Abstract
Synchronized ferroptosis contributes to nephron loss in acute kidney injury (AKI). However, the propagation signals and the underlying mechanisms of the synchronized ferroptosis for renal tubular injury remain unresolved. Here we report that platelet-activating factor (PAF) and PAF-like phospholipids (PAF-LPLs) mediated synchronized ferroptosis and contributed to AKI. The emergence of PAF and PAF-LPLs in ferroptosis caused the instability of biomembranes and signaled the cell death of neighboring cells. This cascade could be suppressed by PAF-acetylhydrolase (II) (PAFAH2) or by addition of antibodies against PAF. Genetic knockout or pharmacological inhibition of PAFAH2 increased PAF production, augmented synchronized ferroptosis and exacerbated ischemia/reperfusion (I/R)-induced AKI. Notably, intravenous administration of wild-type PAFAH2 protein, but not its enzymatically inactive mutants, prevented synchronized tubular cell death, nephron loss and AKI. Our findings offer an insight into the mechanisms of synchronized ferroptosis and suggest a possibility for the preventive intervention of AKI.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All data supporting the findings of this study are available from the corresponding authors upon reasonable request. In molecular dynamics simulations, the initial PAF coordinates were acquired from the PubChem database (ID: 108156), and the PAF force field parameters were borrowed from CGenFF. Source data are provided with this paper.
Change history
05 February 2024
A Correction to this paper has been published: https://doi.org/10.1038/s41589-024-01566-9
References
Dixon, S. J. et al. Ferroptosis: an iron-dependent form of nonapoptotic cell death. Cell 149, 1060–1072 (2012).
Stockwell, B. R. et al. Ferroptosis: a regulated cell death nexus linking metabolism, redox biology, and disease. Cell 171, 273–285 (2017).
Stockwell, B. R. Ferroptosis turns 10: emerging mechanisms, physiological functions, and therapeutic applications. Cell 185, 2401–2421 (2022).
Yang, W. S. & Stockwell, B. R. Synthetic lethal screening identifies compounds activating iron-dependent, nonapoptotic cell death in oncogenic-RAS-harboring cancer cells. Chem. Biol. 15, 234–245 (2008).
Yang, W. S. et al. Regulation of ferroptotic cancer cell death by GPX4. Cell 156, 317–331 (2014).
Jiang, X., Stockwell, B. R. & Conrad, M. Ferroptosis: mechanisms, biology and role in disease. Nat. Rev. Mol. Cell Biol. 22, 266–282 (2021).
Jiang, L. et al. Ferroptosis as a p53-mediated activity during tumour suppression. Nature 520, 57–62 (2015).
Zou, Y. et al. A GPX4-dependent cancer cell state underlies the clear-cell morphology and confers sensitivity to ferroptosis. Nat. Commun. 10, 1617 (2019).
Singhal, R. et al. HIF-2α activation potentiates oxidative cell death in colorectal cancers by increasing cellular iron. J. Clin. Invest. 131, e143691 (2021).
Sun, X. et al. Activation of the p62-Keap1-NRF2 pathway protects against ferroptosis in hepatocellular carcinoma cells. Hepatology 63, 173–184 (2016).
Linkermann, A. et al. Synchronized renal tubular cell death involves ferroptosis. Proc. Natl Acad. Sci. USA 111, 16836–16841 (2014).
Linkermann, A., Stockwell, B. R., Krautwald, S. & Anders, H. J. Regulated cell death and inflammation: an auto-amplification loop causes organ failure. Nat. Rev. Immunol. 14, 759–767 (2014).
Riegman, M. et al. Ferroptosis occurs through an osmotic mechanism and propagates independently of cell rupture. Nat. Cell Biol. 22, 1042–1048 (2020).
Riegman, M., Bradbury, M. S. & Overholtzer, M. Population dynamics in cell death: mechanisms of propagation. Trends Cancer 5, 558–568 (2019).
Belavgeni, A., Meyer, C., Stumpf, J., Hugo, C. & Linkermann, A. Ferroptosis and necroptosis in the kidney. Cell Chem. Biol. 27, 448–462 (2020).
