Abstract
Ribosomes are complex ribozymes that interpret genetic information by translating messenger RNA (mRNA) into proteins. Natural variation in ribosome composition has been documented in several organisms and can arise from several different sources. A key question is whether specific control over ribosome heterogeneity represents a mechanism by which translation can be regulated. We used RiboMeth-seq to demonstrate that differential 2′-O-methylation of ribosomal RNA (rRNA) represents a considerable source of ribosome heterogeneity in human cells, and that modification levels at distinct sites can change dynamically in response to upstream signaling pathways, such as MYC oncogene expression. Ablation of one prominent methylation resulted in altered translation of select mRNAs and corresponding changes in cellular phenotypes. Thus, differential rRNA 2′-O-methylation can give rise to ribosomes with specialized function. This suggests a broader mechanism where the specific regulation of rRNA modification patterns fine tunes translation.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
All RNA sequencing data from this study have been deposited in the GEO under series accession numbers GSE153476 (for RNA-seq data) and GSE76393 (for RiboMeth-seq data). The MS proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE repository with the dataset identifier PXD019449. Source data are provided with this paper.
Code availability
Python code for analysis of RiboMeth-seq data is available in the GitHub repository https://github.com/lundlab/RiboMeth-seq-public
References
Kurylo, C. M. et al. Endogenous rRNA sequence variation can regulate stress response gene expression and phenotype. Cell Rep. 25, 236–248.e6 (2018).
Emmott, E., Jovanovic, M. & Slavov, N. Ribosome stoichiometry: from form to function. Trends Biochem. Sci. 44, 95–109 (2019).
Krogh, N. et al. Profiling of 2′-O-Me in human rRNA reveals a subset of fractionally modified positions and provides evidence for ribosome heterogeneity. Nucleic Acids Res. 44, 7884–7895 (2016).
Taoka, M. et al. Landscape of the complete RNA chemical modifications in the human 80S ribosome. Nucleic Acids Res. 520, 640 (2018).
Parks, M. M. et al. Variant ribosomal RNA alleles are conserved and exhibit tissue-specific expression. Sci. Adv. 4, eaao0665 (2018).
Gay, D. M., Lund, A. H. & Jansson, M. D. Translational control through ribosome heterogeneity and functional specialization. Trends Biochem. Sci. https://doi.org/10.1016/j.tibs.2021.07.001 (2021).
Ferretti, M. B. & Karbstein, K. Does functional specialization of ribosomes really exist? RNA 25, 521–538 (2019).
Dinman, J. D. Pathways to specialized ribosomes: the Brussels lecture. J. Mol. Biol. 428, 2186–2194 (2016).
Sloan, K. E. et al. Tuning the ribosome: the influence of rRNA modification on eukaryotic ribosome biogenesis and function. RNA Biol. https://doi.org/10.1080/15476286.2016.1259781 (2016).
Kiss-László, Z., Henry, Y. & Bachellerie, J. P. Site-specific ribose methylation of preribosomal RNA: a novel function for small nucleolar RNAs. Cell 85, 1077–1088 (1996).
Liang, X.-H., Liu, Q. & Fournier, M. J. rRNA modifications in an intersubunit bridge of the ribosome strongly affect both ribosome biogenesis and activity. Mol. Cell 28, 965–977 (2007).
Polikanov, Y. S., Melnikov, S. V., Söll, D. & Steitz, T. A. Structural insights into the role of rRNA modifications in protein synthesis and ribosome assembly. Nat. Struct. Mol. Biol. 22, 342–344 (2015).
Baudin-Baillieu, A. et al. Nucleotide modifications in three functionally important regions of the Saccharomyces cerevisiae ribosome affect translation accuracy. Nucleic Acids Res. 37, 7665–7677 (2009).
Birkedal, U. et al. Profiling of ribose methylations in RNA by high-throughput sequencing. Angew. Chem. Int. Ed. Engl. 54, 451–455 (2015).
Krogh, N. & Nielsen, H. Sequencing-based methods for detection and quantitation of ribose methylations in RNA. Methods 156, 5–15 (2019).
Krogh, N. et al. Profiling of ribose methylations in ribosomal RNA from diffuse large B-cell lymphoma patients for evaluation of ribosomes as drug targets. Nucleic Acids Res. Cancer https://doi.org/10.1093/narcan/zcaa035 (2020).
Marcel, V. et al. Ribosomal RNA 2′ O-methylation as a novel layer of inter-tumour heterogeneity in breast cancer. Nucleic Acids Res. Cancer https://doi.org/10.1093/narcan/zcaa036 (2020).
van Riggelen, J., Yetil, A. & Felsher, D. W. MYC as a regulator of ribosome biogenesis and protein synthesis. Nat. Rev. 10, 301–309 (2010).
