase and deubiquitinating activities. Cell 127, 1401–1413 (2006).
Hanna, J., Waterman, D., Boselli, M. & Finley, D. Spg5 protein regulates the proteasome in quiescence. J. Biol. Chem. 287, 34400–34409 (2012).
Hanna, J. et al. Cuz1/Ynl155w, a zinc-dependent ubiquitin-binding protein, protects cells from metalloid-induced proteotoxicity. J. Biol. Chem. 289, 1876–1885 (2014).
Sá-Moura, B. et al. A conserved protein with AN1 zinc finger and ubiquitin-like domains modulates Cdc48 (p97) function in the ubiquitin-proteasome pathway. J. Biol. Chem. 288, 33682–33696 (2013).
Lee, D., Takayama, S. & Goldberg, A. L. ZFAND5/ZNF216 is an activator of the 26S proteasome that stimulates overall protein degradation. Proc. Natl Acad. Sci. USA 115, E9550–E9559 (2018).
Shi, Y. et al. Rpn1 provides adjacent receptor sites for substrate binding and deubiquitination by the proteasome. Science 351, 6275 (2016).
Chu-Ping, M., Slaughter, C. A. & DeMartino, G. N. Purification and characterization of a protein inhibitor of the 20S proteasome (macropain). Biochim. Biophys. Acta, Protein Struct. Mol. Enzymol. 1119, 303–311 (1992).
Hatanaka, A. et al. Fub1p, a novel protein isolated by boundary screening, binds the proteasome complex. Genes Genet. Syst. 86, 305–314 (2011).
Zaiss, D. M. W., Standera, S., Holzhütter, H., Kloetzel, P.-M. & Sijts, A. J. A. M. The proteasome inhibitor PI31 competes with PA28 for binding to 20S proteasomes. FEBS Lett. 457, 333–338 (1999).
McCutchen-Maloney, S. L. et al. cDNA cloning, expression, and functional characterization of PI31, a proline-rich inhibitor of the proteasome. J. Biol. Chem. 275, 18557–18565 (2000).
Yashiroda, H. et al. N-terminal α7 deletion of the proteasome 20S core particle substitutes for yeast PI31 function. Mol. Cell. Biol. 35, 141–152 (2015).
Velichutina, I., Connerly, P. L., Arendt, C. S., Li, X. & Hochstrasser, M. Plasticity in eucaryotic 20S proteasome ring assembly revealed by a subunit deletion in yeast. EMBO J. 23, 500–510 (2004).
Emori, Y. et al. Molecular cloning and functional analysis of three subunits of yeast proteasome. Mol. Cell. Biol. 11, 344–353 (1991).
Sadre-Bazzaz, K., Whitby, F. G., Robinson, H., Formosa, T. & Hill, C. P. Structure of a Blm10 complex reveals common mechanisms for proteasome binding and gate opening. Mol. Cell 37, 728–735 (2010).
Rêgo, A. T. & da Fonseca, P. C. A. Characterization of fully recombinant human 20S and 20S-PA200 proteasome complexes. Mol. Cell 76, 138–147.e5 (2019).
Guerra-Moreno, A. & Hanna, J. Tmc1 is a dynamically regulated effector of the Rpn4 proteotoxic stress response. J. Biol. Chem. 291, 14788–14795 (2016).
Edskes, H. K., Stroobant, E. E., DeWilde, M. P., Bezsonov, E. E. & Wickner, R. B. Proteasome control of [URE3] prion propagation by degradation of anti-prion proteins Cur1 and Btn2 in Saccharomyces cerevisiae. Genetics 218, iyab037 (2021).
Blackburn, C. et al. Characterization of a new series of non-covalent proteasome inhibitors with exquisite potency and selectivity for the 20S β5-subunit. Biochem. J. 430, 461–476 (2010).
Huber, E. M. et al. Systematic analyses of substrate preferences of 20S proteasomes using peptidic epoxyketone inhibitors. J. Am. Chem. Soc. 137, 7835–7842 (2015).
Köhler, A. et al. The axial channel of the proteasome core particle is gated by the Rpt2 ATPase and controls both substrate entry and product release. Mol. Cell 7, 1143–1152 (2001).
Kirk, R. et al. Structure of a conserved dimerization domain within the F-box protein Fbxo7 and the PI31 proteasome inhibitor. J. Biol. Chem. 283, 22325–22335 (2008).
Kao, A. et al. Development of a novel cross-linking strategy for fast and accurate identification of cross-linked peptides of protein complexes. Mol. Cell. Proteomics 10, M110.002170 (2011).
Bode, W. & Huber, R. Natural protein proteinase inhibitors and their interaction with proteinases. Eur. J. Biochem. 204, 433–451 (1992).