Tonnus, W. et al. Dysfunction of the key ferroptosis-surveilling systems hypersensitizes mice to tubular necrosis during acute kidney injury. Nat. Commun. 12, 4402 (2021).
Kagan, V. E. et al. Oxidized arachidonic and adrenic PEs navigate cells to ferroptosis. Nat. Chem. Biol. 13, 81–90 (2017).
Doll, S. et al. ACSL4 dictates ferroptosis sensitivity by shaping cellular lipid composition. Nat. Chem. Biol. 13, 91–98 (2017).
Anthonymuthu, T. S. et al. Empowerment of 15-lipoxygenase catalytic competence in selective oxidation of membrane ETE-PE to ferroptotic death signals, HpETE-PE. J. Am. Chem. Soc. 140, 17835–17839 (2018).
Shah, R., Shchepinov, M. S. & Pratt, D. A. Resolving the role of lipoxygenases in the initiation and execution of ferroptosis. ACS Cent. Sci. 4, 387–396 (2018).
Yan, B. et al. Membrane damage during ferroptosis is caused by oxidation of phospholipids catalyzed by the oxidoreductases POR and CYB5R1. Mol. Cell 81, 355–369 (2021).
Beharier, O. et al. PLA2G6 guards placental trophoblasts against ferroptotic injury. Proc. Natl Acad. Sci. USA 117, 27319–27328 (2020).
Sun, W. Y. et al. Phospholipase iPLA2β averts ferroptosis by eliminating a redox lipid death signal. Nat. Chem. Biol. 17, 465–476 (2021).
Schopfer, F. J., Cipollina, C. & Freeman, B. A. Formation and signaling actions of electrophilic lipids. Chem. Rev. 111, 5997–6021 (2011).
McIntyre, T. M. Bioactive oxidatively truncated phospholipids in inflammation and apoptosis: formation, targets, and inactivation. Biochim. Biophys. Acta 1818, 2456–2464 (2012).
Marathe, G. K. et al. Bioactive phospholipid oxidation products. Free Radic. Biol. Med. 28, 1762–1770 (2000).
Greenberg, M. E. et al. The lipid whisker model of the structure of oxidized cell membranes. J. Biol. Chem. 283, 2385–2396 (2008).
Fritz, K. S. & Petersen, D. R. An overview of the chemistry and biology of reactive aldehydes. Free Radic. Biol. Med. 59, 85–91 (2013).
Ayala, A., Munoz, M. F. & Arguelles, S. Lipid peroxidation: production, metabolism, and signaling mechanisms of malondialdehyde and 4-hydroxy-2-nonenal. Oxid. Med. Cell. Longev. 2014, 360438 (2014).
Tyurina, Y. Y. et al. Redox lipidomics technology: looking for a needle in a haystack. Chem. Phys. Lipids 221, 93–107 (2019).
Magtanong, L. et al. Exogenous monounsaturated fatty acids promote a ferroptosis-resistant cell state. Cell Chem. Biol. 26, 420–432 (2019).
Doll, S. et al. FSP1 is a glutathione-independent ferroptosis suppressor. Nature 575, 693–698 (2019).
Gaschler, M. M. et al. FINO2 initiates ferroptosis through GPX4 inactivation and iron oxidation. Nat. Chem. Biol. 14, 507–515 (2018).
Friedmann Angeli, J. P. et al. Inactivation of the ferroptosis regulator Gpx4 triggers acute renal failure in mice. Nat. Cell Biol. 16, 1180–1191 (2014).
Lordan, R., Tsoupras, A., Zabetakis, I. & Demopoulos, C. A. Forty years since the structural elucidation of platelet-activating factor (PAF): historical, current, and future research perspectives. Molecules 24, 4414 (2019).
Travers, J. B., Rohan, J. G. & Sahu, R. P. New insights into the pathologic roles of the platelet-activating factor system. Front. Endocrinol. 12, 624132 (2021).
Chammas, R., de Sousa Andrade, L. N. & Jancar, S. Oncogenic effects of PAFR ligands produced in tumours upon chemotherapy and radiotherapy. Nat. Rev. Cancer 17, 253 (2017).
Ishii, S., Nagase, T. & Shimizu, T. Platelet-activating factor receptor. Prostaglandins Other Lipid Mediat. 68-69, 599–609 (2002).
