Abstract
TFIID is a cornerstone of eukaryotic gene regulation. Distinct TFIID complexes with unique subunit compositions exist and several TFIID subunits are shared with other complexes, thereby conveying precise cellular control of subunit allocation and functional assembly of this essential transcription factor. However, the molecular mechanisms that underlie the regulation of TFIID remain poorly understood. Here we use quantitative proteomics to examine TFIID submodules and assembly mechanisms in human cells. Structural and mutational analysis of the cytoplasmic TAF5–TAF6–TAF9 submodule identified novel interactions that are crucial for TFIID integrity and for allocation of TAF9 to TFIID or the Spt-Ada-Gcn5 acetyltransferase (SAGA) co-activator complex. We discover a key checkpoint function for the chaperonin CCT, which specifically associates with nascent TAF5 for subsequent handover to TAF6–TAF9 and ultimate holo-TFIID formation. Our findings illustrate at the molecular level how multisubunit complexes are generated within the cell via mechanisms that involve checkpoint decisions facilitated by a chaperone.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Atomic coordinates and crystallographic structure factors have been deposited in the PDB under accession code 6F3T. Proteomics data have been deposited in the PRIDE database under accession code PXD011293. Source data for Figs. 1, 3–5 and Supplementary Figs. 6, 8 and 9 are available in the online version of the paper. All other data supporting findings in this study are available from the corresponding authors upon reasonable request.
References
Roeder, R. G. Transcriptional regulation and the role of diverse coactivators in animal cells. FEBS Lett. 579, 909–915 (2005).
Levine, M., Cattoglio, C. & Tjian, R. Looping back to leap forward: transcription enters a new era. Cell 157, 13–25 (2014).
Bieniossek, C. et al. The architecture of human general transcription factor TFIID core complex. Nature 493, 699–702 (2013).
Louder, R. K. et al. Structure of promoter-bound TFIID and model of human pre-initiation complex assembly. Nature 531, 604–609 (2016).
Muller, F., Zaucker, A. & Tora, L. Developmental regulation of transcription initiation: more than just changing the actors. Curr. Opin. Genet. Dev. 20, 533–540 (2010).
Helmlinger, D. & Tora, L. Sharing the SAGA. Trends Biochem. Sci. 42, 850–861 (2017).
Baptista, T. et al. SAGA is a general cofactor for RNA polymerase II transcription. Mol. Cell 68, 130–143 (2017).
Warfield, L. et al. Transcription of nearly all yeast RNA polymerase II-transcribed genes is dependent on transcription factor TFIID. Mol. Cell 68, 118–129 (2017).
Spedale, G., Timmers, H. T. & Pijnappel, W. W. ATAC-king the complexity of SAGA during evolution. Genes Dev. 26, 527–541 (2012).
Hernandez, N. TBP, a universal eukaryotic transcription factor? Genes Dev. 7, 1291–1308 (1993).
Clapier, C. R. & Cairns, B. R. The biology of chromatin remodeling complexes. Annu. Rev. Biochem. 78, 273–304 (2009).
Ellis, R. J. Assembly chaperones: a perspective. Phil. Trans. R. Soc. B 368, 20110398 (2013).
Schopf, F. H., Biebl, M. M. & Buchner, J. The HSP90 chaperone machinery. Nat. Rev. Mol. Cell Biol. 18, 345–360 (2017).
Ramos, P. C. & Dohmen, R. J. PACemakers of proteasome core particle assembly. Structure 16, 1296–1304 (2008).
Venkatesh, S. & Workman, J. L. Histone exchange, chromatin structure and the regulation of transcription. Nat. Rev. Mol. Cell Biol. 16, 178–189 (2015).
Lopez, T., Dalton, K. & Frydman, J. The mechanism and function of group II chaperonins. J. Mol. Biol. 427, 2919–2930 (2015).
Pines, A. et al. TRiC controls transcription resumption after UV damage by regulating Cockayne syndrome protein A. Nat. Commun. 9, 1040 (2018).
Muñoz, I. G. et al. Crystal structure of the open conformation of the mammalian chaperonin CCT in complex with tubulin. Nat. Struct. Mol. Biol. 18, 14–19 (2011).
Altelaar, A. F., Munoz, J. & Heck, A. J. Next-generation proteomics: towards an integrative view of proteome dynamics. Nat. Rev. Genet. 14, 35–48 (2013).