Coller, H. A. et al. Expression analysis with oligonucleotide microarrays reveals that MYC regulates genes involved in growth, cell cycle, signaling, and adhesion. Proc. Natl Acad. Sci. USA 97, 3260–3265 (2000).
Makarova, J. A. & Kramerov, D. A. Analysis of C/D box snoRNA genes in vertebrates: the number of copies decreases in placental mammals. Genomics 94, 11–19 (2009).
Zeller, K. I. et al. Global mapping of c-Myc binding sites and target gene networks in human B cells. Proc. Natl Acad. Sci. USA 103, 17834–17839 (2006).
Shi, Z. et al. Heterogeneous ribosomes preferentially translate distinct subpools of mRNAs genome-wide. Mol. Cell 67, 71–83.e7 (2017).
Jack, K. et al. rRNA pseudouridylation defects affect ribosomal ligand binding and translational fidelity from yeast to human cells. Mol. Cell 44, 660–666 (2011).
Wohlgemuth, I., Pohl, C. & Rodnina, M. V. Optimization of speed and accuracy of decoding in translation. EMBO J. 29, 3701–3709 (2010).
Grosjean, H. & Westhof, E. An integrated, structure- and energy-based view of the genetic code. Nucleic Acids Res. 44, 8020–8040 (2016).
Forrest, M. E. et al. Codon and amino acid content are associated with mRNA stability in mammalian cells. PLoS ONE 15, e0228730 (2020).
Hia, F. et al. Codon bias confers stability to human mRNAs. EMBO Rep. 20, e48220 (2019).
Gingold, H. et al. A dual program for translation regulation in cellular proliferation and differentiation. Cell 158, 1281–1292 (2014).
Liu, Q. & Fredrick, K. Intersubunit bridges of the bacterial ribosome. J. Mol. Biol. 428, 2146–2164 (2016).
Behrmann, E. et al. Structural snapshots of actively translating human ribosomes. Cell 161, 845–857 (2015).
Villa, E. et al. Ribosome-induced changes in elongation factor Tu conformation control GTP hydrolysis. Proc. Natl Acad. Sci. USA 106, 1063–1068 (2009).
Paci, M. & Fox, G. E. Centers of motion associated with EF-Tu binding to the ribosome. RNA Biol. 13, 524–530 (2016).
Fagan, C. E. et al. Reorganization of an intersubunit bridge induced by disparate 16S ribosomal ambiguity mutations mimics an EF-Tu-bound state. Proc. Natl Acad. Sci. USA 110, 9716–9721 (2013).
McClory, S. P., Leisring, J. M., Qin, D. & Fredrick, K. Missense suppressor mutations in 16S rRNA reveal the importance of helices h8 and h14 in aminoacyl-tRNA selection. RNA 16, 1925–1934 (2010).
Hoffer, E. D., Maehigashi, T., Fredrick, K. & Dunham, C. M. Ribosomal ambiguity (ram) mutations promote the open (off) to closed (on) transition and thereby increase miscoding. Nucleic Acids Res. 47, 1557–1563 (2019).
Hanson, G. & Coller, J. Codon optimality, bias and usage in translation and mRNA decay. Nat. Rev. 19, 20–30 (2018).
Schuller, A. P., Wu, C. C.-C., Dever, T. E., Buskirk, A. R. & Green, R. eIF5A functions globally in translation elongation and termination. Mol. Cell 66, 194–205.e5 (2017).
Juszkiewicz, S. & Hegde, R. S. Initiation of quality control during poly(A) translation requires site-specific ribosome ubiquitination. Mol. Cell 65, 743–750.e4 (2017).
Liang, X.-H., Liu, Q. & Fournier, M. J. Loss of rRNA modifications in the decoding center of the ribosome impairs translation and strongly delays pre-rRNA processing. RNA 15, 1716–1728 (2009).
Esguerra, J., Warringer, J. & Blomberg, A. Functional importance of individual rRNA 2′-O-ribose methylations revealed by high-resolution phenotyping. RNA 14, 649–656 (2008).
Mills, E. W. & Green, R. Ribosomopathies: there’s strength in numbers. Science 358, eaan2755 (2017).
Imami, K. et al. Phosphorylation of the ribosomal protein RPL12/uL11 affects translation during mitosis. Mol. Cell 72, 84–98.e9 (2018).
McMahon, M. et al. A single H/ACA small nucleolar RNA mediates tumor suppression downstream of oncogenic RAS. Elife 8, E8603 (2019).
Leppek, K. et al. Gene- and species-specific Hox mRNA translation by ribosome expansion segments. Mol. Cell https://doi.org/10.1016/j.molcel.2020.10.023 (2020).
