Abstract
Mapping a genetic perturbation to a change in phenotype is at the core of biological research. Advances in microscopy have transformed these studies, but they have largely been confined to examining a few strains or cell lines at a time. In parallel, there has been a revolution in creating synthetic libraries of genetically altered cells with relative ease. Here we describe methods that combine these powerful tools to perform live-cell imaging of pool-generated strain libraries for improved biological discovery.
Similar content being viewed by others
Main
There are currently no general models that can reliably predict the phenotypic impact of a specific genetic change, and therefore broad screens of genetic perturbations (see definition in Box 1) will be with us for the foreseeable future. Such screens come in two flavors: those that aim to figure out how cells work and those that aim to build new genetic circuits or protein functions for medical or biotechnological applications; however, the challenge of mapping genotype to phenotype is usually similar. Typical approaches include (1) knocking in/out each gene and detecting changes in the phenotype of interest, (2) altering the regulatory or coding sequence of a specific gene and monitoring the resulting change in expression dynamics or function, or (3) labeling all proteins or genomic loci and studying how these move or localize in response to an environmental signal.
Classic genetic screening techniques, such as randomly mutating cells and seeing what survives, are rapidly being replaced by newer strategies. In the last ten years, our capability to make precise genomic changes has transcended our wildest expectations, mainly owing to CRISPR–Cas9 (refs. 1,2,3). At the same time, microscopy has experienced a revolution in what temporal and spatial resolution can be achieved in living cells4,5. In this Perspective, we will focus on different approaches that combine these methods to study the impact of specific genetic changes by high-throughput live-cell imaging. To keep this piece focused, we have restricted ourselves to the imaging of synthetic libraries, which excludes the large body of literature that deals with imaging of the natural cell-to-cell variation in tissues, such as spatial transcriptomics6 or lineage tracing7, although these methods are closely related to our scope.
Arrayed libraries studied with imaging
The first examples of imaging-based screens used arrayed libraries (Box 1), where genetically modified cell strains (Box 1) were stored in physical isolation (for example, in a series of 96- or 384-well plates; Fig. 1a). Sampling the libraries in a way that preserves this order allows high-content time-lapse microscopy of many strains in one experiment.
a, Arrayed libraries have a high cost in labor to construct and spot them on plates for phenotyping, but each strain in the library can be phenotyped completely in any modality (imaging, sequencing, etc.). b, Pooled libraries where fluorescence intensity serves as the readout. Cells can be sorted by flow cytometry into bins on the basis of fluorescence intensity, and each bin is then barcoded and sequenced. The result is histograms of intensity for each perturbation in the library. c, In a competition screen, a pool-generated library has some selection pressure applied, and the populations before and after application of the pressure are then identified by sequencing and compared to determine enrichment or depletion. d, Another sequencing option is to isolate individual cells with droplet microfluidics and perform scRNA-seq, with the result being data that map a perturbation to changes in a transcriptional profile. e,f, Cells can be phenotyped on the fly, with a handful of strains of interest isolated for downstream analysis (e), or all strains can be phenotyped and fixed in place for genotyping (f). The former has the advantage of not requiring a barcode, making library construction simpler, whereas the latter approach allows larger-scale mapping of genotype to phenotype.
The need for large-scale phenotypic screens was already apparent when the technology for whole-genome sequencing became available8. Sequencing the yeast genome9 allowed for genome-scale targeted studies to replace random mutagenesis in these organisms. An international multi-laboratory consortium generated both haploid and diploid knockouts of 2,026 open reading frames (ORFs) in the yeast genome10, generating an impressive resource that allowed for repeated pooled experiments or individual study of each knockout. A proof-of-principle demonstration of a strategy to replace genes with a unique 20-nucleotide genetic barcode (Box 1) was also described11. Taking advantage of the ability to cross haploid strains, arrayed libraries of double knockouts were also constructed12,13.
In two studies, libraries were constructed that would allow phenotyping by subcellular localization. One was a mixed library of plasmids and transposon-generated epitope tags of ORFs14. In the other, chromosomal green fluorescent protein (GFP) fusions to the end of all yeast ORFs15 were created. The latter approach allows for live-cell imaging. While the authors crossed in a red fluorescent protein (RFP) fusion with a defined spatial pattern as a landmark to aid analysis, cells were manually scored for each strain until over a decade later when machine learning took over16.
On the bacterial side, Taniguchi et al. quantified the expression of >1,000 fluorescent protein (FP) fusions by live-cell single-molecule microscopy in microfluidic chips. The imaging was followed by smFISH (see Box 1) against the FP transcript, thus quantifying both transcription and translation levels in the same cell17. Alternatively, it is possible to spot individual bacterial clones from an arrayed library on agarose18 or agar pads19,20. This approach was applied to the ASKA library of FP fusions21 to generate high-resolution space–time maps of protein locations18.
While the scale of the arrayed libraries and the data produced from these studies are impressive, the methods themselves are limited by the labor involved in performing the screens. Another general drawback of arrayed libraries is that the strains have to be cultured separately, which makes it hard to perform experiments under identical conditions. This may, in turn, limit which phenotypic differences can be resolved.
