Abstract
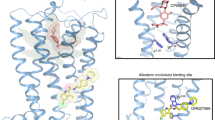
Given the promising clinical value of allosteric modulators of G protein-coupled-receptors (GPCRs), mechanistic understanding of how these modulators alter GPCR function is of significance. Here, we report the crystallographic and cryo-electron microscopy structures of the cannabinoid receptor CB1 bound to the positive allosteric modulator (PAM) ZCZ011. These structures show that ZCZ011 binds to an extrahelical site in the transmembrane 2 (TM2)-TM3-TM4 surface. Through (un)biased molecular dynamics simulations and mutagenesis experiments, we show that TM2 rearrangement is critical for the propagation of allosteric signals. ZCZ011 exerts a PAM effect by promoting TM2 rearrangement in favor of receptor activation and increasing the population of receptors that adopt an active conformation. In contrast, ORG27569, a negative allosteric modulator (NAM) of CB1, also binds to the TM2-TM3-TM4 surface and exerts a NAM effect by impeding the TM2 rearrangement. Our findings fill a gap in the understanding of CB1 allosteric regulation and could guide the rational design of CB1 allosteric modulators.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Structural data have been deposited in the Protein Data Bank (PDB) with coordinate accession numbers 7FEE (crystal structure of the CB1 bound with CP55940 and ZCZ011) and 7WV9 (cryo-EM structure of the CP55940–ZCZ011-bound CB1–Gi complex), and the Electron Microscopy Data Bank (EMDB), accession number EMD-32850, is provided for the cryo-EM structure. All other data generated or analyzed during this study are included in this published article (and its Supplementary information files) or are available from the corresponding author on reasonable request. Source data are provided with this paper.
References
Hauser, A. S., Attwood, M. M., Rask-Andersen, M., Schioth, H. B. & Gloriam, D. E. Trends in GPCR drug discovery: new agents, targets and indications. Nat. Rev. Drug Discov. 16, 829–842 (2017).
Slosky, L. M., Caron, M. G. & Barak, L. S. Biased allosteric modulators: new frontiers in GPCR drug discovery. Trends Pharmacol. Sci. 42, 283–299 (2021).
Bueno, A. B. et al. Structural insights into probe-dependent positive allosterism of the GLP-1 receptor. Nat. Chem. Biol. 16, 1105–1110 (2020).
Chen, S. et al. Human substance P receptor binding mode of the antagonist drug aprepitant by NMR and crystallography. Nat. Commun. 10, 638 (2019).
Zou, S. & Kumar, U. Cannabinoid receptors and the endocannabinoid system: signaling and function in the central nervous system. Int. J. Mol. Sci. 19, 833 (2018).
Lu, D., Immadi, S. S., Wu, Z. & Kendall, D. A. Translational potential of allosteric modulators targeting the cannabinoid CB1 receptor. Acta Pharmacol. Sin. 40, 324–335 (2019).
Lane, J. R., May, L. T., Parton, R. G., Sexton, P. M. & Christopoulos, A. A kinetic view of GPCR allostery and biased agonism. Nat. Chem. Biol. 13, 929–937 (2017).
Foster, D. J. & Conn, P. J. Allosteric modulation of GPCRs: new insights and potential utility for treatment of schizophrenia and other CNS disorders. Neuron 94, 431–446 (2017).
Morales, P., Goya, P., Jagerovic, N. & Hernandez-Folgado, L. Allosteric modulators of the CB1 cannabinoid receptor: a structural update review. Cannabis Cannabinoid Res. 1, 22–30 (2016).
Shao, Z. et al. Structure of an allosteric modulator bound to the CB1 cannabinoid receptor. Nat. Chem. Biol. 15, 1199–1205 (2019).
Latorraca, N. R., Venkatakrishnan, A. J. & Dror, R. O. GPCR dynamics: structures in motion. Chem. Rev. 117, 139–155 (2017).
Ignatowska-Jankowska, B. M. et al. A cannabinoid CB 1 receptor-positive allosteric modulator reduces neuropathic pain in the mouse with no psychoactive effects. Neuropsychopharmacology 40, 2948–2959 (2015).
Cheng, R. K. Y. et al. Structural insight into allosteric modulation of protease-activated receptor 2. Nature 545, 112–115 (2017).
Srivastava, A. et al. High-resolution structure of the human GPR40 receptor bound to allosteric agonist TAK-875. Nature 513, 124–127 (2014).
Hua, T. et al. Crystal structure of the human cannabinoid receptor CB1. Cell 167, 750–762.e14 (2016).