Wolf, S., Sameshima, M., Liebhaber, S. A. & Schlessinger, D. Regulation of ribosomal ribonucleic acid levels in growing, 3H-arrested, and crisis-phase WI-38 human diploid fibroblasts. Biochemistry 19, 3484–3490 (1980).
Bunz, F. et al. Requirement for p53 and p21 to sustain G2 arrest after DNA damage. Science 282, 1497–1501 (1998).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Bertero, A. et al. Optimized inducible shRNA and CRISPR/Cas9 platforms for in vitro studies of human development using hPSCs. Development 143, 4405–4418 (2016).
Lestrade, L. snoRNA-LBME-db, a comprehensive database of human H/ACA and C/D box snoRNAs. Nucleic Acids Res. 34, D158–D162 (2006).
Hebras, J., Krogh, N., Marty, V., Nielsen, H. & Cavaillé, J. Developmental changes of rRNA ribose methylations in the mouse. RNA Biol. 17, 150–164 (2020).
Douthwaite, S. & Kirpekar, F. Identifying modifications in RNA by MALDI mass spectrometry. Methods Enzymol. 425, 3–20 (2007).
Jansson, M. D., Damas, N. D., Lees, M., Jacobsen, A. & Lund, A. H. miR-339-5p regulates the p53 tumor-suppressor pathway by targeting MDM2. Oncogene 34, 1908–1918 (2015).
Willems, E., Leyns, L. & Vandesompele, J. Standardization of real-time PCR gene expression data from independent biological replicates. Anal. Biochem. 379, 127–129 (2008).
Cunningham, J. T., Pourdehnad, M., Stumpf, C. R. & Ruggero, D. Investigating Myc-dependent translational regulation in normal and cancer cells. Methods Mol. Biol. 1012, 201–212 (2013).
Ingolia, N. T., Ghaemmaghami, S., Newman, J. R. S. & Weissman, J. S. Genome-wide analysis in vivo of translation with nucleotide resolution using ribosome profiling. Science 324, 218–223 (2009).
Calviello, L. et al. Detecting actively translated open reading frames in ribosome profiling data. Nat. Methods 13, 165–170 (2016).
Ernlund, A. W., Schneider, R. J. & Ruggles, K. V. RIVET: comprehensive graphic user interface for analysis and exploration of genome-wide translatomics data. BMC Genomics 19, 809 (2018).
Liao, Y., Wang, J., Jaehnig, E. J., Shi, Z. & Zhang, B. WebGestalt 2019: gene set analysis toolkit with revamped UIs and APIs. Nucleic Acids Res. 47, W199–W205 (2019).
Weingarten-Gabbay, S. et al. Comparative genetics. Systematic discovery of cap-independent translation sequences in human and viral genomes. Science 351, aad4939 (2016).
Ramírez, F. et al. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res. 44, W160–W165 (2016).
Zerbino, D. R., Johnson, N., Juettemann, T., Wilder, S. P. & Flicek, P. WiggleTools: parallel processing of large collections of genome-wide datasets for visualization and statistical analysis. Bioinformatics 30, 1008–1009 (2014).
Zhang, J. et al. Measuring energy metabolism in cultured cells, including human pluripotent stem cells and differentiated cells. Nat. Protoc. 7, 1068–1085 (2012).
Belin, S. et al. Dysregulation of ribosome biogenesis and translational capacity is associated with tumor progression of human breast cancer cells. PLoS ONE 4, e7147 (2009).
Kulak, N. A., Pichler, G., Paron, I., Nagaraj, N. & Mann, M. Minimal, encapsulated proteomic-sample processing applied to copy-number estimation in eukaryotic cells. Nat. Methods 11, 319–324 (2014).
Rappsilber, J., Mann, M. & Ishihama, Y. Protocol for micro-purification, enrichment, pre-fractionation and storage of peptides for proteomics using StageTips. Nat. Protoc. 2, 1896–1906 (2007).
Thompson, A. et al. Tandem mass tags: a novel quantification strategy for comparative analysis of complex protein mixtures by MS/MS. Anal. Chem. 75, 1895–1904 (2003).
Zhu, Y. et al. DEqMS: a method for accurate variance estimation in differential protein expression analysis. Mol. Cell. Proteom. 19, 1047–1057 (2020).
Acknowledgements
The Lund laboratory is supported by grants from the Danish Council for Independent Research (Sapere Aude program 418 3-00179B); the Novo Nordisk Foundation (NNF18OC0030656); the Lundbeck Foundation (R198-2015-174) and the Danish Cancer Society (R204-A12532). Furthermore, this project has received funding from the European Union’s Horizon 2020 research and innovation programme under the Marie Skłodowska-Curie grant agreement no 801481.