Pool-synthesized cell libraries
Pool-generated (pooled) libraries (Box 1) present an alternative approach to arrayed libraries. Early examples were created by randomly mutagenizing yeast with transposons, wherein the strains were screened on the basis of fitness22 or even subcellular localization of a transposon-generated fusion epitope for immunofluorescence23 (although, for the latter, we note that when cells were imaged in pools only population-level statistics about spatial patterns could be gathered).
Targeted pooled approaches to scale up strain generation are based on leveraging designable DNA oligonucleotide pools24. Pooled library synthesis makes it significantly easier and more affordable to generate many strains than with the arrayed approach, but the genetic identity of each cell is unknown until the individual cells are genotyped. At present, libraries of hundreds of thousands of designed oligonucleotide sequences, up to 200 nucleotides in length, can be generated for US$10,000–40,000. Smaller libraries (~10,000 oligonucleotides) cost approximately US$1,000, which makes approaches based on this technology affordable. Currently, the most common approaches are to (1) make genome-wide alterations or perturbations using CRISPR-based technology with pools of guide RNAs (gRNAs) or (2) focus on depth by varying a specific genomic locus or a mobile genetic vector. The former allows for wide screens to find targets for follow-up studies25. The latter is typically used to draw precise conclusions about the effect of variation in a specific sequence, such as the contribution of each nucleotide to protein–DNA binding26, or for optimization of protein properties (for example, fluorescent proteins (FPs)27,28, recombination machinery29 and the SARS-CoV-2 receptor-binding domain30).
In terms of altering a specific DNA locus, Kinney et al.26 developed a pioneering assay in which they built a library of bacterial strains with different promoter regions in front of a gene for an FP. By sorting the library into different bins using flow cytometry and sequencing the promoter regions of the cells in each bin, they could precisely quantify the contribution of any base in each sequence position to the promoter activity (Fig. 1b). This simple but elegant screen works because the phenotypic readout is fluorescence level. To quantify expression of any gene of interest, the approach has been extended and generalized by replacing flow cytometry with single-cell RNA sequencing (scRNA-seq) and fluorescence intensity with transcript expression levels31.
A similar experimental workflow was used to measure expression of an FP under the regulation, first, of 75 transcription factors in yeast32 and, later, of all combinations of 114 promoters with 111 ribosome-binding sites in Escherichia coli33. Johns et al.34 extended the concept to a wide range of organisms by barcoding the expression of >29,000 regulatory regions from 184 different bacterial species. They determined both the transcriptional efficiency, using the RNA-to-DNA ratio for each barcode, and the translational efficiency, using SORT-seq. In all cases, barcoding of bins leverages the power of next-generation sequencing to analyze pool-generated libraries of variants in one experiment.
Straightforward screens can also be made when the phenotypic readout is the fitness of a strain in a selective environment (Fig. 1c). In such experiments, the phenotype is typically the relative frequency of each genetic barcode (and thus each genotype) in the population before and after the fitness competition. This approach was the basis for the first generation of CRISPR libraries to multiplex mapping of a perturbation to the corresponding fitness phenotype for knockout35,36,37, knockdown38,39 and activation38,40 in mammalian cell lines. Before CRISPR, conceptually similar fitness screens were made with transposons41,42,43,44 or RNA interference45.
Going beyond counting the frequency of certain barcodes in the population, single-cell sequencing can be used in multiple ways to assess the phenotype of individually perturbed cells (Fig. 1d), to determine the transcriptional state of each individual cell (e.g., droplet-based scRNA-seq46,47,48) or the state of the chromosome (e.g., scHi-C49 or scATAC-seq50). A combination of both approaches has also been demonstrated (scNMT-seq51 and sci-CAR52). Pool-generated CRISPR libraries phenotyped by single-cell sequencing have been used in contexts ranging from immortalized mammalian cell lines53,54,55 to primary immune cells from Cas9-transgenic mice55 and even in cells collected from Cas9- and GFP-transgenic mice transduced with single guide RNA (sgRNA) vectors and then injected into a new host mouse56.
Screening pooled libraries with live-cell imaging
The major limitation of phenotyping based on sequencing is that the cells are lysed in the process, and, as a consequence, all spatial, morphological and dynamic information is lost. This is unfortunate because many of the phenotypes of interest to cell biologists and microbiologists require this type of data.
In the following section, we discuss approaches for microscopy-based screening of pooled cell libraries, which allow for high-resolution imaging and time-lapse microscopy. These methods overcome most of the challenges of the arrayed imaging screens, such as the amount of work needed and the challenge of maintaining identical experimental conditions for all strains. However, because all strains are handled the same, one may need to run the whole experiment under different conditions to maintain the dynamic range. For example, if a reporter protein is expressed at very different levels across a pooled library, imaging conditions that are optimal for capturing variation in intensity for one strain may result in saturated pixel values for another strain with a higher expression level.
First, we will describe some selected recent methods, divided by the approach to genotyping: selection of a few cells with desired phenotypes (Fig. 1e) or in situ genotyping of the entire library (Fig. 1f). Next, we will discuss the relative advantages of the different approaches.