Hua, T. et al. Activation and signaling mechanism revealed by cannabinoid receptor-Gi complex structures. Cell 180, 655–665.e18 (2020).
Zhou, Q. et al. Common activation mechanism of class A GPCRs. eLife 8, e50279 (2019).
Hilger, D. et al. Structural insights into differences in G protein activation by family A and family B GPCRs. Science 369, eaba3373 (2020).
Liu, K. et al. Structural basis of CXC chemokine receptor 2 activation and signalling. Nature 585, 135–140 (2020).
Shao, Z. et al. High-resolution crystal structure of the human CB1 cannabinoid receptor. Nature 540, 602–606 (2016).
Díaz, Ó., Dalton, J. A. & Giraldo, J. Revealing the mechanism of agonist-mediated cannabinoid receptor 1 (CB1) activation and phospholipid-mediated allosteric modulation. J. Med. Chem. 62, 5638–5654 (2019).
Grahl, A., Abiko, L. A., Isogai, S., Sharpe, T. & Grzesiek, S. A high-resolution description of β1-adrenergic receptor functional dynamics and allosteric coupling from backbone NMR. Nat. Commun. 11, 2216 (2020).
Bonomi, M. & Parrinello, M. Enhanced sampling in the well-tempered ensemble. Phys. Rev. Lett. 104, 190601 (2010).
Galdadas, I. et al. Structural basis of the effect of activating mutations on the EGF receptor. eLife 10, e65824 (2021).
Lovera, S. et al. The different flexibility of c-Src and c-Abl kinases regulates the accessibility of a druggable inactive conformation. J. Am. Chem. Soc. 134, 2496–2499 (2012).
Zanetti-Domingues, L. C. et al. The architecture of EGFR’s basal complexes reveals autoinhibition mechanisms in dimers and oligomers. Nat. Commun. 9, 4325 (2018).
Mattedi, G., Acosta-Gutiérrez, S., Clark, T. & Gervasio, F. L. A combined activation mechanism for the glucagon receptor. Proc. Natl Acad. Sci. USA 117, 15414–15422 (2020).
Mattedi, G., Deflorian, F., Mason, J. S., de Graaf, C. & Gervasio, F. L. Understanding ligand binding selectivity in a prototypical GPCR family. J. Chem. Inf. Model. 59, 2830–2836 (2019).
Raniolo, S. & Limongelli, V. Ligand binding free-energy calculations with funnel metadynamics. Nat. Protoc. 15, 2837–2866 (2020).
Tiwary, P. & Parrinello, M. A time-independent free energy estimator for metadynamics. J. Phys. Chem. B 119, 736–742 (2015).
Taylor, B. C., Lee, C. T. & Amaro, R. E. Structural basis for ligand modulation of the CCR2 conformational landscape. Proc. Natl Acad. Sci. USA 116, 8131–8136 (2019).
Lu, S. et al. Activation pathway of a G protein-coupled receptor uncovers conformational intermediates as targets for allosteric drug design. Nat. Commun. 12, 4721 (2021).
Kato, H. E. et al. Conformational transitions of a neurotensin receptor 1-Gi1 complex. Nature 572, 80–85 (2019).
Dror, R. O. et al. Activation mechanism of the β2-adrenergic receptor. Proc. Natl Acad. Sci. USA 108, 18684–18689 (2011).
Manglik, A. et al. Structural insights into the dynamic process of β2-adrenergic receptor signaling. Cell 161, 1101–1111 (2015).
Bolhuis, P. G. Kinetic pathways of β-hairpin (un)folding in explicit solvent. Biophys. J. 88, 50–61 (2005).
Bussi, G., Gervasio, F. L., Laio, A. & Parrinello, M. Free-energy landscape for β hairpin folding from combined parallel tempering and metadynamics. J. Am. Chem. Soc. 128, 13435–13441 (2006).
Tao, Q. & Abood, M. E. Mutation of a highly conserved aspartate residue in the second transmembrane domain of the cannabinoid receptors, CB1 and CB2, disrupts G-protein coupling. J. Pharmacol. Exp. Ther. 285, 651 (1998).
Wingler, L. M. et al. Angiotensin and biased analogs induce structurally distinct active conformations within a GPCR. Science 367, 888–892 (2020).
Varma, N. et al. Crystal structure of jumping spider rhodopsin-1 as a light sensitive GPCR. Proc. Natl Acad. Sci. USA 116, 14547 (2019).