Author information
Authors and Affiliations
Contributions
M.D.J. and A.H.L. conceived the study. M.D.J., S.J.H., N.K., E.J., J.V.A. and P.M. designed and performed the experiments. K.A., M.D.J. and D.T. analyzed RNA sequencing data and other computational analyses. E.M.S. and M.D.J. acquired and analyzed proteomic data. K.L.A. and H.N. provided methodological and conceptual contributions. M.D.J. wrote the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Structural & Molecular Biology thanks U. Thomas Meier and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available. Anke Sparmann and Carolina Perdigoto were the primary editors on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Detection and quantification of rRNA 2’-O-me across multiple cell types.
a-g, RiboMeth-seq (RMS) scores representing fraction of 2’-O-methylation at each site in 18S (left) and 28S rRNAs (right) present in total RNA purified from the indicated cell lines. Nucleotide positions in respective rRNAs are indicated. Points represent mean RMS scores of n = 3 sequenced libraries from individual cell cultures. Error bars represent ± s.d.. Note that (f) and (g) are derived from datasets we published previously (Krogh and Jansson et al.), included here for comparison.
Extended Data Fig. 2 Variation in 2’-O-me levels sites exists at specific sites in cell model systems.
a, MS analysis of 28S:U2031 from fragment generated by RNase T1 digestion. Both methylated and unmethylated fragments are present in the spectra, as expected from the RMS score. The sequence of the fragment is shown above the spectra, with an indication of the expected RNase cleavage sites. All identified fragments are labelled in red. Arrows mark 28S:U2031 position and spectra peaks. Observed mass is shown below each fragment. b, As (a) for MS analysis of 28S:G3606 from fragment generated by RNase A digestion. Theoretical mass (in brackets) is shown for fragment containing Gm3606. c, Schematic showing base pairing of SNORD123 and target region in 28S rRNA containing U2031. d, Comparison of RiboMeth-seq (RMS) scores representing fraction of 18S rRNA molecules 2’-O-methylated at each site in 18S rRNA from HelaS3 and BJhTERT cells. Points denote mean RMS score (n = 3 libraries from individual cultures) per site. Horizontal and vertical error bars indicate ±s.d. for each condition. Sites exhibiting significant change are labelled with nucleotide position (magenta). e, RMS scores for 2’-O-me at 18S:174 and 18S:G683 in HelaS3 and BJhTERT cells. Columns indicate mean RMS score for each condition of n = 3 libraries from individual cultures, points (magenta) denote each value separately. Error bars represent ±s.d. P values (Welch’s t-test, two-tailed) are shown above brackets. f, As (d), for 28S rRNA. g, As (e), for 2’-O-me at 28S:U2031. h, Expression of MYC mRNA in BJMYC cells. RT-qPCR analysis where MYC expression is not induced, or induced for 72 or 120 h as indicated. Columns indicate mean (n = 3 independant experiments) fold-change relative to the control condition, normalized to ACTB expression. Error bars indicate 95% confidence intervals. P values (Welch’s t-test, two-tailed) are shown for pairwise comparisons, indicated by brackets. i, Western blot analysis in BJMYC cells, where MYC expression is not induced, or induced for 72 or 120 h as indicated. Protein levels of MYC, FBL, p53, MDM2, p21 and p16 are shown. Vinculin and GAPDH are included as loading controls. Positions of molecular weight markers (kDa) are indicated. Data is representative of n = 2 independent experiments. j, As (h), for expression of cell-cycle inhibitors CDKN2A (p16) and CDKN1A (p21) in BJMYC cells. k, Cell cycle profiles of BJMYC cells where MYC expression is not induced, or induced for 72 or 120 h as indicated. FACS analysis of propidium iodide (PI) stained cells, showing proportion of cells (count) against PI intensity (FL-2A). Representative data (n = 2 independent experiments) is shown. l, As (h), for expression of MYC target genes NPM1 and FBL in BJMYC cells. m, As (d), for 28S rRNA from BJMYC cells, where MYC expression is induced for 120 h or not induced. n, As (e), for 2’-O-me at 28S:U203 and 28S:U2402 in BJMYC cells, without MYC induction or with MYC induction for 72 or 120 h. o, Sequence alignment of SNORD45C guide regions with putative target sequences in 18S rRNA. Position of methylation sites at A159 (top) and C174 (bottom) is indicated by *.