Selection of a few cells out of many for genotyping
Some of the earliest methods to visually select individual cells from a heterogeneous population used photoinducible chemistry (which fits naturally with microscopy for phenotyping) to mark cells of interest. Photostick57 was one of the first such methods (Fig. 2a). In this approach, a small molecule cross-links the selected cells to the imaging substrate upon light exposure. Non-selected cells are washed away, and the remaining cells are identified, for example, by sequencing. The authors used the method to successfully engineer hippocampal neurons with a specific firing pattern.
a, Cells are selectively attached to the substrate using photochemistry57. b, Photoconvertible molecules in or adjacent to cells are photoconverted, and the cells are sorted by flow cytometry59,61. c, Selected cells are mechanically moved to new locations64,65. d, Cells are grown and phenotyped in a microfluidic chip and moved to an exit channel by optical tweezers70. e, Cells are phenotyped on a culture dish or slide and then genotyped in situ27,80,81. f, Cells are grown and phenotyped in a microfluidic chip that allows for in situ genotyping76,77,79.
Rather than sticking the cells in place, photo-cross-linking can be used to fluorescently label cells, which can then be sorted with flow cytometry. For example, biotin-4-fluorescein was photo-cross-linked to selected cells58, and, similarly, photoconvertible quantum dots were attached to cells before selective photoactivation59. The demonstrated throughput of these methods is, however, limited to several hundred selected cells.
A next step along these lines is to have cells constitutively express a photoconvertible FP, convert the FP in cells that display a desired phenotype and then use flow cytometry to sort out cells of interest (Fig. 2b). This enabled selection of cells from tissues60 or from a large heterogeneous library61. The latter approach was used to study the efficiency of different nuclear localization signal peptides. In recent studies, where machine learning was used for characterization of phenotypes and automation for selecting and photoactivating cells, it was possible to scale the approach to thousands of gRNAs and millions of cells62. Yan et al.63 also showed that they can sort out many populations of cells in the same experiment by different degrees of photoactivation.
Multiple rounds of library imaging and selection of adherent cells were used by Piatkevich et al.64 to evolve proteins on the basis of complex criteria (Fig. 2c). In each round, a computer vision-guided automated micropipette was used to screen 300,000 cells expressing different protein constructs in ~4 h. In particular, Piatkevich et al. evolved a genetically encoded fluorescent voltage indicator, simultaneously optimizing its brightness and membrane localization.
A similar approach was taken by Wheeler et al.65. The researchers seeded a pool-generated CRISPR-edited human cell library at low density in polydimethylsiloxane (PDMS)/magnetic microwells to ensure one founder cell per microwell. The cells were observed by confocal imaging, and the microwells with interesting phenotypes were manually removed with a motorized microneedle for further studies65. In the screen, the authors identified RNA-binding proteins related to the stress-induced formation of punctate protein–RNA assemblies.
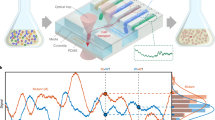
Microfluidics approaches66,67,68,69 are rapidly becoming the standard solution for bacterial imaging, as they allow for exponential growth over many generations, excellent imaging conditions, high reproducibility and highly controlled medium switches. A high-precision method for selecting individual strains from a microfluidic chip was presented by Luro et al.70 (Fig. 2d), who performed targeted mutagenesis on a genetic oscillator71. The researchers then loaded the resulting pool of strains into a microfluidic chip where they could be phenotyped for hours. Strains presenting desired characteristics (that is, more robust oscillations) were identified and individually selected with optical tweezers.
In situ genotyping of all cells in a screen
Selection-based methods are favorable when the goal is to extract a few interesting strains from a large pool. If, on the other hand, the aim is to map each genotype to its resulting phenotype, such methods quickly become impractical because of the laborious process of picking individual cells.
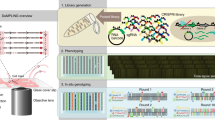
Imaging-based phenotyping of a pool-synthesized library followed by in situ genotyping72,73,74,75 of the whole library was described in 2014 (ref. 76); however, practical implementation of the approach was first demonstrated in two studies published in 2017 (refs. 27,77). Emanuel et al.27 developed a method to screen for novel FPs (Fig. 2e), where a large library of mutated FPs were expressed from plasmids in bacterial cells and imaged on a coverslip. The cells were fixed, and the fluorescence properties of the proteins were connected to the corresponding genotype through an expressed barcode RNA identified by multiplexed FISH74. The coverslip format allows for the screening of large libraries, but for a limited time span because the bacteria are not kept in a state of exponential growth. Simultaneously, our group77 implemented a microfluidic culture system that allows single-molecule microscopy in bacterial cells growing exponentially for many generations (Fig. 2f), named Dynamic u‐fluidic Microscopy‐based Phenotyping of a Library before In situ Genotyping (DuMPLING). The microfluidic design also facilitates direct spatial mapping between the phenotyped cell and the RNA FISH-based genotype barcodes. This proof-of-principle demonstration was implemented on a very small CRISPR interference (CRISPRi78; Box 1) library constructed with barcoded plasmids. CRISPRi makes it possible to target genes anywhere on the chromosome while the gRNA is expressed from one position. This simplifies in situ genotyping enormously because the genetic alteration is in the same place in all strains.