Ahrens, C. H., Brunner, E., Qeli, E., Basler, K. & Aebersold, R. Generating and navigating proteome maps using mass spectrometry. Nat. Rev. Mol. Cell Biol. 11, 789–801 (2010).
Wright, K. J., Marr, M. T. II & Tjian, R. TAF4 nucleates a core subcomplex of TFIID and mediates activated transcription from a TATA-less promoter. Proc. Natl Acad. Sci. USA 103, 12347–12352 (2006).
Trowitzsch, S. et al. Cytoplasmic TAF2-TAF8-TAF10 complex provides evidence for nuclear holo-TFIID assembly from preformed submodules. Nat. Commun. 6, 6011 (2015).
Gupta, K. et al. Architecture of TAF11/TAF13/TBP complex suggests novel regulation properties of general transcription factor TFIID. eLife 6, e30395 (2017).
Scheer, E., Delbac, F., Tora, L., Moras, D. & Romier, C. TFIID TAF6–TAF9 complex formation involves the HEAT repeat-containing C-terminal domain of TAF6 and is modulated by TAF5 protein. J. Biol. Chem. 287, 27580–27592 (2012).
Gangloff, Y. G., Romier, C., Thuault, S., Werten, S. & Davidson, I. The histone fold is a key structural motif of transcription factor TFIID. Trends Biochem. Sci. 26, 250–257 (2001).
van Nuland, R. et al. Quantitative dissection and stoichiometry determination of the human SET1/MLL histone methyltransferase complexes. Mol. Cell. Biol. 33, 2067–2077 (2013).
Schwanhausser, B. et al. Global quantification of mammalian gene expression control. Nature 473, 337–342 (2011).
Bhattacharya, S., Takada, S. & Jacobson, R. H. Structural analysis and dimerization potential of the human TAF5 subunit of TFIID. Proc. Natl Acad. Sci. USA 104, 1189–1194 (2007).
Xie, X. et al. Structural similarity between TAFs and the heterotetrameric core of the histone octamer. Nature 380, 316–322 (1996).
Miyata, Y., Shibata, T., Aoshima, M., Tsubata, T. & Nishida, E. The molecular chaperone TRiC/CCT binds to the Trp-Asp 40 (WD40) repeat protein WDR68 and promotes its folding, protein kinase DYRK1A binding, and nuclear accumulation. J. Biol. Chem. 289, 33320–33332 (2014).
Cuéllar, J. et al. The structure of CCT–Hsc70NBD suggests a mechanism for Hsp70 delivery of substrates to the chaperonin. Nat. Struct. Mol. Biol. 15, 858–864 (2008).
Han, Z. et al. Structural basis for the specific recognition of methylated histone H3 lysine 4 by the WD-40 protein WDR5. Mol. Cell 22, 137–144 (2006).
Haffke, M., Viola, C., Nie, Y. & Berger, I. Tandem recombineering by SLIC cloning and Cre–LoxP fusion to generate multigene expression constructs for protein complex research. Methods Mol. Biol. 1073, 131–140 (2013).
Fitzgerald, D. J. et al. Protein complex expression by using multigene baculoviral vectors. Nat. Methods 3, 1021–1032 (2006).
Kabsch, W. XDS. Acta Crystallogr. D 66, 125–132 (2010).
Kabsch, W. Integration, scaling, space-group assignment and post-refinement. Acta Crystallogr. D 66, 133–144 (2010).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Cowtan, K. The Buccaneer software for automated model building. 1. Tracing protein chains. Acta Crystallogr. D 62, 1002–1011 (2006).
Terwilliger, T. SOLVE and RESOLVE: automated structure solution, density modification and model building. J. Synchrotron Radiat. 11, 49–52 (2004).
Terwilliger, T. C. Automated structure solution, density modification and model building. Acta Crystallogr. D 58, 1937–1940 (2002).
Kelley, L. A. & Sternberg, M. J. Protein structure prediction on the Web: a case study using the Phyre server. Nat Protoc. 4, 363–371 (2009).
Vagin, A. & Teplyakov, A. Molecular replacement with MOLREP. Acta Crystallogr. D 66, 22–25 (2010).
Langer, G., Cohen, S. X., Lamzin, V. S. & Perrakis, A. Automated macromolecular model building for X-ray crystallography using ARP/wARP version 7. Nat Protoc. 3, 1171–1179 (2008).
Carey, M. F., Peterson, C. L. & Smale, S. T. Dignam and Roeder nuclear extract preparation. Cold Spring Harbor Protoc. 2009, pdb.prot5330 (2009).