Extended Data Fig. 3 Development and analysis of a model to elucidate the role of Cm174 in ribosome regulation.
a, Comparison of RiboMeth-seq (RMS) scores representing fraction of 18S rRNA molecules 2’-O-methylated at each site in HelaS3 and BJMYC cells, where MYC expression is induced for 120 h or not induced. Points denote mean RMS score (n = 3 libraries from individual cultures) per site. Horizontal and vertical error bars indicate ±s.d. for each condition. Sites exhibiting significant change are labelled with nucleotide position (magenta). b, RMS scores for 2’-O-me at 18S:174 in HelaS3 and BJMYC cells, where MYC expression is induced for 120 h or not induced. Columns indicate mean RMS score for each condition of n = 3 libraries from individual cultures, points (magenta) denote each value separately. Error bars represent ±s.d. P values (Welch’s t-test, two-tailed) are shown above brackets. c, As (a), for 28S rRNA. d, As (B) for 2’-O-me at 28S:U2031 and 28S:U2402. e, siRNA-mediated knock-down of MYC in HelaS3 (WT) cells. Expression of MYC, and canonical target genes NPM1 and FBL, as well as SNORD45C, SNORD45A and host gene RABGGTB, as determined by RT-qPCR analysis. Columns indicate mean (n = 3 independent experiments) fold-change relative to the control condition, normalized to U6 expression. Error bars indicate 95% confidence intervals. P values (Welch’s t-test, two-tailed) are shown for pairwise comparisons, indicated by brackets. f, RiboMeth-seq (RMS) scores representing fraction of 2’-O-methylation at each site in 18S (left) and 28S rRNAs (right) from HelaS345CKO cells. Nucleotide positions in respective rRNAs are indicated. RMS scores are means of n = 3 sequenced libraries from individual cell cultures. Error bars represent ± s.d. g, Expression of SNORD45C, SNORD45A and host gene RABGGTB in HelaS3 (WT) and HelaS345CKO (45KO) cells, as determined by RT-qPCR analysis. Columns indicate mean (n = 3 independent experiments) fold-change relative to the control condition, normalized to U6 expression. Error bars indicate 95% confidence intervals. P values (Welch’s t-test, two-tailed) are shown for pairwise comparisons, indicated by brackets. Note, due to high sequence similarity between SNORD45C and -A, a residual signal for SNORD45C is apparent in 45KO. h, As (a), for 28S rRNA in HelaS3 and HelaS345CKO cells. No sites exhibit significant change. i, Analysis of rRNA biogenesis pathways in HelaS3GFP (WT), HelaS345CKO-GFP (KO) and HelaS345CKO-GFP+45C (OE) cells, derived from parental HelaS3 and HelaS345CKO (please refer to section ‘Cm174 affects cellular phenotypes’ in the main text where they are formally introduced and described in more detail). Schematic (left) showing rRNA processing steps and location of probes ‘a’ and ‘b’. Levels of pre-rRNA and processing intermediates as assessed by northern blot. Data shown is representative of n = 2 independent experiments. j, Percentage reads mapping to 5′ untranslated regions (5′ UTR), coding sequences (CDS) or 3′ untranslated regions (3′ UTR) of protein-coding genes, for each replicate sequencing library. Both total RNA (left) and ribosome protected fragment (right) derived reads are shown. k, Periodicity of ribosome protected fragment (RPF) reads mapped to mRNA transcripts. Metagene analysis shows normalized mean counts, at single-nucleotide resolution, representing ribosome P-site occupancy relative to start (left) and stop (right) codons from HelaS3 (WT, black) or HelaS345CKO (KO, red) libraries (n = 3 libraries from individual cultures). Number of transcripts analyzed after extreme outlier removal is given by ‘n’. l, Correlation between reads mapped per mRNA transcript in HelaS3 (WT) and HelaS345CKO (KO) RPF libraries (n = 3 libraries from individual cultures). Normalized (CPM) mean counts are plotted. Color scale indicates plotting density. ‘n’ denotes number of transcripts analyzed. Spearman’s rho and associated P value (algorithm AS 89) are shown. m, As (k), for reads mapping to mRNA from total RNA libraries. n, As (k), for correlation between reads mapping to mRNA transcripts from RPF and total RNA libraries in HelaS3 (WT) cells. o, As (m), for HelaS345CKO (KO) cells. p, Differential ribosome occupancy on mRNA transcripts in HelaS345CKO compared to HelaS3 cells. Log2 fold-change in read counts derived from analysis of RPF libraries (Log2FC RPF) and corresponding -Log10 of Benjamini-Hochberg Padj values are plotted. Transcripts changing significantly between conditions are colored (purple). ‘n’ denotes total number of transcripts analyzed. The number of transcripts showing reduced (DN) or increased (UP) translation is also shown. q, As (o), for mRNA transcripts displaying altered expression level (Log2FC mRNA), as measured by analysis of read counts mapped to mRNA transcripts derived from total RNA libraries. Those changing significantly between conditions are colored (cyan). r, Gene ontology analysis of mRNA transcripts displaying elevated expression in HelaS345CKO cells. GO categories (FDR < 0.05) for biological process are labelled. Number of genes overlapping with each biological process GO category is indicated by the color scale gradient (count). No GO terms were found to be overrepresented for transcripts with reduced expression levels (at FDR < 0.05).
