Abstract
In response to environmental changes, cells flexibly and rapidly alter gene expression through translational controls. In plants, the translation of NIP5;1, a boric acid diffusion facilitator, is downregulated in response to an excess amount of boric acid in the environment through upstream open reading frames (uORFs) that consist of only AUG and stop codons. However, the molecular details of how this minimum uORF controls translation of the downstream main ORF in a boric acid-dependent manner have remained unclear. Here, by combining ribosome profiling, translation complex profile sequencing, structural analysis with cryo-electron microscopy and biochemical assays, we show that the 80S ribosome assembled at AUG-stop migrates into the subsequent RNA segment, followed by downstream translation initiation, and that boric acid impedes this process by the stable confinement of eukaryotic release factor 1 on the 80S ribosome on AUG-stop. Our results provide molecular insight into translation regulation by a minimum and environment-responsive uORF.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The ribosome profiling data (GEO GSE189222) and TCP-seq data (GEO GSE223755) were deposited in the National Center for Biotechnology Information. The cryo-EM maps and structural coordinates of the wheat ribosomes generated for this study were deposited in the Electron Microscopy Data Bank and PDB and are available under the following accession codes: high boric acid 80S complex (EMD-35634, PDB 8IP8), high boric acid 40S complex (EMD-35635, PDB 8IP9), high boric acid 40S complex with eIF2 binding (EMD-35636), high boric acid/CHX complex (EMD-35637, PDB 8IPA) and low boric acid/CHX complex (EMD-35638, PDB 8IPB). Source data are provided with this paper.
Code availability
The code for the deep-sequencing data analysis has been deposited in Zenodo (https://doi.org/10.5281/zenodo.8369134).
References
Shu, X. E., Swanda, R. V. & Qian, S.-B. Nutrient control of mRNA translation. Annu. Rev. Nutr. 40, 51–75 (2020).
Hinnebusch, A. G., Ivanov, I. P. & Sonenberg, N. Translational control by 5′-untranslated regions of eukaryotic mRNAs. Science 352, 1413–1416 (2016).
Johnstone, T. G., Bazzini, A. A. & Giraldez, A. J. Upstream ORFs are prevalent translational repressors in vertebrates. EMBO J. 35, 706–723 (2016).
Zhang, H. et al. Determinants of genome-wide distribution and evolution of uORFs in eukaryotes. Nat. Commun. 12, 1076 (2021).
Hellens, R. P., Brown, C. M., Chisnall, M. A. W., Waterhouse, P. M. & Macknight, R. C. The emerging world of small ORFs. Trends Plant Sci. 21, 317–328 (2016).
Morris, D. R. & Geballe, A. P. Upstream open reading frames as regulators of mRNA translation. Mol. Cell. Biol. 20, 8635–8642 (2000).
Hinnebusch, A. G. Translational regulation of GCN4 and the general amino acid control of yeast. Annu. Rev. Microbiol. 59, 407–450 (2005).
Tanaka, M. et al. Boron-dependent degradation of NIP5;1 mRNA for acclimation to excess boron conditions in Arabidopsis. Plant Cell 23, 3547–3559 (2011).
Tanaka, M. et al. The minimum open reading frame, AUG-stop, induces boron-dependent ribosome stalling and mRNA degradation. Plant Cell 28, 2830–2849 (2016).
Sotta, N. et al. Global analysis of boron-induced ribosome stalling reveals its effects on translation termination and unique regulation by AUG-stops in Arabidopsis shoots. Plant J. 106, 1455–1467 (2021).
Ingolia, N. T., Ghaemmaghami, S., Newman, J. R. & Weissman, J. S. Genome-wide analysis in vivo of translation with nucleotide resolution using ribosome profiling. Science 324, 218–223 (2009).
Iwasaki, S. & Ingolia, N. T. The growing toolbox for protein synthesis studies. Trends Biochem. Sci. 42, 612–624 (2017).
Fujita, T., Kurihara, Y. & Iwasaki, S. The plant translatome surveyed by ribosome profiling. Plant Cell Physiol. 60, 1917–1926 (2019).
Tsednee, M., Tanaka, M., Kasai, K. & Fujiwara, T. Boron-dependent regulation of translation through AUGUAA sequence in yeast. Yeast 37, 638–646 (2020).
Loomis, W. D. & Durst, R. W. Chemistry and biology of boron. Biofactors 3, 229–239 (1992).