Cao, C. et al. Structural basis for signal recognition and transduction by platelet-activating-factor receptor. Nat. Struct. Mol. Biol. 25, 488–495 (2018).
Dong, L., Li, Y. & Wu, H. Platelet activating-factor acetylhydrolase II: a member of phospholipase A2 family that hydrolyzes oxidized phospholipids. Chem. Phys. Lipids 239, 105103 (2021).
Klems, M. M. & Bahnson, B. J. Trafficking and oligomeric regulation of platelet-activating factor acetylhydrolase type II. Enzymes 38, 55–69 (2015).
Foulks, J. M. et al. PAF-acetylhydrolase expressed during megakaryocyte differentiation inactivates PAF-like lipids. Blood 113, 6699–6706 (2009).
Adibekian, A. et al. Optimization and characterization of a triazole urea inhibitor for platelet-activating factor acetylhydrolase type 2 (PAFAH2). In Probe Reports from the NIH Molecular Libraries Program (National Center for Biotechnology Information, 2010).
Yang, W. S. et al. Peroxidation of polyunsaturated fatty acids by lipoxygenases drives ferroptosis. Proc. Natl Acad. Sci. USA 113, E4966–E4975 (2016).
Ayee, M. A. A. et al. Molecular-scale biophysical modulation of an endothelial membrane by oxidized phospholipids. Biophys. J. 112, 325–338 (2017).
Wang, B. & Tontonoz, P. Phospholipid remodeling in physiology and disease. Annu. Rev. Physiol. 81, 165–188 (2019).
Adibekian, A., Hsu, K. L., Speers, A. E., Monillas, E. S. & Rosen, H. Optimization and characterization of a triazole urea inhibitor for platelet-activating factor acetylhydrolase type 2 (PAFAH2). In Probe Reports from the NIH Molecular Libraries Program (National Center for Biotechnology Information, 2010).
Ke, Y. et al. Elevated truncated oxidized phospholipids as a factor exacerbating ALI in the aging lungs. FASEB J. 33, 3887–3900 (2019).
Zhu, H. et al. Metabolomic profiling of single enlarged lysosomes. Nat. Methods 18, 788–798 (2021).
Jo, S., Kim, T., Iyer, V. G. & Im, W. CHARMM-GUI: a web-based graphical user interface for CHARMM. J. Comput. Chem. 29, 1859–1865 (2008).
Huang, J. et al. CHARMM36m: an improved force field for folded and intrinsically disordered proteins. Nat. Methods 14, 71–73 (2016).
Vanommeslaeghe, K. et al. CHARMM general force field: a force field for drug-like molecules compatible with the CHARMM all-atom additive biological force fields. J. Comput. Chem. 31, 671–690 (2010).
Abraham, M. J. et al. GROMACS: high performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1–2, 19–25 (2015).
Pluhackova, K. et al. A critical comparison of biomembrane force fields: structure and dynamics of model DMPC, POPC, and POPE bilayers. J. Phys. Chem. B 120, 3888–3903 (2016).
Tahir, M. A., Van Lehn, R. C., Choi, S. H. & Alexander-Katz, A. Solvent-exposed lipid tail protrusions depend on lipid membrane composition and curvature. Biochim. Biophys. Acta 1858, 1207–1215 (2016).
Kumar, S., Rosenberg, J. M., Bouzida, D., Swendsen, R. H. & Kollman, P. A. The weighted histogram analysis method for free-energy calculations on biomolecules. I. The method. J. Comput. Chem. 13, 1011–1021 (1992).
Acknowledgements
This research was funded by the National Key R&D Program of China (2019YFA0508601) and the National Natural Science Foundation of China (32230046) to Q.C.; the National Natural Science Foundation of China (32170780 and 31970716) to Y.L.; the National Natural Science Foundation of China (81873496) to X.L.; and Zhejiang Provincial Natural Science Foundation of China (LZ24C050003) and the Information Technology Center and State Key Lab of Computer-Aided Design (CAD) & Computer Graphics (CG) of Zhejiang University, the National Key R&D Program of China (2021YFF1200404), the National Natural Science Foundation of China (32371300) and the Fundamental Research Funds for the Central Universities of China (K20220228) to Y.W.