We later used the microfluidic format to identify genes related to synchronization of the division and replication cycles in E. coli79. In this study, a pooled CRISPRi library was used to monitor the effect of different gene knockdowns on DNA replication by time-lapse imaging of replication forks throughout multiple division cycles in hundreds of different strains. As in the previous study77, phenotypes were mapped to genotypes in situ by sequential FISH probing of an RNA barcode. The structure of the microfluidic system accommodates many physically isolated strains in the same field of view, making it possible to perform time-lapse microscopy on tens of thousands of bacteria with 1-min time resolution.
Implementation of a large-scale imaging-based pooled screen in human cells was performed by Feldman et al.80, who studied knockouts of 1,000 different genes with 4,000 distinct barcodes (Fig. 2e). In total, 20 million cells were analyzed across these screens, where cell nuclei were tracked using a DNA stain and the nuclear translocation of p65–mNeonGreen was assessed at each time point. Following live-cell phenotyping, cells were fixed and the identity of the disrupted gene was determined by in situ sequencing of the sgRNA sequences, as well as barcodes, using an extension of the gap-fill padlock rolling circle amplification approach72. The gap-fill approach requires that the ends of the hybridization probe bind on each side of the barcode (the remainder of the probe sequence loops out and is not hybridized), such that the polymerization reaction can integrate the barcode sequence into the circular template. The template is next amplified by rolling circle amplification. Another example of an application in eukaryotes was demonstrated by Wang et al.81, who studied the effect of 54 CRISPRi knockdowns on RNA localization to nuclear compartments. In this study, fixed cells were used for phenotyping with FISH probes and antibodies, and the genotypes were assessed by multiplexed FISH.
When to use what: comparing the strengths and weaknesses of the different methods
In the following subsections, we contrast the relative strengths and weaknesses of different imaging-based methods and give practical guidance on how to pick the appropriate approach for different applications. Please also see the roadmap in Fig. 3.
Pooled library generation
The relative ease and flexibility of library generation for selection-based methods follow from the fact that the cells can be cultured and manipulated downstream of the imaging step (Fig. 2c,d). In this way, essentially any DNA variation can be identified by nucleic acid sequencing. In principle, even a mixed population of unknown cell types can be analyzed, for example, in an environmental sample. In contrast, if the experiment requires identification of all the different clones in the library, genotyping in situ is usually necessary and generally requires a barcode nucleic acid sequence that is separate from the sequence that defines the variation or perturbation (Fig. 2e,f). Most applications use an RNA barcode so that many copies of the barcode can be generated before fixing the cells (although we note exciting recent work that allows for amplification of a desired sequence after fixation82).
A common problem for methods with barcode sequences physically distant from the genetic perturbation is the formation of mismatched barcodes (chimeras). In the CRISPRi setting, for example, there is generally a short sequence that acts as a linker between the promoter for the barcode RNA and the sgRNA gene. This linker sequence is ripe for recombination, for example, during amplification of the oligonucleotide pool (that is, chimera formation during PCR). A solution to the issue of barcode–perturbation mismatch was proposed in the context of CRISPRi screening with scRNA-seq readout54. The authors devised a construct with two promoters upstream of the sgRNA sequence resulting in two transcripts with different functions: (1) a CRISPR gRNA that is generated by RNA polymerase III and (2) a polyadenylated transcript that is generated by RNA polymerase II, which can be captured for identification by scRNA-seq.
Phenotyping: what is the time scale of the process?
For phenotypes that can be determined by a single image, like cell morphology or spatial distribution of molecules, fixed cells on a coverslip offer an easy and viable approach. However, as has been discussed elsewhere, static distributions can result in an ambiguous picture of the underlying mechanism83. In such cases, it is essential that the same cell be observed at multiple time points. For short time scales (shorter than a cell division), cells adhered to or sandwiched on a coverslip may be sufficient80. However, a microfluidic approach is indispensable when studying processes on a longer time scale, that is, more than one cell generation70,77,79.
Connecting phenotypes to genotypes: breadth versus depth
Selection-based methods have the advantage that the identified cells can be separated and cultured for downstream analysis (such as scRNA-seq or Hi-C to assay the state of the transcriptome or chromosome conformation, respectively) and to make stocks for later use (Fig. 2c,d). This naturally enables a more complete and deeper view of the phenotype resulting from each selected perturbation. In situ identification methods have not yet been successfully combined with other phenotyping assays downstream of the imaging phase, but, theoretically, they are compatible with transcriptome-scale in situ methods such as seqFISH84,85 and MERFISH86.
The obvious limitation of the selection approaches is throughput. The selected cells must be isolated either one at a time, for example, by optical tweezers70, or in a batch, for example, by photoactivation and sorting by flow cytometry. By contrast, in situ genotyping methods generally reveal the genotype of all cells that are imaged (Fig. 2e,f), giving far greater breadth to the results.
A method to increase throughput of selection-based genotyping is imaging-activated cell sorting, where cells briefly pass the microscope (for example, see refs. 87,88). However, imaging resolution has remained insufficient for phenotyping beyond distinguishing coarse differences in cell types. A recent microfluidic solution for improved image resolution uses a PDMS valve to transiently press cells against a coverslip. The trapped cells are then either kept or discarded89. While this approach allows for single-molecule fluorescence microscopy, the throughput is relatively low, and, as several cells are trapped together, additional selection steps would likely be required for most applications.
