Abstract
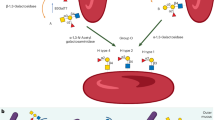
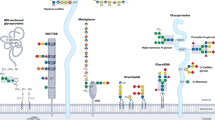
Bifidobacteria are early colonizers of the human gut and play central roles in human health and metabolism. To thrive in this competitive niche, these bacteria evolved the capacity to use complex carbohydrates, including mammalian N-glycans. Herein, we elucidated pivotal biochemical steps involved in high-mannose N-glycan utilization by Bifidobacterium longum. After N-glycan release by an endo-β-N-acetylglucosaminidase, the mannosyl arms are trimmed by the cooperative action of three functionally distinct glycoside hydrolase 38 (GH38) α-mannosidases and a specific GH125 α-1,6-mannosidase. High-resolution cryo-electron microscopy structures revealed that bifidobacterial GH38 α-mannosidases form homotetramers, with the N-terminal jelly roll domain contributing to substrate selectivity. Additionally, an α-glucosidase enables the processing of monoglucosylated N-glycans. Notably, the main degradation product, mannose, is isomerized into fructose before phosphorylation, an unconventional metabolic route connecting it to the bifid shunt pathway. These findings shed light on key molecular mechanisms used by bifidobacteria to use high-mannose N-glycans, a perennial carbon and energy source in the intestinal lumen.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Atomic coordinates and structure factors were deposited to the Protein Data Bank (PDB) with the following accession codes: PDB 7UFR/Electron Microscopy Data Bank (EMDB) 26478 (Bl_Man38A), PDB 7UFS/EMDB 26479 (Bl_Man38B), PDB 7UFT/EMDB 26480 (Bl_Man38C) and PDB 7UFU/EMDB 26481 (Bl_Man38AD387A–MAN). Other data generated or analyzed during this study are included in the published article and its Supplementary Information files. Source data are provided with this paper.
References
Fanning, S. et al. Bifidobacterial surface-exopolysaccharide facilitates commensal–host interaction through immune modulation and pathogen protection. Proc. Natl Acad. Sci. USA 109, 2108–2113 (2012).
Andlid, T. A., D’Aimmo, M. R. & Jastrebova, J. in The Bifidobacteria and Related Organisms (eds Mattarelli, P., Biavati, B., Holzapfel, W. H. & Wood, B. J. B.) 195–212 (Elsevier, 2018).
Moya-Pérez, A., Perez-Villalba, A., Benítez-Páez, A., Campillo, I. & Sanz, Y. Bifidobacterium CECT 7765 modulates early stress-induced immune, neuroendocrine and behavioral alterations in mice. Brain Behav. Immun. 65, 43–56 (2017).
Luck, B. et al. Bifidobacteria shape host neural circuits during postnatal development by promoting synapse formation and microglial function. Sci. Rep. 10, 7737 (2020).
Milani, C. et al. Genomics of the genus Bifidobacterium reveals species-specific adaptation to the glycan-rich gut environment. Appl. Environ. Microbiol. 82, 980–991 (2016).
Jung, D. H. et al. The presence of resistant starch-degrading amylases in Bifidobacterium adolescentis of the human gut. Int. J. Biol. Macromol. 161, 389–397 (2020).
la Rosa, S. L. et al. Wood-derived dietary fibers promote beneficial human gut microbiota. mSphere 4, e00554-18 (2019).
Yamada, C. et al. Molecular insight into evolution of symbiosis between breast-fed infants and a member of the human gut microbiome Bifidobacterium longum. Cell Chem. Biol. 24, 515–524 (2017).
Katoh, T. et al. Enzymatic adaptation of Bifidobacterium bifidum to host glycans, viewed from glycoside hydrolases and carbohydrate-binding modules. Microorganisms 8, 481 (2020).
Higel, F., Seidl, A., Sörgel, F. & Friess, W. N-Glycosylation heterogeneity and the influence on structure, function and pharmacokinetics of monoclonal antibodies and Fc fusion proteins. Eur. J. Pharm. Biopharm. 100, 94–100 (2016).
Schjoldager, K. T., Narimatsu, Y., Joshi, H. J. & Clausen, H. Global view of human protein glycosylation pathways and functions. Nat. Rev. Mol. Cell Biol. 21, 729–749 (2020).