Spruijt, C. G., Baymaz, H. I. & Vermeulen, M. Identifying specific protein–DNA interactions using SILAC-based quantitative proteomics. Methods Mol. Biol. 977, 137–157 (2013).
Low, T. Y. et al. Quantitative and qualitative proteome characteristics extracted from in-depth integrated genomics and proteomics analysis. Cell Rep. 5, 1469–1478 (2013).
Acknowledgements
We thank S. Trowitzsch (Goethe University Frankfurt), P. Legrand and A. Thompson (Synchrotron SOLEIL), W. Vonk (Princess Maxima Centre Utrecht), R. Baas (Netherlands Cancer Institute Amsterdam) and M. Vermeulen (Radboud University Nijmegen) for assistance and reagents. We greatly appreciate discussion with R. Sawakar (MPI for Immunobiology and Epigenetics). This research was supported by the Netherlands Organization for Scientific Research (NWO) grants 022.004.019 (S.V.A.), ALW820.02.013 (H.T.M.T.) and 184.032.201 Proteins@Work (E.C., T.Y.L., H.R.V. and A.J.R.H.), a Kékulé fellowship from the Fonds der Chemischen Industrie (M.H.), a European Research Council Advanced grant ERC-2013-340551 (L.T.), Agence Nationale de Recherche research grants ANR-10-IDEX-0002-02 and ANR-10-LABX-0030-INRT (L.T.) and a Wellcome Trust Senior Investigator Award 106115/Z/14/Z (I.B.). This work used the platforms of the Grenoble Instruct-ERIC Center (ISBG UMS 3518 CNRS-CEA-UGA-EMBL) with support from FRISBI (ANR-10-INBS-05-02) and GRAL (ANR-10-LABX-49-01) within the Grenoble Partnership for Structural Biology (PSB). This research received support from BrisSynBio, a BBSRC/EPSRC Research Centre for synthetic biology at the University of Bristol (BB/L01386X/1).
Author information
Authors and Affiliations
Contributions
H.T.M.T. and I.B. conceived the study with input from S.V.A., M.H., L.T. and A.J.R.H. S.V.A. carried out all cell biology experiments, assisted by M.M. and E.S. The majority of the quantitative proteomics analyses were carried out by E.C. and T.Y.L., assisted by S.V.A. R.M.v.E. and H.R.V. carried out proteomics analyses of the pulse–chase experiment. M.H. carried out all recombinant protein work, crystallization and structural analysis, assisted by L.S., K.G. and with input from I.B. S.V.A., M.H., L.T., A.J.R.H., H.T.M.T. and I.B. designed experiments, interpreted data and wrote the manuscript together with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Expression analysis of WT GFP-TAF5, GFP-TAF6 and GFP-TAF9 inducible cell lines.
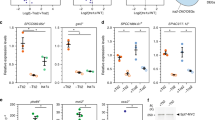
a, Expression of GFP-TAF5 and GFP-TAF6 proteins relative to their endogenous counterparts. Technical replicates n = 2. b, Relative expression levels of GFP-TAF5, GFP-TAF6 and GFP-TAF9 by immunoblotting. Technical replicates n = 2. Uncropped blot/gel images are shown in Supplementary Data 1. c, GFP mRNA analysis of GFP-TAF5, GFP-TAF6 and GFP-TAF9. Technical replicates n = 3.
Supplementary Figure 2 Limited proteolysis of the full-length TAF5–TAF6–TAF9 complex.
a, Proteolysis time course of full-length TAF5–TAF6–TAF9 at a 1:250 (wt/wt) ratio with α-chymotrypsin on ice. b, Size-exclusion chromatography profile on a Superdex S200 10/300GL column of preparative proteolysis with α-chymotrypsin at a 1:200 (wt/wt) ratio on ice for 120 min and SDS–PAGE analysis of peak fractions. Fragments highlighted in SDS–PAGE were analyzed by N-terminal protein sequencing with Edman degradation and LC–ESI/MS. c, The identified N- and C-terminal domain boundaries obtained by limited proteolysis. Uncropped blot/gel images are shown in Supplementary Data 1.