Extended Data Fig. 4 mRNA codon composition determines translational regulation by Cm174 methylated ribosomes.
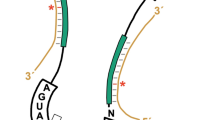
a, Gene set enrichment analysis comparing genes that exhibit Cm174-dependent translational regulation to a high-confidence set harboring IRES elements. Number of genes in both sets is given by ‘n’. Values obtained for FDR q-value, nominal (NOM) p-value, FWER p-value and normalized enrichment score (NES) are indicated. b, Luciferase reporter assays for IRES versus 5’cap-mediated translation initiation in HelaS3 (WT) compared to HelaS345CKO (KO) cells. MYC and IGFR1 IRES elements were tested. Columns indicate means for each condition (n = 3 independent experiments), points (black) denote each value separately. P values (Welch’s t-test, two-tailed) are shown, indicated by brackets. c, Principal component analysis of codon usage frequency in transcripts exhibiting increased (TC-UP, red), decreased (TC-DN, blue), and not-significantly altered (NS, grey) expression in HelaS345CKO cells. d, Usage frequencies for each codon in mRNA transcripts exhibiting increased (TC-UP), decreased (TC-DN) expression in HelaS345CKO cells. Median frequencies for all transcripts are plotted in each case. Codons are color coded by amino acid as indicated. Codons changing with Log2FC > 1, minimum frequency of 0.02 and Padj value < 0.001 (Bonferroni corrected Wilcoxon tests, unpaired, two-sided) are annotated by codon sequence. e, Fraction of G or C nucleotides present in codons comprising mRNA transcripts in each mRNA expression regulation category, down-regulated (TC-DN, blue), not-significantly regulated (NS, grey) or Up-regulated (TC-UP, red). Comparisons for entire codons (codon) or first, second and third codon positions (pos 1-3) are shown separately. Padj values for comparisons are shown above brackets. f, Codon usage frequencies in mRNA transcripts exhibiting increased (TL-UP) or decreased (TL-DN) translation in HelaS345CKO cells. Median frequency of each codon is plotted in each case. Colors denote groups of codons enriched in mRNAs associated with indicated meta-GO categories defined in a previous study: DIFF (differentiation), PROLIF (proliferation), OTHER (not related to proliferation or differentiation), and NA (codons without a cognate anti-codon match). g, Conservation of h8 and h14. Schematic showing sequence and base-pairing interactions of h8 and h14 in E.coli, human and yeast 18S rRNA. Conserved residues (red) and 2’-O-methylated residues (bold font) are indicated. Position of Cm174 in human is highlighted by an arrow.
Extended Data Fig. 5 Cm174 ribosome methylation influences cell proliferation and metabolism.
a, Cell proliferation in HelaS3 (WT, black) and HelaS345CKO (KO, red) cells, measured by crystal violet assay at specified times. Points represent mean of n = 3 independent experiments. Error bars represent ± s.d. P values (Welch’s t-test, two-tailed), indicated by brackets are shown comparisons at days 3 and 4. b, Cell cycle phase analysis in HelaS3 (WT) and HelaS345CKO (45CKO) cells. Incorporation of EdU (+EdU), as determined by FACS. Cell seeding was optimized to ensure approximately equal densities at harvest, in order to reduce the effect of differing proliferation rates on cell cycle phase duration. Percentage of cells in gates representing cell cycle phases G1, S, G2/M is shown. Bottom panels show control samples not pulsed with EdU (-EdU). Data shown is representative of n = 3 independent experiments. c, Quantification of cell cycle phase analysis, as described in (b). Columns represent mean percentage cells in each phase from n = 3 independent experiments. Error bars represent ±SEM. P values (two-way ANOVA with Sidak correction for multiple comparisons) are indicated by brackets. d, RiboMeth-seq (RMS) scores representing fraction of 2’-O-methylation at each site in 18S (left) and 28S rRNAs (right) HelaS3GFP-45CKO control cell line, where SNORD45C is not present within the EGFP intron. Nucleotide positions in respective rRNAs are indicated. Points denote mean RMS score (n = 3 libraries from individual cultures) per site. Error bars represent ± s.d. e, As (d), for HelaS3GFP-45CKO+45C cell line, where SNORD45C is inserted inside the EGFP intron. f, Comparison of RiboMeth-seq (RMS) scores in 28S rRNA from HelaS3GFP-45CKO and HelaS3GFP-45CKO+45C cells. Points denote mean RMS score (n = 3 libraries from individual cultures) per site. Horizontal and vertical error bars indicate ±s.d. for each condition. No sites exhibit significant change. g, Expression of SNORD45C, SNORD45A and host gene RABGGTB in HelaS3GFP (WT), HelaS3GFP-45CKO (45KO) and HelaS3GFP-45CKO+45C (45KO + 45C) cells, as determined by