Bjursell, M. K., Martens, E. C. & Gordon, J. I. Functional genomic and metabolic studies of the adaptations of a prominent adult human gut symbiont, Bacteroides thetaiotaomicron, to the suckling period. J. Biol. Chem. 281, 36269–36279 (2006).
Martens, E. C., Koropatkin, N. M., Smith, T. J. & Gordon, J. I. Complex glycan catabolism by the human gut microbiota: the Bacteroidetes Sus-like paradigm. J. Biol. Chem. 284, 24673–24677 (2009).
Hemsworth, G. R., Déjean, G., Davies, G. J. & Brumer, H. Learning from microbial strategies for polysaccharide degradation. Biochem. Soc. Trans. 44, 94–108 (2016).
Foley, M. H., Cockburn, D. W. & Koropatkin, N. M. The Sus operon: a model system for starch uptake by the human gut Bacteroidetes. Cell. Mol. Life Sci. 73, 2603–2617 (2016).
Cuskin, F. et al. Human gut Bacteroidetes can utilize yeast mannan through a selfish mechanism. Nature 517, 165–169 (2015).
Robb, M. et al. Molecular characterization of N-glycan degradation and transport in Streptococcus pneumoniae and its contribution to virulence. PLoS Pathog. 13, e1006090 (2017).
Dupoiron, S. et al. The N-glycan cluster from Xanthomonas campestris pv. campestris: a toolbox for sequential plant N-glycan processing. J. Biol. Chem. 290, 6022–6036 (2015).
Briliūtė, J. et al. Complex N-glycan breakdown by gut Bacteroides involves an extensive enzymatic apparatus encoded by multiple co-regulated genetic loci. Nat. Microbiol. 4, 1571–1581 (2019).
Trastoy, B. et al. Structural basis of mammalian high-mannose N-glycan processing by human gut Bacteroides. Nat. Commun. 11, 889 (2020).
Higgins, M. A. et al. N-Glycan degradation pathways in gut- and soil-dwelling Actinobacteria share common core genes. ACS Chem. Biol. 16, 701–711 (2021).
Reichenbach, T. et al. Structural and biochemical characterization of the Cutibacterium acnes exo-β-1,4-mannosidase that targets the N-glycan core of host glycoproteins. PLoS ONE 13, e0204703 (2018).
Cordeiro, R. L. et al. N-Glycan utilization by Bifidobacterium gut symbionts involves a specialist β-mannosidase. J. Mol. Biol. 431, 732–747 (2019).
Garrido, D. et al. Endo-β-N-acetylglucosaminidases from infant gut-associated bifidobacteria release complex N-glycans from human milk glycoproteins. Mol. Cell Proteomics 11, 775–785 (2012).
Schell, M. A. et al. The genome sequence of Bifidobacterium longum reflects its adaptation to the human gastrointestinal tract. Proc. Natl Acad. Sci. USA 99, 14422–14427 (2002).
Trombetta, E. S., Simons, J. F. & Helenius, A. Endoplasmic reticulum glucosidase II is composed of a catalytic subunit, conserved from yeast to mammals, and a tightly bound noncatalytic HDEL-containing subunit. J. Biol. Chem. 271, 27509–27516 (1996).
Parche, S. et al. Sugar transport systems of Bifidobacterium longum NCC2705. J. Mol. Microbiol. Biotechnol. 12, 9–19 (2006).
Caescu, C. I., Vidal, O., Krzewinski, F., Artenie, V. & Bouquelet, S. Bifidobacterium longum requires a fructokinase (Frk; ATP:d-fructose 6-phosphotransferase, EC 2.7.1.4) for fructose catabolism. J. Bacteriol. 186, 6515–6525 (2004).
Fushinobu, S. Unique sugar metabolic pathways of bifidobacteria. Biosci. Biotechnol. Biochem. 74, 2374–2384 (2010).
Gregg, K. J. et al. Analysis of a new family of widely distributed metal-independent α-mannosidases provides unique insight into the processing of N-linked glycans. J. Biol. Chem. 286, 15586–15596 (2011).
Shah, N., Kuntz, D. A. & Rose, D. R. Golgi α-mannosidase II cleaves two sugars sequentially in the same catalytic site. Proc. Natl Acad. Sci. USA 105, 9570–9575 (2008).