Ahmed, I., Yokota, A. & Fujiwara, T. A novel highly boron tolerant bacterium, Bacillus boroniphilus sp. nov., isolated from soil, that requires boron for its growth. Extremophiles 11, 217–224 (2007).
O’Neill, M. A., Eberhard, S., Albersheim, P. & Darvill, A. G. Requirement of borate cross-linking of cell wall rhamnogalacturonan II for Arabidopsis growth. Science 294, 846–849 (2001).
Quax, T. E. F., Claassens, N. J., Söll, D. & van der Oost, J. Codon bias as a means to fine-tune gene expression. Mol. Cell 59, 149–161 (2015).
Richter, J. D. & Coller, J. Pausing on polyribosomes: make way for elongation in translational control. Cell 163, 292–300 (2015).
Buskirk, A. R. & Green, R. Ribosome pausing, arrest and rescue in bacteria and eukaryotes. Philos. Trans. R. Soc. Lond. B Biol. Sci. 372, 20160183 (2017).
Ingolia, N. T., Hussmann, J. A. & Weissman, J. S. Ribosome profiling: global views of translation. Cold Spring Harb. Perspect. Biol. 11, a032698 (2019).
Collart, M. A. & Weiss, B. Ribosome pausing, a dangerous necessity for co-translational events. Nucleic Acids Res. 48, 1043–1055 (2020).
Dana, A. & Tuller, T. Determinants of translation elongation speed and ribosomal profiling biases in mouse embryonic stem cells. PLoS Comput. Biol. 8, e1002755 (2012).
Pop, C. et al. Causal signals between codon bias, mRNA structure, and the efficiency of translation and elongation. Mol. Syst. Biol. 10, 770 (2014).
Ibrahim, F., Maragkakis, M., Alexiou, P. & Mourelatos, Z. Ribothrypsis, a novel process of canonical mRNA decay, mediates ribosome-phased mRNA endonucleolysis. Nat. Struct. Mol. Biol. 25, 302–310 (2018).
Sharma, A. K. et al. A chemical kinetic basis for measuring translation initiation and elongation rates from ribosome profiling data. PLoS Comput. Biol. 15, e1007070 (2019).
Preis, A. et al. Cryoelectron microscopic structures of eukaryotic translation termination complexes containing eRF1–eRF3 or eRF1–ABCE1. Cell Rep. 8, 59–65 (2014).
Brown, A., Shao, S., Murray, J., Hegde, R. S. & Ramakrishnan, V. Structural basis for stop codon recognition in eukaryotes. Nature 524, 493–496 (2015).
Matheisl, S., Berninghausen, O., Becker, T. & Beckmann, R. Structure of a human translation termination complex. Nucleic Acids Res. 43, 8615–8626 (2015).
Shao, S. et al. Decoding mammalian ribosome–mRNA states by translational GTPase complexes. Cell 167, 1229–1240 (2016).
Gunišová, S., Hronová, V., Mohammad, M. P., Hinnebusch, A. G. & Valášek, L. S. Please do not recycle! Translation reinitiation in microbes and higher eukaryotes. FEMS Microbiol. Rev. 42, 165–192 (2018).
Chen, J. et al. Coupling of mRNA structure rearrangement to ribosome movement during bypassing of non-coding regions. Cell 163, 1267–1280 (2015).
Guydosh, N. R. & Green, R. Dom34 rescues ribosomes in 3′ untranslated regions. Cell 156, 950–962 (2014).
Mills, E. W., Wangen, J., Green, R. & Ingolia, N. T. Dynamic regulation of a ribosome rescue pathway in erythroid cells and platelets. Cell Rep. 17, 1–10 (2016).
Skabkin, M. A., Skabkina, O. V., Hellen, C. U. & Pestova, T. V. Reinitiation and other unconventional posttermination events during eukaryotic translation. Mol. Cell 51, 249–264 (2013).
Saito, K., Green, R. & Buskirk, A. R. Ribosome recycling is not critical for translational coupling in Escherichia coli. eLife 9, e59974 (2020).
Young, D. J., Guydosh, N. R., Zhang, F., Hinnebusch, A. G. & Green, R. Rli1/ABCE1 recycles terminating ribosomes and controls translation reinitiation in 3′ UTRs in vivo. Cell 162, 872–884 (2015).