RT-qPCR analysis. Columns indicate mean (n = 3 independant experiments) fold-change relative to the control condition, normalized to invariant SNORD46 expression. Error bars indicate 95% confidence intervals. P values (Welch’s t-test, two-tailed) are shown for pairwise comparisons, indicated by brackets. h, Oxygen consumption rates (OCR) in HelaS3 (WT) compared to HelaS345CKO (45CKO) cells, measured by Seahorse XFe96 analyzer mito-stress test assay. Trace displays OCR over time, at basal levels and following injection of with compounds modulating cellular respiration (oligomycin, FCCP and a combination of rotenone and antimycin A) at the indicated time points. Representative data from n = 4 independent experiments is shown. Error bars represent ± SEM. i, Respiration and key parameters of mitochondrial function in HelaS3 (WT) compared to HelaS345CKO (45CKO) cells. Quantification of OCR as described in (A) measuring basal respiration, ATP synthesis, proton leak and uncoupled respiration. Columns indicate mean OCR for each condition (n = 4 independent experiments), points (black) denote each value separately. Error bars represent ± SEM. P values (Welch’s t-test, two-tailed) for comparisons are shown, indicated by brackets. j, As (h), for HelaS3GFP (WT-GFP), HelaS345CKO-GFP (45CKO-GFP) and HelaS345CKO-GFP+45C (45CKO-GFP+45C) cells. Representative data from n = 5 independent experiments is shown.
Extended Data Fig. 6 Ribosome regulation by Cm174 alters the levels of proteins involved in distinct cellular processes and affects translation of codons with differing GC content.
a, Overview of protein-level quantitation values between replicate mass spectrometry analyses. Coefficient of variation (CV) is below 20% for >96% of protein quantitation values in for both HelaS3 (WT, left) and HelaS345CKO (KO, right) samples (n = 4 per condition), indicating high reproducibility between replicates (n = 4 independent protein samples per condition). b, Correlation between mRNA translation and protein abundance in HelaS3 (WT) cells. Points represent mean total RNA (RNA) read count per million mapped reads (CPM) for each transcript (n = 3 libraries from independent cultures per condition, from ribosome profiling) and corresponding mean protein abundance for the same gene (n = 4 independent protein samples per condition, from MS analysis). Color scale indicates plotting density. ‘n’ denotes number of genes/proteins. Spearman’s rho and associated P value (algorithm AS 89) are shown. c, as (b), for HelaS345CKO (45CKO) cells. d, Gene ontology analysis of overlap between PA-DN and TL-DN sets. Top 10 GO categories for biological process are labelled. Number of genes overlapping with each GO category is indicated by the color scale gradient (count). e, As (d), for overlap between PA-UP and TL-UP sets. f, Principal component analysis of codon usage frequency in transcripts exhibiting significant Log2FC changes in both proteomic and ribosome profiling analyses of HeLaS3 and HelaS345CKO cells. Increased (PA-UP & TL-UP, red), decreased (PA-DN & TL-DN, blue), not-significantly altered (NS, grey), and opposite (magenta) change categories are indicated. g, Usage frequencies for each codon in transcripts included in PA-UP & TL-UP versus PA-DN & TL-DN sets. Median frequencies are plotted in each case. Codons are color coded by amino acid as indicated. Codons changing with Log2FC >1, minimum frequency of 0.02 and Padj value < 0.001 (Bonferroni corrected Wilcoxon tests, unpaired, two-sided) are annotated by codon sequence. h, Fraction of G or C nucleotides present in codons comprising mRNA transcripts in included in PA-DN & TL-DN (blue), PA-UP & TL-UP (red), or not-significantly altered (NS, grey) sets. Comparisons for entire codons (codon) or first, second and third codon positions (pos 1-3) are shown separately. Padj values (Bonferroni corrected Wilcoxon tests, unpaired, two-sided) for comparisons are given above brackets. i, High content imaging assay of codon reporter cell lines (replicate 1), containing either AT or GC rich codon inserts as indicated. mTagBFP2:mCherry ratio, after expression of the reporter cassette was induced for 24 h. The center of the box represents the median; the top and the bottom edges of the box represent the third and first quartiles, respectively. Whiskers represent minima and maxima, excluding outliers beyond 1.5× IQR. Bonferroni Padj values derived from Wilcoxon tests, two-tailed for each pairwise comparison indicated by brackets are given. See Supplementary Table 7 for all possible statistical comparisons. The number of double-positive mTagBFP2 and mCherry cells assayed in each condition, from left to right: n = 7667, 1828, 1839, 9081, 3545, 2914.