Nielsen, J. W. et al. Metal-ion dependent catalytic properties of sulfolobus solfataricus class II α-mannosidase. Biochemistry 51, 8039–8046 (2012).
Sonnenburg, J. L., Chen, C. T. L. & Gordon, J. I. Genomic and metabolic studies of the impact of probiotics on a model gut symbiont and host. PLoS Biol. 4, 2213–2226 (2006).
Bertipaglia, C. et al. Higher-order assemblies of oligomeric cargo receptor complexes form the membrane scaffold of the Cvt vesicle. EMBO Rep. 17, 1044–1060 (2016).
Zhang, J., Wang, Y. Y., Du, L. L. & Ye, K. Cryo-EM structure of fission yeast tetrameric α-mannosidase Ams1. FEBS Open Bio 10, 2437–2451 (2020).
Suits, M. D. L. et al. Structure and kinetic investigation of Streptococcus pyogenes family GH38 α-mannosidase. PLoS ONE 5, e9006 (2010).
Suzuki, T. et al. Man2C1, an α-mannosidase, is involved in the trimming of free oligosaccharides in the cytosol. Biochem. J. 400, 33–41 (2006).
Heikinheimo, P. et al. The structure of bovine lysosomal α-mannosidase suggests a novel mechanism for low-pH activation. J. Mol. Biol. 327, 631–644 (2003).
Howard, E. et al. Structural basis of outstanding multivalent effects in jack bean α-mannosidase inhibition. Angew. Chem. Int. Ed. Engl. 57, 8002–8006 (2018).
Yamagishi, M., Ishimizu, T., Natsuka, S. & Hase, S. Co(II)-regulated substrate specificity of cytosolic α-mannosidase. J. Biochem. 132, 253–256 (2002).
Chaudet, M. M. & Rose, D. R. Suggested alternative starch utilization system from the human gut bacterium Bacteroides thetaiotaomicron. Biochem. Cell Biol. 94, 241–246 (2016).
Tan, K. et al. Novel α-glucosidase from human gut microbiome: substrate specificities and their switch. FASEB J. 24, 3939–3949 (2010).
Ikegaya, M. et al. Structural basis of the strict specificity of a bacterial GH31 α-1,3-glucosidase for nigerooligosaccharides. J. Biol. Chem. 298, 101827 (2022).
Park, D. et al. Enterocyte glycosylation is responsive to changes in extracellular conditions: implications for membrane functions. Glycobiology 27, 847–860 (2017).
Bilyy, R. O. et al. Macrophages discriminate glycosylation patterns of apoptotic cell-derived microparticles. J. Biol. Chem. 287, 496–503 (2012).
Egan, M. & Van Sinderen, D. in The Bifidobacteria and Related Organisms (eds Mattarelli, P., Biavati, B., Holzapfel, W. H. & Wood, B. J. B.) 145–164 (Elsevier, 2018).
Itoh, T., Mikami, B., Hashimoto, W. & Murata, K. Crystal structure of YihS in complex with d-mannose: structural annotation of Escherichia coli and Salmonella enterica yihS-encoded proteins to an aldose–ketose isomerase. J. Mol. Biol. 377, 1443–1459 (2008).
Price, M. N., Dehal, P. S. & Arkin, A. P. FastTree 2—approximately maximum-likelihood trees for large alignments. PLoS ONE 5, e9490 (2010).
Gilchrist, C. L. M. et al. cblaster: a remote search tool for rapid identification and visualization of homologous gene clusters. Bioinform. Adv. 1, vbab016 (2021).
Dische, Z. & Borenfreund, E. A new spectrophotometric method for the detection and determination of keto sugars and trioses. J. Biol. Chem. 192, 583–587 (1951).
van Heel, M. et al. in International Tables for Crystallography, Volume F, 2nd edition, Crystallography of Biological Macromolecules (eds Arnold, E., Himmel, D. M. & Rossmann, M. G.) 624–628 (Wiley, 2012).
Grant, T., Rohou, A. & Grigorieff, N. CisTEM, user-friendly software for single-particle image processing. eLife 7, e35383 (2018).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D Biol. Crystallogr. 66, 12–21 (2010).
Trott, O. & Olson, A. J. Autodock vina: improving the speed and accuracy of docking. J. Comput. Chem. 31, 455–461 (2019).
