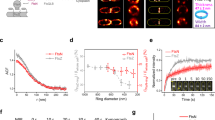
Abstract
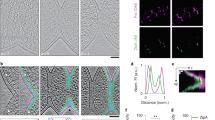
Members of the Corynebacterineae, including Corynebacterium and Mycobacterium, have an atypical cell envelope characterized by an additional mycomembrane outside of the peptidoglycan layer. How this multilayered cell envelope is assembled remains unclear. Here, we tracked the assembly dynamics of different envelope layers in Corynebacterium glutamicum and Mycobacterium smegmatis by using metabolic labeling and found that the septal cell envelope is assembled sequentially in both species. Additionally, we demonstrate that in C. glutamicum, the peripheral peptidoglycan layer at the septal junction remains contiguous throughout septation, forming a diffusion barrier for the fluid mycomembrane. This diffusion barrier is resolved through perforations in the peripheral peptidoglycan, thus leading to the confluency of the mycomembrane before daughter cell separation (V snapping). Furthermore, the same junctional peptidoglycan also serves as a mechanical link holding the daughter cells together and undergoes mechanical fracture during V snapping. Finally, we show that normal V snapping in C. glutamicum depends on complete assembly of the septal cell envelope.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Code availability
All MATLAB scripts used to analyze microscopy images are available upon request.
Data availability
All data generated or analyzed during this study are included in this published article (and its supplementary information files) or are available from the corresponding author on reasonable request.
References
Silhavy, T. J., Kahne, D. & Walker, S. The bacterial cell envelope. Cold Spring Harb. Perspect. Biol. 2, a000414 (2010).
Hoffmann, C., Leis, A., Niederweis, M., Plitzko, J. M. & Engelhardt, H. Disclosure of the mycobacterial outer membrane: cryo-electron tomography and vitreous sections reveal the lipid bilayer structure. Proc. Natl Acad. Sci. USA 105, 3963–3967 (2008).
Zuber, B. et al. Direct visualization of the outer membrane of mycobacteria and corynebacteria in their native state. J. Bacteriol. 190, 5672–5680 (2008).
Marrakchi, H., Lanéelle, M. A. & Daffé, M. Mycolic acids: structures, biosynthesis, and beyond. Chem. Biol. 21, 67–85 (2014).
Rodriguez-Rivera, F. P., Zhou, X., Theriot, J. A. & Bertozzi, C. R. Visualization of mycobacterial membrane dynamics in live cells. J. Am. Chem. Soc. 139, 3488–3495 (2017).
Draper, P. The outer parts of the mycobacterial envelope as permeability barriers. Front. Biosci. 3, D1253–D1261 (1998).
Jackson, M. et al. Inactivation of the antigen 85C gene profoundly affects the mycolate content and alters the permeability of the Mycobacterium tuberculosis cell envelope. Mol. Microbiol. 31, 1573–1587 (1999).
Zhang, Y. The magic bullets and tuberculosis drug targets. Annu. Rev. Pharmacol. Toxicol. 45, 529–564 (2005).
Janin, Y. L. Antituberculosis drugs: ten years of research. Bioorg. Med. Chem. 15, 2479–2513 (2007).
Jankute, M., Cox, J. A., Harrison, J. & Besra, G. S. Assembly of the mycobacterial cell wall. Annu. Rev. Microbiol. 69, 405–423 (2015).
Daniel, R. A. & Errington, J. Control of cell morphogenesis in bacteria: two distinct ways to make a rod-shaped cell. Cell 113, 767–776 (2003).
Kuru, E. et al. In situ probing of newly synthesized peptidoglycan in live bacteria with fluorescent d-amino acids. Angew. Chem. Int. Edn. Engl. 51, 12519–12523 (2012).
Letek, M. et al. DivIVA is required for polar growth in the MreB-lacking rod-shaped actinomycete Corynebacterium glutamicum. J. Bacteriol. 190, 3283–3292 (2008).
Meniche, X. et al. Subpolar addition of new cell wall is directed by DivIVA in mycobacteria. Proc. Natl Acad. Sci. USA 111, E3243–E3251 (2014).
Carel, C. et al. Mycobacterium tuberculosis proteins involved in mycolic acid synthesis and transport localize dynamically to the old growing pole and septum. PLoS One 9, e97148 (2014).
Hill, H. W. Branching in bacteria with special reference to B. diphtheriae. J. Med. Res. 7, 115–127 (1902).
Dahl, J. L. Electron microscopy analysis of Mycobacterium tuberculosis cell division. FEMS Microbiol. Lett. 240, 15–20 (2004).