Author information
Authors and Affiliations
Contributions
Q.C. and Q.Z. conceived the project and designed experiments. Q.C., Q.Z., T.S., J.C., Y.W. and A.C.M. wrote the manuscript. Q.Z. performed most of the cell experiments, with assistance from W.L., J.G. and H. Zheng. Q.Z. and T.S. performed mouse experiments, with assistance from J.G., J.L., C. Mu, X. Liu, Y.G., Z.L. and Y.P. Y.W. and J.C. performed the molecular dynamics simulations and analyzed the simulation data. F.Y. performed the TEM and super-high-resolution microscopy. C. Lu, S.M.L. and G.S. performed quantitative lipidomic analysis of PAFs. C. Miao, H.G., H. Zhu and W.X. performed the lipidomic analysis of cell culture medium by using a Q Exactive Plus MS instrument. M.S. and C. Li synthesized the inhibitor ML225. W.T. and Y.Z. helped with the micro-injection experiments. X.B. helped with the liposome experiments. Y.L., X. Liao, Z.Y., A.C.M. and G.C. provided advice on the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Chemical Biology thanks Kristyna Puhackova, Wulf Tonnus and the other, anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Different kinds of PAFs cause different levels of lethality.
(A, B) Cell viability assay in U-2 OS cells treated with gradient concentrations of PAF C16 (A) or PAF C18 (C) for 12 hours. (C, D) Cell viability assay in HT1080 (C) or HUVEC (D) cells subjected to M-PAFs with different lengths of sn-1: Methylcarbamyl PAF C16 and Methylcarbamyl PAF C8. (E, F) Cell viability assay in HT1080 (E) or HUVEC (F) cells subjected to indicated PAFs with different lengths of sn-2. (G, H) Cell viability assay in HT1080 (G) or HUVEC (H) cells subjected to PAFs with different head-groups in the sn-3 position: Pyrrolidino PAF C-16 and Hexanolamino PAF C-16. For A-H, data show mean ± s.d. of three independent experiments.
Extended Data Fig. 2 The cell death induced by PAFs has characteristics of ferroptosis.
(A) Cell viability of HT1080 treated with M-PAF C16 (75 μM) or PAF C18 (100 μM) and different inhibitors. (B-D) Cell viability of U-2 OS(B), HK2(C) and HUVEC(D) cells treated with PAF C16 (100 μM) or PAF C18 (100 μM) and different inhibitors. The concentration of inhibitors: Fer-1 (2 μM), DFO (50 μM), Nec-1 (20 μM), Nec-1S (20 μM), Z-VAD-FMK (50 μM). (E) Super-high resolution microscopy images of HT1080 cells treated with control or PAF C16 (75 μM). Scale bar, 2 μm. (F-H) Flow cytometry analysis of fluorescence of BODIPY-C11 when HT1080 cells subjected to PAF C16, PAF C18 or M-PAF C16 with or without pretreatment of ferroptosis inhibitors. (I, J) Quantification of the mean fluorescence intensity of Liperfluo (I) and MitoPeDPP (J) in HT1080 cells treated with control, PAF C16 (75 μM) or RSL3 (0.2 μM). (K, L) Flow cytometry analysis of Mito-FerroGreen staining in HT1080 cells treated with PAF C16 (75 μM) for indicated time. (M) ATP levels in HT1080 cells treated with PAF C16 (75 μM). (N-P) Flow cytometry analysis of TMRE in HT1080 cells treated with PAF C16, M-PAF C16 or 2-O-methyl PAF C-16. (Q) The relative mRNA levels were measured by qRT-PCR in HT1080 cells treated with PAF C16 (75 μM). Data show mean ± s.d. of three independent experiments. For A-D, I-J, L-M, data show mean ± s.d. of three independent experiments. Unpaired two-sided Student’s t-test.
Extended Data Fig. 3 FINs and iron salts sensitize cells to PAF induced ferroptosis.