D’Antona, A. M., Ahn, K. H. & Kendall, D. A. Mutations of CB1 T210 produce active and inactive receptor forms: correlations with ligand affinity, receptor stability, and cellular localization. Biochemistry 45, 5606–5617 (2006).
Lu, J. et al. Structural basis for the cooperative allosteric activation of the free fatty acid receptor GPR40. Nat. Rev. Mol. Cell Biol. 24, 570–577 (2017).
Kruse, A. C. et al. Activation and allosteric modulation of a muscarinic acetylcholine receptor. Nature 504, 101–106 (2013).
Maeda, S., Qu, Q., Robertson, M. J., Skiniotis, G. & Kobilka, B. K. Structures of the M1 and M2 muscarinic acetylcholine receptor/G-protein complexes. Science 364, 552 (2019).
Liu, X. et al. Mechanism of β2AR regulation by an intracellular positive allosteric modulator. Science 364, 1283–1287 (2019).
Lin, S. et al. Structures of Gi-bound metabotropic glutamate receptors mGlu2 and mGlu4. Nature 594, 583–588 (2021).
Shen, C. et al. Structural basis of GABAB receptor–Gi protein coupling. Nature 594, 594–598 (2021).
Xiao, P. et al. Ligand recognition and allosteric regulation of DRD1-Gs signaling complexes. Cell 184, 943–956.e18 (2021).
Qi, X., Friedberg, L., De Bose-Boyd, R., Long, T. & Li, X. Sterols in an intramolecular channel of Smoothened mediate Hedgehog signaling. Nat. Chem. Biol. 16, 1368–1375 (2020).
Mao, C. et al. Cryo-EM structures of inactive and active GABAB receptor. Cell Res. 30, 564–573 (2020).
Yan, W. et al. Structure of the human gonadotropin-releasing hormone receptor GnRH1R reveals an unusual ligand binding mode. Nat. Commun. 11, 5287 (2020).
Collaborative Computational Project, No. 4 The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D Biol. Crystallogr . 50, 760–763 (1994).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Koehl, A. et al. Structure of the µ-opioid receptor–Gi protein complex. Nature 558, 547–552 (2018).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010).
Olsen, R. H. et al. TRUPATH, an open-source biosensor platform for interrogating the GPCR transducerome. Nat. Chem. Biol. 16, 841–849 (2020).
Huang, J. & MacKerell, A. D. Jr. CHARMM36 all-atom additive protein force field: validation based on comparison to NMR data. J. Comput. Chem. 34, 2135–2145 (2013).
Abraham, M. J. et al. GROMACS: high performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1-2, 19–25 (2015).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (31972916, 81930125, 82130104, 32100988), Science and Technology Department of Sichuan Province 2020YJ0208 (Z.S.), 1.3.5 project for disciplines of excellence, West China Hospital, Sichuan University (ZYYC20023, ZYGD18001, ZYXY21001). Science and Technology Department of Chengdu (2019-YF05-00294-SN to Z.S.). We thank staff of the BL18U beamline at National Center for Protein Sciences Shanghai (NCPSS) and BL32XU beamline of Spring-8. The diffraction data collection was performed at the BL32XU of Spring-8 with approval of the Japan Synchrotron Radiation Research Institute (JASRI) (proposal number 2019B2705). We thank J. Giraldo for providing MD-generated receptor conformations at the beginning of the project. This research used resources from the Duyu High Performance Computing Center, Sichuan University, and Big Data Platform at West China Hospital of Sichuan University (WCH-BDP).
Author information
Authors and Affiliations
Contributions
Z.S. initiated the project for allostery of CB1. S.Y. and Z.S. conceived and supervised the project. S.Y. and X.Y. designed molecular dynamics simulations. X.Y. performed simulations with the assistance of Y.W. and J.Z. X.Y. analyzed simulations with the assistance of K.L., M.W. and A.X. X.W. performed gene expression and protein purification, crystallization and diffraction data collection with assistance of G.L., Y.Z. and J.L. X.W. determined and analyzed the crystal structures with the assistance of L.C. X.W. designed the expression constructs, purified the ZCZ011–CP55940–CB1–Gi complexes and prepared the final samples for cryo-EM experiments. Z.X. determined the cryo-EM structure with the assistance of X.W. C.W. and Z.S. designed the cellular assays and analyzed results. S.Y., Z.S. with the assistance of X.Y., W.Y. and Z.X. wrote and revised the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Chemical Biology thanks Francesco Gervasio, Aashish Manglik and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Purification and crystallization of CB1-CP55940-ZCZ011 ternary complex.