Archer, S. K., Shirokikh, N. E., Beilharz, T. H. & Preiss, T. Dynamics of ribosome scanning and recycling revealed by translation complex profiling. Nature 535, 570–574 (2016).
Shirokikh, N. E., Archer, S. K., Beilharz, T. H., Powell, D. & Preiss, T. Translation complex profile sequencing to study the in vivo dynamics of mRNA–ribosome interactions during translation initiation, elongation and termination. Nat. Protoc. 12, 697–731 (2017).
Bohlen, J., Fenzl, K., Kramer, G., Bukau, B. & Teleman, A. A. Selective 40S footprinting reveals cap-tethered ribosome scanning in human cells. Mol. Cell 79, 561–574 (2020).
Wagner, S. et al. Selective translation complex profiling reveals staged initiation and co-translational assembly of initiation factor complexes. Mol. Cell 79, 546–560 (2020).
Ichihara, K. et al. Combinatorial analysis of translation dynamics reveals eIF2 dependence of translation initiation at near-cognate codons. Nucleic Acids Res. 49, 7298–7317 (2021).
Wagner, S. et al. Selective footprinting of 40S and 80S ribosome subpopulations (Sel-TCP-seq) to study translation and its control. Nat. Protoc. 17, 2139–2187 (2022).
Dingwall, C., Lomonossoff, G. P. & Laskey, R. A. High sequence specificity of micrococcal nuclease. Nucleic Acids Res. 9, 2659–2674 (1981).
Hwang, J.-Y. & Buskirk, A. R. A ribosome profiling study of mRNA cleavage by the endonuclease RelE. Nucleic Acids Res. 45, 327–336 (2016).
Waterhouse, A. et al. SWISS-MODEL: homology modelling of protein structures and complexes. Nucleic Acids Res. 46, W296–W303 (2018).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Armache, J.-P. et al. Cryo-EM structure and rRNA model of a translating eukaryotic 80S ribosome at 5.5-Å resolution. Proc. Natl Acad. Sci. USA 107, 19748–19753 (2010).
Armache, J.-P. et al. Localization of eukaryote-specific ribosomal proteins in a 5.5-Å cryo-EM map of the 80S eukaryotic ribosome. Proc. Natl Acad. Sci. USA 107, 19754–19759 (2010).
Boccaletto, P. et al. MODOMICS: a database of RNA modification pathways. 2017 update. Nucleic Acids Res. 46, D303–D307 (2018).
Polacek, N. et al. The critical role of the universally conserved A2602 of 23S ribosomal RNA in the release of the nascent peptide during translation termination. Mol. Cell 11, 103–112 (2003).
Youngman, E. M., Brunelle, J. L., Kochaniak, A. B. & Green, R. The active site of the ribosome is composed of two layers of conserved nucleotides with distinct roles in peptide bond formation and peptide release. Cell 117, 589–599 (2004).
Garreau de Loubresse, N. et al. Structural basis for the inhibition of the eukaryotic ribosome. Nature 513, 517–522 (2014).
Lawson, M. R. et al. Mechanisms that ensure speed and fidelity in eukaryotic translation termination. Science 373, 876–882 (2021).
Huang, S. et al. Ataluren binds to multiple protein synthesis apparatus sites and competitively inhibits release factor-dependent termination. Nat. Commun. 13, 2413 (2022).
Meydan, S. & Guydosh, N. R. Disome and trisome profiling reveal genome-wide targets of ribosome quality control. Mol. Cell 79, 588–602 (2020).
Han, P. et al. Genome-wide survey of ribosome collision. Cell Rep. 31, 107610 (2020).
Skabkin, M. A. et al. Activities of ligatin and MCT-1/DENR in eukaryotic translation initiation and ribosomal recycling. Genes Dev. 24, 1787–1801 (2010).
Dmitriev, S. E. et al. GTP-independent tRNA delivery to the ribosomal P-site by a novel eukaryotic translation factor. J. Biol. Chem. 285, 26779–26787 (2010).
Schleich, S. et al. DENR–MCT-1 promotes translation re-initiation downstream of uORFs to control tissue growth. Nature 512, 208–212 (2014).
Lomakin, I. B. et al. Crystal structure of the human ribosome in complex with DENR–MCT-1. Cell Rep. 20, 521–528 (2017).