Roe, D. R. & Cheatham, T. E. PTRAJ and CPPTRAJ: software for processing and analysis of molecular dynamics trajectory data. J. Chem. Theory Comput. 9, 3084–3095 (2013).
Acknowledgements
We thank R. dos Santos Bezerra for support with some bioinformatics analyses. We thank M. L. Sforca and S. A. Rocco for conducting and analyzing the NMR experiments. We thank L. M. Z. Murakami for support in characterization of isomerase activity. We acknowledge the National Nanotechnology Laboratory (LNNano) for the provision of time (proposal TEM-C2-26053 and TEM-C2-26816) on the transmission electron microscopes JEM 1400 Plus (Jeol), Talos F200C (Thermo Fisher Scientific), Talos Arctica G2 (Thermo Fisher Scientific) and Titan Krios G3i (Thermo Fisher Scientific). We acknowledge the Brazilian Biorenewables National Laboratory (LNBR) for the use of the characterization of macromolecules facility (proposals 26914, 27586 and 20220639) and for the use of the metabolomics facility (proposal 20220817). We acknowledge the Brazilian Biosciences National Laboratory (LNBio) for the use of the NMR facility (proposal 20220793). LNNano, LNBR and LNBio are operated by the Brazilian National Center for Research in Energy and Materials for the Brazilian Ministry for Science, Technology and Innovations (MCTIC). We acknowledge the National Laboratory for Scientific Computing (LNCC/MCTI, Brazil) for providing high-performance computing resources of the SDumont supercomputer (http://sdumont.lncc.br), which have contributed to the research results reported within this paper (proposals 212394 and 221167). This research was supported by grants from Fundação de Amparo à Pesquisa do Estado de São Paulo (grant number 2015/26982-0 to M.T.M., 2017/15340-2 to R.V.P. and 2016/00740-2 to R.L.C.) and Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq; grant number 306135/2016-7 and 305013/2020-3 to M.T.M.) and Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES). This work was also supported in part by NIH grant R01AI149297.
Author information
Authors and Affiliations
Contributions
R.L.C., C.R.S., P.O.G. and M.T.M. designed the study and wrote the original draft. R.L.C., C.R.S., M.E.G., E.J.S., P.O.G. and M.T.M. revised and contributed to the final version of the paper. R.L.C. and G.F.P. performed the bioinformatics analyses. M.A.B.M. and F.M.C. performed the molecular dynamic simulations. T.B.L., R.A.S.P. and F.C.G. performed the MS analyses. R.L.C., M.N.D., R.Y.M. and F.S. expressed and purified the enzymes and performed the enzymatic assays. C.L. and L.-X.W. produced the carbohydrates Man9GlcNAc2-Asn and Man5GlcNAc2-Asn. R.L.C., M.N.D., M.E.G., E.J.S., P.O.G. and M.T.M. analyzed the functional data. A.C.B., M.A.d.F., M.v.H. and R.V.P. prepared grids and collected and processed the cryo-EM data. R.L.C. and C.R.S. modeled and refined the atomic models. R.L.C., C.R.S., P.O.G. and M.T.M. performed the structural analyses. All authors analyzed the results and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Chemical Biology thanks David Cannella, Lucy Crouch and Jose Munoz for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Genomic organization of the predicted N-glycan utilization loci (N-GUL) from B. longum NCC2705.
a, Genes that compose N-GUL represented by arrows and their respective gene products. a = biochemically characterized in this work, b = predicted, c = characterized in previous works23,24,28. The symbol * represents transcriptional regulators (bl1336 – encoding a LacI transcriptional regulator and bl1340 – encoding a ROK transcriptional regulator). b, A schematic representation of a high-mannose N-glycan indicating its subunits (geometric forms), glycosidic linkages (gray symbols) and the cleavage sites (scissors) recognized by the GH5 β-mannosidase23 and GH85 ENGase24 enzymes previously characterized. Predictions of uncharacterized enzymes were performed by sequence and hidden-Markov similarities analyses (Supplementary Tables 1, 2). c, Representation of Man9GlcNAc. d, Representation of Man5GlcNAc.