Genotyping in situ: sequencing or hybridization
A division within in situ identification approaches is the method of genotyping: sequencing or hybridization. The sequencing methods are generally extensions or adaptations of sequencing-by-ligation protocols72,73, whereas the hybridization methods are adaptations of combinatorial RNA FISH74,75. The advantage of the sequencing methods is that the barcode can be very compact, and diversity scales as 4(barcode length). For some library designs, such as in an operator library, it is even possible to read out the genetic variation directly. However, these methods require multiple enzymatic steps in situ, which has proven to be difficult in microfluidic settings77. Hybridization methods are generally less experimentally challenging. While FISH methods require longer target regions (typically 15–20 nucleotides per probe), they have been demonstrated to scale up to 60,000 variants, with a million barcodes being a viable extension29.
Future directions
The methods presented in this piece have three methodological concepts in common: library construction, genotyping and phenotyping. While great advances have been made, some aspects of the current methods have wide space for improvement. The tools for cloning and genome engineering are rapidly progressing and moving past most conceivable obstacles, even in a pooled format. In eukaryotes, new tools are moving library-scale gene editing with CRISPR toward reliable, specific and targeted gene edits (for example, by improving template delivery90,91). Methods for robust in situ identification of chromosomal barcodes at the single-cell level have been limiting, but important progress in this direction has been reported82.
Methods for imaging-based phenotyping have few limitations in enabling the study of intracellular dynamic processes in perturbed libraries beyond the usual microscopy caveats of labeling, resolution and cell toxicity. A logical phenotyping modality to combine with large knockout screens is high-dimensional unbiased imaging of cellular morphologies, such as Cell Painting92. Imaging-based screens can also be taken into the dizzyingly large combinatorial space of drug combinations93 or interactions of different combinations of cell types94,95.
The major limitation of live-cell phenotyping of libraries is throughput: it is not possible to move the microscope stage and image with sufficient speed to capture more than a few frames per second, which introduces a trade-off between the number of strains imaged and time resolution. Large field-of-view setups96 will be at least part of the solution. However, in the near future, it is likely that these experimental challenges will be small compared to those related to handling and analyzing the data. As an example, we are already acquiring 1 TB of relevant physiological data per day in DuMPLING screens79. The staggering volumes of high-quality data are an obvious match for machine learning approaches, as has been covered elsewhere97,98. As an early example, a support vector machine was used to derive meaningful phenotypic information from fluorescence microscopy data for 20 million cells representing an arrayed library of 5,000 genetically altered yeast strains99.
As with all machine learning applications, image analysis may be sensitive to biases in manually curated reference datasets; thus, in building standardized tools, it is important to test for such subjectivity in the final results100. Much work is still needed to avoid the pitfall where each laboratory designs a tool for their own dataset, making standardization of analysis nearly impossible. Starfish is an effort by the Chan–Zuckerberg Institute to combat this by standardizing dot detection in single-molecule spatial transcriptomics experiments101, which is relevant for genotyping in situ and likely will produce tools that are useful for phenotyping where dot detection is involved. Another institutional effort is CellProfiler, an endeavor to fill the space of general phenotyping tools102.
Discussion
We will soon be at a state in biological research where our capacity for making genetic perturbation and detailed imaging of complex phenotypes in individual cells will have few real limitations, except for practicalities such as cost, storage space, and imaging and analysis speed. For a long time, biological research has been akin to trying to understand how a commercial airliner works by removing one part at a time and observing when it crashes. We now have the tools to gently turn the knobs in the cockpit and at the same time monitor the flaps on the wings. However, because the number of possible genetic alterations, even in a small bacterial genome, massively exceeds the number of atoms in the universe, and the number of phenotypes that can be studied is even more bewildering, the real challenge still lies in making clever and specific experimental designs and developing the tools to analyze the data generated. So, in conclusion, focus on the biological question of interest, look at the relevant flap and turn the right knobs gently.
References
Cong, L. E. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Mali, P. et al. RNA-guided human genome engineering via Cas9. Science 339, 823–826 (2013).
Elf, J. & Barkefors, I. Single-molecule kinetics in living cells. Annu. Rev. Biochem. 88, 635–659 (2019).
Liu, T.-L. et al. Observing the cell in its native state: imaging subcellular dynamics in multicellular organisms. Science https://doi.org/10.1126/science.aaq1392 (2018).
Liao, J., Lu, X., Shao, X., Zhu, L. & Fan, X. Uncovering an organ’s molecular architecture at single-cell resolution by spatially resolved transcriptomics. Trends Biotechnol. https://doi.org/10.1016/j.tibtech.2020.05.006 (2020).
Wagner, D. E. & Klein, A. M. Lineage tracing meets single-cell omics: opportunities and challenges. Nat. Rev. Genet. 21, 410–427 (2020).
Oliver, S. G. From DNA sequence to biological function. Nature 379, 597–600 (1996).
Goffeau, A. et al. Life with 6000 genes. Science 274, 563–567 (1996).
Winzeler, E. A. et al. Functional characterization of the S. cerevisiae genome by gene deletion and parallel analysis. Science 285, 901–906 (1999).