Schleich, S., Acevedo, J. M., Clemm von Hohenberg, K. & Teleman, A. A. Identification of transcripts with short stuORFs as targets for DENR•MCTS1-dependent translation in human cells. Sci. Rep. 7, 3722 (2017).
Weisser, M. et al. Structural and functional insights into human re-initiation complexes. Mol. Cell 67, 447–456 (2017).
Young, D. J. et al. Tma64/eIF2D, Tma20/MCT-1, and Tma22/DENR recycle post-termination 40S subunits in vivo. Mol. Cell 71, 761–774 (2018).
Bohlen, J. et al. DENR promotes translation reinitiation via ribosome recycling to drive expression of oncogenes including ATF4. Nat. Commun. 11, 4676 (2020).
Simms, C. L., Yan, L. L. & Zaher, H. S. Ribosome collision is critical for quality control during no-go decay. Mol. Cell 68, 361–373 (2017).
D’Orazio, K. N. et al. The endonuclease Cue2 cleaves mRNAs at stalled ribosomes during no go decay. eLife 8, e49117 (2019).
Ikeuchi, K. et al. Collided ribosomes form a unique structural interface to induce Hel2-driven quality control pathways. EMBO J. 38, e100276 (2019).
Glover, M. L. et al. NONU-1 encodes a conserved endonuclease required for mRNA translation surveillance. Cell Rep. 30, 4321–4331 (2020).
Tomomatsu, S. et al. Two modes of Cue2-mediated mRNA cleavage with distinct substrate recognition initiate no-go decay. Nucleic Acids Res. 51, 253–270 (2023).
Miyake, T. et al. Minimal upstream open reading frame of Per2 mediates phase fitness of the circadian clock to day/night physiological body temperature rhythm. Cell Rep. 42, 112157 (2023).
Mito, M., Mishima, Y. & Iwasaki, S. Protocol for disome profiling to survey ribosome collision in humans and zebrafish. STAR Protoc. 1, 100168 (2020).
Matsuura-Suzuki, E. et al. METTL18-mediated histidine methylation of RPL3 modulates translation elongation for proteostasis maintenance. eLife 11, e72780 (2022).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Prongidi-Fix, L. et al. Rapid purification of ribosomal particles assembled on histone H4 mRNA: a new method based on mRNA–DNA chimaeras. Biochem. J. 449, 719–728 (2013).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. eLife 7, e42166 (2018).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Croll, T. I. ISOLDE: a physically realistic environment for model building into low-resolution electron-density maps. Acta Crystallogr. D Biol. Crystallogr. 74, 519–530 (2018).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D Biol. Crystallogr. 66, 12–21 (2010).
Goddard, T. D. et al. UCSF ChimeraX: meeting modern challenges in visualization and analysis. Protein Sci. 27, 14–25 (2018).
Jester, B. tRNA northern analysis. Protocol Exchange https://doi.org/10.1038/protex.2011.223 (2011).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Machida, K. et al. A translation system reconstituted with human factors proves that processing of encephalomyocarditis virus proteins 2A and 2B occurs in the elongation phase of translation without eukaryotic release factors. J. Biol. Chem. 289, 31960–31971 (2014).
Acknowledgements
This work was supported, in part, by the Ministry of Education, Culture, Sports, Science and Technology of Japan (JP19H05637 and JP18H05490 (T.F.), JP18K06278 (M.T.), JP19H02959 and JP20H05784 (S.I.) and JP19H03172 and JP21H05281 (T.I.)), the Japan Science and Technology Agency (JPMJPR20EG; T.Y.), the Japan Agency for Medical Research and Development (AMED; JP23gm1410001; S.I. and T.I.) and RIKEN (Pioneering project ‘Biology of Intracellular Environments’; M.S., S.I. and T.I.). N.S. was a JSPS Overseas Research Fellow. H. Saito was a Junior Research Associate of RIKEN. This work was also supported by the Platform Project for Supporting Drug Discovery and Life Science Research (Basis for Supporting Innovative Drug Discovery and Life Science Research from AMED; JP20am0101082, support number 0245), Support Unit for Bio-Material Analysis, RIKEN CBS Research Resources Division for Sanger sequencing and the supercomputer HOKUSAI Sailing Ship at RIKEN for computations. The cryo-EM with the CRYO ARM 300 II (powered by AMED research grant JP20am0101095) was provided by the Advanced Research Center for Innovations in Next-Generation Medicine, Tohoku University. Some of the cryo-EM experiments were performed at the RIKEN Yokohama cryo-EM facility (Yokohama).