Extended Data Fig. 2 Evolutionary analysis of Bifidobacterium strains.
a, Phylogenetic tree of Bifidobacterium genomes. *1: B. ruminantium, B. pseudocatenulatum, B. catenulatum, and B. moukalabense; *2: B. jacchi, B. scardovii, B. samirii, B. biavatii, B. ramosum, B. hapali, B. aerophilum, and B. leontopitheci; *3: B. merycicum, B. callitrichos, B. platyrrhinorum, B. aesculapii, B. parmae, B. stellenboschens, B. eulemuris, B. lemurum, B. scaligerum, B. callitrichidarum, B. myosotis, B. reuteri, B. cebidarum, B. felsineum, B. imperatoris, and B. saguini. b, Examples of N-GUL gene organizations that can be found in intra- and interspecies.
Extended Data Fig. 3 Man9GlcNAc breakdown by the action of Bl_Man38A and Bl_Man38C.
a, Negative control of Man9GlcNAc with no enzyme. b, Only Bl_Man38A. c, Only Bl_Man38C. d, Both Bl_Man38A and Bl_Man38C added into the reaction. Note that the combined reaction led to complete depletion of the substrate, forming mainly the final product Man1GlcNAc and residual amounts of Man3GlcNAc.
Extended Data Fig. 4 Bl_Man125 displayed no activity over Man5GlcNAc and Man9GlcNAc substrates.
Biochemical profile of Bl_Man125 using Man5GlcNAc (a, b) and Man9GlcNAc (c, d) as putative substrates in 60-min reactions. a, Negative control of Man5GlcNAc with no enzyme. b, Man5GlcNAc with Bl_Man125. c, Negative control of Man9GlcNAc with no enzyme. d, Man9GlcNAc with Bl_Man125.
Extended Data Fig. 5 The complementary activity of Bl_Man125 with Bl_Man38A-C enzymes on Man5GlcNAc.
Reactions combining the GH125 α-1,6-mannosidase and the GH38 enzymes using Man5GlcNAc to evaluate a putative booster effect in the degradation of small oligosaccharides. a, Negative control (no enzyme). b, Bl_Man125, c, Bl_Man38A, d, Bl_Man38A and Bl_Man125, e, Bl_Man38B, f, Bl_Man38B and Bl_Man125, g, Bl_Man38C, h, Bl_Man38C and Bl_Man125, i, Bl_Man38A and Bl_Man38C, j, Bl_Man38A, Bl_Man38C and Bl_Man125. Note that a remarkable difference is observed for Bl_Man38B, in which peaks corresponding to Man3GlcNAc and Man2GlcNAc are depleted in the presence of Bl_Man125.
Extended Data Fig. 6 The distinct oligomeric states observed in the GH38 family.
Oligomeric arrangements described for the members of the GH38 family, represented by the structures under the PDB code 1HTY, 6LZ1, 1O7D and 2WYH. Blue protomers were structurally aligned, maintaining the same position and dimension for all the structures. The r.m.s.d. values of the structural comparisons of these enzymes with Bl_Man38A, Bl_Man38B and Bl_Man38C are provided in the Supplementary Table 7.
Extended Data Fig. 7 Maximum likelihood phylogeny of characterized members of the GH38 family.
Red circles indicate the sequences containing at least one structure deposited in the Protein Data Bank, with their respective codes assigned. The yellow clade indicates bacterial GH38 α-mannosidases that oligomerizes as globular dimers and is represented by the structure under the PDB code 2WYH. The purple clades correspond to lysosomal and vacuolar (for plants) family members, which oligomerize as elongated dimers as shown for the structure under the PDB code 1O7D. The green clade corresponds to monomeric enzymes that are anchored to Golgi apparatus membrane, represented by the structure under the PDB code 1HTY. The red clades comprise the tetrameric enzymes, grouping cytosolic, vacuolar (for yeasts) and bifidobacterial members in the same phylogenetic branch. Grey clades do not have structural data available to infer the oligomeric state of their members. A GH92 α-mannosidase was used as the outgroup.
Extended Data Fig. 8 Functional comparison between the mutant Bl_Man38CF726W and the wild-type Bl_Man38C.