Supplementary Figure 3 Purified TAF5–TAF6–TAF9 complexes for crystallization trials.
a, Constructs in which TAF5 is lacking the N-terminal LisH domain. b, Constructs in which TAF5 retains the N-terminal LisH domain. SDS–PAGE analysis of purified complexes is shown on the left. TAF5-LisH corresponds to TAF589–800, TAF5-NTD to TAF5194–800, TAF6-HEAT to TAF61–480, TAF6-HFDL to TAF61–92RLRRRAH, TAF6-HFDS to TAF61–92, TAF9-HFDL to TAF91–139, and TAF9-HFDS to TAF91–120. Size-exclusion chromatography profiles of Superdex S200 PC3.2/300 chromatography are shown on the right. Elution volumes of molecular weight standards are indicated by triangles. Uncropped blot/gel images are shown in Supplementary Data 1.
Supplementary Figure 4 Crystal structure determination and electron density map quality of the TAF5–TAF6–TAF9 complex.
a, Optimized native crystals of TAF5–TAF6–TAF9 complex grew to 300 μm in size. b, Crystals soaked with Ta6Br12 cluster for SAD phasing experiments. Note the green color due to incorporation of the Ta6Br12 cluster. c, Diffraction pattern of native TAF5–TAF6–TAF9 complex crystals extending beyond a resolution of 2.7 Å. d, Ta6Br12-soaked crystals diffracted beyond a resolution of 3.8 Å. e–g, Quality of the initial experimental electron density map (e) improved significantly by density modification without (f) and including (g) NCS averaging. The top panel shows the central α-helix of the TAF6 HFD; the bottom panel shows a β-sheet of the TAF5 WD40 repeat. h,i, Quality of the final 2Fo – Fc electron density map, contoured at 1.5σ. A region of the TAF6–TAF9 HFD heterodimer (h) and a β-strand of the TAF5-WD40 repeat are shown (i).
Supplementary Figure 5 TAF5 NTD structures.
a, Superposition of the TAF5-NTD in the TAF5–TAF6–TAF9 crystal structure and of TAF5-NTD crystallized in isolation (PDB 2NXP; colored in gray). b, Detailed view of the side chain orientations of L298, R301 and K304. Helix α1, which is present in the isolated TAF5-NTD structure but could not be traced in the TAF5–TAF6–TAF9 complex structure is shown as a ribbon (colored in red). Note the clash between the reoriented helix α7 (residues L298 to R301) and the location of helix α1 in the isolated TAF5-NTD structure. Helices α3 and α4 are not shown for clarity.
Supplementary Figure 6 Probing the TAF5-TAF9 interface.
a, Enrichment represented in volcano plots reveals loss of TFIID and SAGA subunits with both GFP-TAF9m1 (left) and GFP-TAF9m3 (right). GFP-TAF9m2 (middle) is defective in TFIID formation only. Subunits are colored as in Fig. 1f. Each data point is plotted as the average of technical triplicates; n = 2 independent experimental replicates for the representative GFP-TAF9m1 sample. Dashed red lines denote the threshold between background and significant enrichment (two-tailed t test; FDR = 1%; S0 =1). b, Mutation of all three regions involved in forming the TAF5 interface disrupts all interactions of GFP-TAF9 except with TAF6. c, Expression of GFP-TAF9 mutant proteins compared to GFP-TAF9 WT by immunoblot. d, Confocal fluorescence microscopy shows nuclear localization of all GFP-TAF9 proteins; scale bar, 10 μm. e, Volcano plot analysis indicates that GFP-TAF5m1 (left), GFP-TAF5m2 (middle) and GFP-TAF5m3 (right) co-purify all TFIID subunits. Each data point is plotted as the average of technical triplicates. f, Relative enrichment of the co-purified TFIID subunits compared to GFP-TAF5 wild-type indicating reduced assembly of the mutants into TFIID. Each bar represents an average of technical triplicates. Error bars, s.d. of the mean. g, Expression of GFP-TAF5 mutant proteins compared to GFP-TAF5 WT by immunoblot. Source data for f is available online. Uncropped blot/gel images are shown in Supplementary Data 1.
Supplementary Figure 7 Localization and local residue environment of TAF5 mutations.
a, Overview of TA5 in cartoon representation with the WD40 repeat domain colored in blue and the NTD in cyan. Mutated residues in TAF5m1–TAF5m4 are shown as sticks in the same color code as in Fig. 2. b–f, Local residue environment around the mutated residues. Neighboring residues in the WD40 repeat domain and the NTD of TAF5 are shown within a 4-Å radius of the mutations in TAF5m1 (b), TAF5m2 (c), TAF5m3 (d) and TAF5m4 (e,f). Polar contacts are shown as black dashed lines, and only residues involved in polar contacts with the mutated residues are labeled with residue numbers for clarity. The majority of the residues mutated in TAF5 are located in loop regions and have polar contacts via or to backbone atoms. Side chain mutations in TAF5m1–TAF5m4 thus most likely do not affect the protein folding of TAF5.