(a) Snake representation of CB1 crystallization construct used in this study. (b) First derivative analysis of DSF data for purified CB1 constructs with CP55940-ZCZ011. PGS is a fusion protein. 5 M means S2033.39K, T2103.46A, E2735.37K, T2835.47V and R3406.32E mutations in CB1 receptor. (c) Superdex 200 gel-filtration trace of CB1 receptor in complex with CP55940 and ZCZ011. SDS-PAGE is for crystallization protein sample validation. Samples are prepared and repeated independently over three times (n = 3). (d) Crystal images for CB1-CP55940-ZCZ011 complex. The left image shown in visible light and the right shown under polarized light. Crystals can be formed over three independent experiments (n = 3). (e) Lattice packing of structure of CB1-CP55940-ZCZ011 complex.
Extended Data Fig. 2 PAM binding site analysis.
(a) The 2Fo-Fc map of PAM ZCZ011 and surrounding residues within 4 Å are contoured at 1.5 σ level. (b) Measurement of expression levels of the CB1 WT and indicated mutants by ELISA experiments. The mutant receptor expression was normalized to comparable levels as wild-type receptor expression. Values are means ± SEM (n = 4) from four independent experiments performed in triplicates. ns, no significance, *p < 0.1, **p < 0.01, ***p < 0.001 (one−way analysis of variance [ANOVA] followed by the Dunnett’s test, compared with the response of WT, p = 0.66, p = 0.09, p = 0.39, p < 0.001, p = 0.91, p = 0.19, p = 0.12, p < 0.01, p = 0.99, p < 0.001, p < 0.001, p = 0.18, p = 0.05, p = 0.14, p = 0.13, p = 0.08, p > 0.99, p < 0.001, p < 0.01 from left to right). (c) Effects of the S1732.60A in CB1 on CP55940 induced cAMP inhibition. Data are presented as the means ± SEM (n = 3) of three independent experiments performed in triplicate. (d) The inhibitory effect of CP55940 on forskolin-induced cAMP accumulation assay in the presence of or the absence of 0.5 μM PAM ZCZ011. F1913.27L, S1993.35T and V2494.58Y mutations decreased PAM potency significantly. Bars represent differences in calculated the ratio of the potency of CP55940 (EC50 [half maximal effective concentration]) in the presence of ZCZ011 or not for each mutation relative to WT of CB1. ns means not significant, **p < 0.01, ***p < 0.001 (one-way analysis of variance [ANOVA] followed by the Dunnett’s test, compared with the response of WT, p < 0.01, p = 0.91, p > 0.99, p < 0.001, p < 0.001 from left to right). All data are presented as the means ± SEM; n = 3.
Extended Data Fig. 3 Comparison of allosteric sites of CB1 with that in other GPCRs (PAR2 and GPR40).
(a) Analysis of sequence conservation in PAM site of CB1. (b) Structural comparison of PAM bound CB1 with allosteric modulator AZ3451 bound PAR2 (5NDZ). (c) Structural comparison of PAM bound CB1 with TAK-875 bound GPR40 (4PHU).
Extended Data Fig. 5 Conformational changes in CB1 activation.
(a) Comparison of the active state CB1 bound agonist AM841 (salmon, PDB: 6KPG) with the inactive state CB1 bound antagonist AM6538 (slate, PDB: 5TGZ) reveals that the intracellular end of the TM6 helix moves outward significantly. (b) The upward rearrangement of TM3 upon CB1 activation (salmon, PDB: 6KPG). (c) The extracellular end of TM2 has inward movement, and its intracellular end is found to have downward rotation along the helical axis when comparing with inactive state of CB1 (slate, PDB: 5TGZ). In particular, F1552.42 undergoes a major conformational transition during receptor activation. (d) Structural superimposition reveals that the extracellular ends of TM1 and TM2 move into the core of helical bundles.
Extended Data Fig. 6 Structural comparison of in active and inactive state of GPCRs.