Shoemaker, D. D. et al. Quantitative phenotypic analysis of yeast deletion mutants using a highly parallel molecular bar-coding strategy. Nat. Genet. 14, 450–456 (1996).
Costanzo, M. et al. The genetic landscape of a cell. Science 327, 425–431 (2010).
Tong, A. H. Y. Systematic genetic analysis with ordered arrays of yeast deletion mutants. Science https://doi.org/10.1126/science.1065810 (2001).
Kumar, A. et al. Subcellular localization of the yeast proteome. Genes Dev. 16, 707–719 (2002).
Huh, W.-K. et al. Global analysis of protein localization in budding yeast. Nature 425, 686–691 (2003).
Chong, Y. T. et al. Yeast proteome dynamics from single cell imaging and automated analysis. Cell 161, 1413–1424 (2015).
Taniguchi, Y. et al. Quantifying E. coli proteome and transcriptome with single-molecule sensitivity in single cells. Science 329, 533–538 (2010).
Kuwada, N. J., Traxler, B. & Wiggins, P. A. Genome-scale quantitative characterization of bacterial protein localization dynamics throughout the cell cycle. Mol. Microbiol. 95, 64–79 (2015).
Shi, H., Colavin, A., Lee, T. K. & Huang, K. C. Strain Library Imaging Protocol for high-throughput, automated single-cell microscopy of large bacterial collections arrayed on multiwell plates. Nat. Protoc. 12, 429–438 (2017).
Peters, J. M. et al. A comprehensive, CRISPR-based functional analysis of essential genes in bacteria. Cell 165, 1493–1506 (2016).
Kitagawa, M. et al. Complete set of ORF clones of Escherichia coli ASKA library (A Complete Set of E. coli K-12 ORF Archive): unique resources for biological research. DNA Res. 12, 291–299 (2005).
Smith, V., Chou, K. N., Lashkari, D., Botstein, D. & Brown, P. O. Functional analysis of the genes of yeast chromosome V by genetic footprinting. Science https://doi.org/10.1126/science.274.5295.2069 (1996).
Burns, N. et al. Large-scale analysis of gene expression, protein localization, and gene disruption in Saccharomyces cerevisiae. Genes Dev. 8, 1087–1105 (1994).
Kinney, J. B. & McCandlish, D. M. Massively parallel assays and quantitative sequence–function relationships. Annu. Rev. Genomics Hum. Genet. https://doi.org/10.1146/annurev-genom-083118-014845 (2019).
Doench, J. G. Am I ready for CRISPR? A user’s guide to genetic screens. Nat. Rev. Genet. 19, 67–80 (2018).
Kinney, J. B., Murugan, A., Callan, C. G.Jr. & Cox, E. C. Using deep sequencing to characterize the biophysical mechanism of a transcriptional regulatory sequence. Proc. Natl Acad. Sci. USA 107, 9158–9163 (2010).
Emanuel, G., Moffitt, J. R. & Zhuang, X. High-throughput, image-based screening of pooled genetic-variant libraries. Nat. Methods 14, 1159–1162 (2017).
Yoshioka-Kobayashi, K. et al. Coupling delay controls synchronized oscillation in the segmentation clock. Nature 580, 119–123 (2020).
Wannier, T. M. et al. Improved bacterial recombineering by parallelized protein discovery. Proc. Natl Acad. Sci. USA https://doi.org/10.1073/pnas.2001588117 (2020).
Starr, T. N. et al. Deep mutational scanning of SARS-CoV-2 receptor binding domain reveals constraints on folding and ACE2 binding. Cell 182, 1295–1310.e20 (2020).
Ireland, W. T. et al. Deciphering the regulatory genome of Escherichia coli, one hundred promoters at a time. eLife https://doi.org/10.7554/eLife.55308 (2020).
Sharon, E. et al. Inferring gene regulatory logic from high-throughput measurements of thousands of systematically designed promoters. Nat. Biotechnol. 30, 521–530 (2012).
Kosuri, S. et al. Composability of regulatory sequences controlling transcription and translation in Escherichia coli. Proc. Natl Acad. Sci. USA 110, 14024–14029 (2013).
Johns, N. I. et al. Metagenomic mining of regulatory elements enables programmable species-selective gene expression. Nat. Methods 15, 323–329 (2018).
Koike-Yusa, H. et al. Genome-wide recessive genetic screening in mammalian cells with a lentiviral CRISPR-guide RNA library. Nat. Biotechnol. 32, 267–273 (2014).
Shalem, O. et al. Genome-scale CRISPR–Cas9 knockout screening in human cells. Science 343, 84–87 (2014).
Zhou, Y. et al. High-throughput screening of a CRISPR/Cas9 library for functional genomics in human cells. Nature 509, 487–491 (2014).
Gilbert, L. A. et al. Genome-scale CRISPR-mediated control of gene repression and activation. Cell 159, 647–661 (2014).
Wang, T., Wei, J. J., Sabatini, D. M. & Lander, E. S. Genetic screens in human cells using the CRISPR–Cas9 system. Science 343, 80–84 (2014).
Konermann, S. et al. Genome-scale transcriptional activation by an engineered CRISPR–Cas9 complex. Nature 517, 583–588 (2015).