Author information
Authors and Affiliations
Contributions
M.T., T.Y., H. Saito, M.S., S.I., T.I. and T.F. designed the concept. M.T., T.Y. and H. Shigematsu established the methodology. M.T. performed western and northern blotting and prepared the cryo-EM samples. T.Y. collected the cryo-EM data and solved the structures. H. Saito performed ribosome profiling and TCP-seq, and M.N. conducted the purification of eRFs and the tRNA aminoacylation assay. M.T., T.Y., H. Saito, K.T., N.S., H. Shigematsu, T.I., M.S. and T.F. performed formal analyses. M.T., T.Y., N.S., S.I., T.I. and T.F. wrote the original draft. M.T., T.Y., H. Saito, M.N., K.T., N.S., H. Shigematsu, M.S., S.I., T.I. and T.F wrote, reviewed and edited the paper. M.T., T.Y., H. Saito and S.I. visualized the results. M.S., S.I., T.I. and T.F. supervised the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Chemical Biology thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
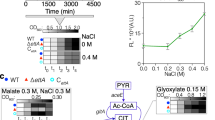
Extended Data Fig. 1 Characterization of ribosome footprints from reporter mRNA translated in vitro.
(a) Relative luminescence for the reporter mRNA bearing the NIP5;1 5′-UTR. In addition to the wild-type NIP5;1 5′-UTR with AUG-UAA, a mutated (AUC-UAA) reporter was tested with the indicated concentrations of boric acid supplementation. Means (points) ± standard deviations (errors) from three replicates are shown. (b and e) Length distribution of ribosome footprints mapped to the main ORF (b) and the inter-ORF region (e). (c and f) Frame position of the 5′ end of footprints (27 nt) mapped to the main ORF (c) or the inter-ORF region (f). Means (bars) from two replicates are shown. (d and g) A and T contents in the main ORF sequence (d) and the inter-ORF sequence (g). (h) Sucrose density gradient centrifugation for crosslinked ribosomal complexes, used for TCP-Seq. (i) The distribution of 5′ ends of footprints in TCP-Seq. For b-c, e, f, and h-i, the data from two biologically independent replicates are shown. RPM, reads per million mapped reads.
Extended Data Fig. 2 Image processing scheme for the cryo-EM reconstruction of the 80S stalled complex on AUG-stop in +B conditions.
(a) Details of the image processing to obtain cryo-EM maps of the 80S stalled on AUG-stop in +B conditions and 40S initiation complexes with or without eIF2 binding. Of the 1,407 K particles initially 2D classified into 80S and 40S subgroups, 367 K particles sorted to 80S structures were auto-refined to obtain the consensus reconstruction. Particles were further 3D classified based on the alignment information from the consensus structure. Afterwards, 246 K particles were sorted to 80S ribosomes in complex with P-tRNA, E-tRNA, mRNA, and eRF1. The focused classification on eRF1 with residual signal subtraction was performed to obtain particles containing eRF1. The final reconstruction with 96 K particles was at a 2.9 Å resolution. In total, 181 K particles corresponding to 40S structures from the 2D classification were auto-refined to obtain the consensus reconstruction. 40S particles were 3D classified based on the alignment information from the consensus structure. Subsequently, 53 K particles were sorted to 40S ribosomes in complex with P-tRNA, mRNA, and eIF2. Focused classification on tRNAi and eIF2 with residual signal subtraction was performed to classify particles into subgroups with or without eIF2 binding. As a result, 41 K particles without eIF2 and 12 K particles with eIF2 were reconstructed at 3.0 Å and 3.7 Å, respectively. (b-c) Resolution (gold-standard FSC) curves of three reconstructions (b) and models vs. cryo-EM maps (c) are shown. (d) Color-coded local resolution distributions of the cryo-EM map of the 80S complex from this dataset.
Extended Data Fig. 3 40S complexes reconstructed from sorted subgroups of the 3D classification.
(a and b) Cryo-EM structures of the wheat 40S ribosome in complex with a tRNAi and an mRNA bearing AUG-stop (a) and that with the additional eIF2 binding (b). The contour level of the density corresponding to eIF2 in b is lowered to show entire structure of eIF2.
Extended Data Fig. 4 Cryo-EM density of tRNA in the P site represents a tRNAi-specific structural feature.