Assays using Man5GlcNAc (a–c) and Man9GlcNAc (d–f) as substrate, in 60-min reactions with no enzyme (a, d), with Bl_Man38C (b, e) and with Bl_Man38CF726W (c, f). *: internal control (xylohexaose). g–i, Kinetic assays with Bl_Man38C and Bl_Man38CF726W on the disaccharides α-1,2-mannobiose (g), α-1,3-mannobiose (h) and α-1,6-mannobiose (i). The reactions were supplemented with 0.1 mM CoCl2. Results are expressed as mean ± SD from three independent experiments (g–i).
Extended Data Fig. 9 Functional comparison between the mutant Bl_Man38AΔLeu164-Pro168 and the wild-type Bl_Man38A.
Assays using Man5GlcNAc (a–c) and Man9GlcNAc (d–f) as substrate, with no enzyme (a, d), with Bl_Man38A (b, e) and with Bl_Man38AΔLeu164-Pro168 (c, f). *: internal control (xylohexaose). The reactions were supplemented with 0.1 mM CoCl2.
Extended Data Fig. 10 Enzymatic characterization of the α-glucosidase Bl_Glc31.
Enzyme activity was analyzed by colorimetric (a, b) and capillary electrophoresis methods (c–j). a, Relative activity of Bl_Glc31 on pNP-α-Glc in function of pH in reactions containing McIlvaine buffer (green) and 200 mM sodium/potassium phosphate (purple) buffers. b, Relative activity of Bl_Glc31 over pNP-α-Glc in 200 mM sodium/potassium phosphate buffer in function of temperature. Data from panels a and b are shown as mean ± SD from three independent experiments (n = 3). c–f, Capillary electrophoresis patterns for fructose (c), glucose (d), maltose (e) and sucrose (f). Note that in the sucrose profile only contaminant glucose is observed, and no sucrose is detected due to its lack of reactivity with APTS. (g–j) Reactions of Bl_Glc31 against maltose and sucrose as putative substrates in optimal conditions of the enzyme (200 mM sodium/potassium phosphate buffer pH 5.5, at 25 °C) for 16 hrs. Due to the long reaction time, negative controls without enzyme were performed to verify possible spontaneous hydrolysis of maltose (g) and sucrose (i). In 4 min of elution, a peak of APTS is seen for all the reactions. G2 = maltose. G1 = glucose.
Supplementary information
Supplementary Information
Supplementary Figs. 1–30, Tables 1–14 and references.
Supplementary Table
Data used for preparing Supplementary Figs. 1, 5, 6, 7, 9, 16 and 23–30.
Supplementary Data 1
Cryo-EM micrograph of Bl_Man38C.
Supplementary Data 2
Cryo-EM micrograph of Bl_Man38B.
Supplementary Data 3
Cryo-EM micrograph of Bl_Man38A.
Supplementary Data 4
Cryo-EM micrograph of the mutant Bl_Man38AD387A.
Source data
Source Data Fig. 1
Activity of Bl_Man38A, Bl_Man38B and Bl_Man38C on native substrates and disaccharides.
Source Data Fig. 4
Cleavage profile of high-mannose N-glycans by Bl_Endo85 and Bl_Glc31.
Source Data Fig. 5
Substrate saturation curve of the ManI Bl_MI.
Source Data Extended Data Fig. 3
Man9GlcNAc breakdown by the action of Bl_Man38A and Bl_Man38C.
Source Data Extended Data Fig. 4
Bl_Man125 displayed no activity over Man5GlcNAc and Man9GlcNAc substrates.
Source Data Extended Data Fig. 5
The complementary activity of Bl_Man125 with Bl_Man38A–Bl_Man38C enzymes on Man5GlcNAc.
Source Data Extended Data Fig. 8
Functional comparison between the mutant Bl_Man38CF726W and the wild-type Bl_Man38C.
Source Data Extended Data Fig. 9
Functional comparison between the mutant Bl_Man38AΔLeu 164–Pro 168 and the wild-type Bl_Man38A.
Source Data Extended Data Fig. 10
Enzymatic characterization of the α-glucosidase Bl_Glc31.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Cordeiro, R.L., Santos, C.R., Domingues, M.N. et al. Mechanism of high-mannose N-glycan breakdown and metabolism by Bifidobacterium longum. Nat Chem Biol 19, 218–229 (2023). https://doi.org/10.1038/s41589-022-01202-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-022-01202-4
This article is cited by
-
A sweet feast
Nature Chemical Biology (2023)