Gawronski, J. D. et al. Tracking insertion mutants within libraries by deep sequencing and a genome-wide screen for haemophilus genes required in the lung. Proc. Natl Acad. Sci. USA 106, 16422–16427 (2009).
Goodman, A. L. et al. Identifying genetic determinants needed to establish a human gut symbiont in its habitat. Cell Host Microbe 6, 279–289 (2009).
Langridge, G. C. et al. Simultaneous assay of every Salmonella typhi gene using one million transposon mutants. Genome Res. 19, 2308–2316 (2009).
van Opijnen, T., Bodi, K. L. & Camilli, A. Tn-seq: high-throughput parallel sequencing for fitness and genetic interaction studies in microorganisms. Nat. Methods 6, 767–772 (2009).
Boutros, M. & Ahringer, J. The art and design of genetic screens: RNA interference. Nat. Rev. Genet. 9, 554–566 (2008).
Klein, A. M. et al. Droplet barcoding for single-cell transcriptomics applied to embryonic stem cells. Cell 161, 1187–1201 (2015).
Macosko, E. Z. et al. Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell 161, 1202–1214 (2015).
Zheng, G. X. Y. et al. Massively parallel digital transcriptional profiling of single cells. Nat. Commun. 8, 14049 (2017).
Nagano, T. et al. Single-cell Hi-C reveals cell-to-cell variability in chromosome structure. Nature 502, 59–64 (2013).
Mezger, A. et al. High-throughput chromatin accessibility profiling at single-cell resolution. Nat. Commun. 9, 3647 (2018).
Clark, S. J. et al. scNMT-seq enables joint profiling of chromatin accessibility DNA methylation and transcription in single cells. Nat. Commun. 9, 781 (2018).
Cao, J. et al. Joint profiling of chromatin accessibility and gene expression in thousands of single cells. Science 361, 1380–1385 (2018).
Adamson, B. et al. A multiplexed single-cell CRISPR screening platform enables systematic dissection of the unfolded protein response. Cell 167, 1867–1882 (2016).
Datlinger, P. et al. Pooled CRISPR screening with single-cell transcriptome readout. Nat. Methods 14, 297–301 (2017).
Dixit, A. et al. Perturb-seq: dissecting molecular circuits with scalable single-cell RNA profiling of pooled genetic screens. Cell 167, 1853–1866.e17 (2016).
Jaitin, D. et al. Dissecting immune circuits by linking CRISPR-pooled screens with single-cell RNA-seq. Cell 167, 1883–1896 (2016).
Chien, M.-P. et al. Photostick: a method for selective isolation of target cells from culture. Chem. Sci. 6, 1701–1705 (2015).
Binan, L. et al. Live single-cell laser tag. Nat. Commun. 7, 11636 (2016).
Kuo, C.-T. et al. Optical painting and fluorescence activated sorting of single adherent cells labelled with photoswitchable Pdots. Nat. Commun. 7, 11468 (2016).
Medaglia, C. et al. Spatial reconstruction of immune niches by combining photoactivatable reporters and scRNA-seq. Science 358, 1622–1626 (2017).
Hasle, N. et al. High-throughput, microscope-based sorting to dissect cellular heterogeneity. Mol. Syst. Biol. 16, e9442 (2020).
Kanfer, G. et al. Image-based pooled whole genome CRISPR screening for Parkin and TFEB subcellular localization. J. Cell Biol. 220, e202006180 (2021).
Yan, X. et al. High-content imaging-based pooled CRISPR screens in mammalian cells. J. Cell Biol. 220, e202008158 (2021).
Piatkevich, K. D. et al. A robotic multidimensional directed evolution approach applied to fluorescent voltage reporters. Nat. Chem. Biol. 14, 352–360 (2018).
Wheeler, E. C. et al. Pooled CRISPR screens with imaging on microraft arrays reveals stress granule-regulatory factors. Nat. Methods https://doi.org/10.1038/s41592-020-0826-8 (2020).
Baltekin, Ö., Boucharin, A., Tano, E., Andersson, D. I. & Elf, J. Antibiotic susceptibility testing in less than 30 min using direct single-cell imaging. Proc. Natl Acad. Sci. USA https://doi.org/10.1073/pnas.1708558114 (2017).
Cookson, S. et al. Monitoring dynamics of single-cell gene expression over multiple cell cycles. Mol. Syst. Biol. 1, 2005.0024 (2005).
Hammar, P. et al. Direct measurement of transcription factor dissociation excludes a simple operator occupancy model for gene regulation. Nat. Genet. 46, 405–408 (2014).
Wang, P. et al. Robust growth of Escherichia coli. Curr. Biol. 20, 1099–1103 (2010).
Luro, S., Potvin-Trottier, L., Okumus, B. & Paulsson, J. Isolating live cells after high-throughput, long-term, time-lapse microscopy. Nat. Methods 17, 93–100 (2020).
Elowitz, M. B. & Leibler, S. A synthetic oscillatory network of transcriptional regulators. Nature 403, 335–338 (2000).
Ke, R. et al. In situ sequencing for RNA analysis in preserved tissue and cells. Nat. Methods 10, 857–860 (2013).
Lee, J. H. et al. Highly multiplexed subcellular RNA sequencing in situ. Science 343, 1360–1363 (2014).