(a) Schematic diagram of tRNAi (left) and an elongator tRNAMet (right) for depicting the differences between the two tRNA species that charge methionine. (b) A close-up view of G31 and C39 base-pairing in the cryo-EM reconstruction. The density fits well to an atomic model of tRNAi, rather than the elongator tRNAMet, which has two pseudouridines in the corresponding nucleotide positions (ψ31 and ψ39).
Extended Data Fig. 5 The GGQ motif on the tip of the M domain of eRF1 in +B conditions is located near the CCA-end of tRNAi.
Isolated cryo-EM densities corresponding to the CCA-end of tRNAi (green) and the GGQ motif of eRF1 (blue) are shown in their surrounding ribosomal environments.
Extended Data Fig. 6 Image processing scheme of the cryo-EM reconstruction of the 80S stalled complex on AUG-stop in +B/ − B conditions with CHX.
(a) Details of image processing to obtain the cryo-EM map of the 80S stalled on AUG-stop in +B conditions with CHX addition. First, 131 K particles were initially 2D classified to obtain 80S particles. The 98 K particles sorted to 80S structures were auto-refined to obtain the consensus reconstruction. Particles were further 3D classified based on the alignment information from the consensus structure. 80S ribosomes in complex with P-tRNA, mRNA, eRF1, and CHX were selected. Focused classification on eRF1 with residual signal subtraction was performed to obtain particles containing eRF1. The final reconstruction with 69 K particles was at a 3.4 Å resolution. (b) Details of image processing to obtain the cryo-EM map of the 80S stalled on AUG-stop in −B conditions with CHX addition. The picked 220 K particles were initially 2D classified to obtain 80S particles, and then 92 K particles sorted to 80S structures were auto-refined to obtain the consensus reconstruction. Particles were further 3D classified based on the alignment information from the consensus structure. 80S ribosomes in complex with P-tRNA, mRNA, and CHX were selected. The focused classification on the fragmented eRF1 density with residual signal subtraction was performed. The obtained 52 K particles containing some density on the A site were reconstructed at a 3.4 Å resolution. (c-d) Resolution (gold-standard FSC) curves of two reconstructions (c) and models vs. cryo-EM maps (d) are shown. (e-f) Color-coded local resolution distributions of the cryo-EM map of the 80S complex from these datasets.
Extended Data Fig. 7 Cycloheximide binding to the E site sterically clashes with A76 of a deacylated tRNA bound to the E site.
(a-b) Cryo-EM densities of cycloheximide (a) and the CCA-end (b) and their corresponding atomic models are shown in the 60S environment. (c) Superimposition of the two models, showing the steric clash of cycloheximide with tRNA.
Extended Data Fig. 8 Purification of recombinant Arabidopsis eRF1-1 and eRF3 proteins.
CBB staining of the indicated recombinant proteins. The expected molecular weights of eRF1-1 and eRF3 are 48.7 kDa and 60.5 kDa, respectively. The lane of purified eRF3 was on the same gel but the unrelated lanes in the middle were omitted without changing the Y-axis position of the bands. The similar staining image was obtained repeatedly (3 replicates).
Extended Data Fig. 9 Characterization of Met-tRNAi hydrolysis on ribosomes stalled on AUG-stop.
(a and b) The same as Fig. 4d but with titrated amounts of eRF1/eRF3 (a) and along the time course (b). For a, 0, 1, 10, and 100 ng of eRF1/eRF3 were used and incubated for 30 min in the presence of boric acid. For b, reactions were incubated for 0, 5, 10, 30 and 60 min with 300 ng eRF1/eRF3 in the presence of boric acid. For a, means (bars) of two biological replicates (points) are shown. For b, means (large points) of two biologically independent replicates (small points) are shown.
Supplementary information
Supplementary Information
Supplementary Fig. 1 and Tables 1 and 2.
Source data
Source Data Fig. 3
Uncropped western blots in Fig. 3d,e.
Source Data Fig. 4
Uncropped northern blots in Fig. 4a,c,d.
Source Data Extended Data Fig. 9
Uncropped northern blot in Extended Data Fig. 9a,b.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Tanaka, M., Yokoyama, T., Saito, H. et al. Boric acid intercepts 80S ribosome migration from AUG-stop by stabilizing eRF1. Nat Chem Biol 20, 605–614 (2024). https://doi.org/10.1038/s41589-023-01513-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-023-01513-0