Supplementary Figure 8 CCT subunits are co-enriched in GFP-TAF5 purifications.
a, GFP-TAF5 WT and mutants co-enrich CCT subunits to a different extent. Volcano plot analysis indicates that TAF5 mutants display increased enrichment of all CCT subunits as compared to WT TAF5, supporting transient association of the chaperonin with WT TAF5. TFIID subunits are colored in green, CCT subunits in magenta. Each data point is plotted as the average of technical triplicates. Dashed red lines denote the threshold between background and significant enrichment (two-tailed t test; FDR = 1%; S0 =1). n = 2 independent experimental replicates for the representative GFP-TAF5m1+2 sample. b, Relative protein abundance plot of CCT subunits indicates the highest enrichment in mutants 1 and 2 of GFP-TAF5 as well as substantial enrichment in purifications on cytoplasmic GFP-TAF5 wild-type. GFP-TAF5m1+2 has been excluded from the graph for representation purpose, as CCT enrichment is at a 1:2 ratio with the bait. Each bar represents an average of technical triplicates. Error bars, s.d. of the mean. Source data for b are available online.
Supplementary Figure 9 Pulse-chase experiments reveal transient CCT-TAF5 association.
a, Schematic representation of the forward (left) and reverse (right) pulse-chase setups. Forward: cells are treated with CHX for 30 min prior to Dox induction. Upon mRNA accumulation, the translation block is released and the newly synthesized GFP-tagged protein is followed in time. Expectations include GFP-TAF5 association with CCT upon protein synthesis stimulation. Reverse: GFP-tagged protein expression is induced for 24 h with Dox, allowing accumulation of GFP-TAF5 in two separate complexes—transiently with CCT for the newly synthesized GFP-TAF5 and stably with TFIID complex. CHX addition to the induced cells blocks synthesis of GFP-TAF5 and therefore its transient association with CCT. b, Analysis by immunoblot of CCT interaction with newly synthesized GFP-TAF5m1+2 and GFP-TAF7 followed in pulse-chase forward and reverse setups. Input represents 5% of the protein sample used in each IP. Uncropped blot/gel images are shown in Supplementary Data 1. c, qMS analysis of pulse-chase forward setup reveals gradual enrichment of TFIID subunits during GFP-TAF5 synthesis. Data are normalized to bait. Each bar represents an average of technical triplicates. Error bars, s.d. of the mean. d, siRNA-mediated TCP1 knockdown affects cellular localization of TAF5m1+2, which shifts to cytoplasmic. GFP-WDR5 localization is unaffected. GADPH siRNA (top) and non-targeting siRNAs (middle) were applied as negative controls. Scale bar, 10 μm. Technical replicates n = 2. Source data for c are available online.Source Data
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–9 and Supplementary Table 1
Supplementary Video 1
Chaperonin CCT binds TAF5 by its WD40 domain. Subsequently, TAF6–TAF98 complex engages CCT-bound TAF5 via the TAF5 NTD, resulting in release of TAF5 from CCT and formation of a stable TAF5–TAF6–TAF9 complex in the cytoplasm
Supplementary Data 1
Uncropped gel/blot images
Rights and permissions
About this article
Cite this article
Antonova, S.V., Haffke, M., Corradini, E. et al. Chaperonin CCT checkpoint function in basal transcription factor TFIID assembly. Nat Struct Mol Biol 25, 1119–1127 (2018). https://doi.org/10.1038/s41594-018-0156-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-018-0156-z
This article is cited by
-
Simple model systems reveal conserved mechanisms of Alzheimer’s disease and related tauopathies
Molecular Neurodegeneration (2023)
-
Release of Histone H3K4-reading transcription factors from chromosomes in mitosis is independent of adjacent H3 phosphorylation
Nature Communications (2023)
-
Hierarchical TAF1-dependent co-translational assembly of the basal transcription factor TFIID
Nature Structural & Molecular Biology (2023)
-
Multi-omics analyses of MEN1 missense mutations identify disruption of menin–MLL and menin–JunD interactions as critical requirements for molecular pathogenicity
Epigenetics & Chromatin (2022)
-
Structure of the human SAGA coactivator complex
Nature Structural & Molecular Biology (2021)