Supplementary information
Supplementary Tables
Supplementary Tables 1–7. Table 1: Evolutionary conservation of 2′-O-me sites. Table 2: Ribosome profiling analysis. Table 3: Codon enrichment in translationally regulated mRNAs. Table 4: MS proteomic analysis. Table 5: Oligonucleotides used in this study. Table 6: List of canonical transcripts and selection strategy. Table 7: Full statistical analysis of codon reporter assays.
Source data
Source Data Fig. 1
Fig. 1a,b. RiboMeth-seq data BJ-MYC − DOX versus BJ-MYC + DOX 120 h for 18S rRNA. Fig. 1e. SYBR green qPCR data BJ-MYC DOX induction.
Source Data Fig. 2
Fig. 2a,b. RiboMeth-seq data HeLaS3-45CKO versus HelaS3-WT for 18S rRNA. Fig. 2d,e. [35S]Methionine/cysteine incorporation densitometry data.
Source Data Fig. 5
Fig. 5b,c. RiboMeth-seq data HeLaS3-45CKO-GFP versus HeLaS3-45CKO-GFP + 45C for 18S rRNA. Fig. 5d. Seahorse data for metabolic analysis of OCR in HeLaS3-WT-GFP, HeLaS3-45CKO-GFP and HelaS3-45CKO-GFP + 45C. Experiments 1–5.
Source Data Extended Data Fig. 1
Fully analyzed RMS data for cell lines shown in Extended Data Fig. 1.
Source Data Extended Data Fig. 2
Extended Data Fig. 2d–g. RiboMeth-seq data BJ-hTERT versus HeLaS3 for 18S and 28S rRNAs. Extended Data Fig. 2h,j,l. TaqMan qPCR data BJ-MYC DOX induction. Extended Data Fig. 2m,n. RiboMeth-seq data BJ-MYC - DOX versus BJ-MYC + DOX 120 h for 28S rRNA.
Source Data Extended Data Fig. 2
Unprocessed western blots BJ-MYC DOX induction.
Source Data Extended Data Fig. 3
Extended Data Fig. 3a–d. RiboMeth-seq data BJ-MYC +120 h versus HeLaS3 for 18S and 28S rRNAs. Extended Data Fig. 3e. TaqMan qPCR data HeLaS3 siRNA MYC 48 h. Extended Data Fig. 3e. SYBR green qPCR data HeLaS3 siRNA MYC 48 h. Extended Data Fig. 3f. Fully analyzed RMS data for HeLaS3-45CKO. Extended Data Fig. 3g. SYBR green qPCR data HeLaS3 versus HeLaS3-45CKO. Extended Data Fig. 3h. RiboMeth-seq data HeLaS3-45CKO versus HelaS3-WT for 28S rRNA.
Source Data Extended Data Fig. 4
Extended Data Fig. 4b. Luciferase firefly/Renilla values for IRES- versus cap-dependent translation reporter assays.
Source Data Extended Data Fig. 5
Extended Data Fig. 5c. Percentage of cells in cell cycle phases from EdU labeling analysis in HeLaS3 and HelaS3-45CKO. Extended Data Fig. 5d. Fully analyzed RMS data for HeLaS3-45CKO-GFP. Extended Data Fig. 5e. Fulyl analyzed RMS data for HeLaS3-45CKO-GFP + 45C. Extended Data Fig. 5f. RiboMeth-seq data HeLaS3-45CKO-GFP versus HelaS3-45CKO-GPP + 45C for 28S rRNA. Extended Data Fig. 5g. SYBR green qPCR data HeLaS3-WT-GFP versus HeLaS3-45CKO-GFP versus HeLaS3-45CKO-GFP + 45C. Extended Data Fig. 5i. Seahorse data for metabolic analysis of OCR in HeLaS3, HeLaS3-45CKO.
Rights and permissions
About this article
Cite this article
Jansson, M.D., Häfner, S.J., Altinel, K. et al. Regulation of translation by site-specific ribosomal RNA methylation. Nat Struct Mol Biol 28, 889–899 (2021). https://doi.org/10.1038/s41594-021-00669-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-021-00669-4
This article is cited by
-
Translation efficiency driven by CNOT3 subunit of the CCR4-NOT complex promotes leukemogenesis
Nature Communications (2024)
-
FUS regulates a subset of snoRNA expression and modulates the level of rRNA modifications
Scientific Reports (2023)
-
Pervasive transcriptome interactions of protein-targeted drugs
Nature Chemistry (2023)
-
A single pseudouridine on rRNA regulates ribosome structure and function in the mammalian parasite Trypanosoma brucei
Nature Communications (2023)
-
RNA modifications in cancer
British Journal of Cancer (2023)