(A-D) Cell viability in HK2 and U2OS cells treated with PAF C16 (75 μM) only, FINs (5 μM RSL3, 20 μM Erastin, 5 μM FINO2, 20 μM iFSP1) only or co-treated with PAF C16 and FINs for 12 h. (E, F) The relative NADP(H) levels in HT1080 cells(E) and HK2 cells(F) treated with Erastin2 (positive control) and PAF C16 (75 μM) for the indicated time. (G, H) The relative glutathione levels of HT1080 cells (G) and HK2 cells (H) treated with Erastin2 (positive control) and PAF C16 (75 μM) for the indicated time. (I) Immunoblots show the knockdown efficiency of ACSL4 siRNAs. Actin was used as a loading control. (J) Dose response of PAF C16 on the viability of HT1080 cells transfected with negative control or ACSL4 siRNAs. Data show mean ±s.d. of three independent experiments ((K) Cell viability in HT1080 cells incubated with PAF C16 (75 μM) for 12 h with or without pretreatment of FAC (1 μg/mL) or FAS (1 μg/mL) for 2 h. For A-H and K, data show mean ±s.d. of three independent experiments. Unpaired two-sided Student’s t-test.
Extended Data Fig. 4 The disturbance in membrane structure induced by PAF.
(A, B) The permeability assay of POPC liposomes (2% NBD-DPPE) containing different percentages of PAF C16 with DMSO or Fer-1 added in this system. Data show mean ± s.d., n = 3 independent experiments. (C, D) The snapshots of locally lipid bilayer of 50% PAF and 50% LPC16 obtained from multi-microsecond all-atom MD simulations. (D) The atomistic structures of POPC, PAF C16 and LPC 16, which are colored in green and silver and orange, respectively. (E) The comparison of water density along the normal direction of three kinds of lipid bilayers indicated. In all water density profiles, the membrane centre is located at 0 nm of the z-axis. (F) The protrusions per lipid for three bilayers compositions. (G) The interaction between the lipid tails and the water was quantified by the number of contacts per frame of each atom with water. (H) The interaction between the lipid tails and the water was quantified by the number of contacts per frame of each lipid with water. For F-H, data are averaged from two independent MD simulation.
Extended Data Fig. 5 Endocytosis is involved in propagation of PAFs in ferroptosis.
(A) PAF ELISA analysis of culture mediums collected from HT1080 cells treated with RSL3 (0.2 μM). (B) The intensity of major PAFs in supernatants from HT1080 cells upon RSL3 treatment. For A, B data show mean ± s.d., n = 4 biologically independent samples. Unpaired two-sided Student’s t-test. (C) The knockdown efficiency of PAFR siRNAs in HT1080 cells. (D, E) Cell viability of HT1080 cells subjected to DMSO or RSL3 (12 h for D, 6 h for E) with or without pretreatment of WEB2086 (10 μM, 12 h, D), Ginkgolide B (100 μM, 12 h, D) or Dynasore (50 μM, 6 h, E). (F) The knockdown efficiency of AP2M1 siRNAs in HT1080 cells. (G) Confocal images of isolated kidney tubules subjected to DMSO or 0.5 μM RSL3 with injection of IgG or PAF antibody accompanied with Dextran Cascade Blue (white arrows show the injection position). Scale bar, 50 μm. (H) The number of dead cells in per 100 μm kidney tubules treated as in G was quantified. (I) Confocal images of U-2 OS cells incubated with POPC or PAF liposome (2% NBD-DPPE, Green). The lysosome is indicated by Lyso-tracker-Red. (J) Cell viability of HT1080 cells subjected to POPC or PAF liposomes for 12 h with pretreatment of different inhibitors. For C-F, H and J, data show mean ± s.d., n = 3 independent experiments. Unpaired two-sided Student’s t-test.
Extended Data Fig. 6 Inactivation of PAFAH2 sensitizes cells to ferroptosis.