Chen, K. H. et al. Spatially resolved, highly multiplexed RNA profiling in single cells. Science 348, aaa6090 (2015).
Lubeck, E., Coskun, A. F., Zhiyentayev, T., Ahmad, M. & Cai, L. Single-cell in situ RNA profiling by sequential hybridization. Nat. Methods 11, 360–361 (2014).
Elf, J., Öhman, O. & Church, G. Phenotypic characterization and in situ genotyping of a library of genetically different cells. US patent 10,570,437 (2014).
Lawson, M. J. et al. In situ genotyping of a pooled strain library after characterizing complex phenotypes. Mol. Syst. Biol. 13, 947 (2017).
Qi, L. S. et al. Repurposing CRISPR as an RNA-guided platform for sequence-specific control of gene expression. Cell 152, 1173–1183 (2013).
Camsund, D. et al. Time-resolved imaging-based CRISPRi screening. Nat. Methods 17, 86–92 (2020).
Feldman, D. et al. Optical pooled screens in human cells. Cell 179, 787–799 (2019).
Wang, C., Lu, T., Emanuel, G., Babcock, H. P. & Zhuang, X. Imaging-based pooled CRISPR screening reveals regulators of lncRNA localization. Proc. Natl Acad. Sci. USA 116, 10842–10851 (2019).
Askary, A. et al. In situ readout of DNA barcodes and single base edits facilitated by in vitro transcription. Nat. Biotechnol. 38, 66–75 (2020).
Potvin-Trottier, L., Luro, S. & Paulsson, J. Microfluidics and single-cell microscopy to study stochastic processes in bacteria. Curr. Opin. Microbiol. 43, 186–192 (2018).
Eng, C.-H. et al. Transcriptome-scale super-resolved imaging in tissues by RNA seqFISH. Nature 568, 235–239 (2019).
Shah, S. et al. Dynamics and spatial genomics of the nascent transcriptome by intron seqFISH. Cell https://doi.org/10.1016/j.cell.2018.05.035 (2018).
Xia, C., Fan, J., Emanuel, G., Hao, J. & Zhuang, X. Spatial transcriptome profiling by MERFISH reveals subcellular RNA compartmentalization and cell cycle-dependent gene expression. Proc. Natl Acad. Sci. USA 116, 19490–19499 (2019).
Nitta, N. et al. Intelligent image-activated cell sorting. Cell 175, 266–276 (2018).
Ota, S. et al. Ghost cytometry. Science 360, 1246–1251 (2018).
Okumus, B. et al. Single-cell microscopy of suspension cultures using a microfluidics-assisted cell screening platform. Nat. Protoc. 13, 170–194 (2018).
Anzalone, A. V. et al. Search-and-replace genome editing without double-strand breaks or donor DNA. Nature 576, 149–157 (2019).
Sharon, E. et al. Functional genetic variants revealed by massively parallel precise genome editing. Cell https://doi.org/10.1016/j.cell.2018.08.057 (2018).
Gustafsdottir, S. M., Gibson, C. C. & Carpenter, A. E. Cell Painting, a high-content image-based assay for morphological profiling using multiplexed fluorescent dyes. Nat. Protoc. https://doi.org/10.1038/nprot.2016.105 (2016).
Kulesa, A. et al. Combinatorial drug discovery in nanoliter droplets. Proc. Natl Acad. Sci. USA 115, 6685–6690 (2018).
Rivron, N. C. et al. Blastocyst-like structures generated solely from stem cells. Nature 557, 106–111 (2018).
Kehe, J. et al. Massively parallel screening of synthetic microbial communities. Proc. Natl Acad. Sci. USA 116, 12804–12809 (2019).
Douglass, K. M. et al. Super-resolution imaging of multiple cells by optimised flat-field epi-illumination. Nat. Photonics 10, 705–708 (2016).
Isozaki, A. et al. AI on a chip. Lab Chip 20, 3074–3090 (2020).
Moen, E. et al. Deep learning for cellular image analysis. Nat. Methods 16, 1233–1246 (2019).
Styles, E. B. et al. Exploring quantitative yeast phenomics with single-cell analysis of DNA damage foci. Cell Syst. https://doi.org/10.1016/j.cels.2016.08.008 (2016).
Segebarth, D. et al. On the objectivity, reliability, and validity of deep learning enabled bioimage analyses. eLife https://doi.org/10.7554/eLife.59780 (2020).
Perkel, J. M. Starfish Enterprise: finding RNA patterns in single cells. Nature 572, 549–551 (2019).
McQuin, C. et al. CellProfiler 3.0: next-generation image processing for biology. PLoS Biol. 16, e2005970 (2018).
Acknowledgements
We thank D. Jones, P. Blainey, B. Huber, M. Polonsky and L. Ombelets for critical and constructive reading of the manuscript. The work was funded by the Knut and Alice Wallenberg Foundation (KAW).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
J.E. has a patent on in situ genotyping after phenotyping (US patent 10,570,437).
Additional information
Peer review information Rita Strack was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Lawson, M., Elf, J. Imaging-based screens of pool-synthesized cell libraries. Nat Methods 18, 358–365 (2021). https://doi.org/10.1038/s41592-020-01053-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-020-01053-8