Schnell, H. M. et al. Structures of chaperone-associated assembly intermediates reveal coordinated mechanisms of proteasome biogenesis. Nat. Struct. Mol. Biol. 28, 418–425 (2021).
Shang, J., Huang, X. & Du, Z. The FP domains of PI31 and Fbxo7 have the same protein fold but very different modes of protein–protein interaction. J. Biomol. Struct. Dyn. 33, 1528–1538 (2015).
Li, X., Thompson, D., Kumar, B. & DeMartino, G. N. Molecular and cellular roles of PI31 (PSMF1) protein in regulation of proteasome function. J. Biol. Chem. 289, 17392–17405 (2014).
Liu, C.-W., Corboy, M. J., DeMartino, G. N. & Thomas, P. J. Endoproteolytic activity of the proteasome. Science 299, 408–411 (2003).
Groll, M. et al. Structure of 20S proteasome from yeast at 2.4Å resolution. Nature 386, 463–471 (1997).
Kulak, N. A., Pichler, G., Paron, I., Nagaraj, N. & Mann, M. Minimal, encapsulated proteomic-sample processing applied to copy-number estimation in eukaryotic cells. Nat. Methods 11, 319–324 (2014).
Cho-Park, P. F. & Steller, H. Proteasome regulation by ADP-ribosylation. Cell 153, 614–627 (2013).
Liu, K. et al. PI31 is an adaptor protein for proteasome transport in axons and required for synaptic development. Dev. Cell 50, 509–524.e10 (2019).
Zhao, L. et al. A rare variant nonparametric linkage method for nuclear and extended pedigrees with application to late-onset Alzheimer disease via WGS data. Am. J. Hum. Genet. 105, 822–835 (2019).
Minis, A. et al. The proteasome regulator PI31 is required for protein homeostasis, synapse maintenance, and neuronal survival in mice. Proc. Natl Acad. Sci. 116, 24639–24650 (2019).
Wani, P. S., Rowland, M. A., Ondracek, A., Deeds, E. J. & Roelofs, J. Maturation of the proteasome core particle induces an affinity switch that controls regulatory particle association. Nat. Commun. 6, 6384 (2015).
Leggett, D. S. et al. Multiple associated proteins regulate proteasome structure and function. Mol. Cell 10, 495–507 (2002).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Wagner, T. et al. SPHIRE-crYOLO is a fast and accurate fully automated particle picker for cryo-EM. Commun. Biol. 2, 218 (2019).
Scheres, S. H. W. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. CryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Morin, A. et al. Collaboration gets the most out of software. eLife 2, e01456 (2013).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr., Sect. D: Biol. Crystallogr. D66, 486–501 (2010).
Croll, T. I. ISOLDE: a physically realistic environment for model building into low-resolution electron-density maps. Acta Crystallogr., Sect. D: Struct. Biol. D74, 519–530 (2018).
Liebschner, D. et al. Macromolecular structure determination using X-rays, neutrons and electrons: recent developments in Phenix. Acta Crystallogr., Sect. D: Struct. Biol. D75, 861–877 (2019).
Pfab, J., Phan, N. M. & Si, D. DeepTracer for fast de novo cryo-EM protein structure modeling and special studies on CoV-related complexes. Proc. Natl Acad. Sci. 118, e2017525118 (2021).
Celniker, G. et al. ConSurf: using evolutionary data to raise testable hypotheses about protein function. Isr. J. Chem. 53, 199–206 (2013).
Ashkenazy, H. et al. ConSurf 2016: an improved methodology to estimate and visualize evolutionary conservation in macromolecules. Nucleic Acids Res. 44, W344–W350 (2016).
Gutierrez, C. B. et al. Developing an acidic residue reactive and sulfoxide-containing MS-cleavable homobifunctional cross-linker for probing protein–protein interactions. Anal. Chem. 88, 8315–8322 (2016).
Gietz, R. D. & Sugino, A. New yeast-Escherichia coli shuttle vectors constructed with in vitro mutagenized yeast genes lacking six-base pair restriction sites. Gene 74, 527–534 (1988).
Acknowledgements
We thank D. Waterman and D. Finley for early contributions to the project, and D. Finley for assistance with chromatography, helpful advice and comments on the manuscript. We thank J. Roelofs for the Pba1/2 antibody. This work was supported by National Institutes of Health (NIH) grants R01-GM144367 (to J.H.), DP5-OD019800 (to J.H.), R01-GM074830 (to L.H.) and R01-GM130144 (to L.H.).