(A) Immunoblots show the level of indicated proteins treated with RSL3. The samples derived from the same experiment and the blots were processed in parallel. (B)The relative intensity (PAFAH2/β-actin) was quantified. (C-E) Cell viability of cells treated with RSL3 or Erastin2 with or without ML225 pretreatment. (F. G) Flow cytometry analysis of fluorescence of BODIPY-C11 in HT1080 cells and HUVEC cells subjected to indicated treatments. (G) Statistics of fluorescence of BODIPY-C11 in cells subjected to indicated treatments. (H) The knockdown efficiency of PAFAH2 siRNAs in HT1080 cells. (I) Cell viability of HT1080 cells transfected with control or PAFAH2 siRNAs with RSL3 treatment. (J, L) Immunoblots show the loss of PAFAH2. The samples derived from the same experiment and the blots were processed in parallel. (K) Cell viability of cas9 control or PAFAH2KO HT1080 cells treated with Staurosporine. (M) Dose response of RSL3 on the viability of cas9 control or PAFAH2KO U-2 OS and HUVEC cells. (N) Flow cytometry analysis of fluorescence of BODIPY-C11 in cas9 control or PAFAH2KO HT1080 cells treated with DMSO or RSL3 (0.2 μM). For B-E,H-I,K and M, data show mean ± s.d., n = 3 independent experiments. For B, G and H, unpaired two-sided Student’s t-test.
Extended Data Fig. 7 PAFAH2 is involved in ischemia reperfusion induced AKI and ferroptosis.
(A-I) C57BL/6 J mice were subjected to sham operation or bilateral kidney ischemia (30 minutes) and then reperfusion. (A, B) Relative mRNA levels of Ptgs2 and Chac1 in renal cortex. Data show mean ± S.D., n = 3 (C) Immunoblots of indicated proteins in renal cortex, Actin was used as a loading control. (D-F) C57BL/6 J mice were pre-treated with vehicle or Ferrostatin-1 (5 mg/kg) and then subjected to bilateral kidney I/R. (D, E) The Levels of BUN and serum creatinine. Data show mean ± s.d., n = 4 in each group. Unpaired two-sided Student’s t-test (F) The damage of renal cortex shown by H&E staining. The immune-histochemical staining of 4-HNE, TfR1 and COX2 in renal cortex slices were shown in sequence. Scale bar, 50 μm. (G) Immunoblots of PAFAH2 protein in renal cortex. Actin was used as a loading control. (H) The immunohistochemical staining of PAFAH2 in renal cortex slices. Scale bar, 50 μm. (I) The percentage of PAFAH2 stained tubules were quantified. Data show mean ± s.d., n = 4. Unpaired two-sided Student’s t-test.
Extended Data Fig. 8 Pharmacological inhibition of PAFAH2 accelerates I/R induced AKI and ferroptosis.
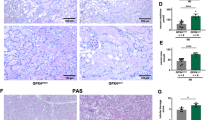
(A-H) C57BL/6 J mice were pretreated with vehicle, ML225 (10 mg/kg), ML225 and Ferrostatin-1 (5 mg/kg) and then subjected to bilateral kidney ischemia (25 min) and reperfusion for 24 h. (A) The damage of renal cortex is shown by PAS staining and H&E staining. The immuno-histochemical staining of 4-HNE, TfR1, COX2 and PAFAH2 of renal cortex slices. The representative TEM images show the mitochondria morphology in renal cortex. Scale bars, 50 µm or 0.5 µm (only in TEM images). (B, C) The Levels of BUN and serum creatinine. Data show mean ± s.d., n = 4 in vehicle+Sham group, n = 5 in other groups. (D) The level of tubular injury. (E-H) Quantification of the percentage of 4-HNE, TfR1, COX2) and PAFAH2 stained tubules in renal cortex. Data show mean ± s.d., n = 4 in D-F, n = 5 in G, n = 7 in H. (I, J) C57BL/6 J mice were subjected to unilateral kidney I/R (25 min) and the kidney were imaged intravitally by two-photon microscope. Scale bar, 50 μm. (J) The percentage of dead tubules. Data show mean ± s.d., n = 4 in vehicle group, n = 3 in other two groups. For B-H and J, unpaired two-sided Student’s t-test.
Extended Data Fig. 9 PAFAH2 deficiency accelerates PAF induced cell death of isolated kidney tubules.