Structural comparison of TM2 conformation in active and inactive GPCRs. (a)The comparison reveals that there is less obvious conformational rearrangement of TM2 in most GPCRs, including aminergic receptors β2AR (active PDB: 6NI3, inactive PDB: 5D5A), M2 (active PDB: 6OIK, inactive PDB: 5ZKC), D2 (active PDB: 6VMS, inactive PDB: 7DFP), 5-HT2B (active PDB: 5TUD, intermedia: 6DRY) and H1 (active PDB: 7DFL, inactive PDB: 3RDZ); Nucleotide receptors A1R (active PDB: 6D9H, inactive PDB: 5N2S), Lipid receptors CB2 (active PDB: 6KPF, inactive PDB: 5ZTY) and EP4 (active PDB: 7D7M, inactive PDB: 5YWY); Chemokine receptor CXCR2 (active PDB: 6LFM, inactive PDB: 6LFL); Peptide receptors OPRD (active PDB: 6PT2, inactive PDB: 4RWD), AT1R (active PDB: 6OS0, inactive PDB: 4ZUD) and MC4 (active PDB: 7AUE, inactive PDB: 6W25). (b) The comparison reveals that there is more obvious conformational rearrangement of TM2 in most GPCRs, including OX2R (active PDB: 7L1V, inactive PDB: 6TPN), 5-HT2A (active PDB: 6WHA, inactive PDB: 6A94), A2A (active PDB: 6GDG, inactive PDB: 6ZDV), OPRM (active PDB: 6DDE, inactive PDB: 4DKL), OPRK (active PDB: 6B73, inactive PDB: 6VI4).
Extended Data Fig. 7 Traces during simulations of CB1-CP55940, starting from the crystal structure (PDB: 6KQI).
(a) RMSD of CP55940 from the initial crystal state during MD simulations of CB1 with bound CP55940. (b) RMSD of Cα atoms of CB1 TM6 from the initial crystal state. (c) RMSD of Cα atoms of TM helices from the initial crystal state. (d) Distance between ionic lock residues in CB1 (R2143.50 and D3386.30).
Extended Data Fig. 8 Representative snapshots during unbiased simulations of CB1-CP55940, starting from the crystal structure (PDB: 6KQI).
(a) Conformation comparison of representative snapshot in trajectory #2 (smudge) and CP99540 bound inactive CB1 (PDB: 6KQI) (gray). (b) Along the MD trajectories of CP55940-bound CB1, a small number of other different binding poses exist for CP55940.
Extended Data Fig. 9 ZCZ011 enhances CB1 binding to Gi protein.
(a) Antagonist AM6538 (blue) -bound CB1 (light blue) (PDB 5TGZ) orthotopic pockets expand and agonist CP55940 (magenta) bound CB1 (green-cyan) orthotopic pockets shrink. (b) ZCZ011 bound CB1 to enhance the interaction between S1522.39 and Gi D350.
Extended Data Fig. 10 CB1 receptor-Gi complex bound to CP55940 and ZCZ011 purification, cryo-EM data collection and cryo-EM map quality.
(a) Size-exclusion chromatography elution profiles of the purified receptor complex. (b) SDS-PAGE analysis of CB1 receptor-Gαo1-Gβγ-scFv16 complexes. Samples are prepared and repeated over three times. (c) Representative cryo-EM image micrographs of CB1 receptor-Gαo1-Gβγ-scFv16 complexes (left panel) and 2D class averages (right panel), from one of the total 5501 movies. (d) Flow chart of cryo-EM data analysis for the densities of ghrelin-ghrelin receptor-Go complex. The final resolution of the density is 3.3 Å. (e) The ‘gold-standard’ FSC curves, with the global resolution defined at the FSC = 0.143 is 3.3 Å for CP55940 and ZCZ011 bound CB1 receptor-Gi complex. (f) Representative cryo-EM density maps and fitted atomic models for all seven transmembrane helixes and the helix 8.
Supplementary information
Supplementary Information
Supplementary Tables 1–3, Figs. 1–19 and Notes 1 and 2.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 1
Unprocessed gel.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 10
Unprocessed gel.
Rights and permissions
About this article
Cite this article
Yang, X., Wang, X., Xu, Z. et al. Molecular mechanism of allosteric modulation for the cannabinoid receptor CB1. Nat Chem Biol 18, 831–840 (2022). https://doi.org/10.1038/s41589-022-01038-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-022-01038-y
This article is cited by
-
G protein-coupled receptors (GPCRs): advances in structures, mechanisms, and drug discovery
Signal Transduction and Targeted Therapy (2024)
-
Effects of CB1 receptor negative allosteric modulator Org27569 on oxycodone withdrawal symptoms in mice
Psychopharmacology (2024)
-
GPR161 structure uncovers the redundant role of sterol-regulated ciliary cAMP signaling in the Hedgehog pathway
Nature Structural & Molecular Biology (2024)
-
Structure, function and drug discovery of GPCR signaling
Molecular Biomedicine (2023)
-
Mechanism of activation and biased signaling in complement receptor C5aR1
Cell Research (2023)