Author information
Authors and Affiliations
Contributions
J.H., B.V., H.M.S., M.B., J.A. and M.K.B. performed the biochemical aspects of the work. R.M.W. and S.R. performed cryo-EM sample preparation, data collection, data processing, model building and refinement, and R.M.W., S.R. and H.M.S. performed the data analysis. F.J. and L.H. performed the crosslinking experiments. H.M.S, J.H., S.R. and R.M.W. prepared the figures. J.H. wrote the paper with assistance from H.M.S., R.M.W. and S.R., as well as input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Structural and Molecular Biology thanks Youdong Mao and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Florian Ullrich, in collaboration with the Nature Structural and Molecular Biology team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Proteasome Activity in Whole Cell Extracts.
a, Proteasome activity in whole cell extracts from wild-type and fub1∆ cells, as determined using the fluorogenic substrate suc-LLVY-AMC. Open circles indicate individual data points from biologic replicates. b, Proteasome levels are comparable in whole cell extracts. Extracts were prepared and analyzed by SDS-PAGE followed by immunoblotting with indicated antibodies. Alpha5 is a CP subunit while Pgk1 represents a loading control. Uncropped images for panel b and data for panel a are available as source data.
Extended Data Fig. 2 Additional Analysis of Fub1 Binding to alpha3Δ CP.
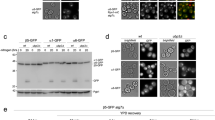
a, Purified CP (1.4 µg) from the indicated strains was analyzed by native gel electrophoresis followed by immunoblotting with antibodies recognizing Fub1 and Alpha5. These results confirm that the Fub1-immunoreactive species do indeed represent Fub1. b, Purified CP (1.1 µg) from the indicated strains was analyzed by native gel electrophoresis followed by immunoblotting with antibodies recognizing Fub1 and Blm10. These results indicate that the two supra-20S species contain Blm10. The identity of the highest molecular weight species, however, remains uncertain as doubly Blm10-capped CP was not visualized by cryo-EM (see Extended Data Fig. 3, below). Similar results were obtained in two independent experiments.
Extended Data Fig. 3 Cryo-EM Classification of CP Species.
Processing scheme for classification and refinement of proteasome species. “Junk” classes are colored grey while identifiable species are colored by species. All 3D classification steps were carried out in cryoSPARC.
Extended Data Fig. 4 Cryo-EM Data Analysis for CP Species.
a, Representative micrograph of proteasome particles embedded in vitreous ice. Scale bar = 500 Å. A total of 12,834 micrographs were collected from a single multi-day experiment. b, Selected 2D class averages of 20 S particles. c, Local resolution slice through the α3Δ + Fub1 containing reconstruction. Local resolution values vary between ~3.0–5.0 Å in this region. CP density around Fub1 is shown for orientation. d, Reconstructions of Fub1-containing (top panels) and Fub1-lacking (bottom panels) CP filtered and colored by local resolution (left), gold-standard Fourier shell correlation (FSC) curves from cryoSPARC (middle), and viewing direction distribution plots (right). Resolution determined at FSC = 0.143.
Extended Data Fig. 5 Further Analysis of β2 Inhibition by Fub1 Peptides.
Mapping of the inhibitory effect of the β2 peptide. The structure of this part of Fub1 is shown above, although the last three residues were not resolved.
Extended Data Fig. 6 Additional Low Resolution Density in the α3Δ + Fub1 Structure. Shown.
Shown is the same low pass filtered map from Fig. 7d. In addition to the capping density shown in Fig. 7d, there is further density from difference maps (shown in green) within the CP barrel that extends through the α-ring and into the β-ring. Difference density has been further gaussian filtered and additional unconnected density omitted for clarity.
Supplementary information
Supplementary Information
Supplementary Tables 1–3 and Supplementary Fig. 1.
Source data
Source Data Fig. 1
Raw data.
Source Data Fig. 1
Unprocessed blots.
Source Data Fig. 2
Unprocessed gel.
Source Data Fig. 5
Raw data.
Source Data Fig. 6
Unprocessed blots.
Source Data Fig. 7
Raw data.
Source Data Extended Data Fig. 1a
Raw data.
Source Data Extended Data Fig. 1b
Unprocessed blot.
Rights and permissions
About this article
Cite this article
Rawson, S., Walsh, R.M., Velez, B. et al. Yeast PI31 inhibits the proteasome by a direct multisite mechanism. Nat Struct Mol Biol 29, 791–800 (2022). https://doi.org/10.1038/s41594-022-00808-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-022-00808-5
This article is cited by
-
Structural basis of human 20S proteasome biogenesis
Nature Communications (2024)
-
Structure of the reduced microsporidian proteasome bound by PI31-like peptides in dormant spores
Nature Communications (2022)