(A, B) Freshly isolated kidney tubules from wild-type or PAFAH2 KO mice were treated with Ethanol or PAF C16 (75 μΜ) and immediately subjected to living cell imaging. The living or dead cells were indicated by Calcein-AM (2 μg/mL) or PI (1 μg/mL) respectively. The number of dead cells in per 100 μm kidney tubules were quantified. Data show mean ± s.d., n = 3 in each group. Unpaired two-sided Student’s t-test.
Extended Data Fig. 10 The recombinant PAFAH2 protein eliminates PAF/PAF LPLs released from RSL3 treated cells.
(A) Images of SDS-PAGE and Coomassie blue staining of purified PAFAH2-WT, PAFAH2-S236A and PAFAH2-3A, which were repeated three times independently with similar results. (B) Lipidomic analysis of the culture medium collected from RSL3-treated cells with or without extracellular PAFAH2 addition (n = 3). Each lipid species was normalized to the corresponding mean value. PAF C16 represents the PAF C16 H + , and PAF C16.1 represents the PAF C16 Na + .
Supplementary information
Supplementary Information
Supplementary Figs. 1–7, Tables 1–4 and Notes 1 and 2.
Supplementary Video 1
The cell death propagation in renal tubules upon the treatment of different cell death inhibitors. Isolated kidney tubules were pre-treated with DMSO or different cell death inhibitors for 1 h and then administered with ethanol (with 1% BSA) or PAF C16 (100 μΜ, with 1% BSA) for 3 h of living cell imaging. The living or dead cells were indicated by the staining of calcein-AM (2 μg ml−1) or PI (1 μg ml−1), respectively.
Supplementary Video 2
MD simulations of biomembranes containing PAF C16 and LPC16. The trajectories of MD simulations of lipid bilayers with different compositions of pure POPC, 50%PAF C16:50%POPC and 50%LPC16:50%POPC.
Supplementary Video 3
PAF C16 propagates ferroptosis. Approximately 10 HT1080 cells were micro-injected with PAF C16, and time series of cell death was indicated by PI staining (1 μg ml−1) immediately for 4 h.
Supplementary Video 4
PAF antibody restrained the renal tubular cell death induced by PAF C16. Isolated kidney tubules were injected with IgG or PAF neutralizing antibody accompanied with dextran-Cascade Blue (1 mg ml−1) in the one end and then injected with ethanol (with 1% BSA) or PAF C16 (with 1% BSA) accompanied with dextran-488 in the other end. The time series of synchronized cell death was recorded by confocal microscope and indicated by PI staining.
Supplementary Video 5
PAF antibody attenuated the renal tubular cell death induced by RSL3. Isolated kidney tubules were injected with IgG or PAF neutralizing antibody accompanied with dextran-Cascade Blue (1 mg ml−1) in the one end and then subjected with treatment of DMSO or RSL3 (0.5 μΜ) for 2 h. The time series of synchronized cell death was recorded by confocal microscope and indicated by PI staining.
Supplementary Video 6
The kidney tubules isolated from PAFAH2-KO mice were more vulnerable to PAF treatment. Freshly isolated kidney tubules from wild-type or PAFAH2 knockout mice were treated with ethanol (with 1% BSA) or PAF C16 (100 μΜ, with 1% BSA) and immediately subjected to living cell imaging. The living or dead cells were indicated by calcein-AM (2 μg ml−1) or PI (1 μg ml−1), respectively.
Supplementary Data 1
Source data for Supplementary Fig. 2.
Supplementary Data 2
Source data for Supplementary Fig. 3.
Supplementary Data 3
Source data for Supplementary Fig. 4.
Supplementary Data 4
Source data for Supplementary Fig. 5.
Supplementary Data 5
Source data for Supplementary Fig. 6.
Supplementary Data 6
Source data for Supplementary Fig. 7.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 4
Unprocessed western blots.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 3
Unprocessed western blots.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 6
Unprocessed western blots.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 7
Unprocessed western blots.
Source Data Extended Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 9
Statistical source data.
Source Data Extended Data Fig. 10
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, Q., Sun, T., Yu, F. et al. PAFAH2 suppresses synchronized ferroptosis to ameliorate acute kidney injury. Nat Chem Biol (2024). https://doi.org/10.1038/s41589-023-01528-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41589-023-01528-7
This article is cited by
-
Chains of death
Nature Chemical Biology (2024)