Letek, M. et al. Cell growth and cell division in the rod-shaped actinomycete Corynebacterium glutamicum. Antonie van Leeuwenhoek 94, 99–109 (2008).
Zhou, X., Halladin, D. K. & Theriot, J. A. Fast mechanically driven daughter cell separation is widespread in actinobacteria. MBio 7, e00952–16 (2016).
Siegrist, M. S., Swarts, B. M., Fox, D. M., Lim, S. A. & Bertozzi, C. R. Illumination of growth, division and secretion by metabolic labeling of the bacterial cell surface. FEMS Microbiol. Rev. 39, 184–202 (2015).
Backus, K. M. et al. Uptake of unnatural trehalose analogs as a reporter for Mycobacterium tuberculosis. Nat. Chem. Biol. 7, 228–235 (2011).
Foley, H. N., Stewart, J. A., Kavunja, H. W., Rundell, S. R. & Swarts, B. M. Bioorthogonal chemical reporters for selective in situ probing of mycomembrane components in mycobacteria. Angew. Chem. Int. Edn. Engl. 55, 2053–2057 (2016).
Kumagai, Y., Hirasawa, T., Hayakawa, K., Nagai, K. & Wachi, M. Fluorescent phospholipid analogs as microscopic probes for detection of the mycolic acid-containing layer in Corynebacterium glutamicum: detecting alterations in the mycolic acid-containing layer following ethambutol treatment. Biosci. Biotechnol. Biochem. 69, 2051–2056 (2005).
Puech, V. et al. Structure of the cell envelope of corynebacteria: importance of the non-covalently bound lipids in the formation of the cell wall permeability barrier and fracture plane. Microbiology 147, 1365–1382 (2001).
Ngo, J. T. et al. Click-EM for imaging metabolically tagged nonprotein biomolecules. Nat. Chem. Biol. 12, 459–465 (2016).
Vijay, S., Anand, D. & Ajitkumar, P. Unveiling unusual features of formation of septal partition and constriction in mycobacteria: an ultrastructural study. J. Bacteriol. 194, 702–707 (2012).
Zhou, X. et al. Mechanical crack propagation drives millisecond daughter cell separation in Staphylococcus aureus. Science 348, 574–578 (2015).
Peyret, J. L. et al. Characterization of the cspB gene encoding PS2, an ordered surface-layer protein in Corynebacterium glutamicum. Mol. Microbiol. 9, 97–109 (1993).
Hansmeier, N. et al. Classification of hyper-variable Corynebacterium glutamicum surface-layer proteins by sequence analyses and atomic force microscopy. J. Biotechnol. 112, 177–193 (2004).
Hansmeier, N. et al. The surface (S)-layer gene cspB of Corynebacterium glutamicum is transcriptionally activated by a LuxR-type regulator and located on a 6kb genomic island absent from the type strain ATCC 13032. Microbiology 152, 923–935 (2006).
Barreau, C., Bimet, F., Kiredjian, M., Rouillon, N. & Bizet, C. Comparative chemotaxonomic studies of mycolic acid-free coryneform bacteria of human origin. J. Clin. Microbiol. 31, 2085–2090 (1993).
Umeda, A. & Amako, K. Growth of the surface of Corynebacterium diphtheriae. Microbiol. Immunol. 27, 663–671 (1983).
Tsuge, Y., Ogino, H., Teramoto, H., Inui, M. & Yukawa, H. Deletion of cgR_1596 and cgR_2070, encoding NlpC/P60 proteins, causes a defect in cell separation in Corynebacterium glutamicum R. J. Bacteriol. 190, 8204–8214 (2008).
Atilgan, E., Magidson, V., Khodjakov, A. & Chang, F. Morphogenesis of the fission yeast cell through cell wall expansion. Curr. Biol. 25, 2150–2157 (2015).
Telenti, A. et al. The emb operon, a gene cluster of Mycobacterium tuberculosis involved in resistance to ethambutol. Nat. Med. 3, 567–570 (1997).
Escuyer, V. E. et al. The role of the embA and embB gene products in the biosynthesis of the terminal hexaarabinofuranosyl motif of Mycobacterium smegmatis arabinogalactan. J. Biol. Chem. 276, 48854–48862 (2001).
Mikusová, K., Slayden, R. A., Besra, G. S. & Brennan, P. J. Biogenesis of the mycobacterial cell wall and the site of action of ethambutol. Antimicrob. Agents Chemother. 39, 2484–2489 (1995).
Radmacher, E. et al. Ethambutol, a cell wall inhibitor of Mycobacterium tuberculosis, elicits l-glutamate efflux of Corynebacterium glutamicum. Microbiology 151, 1359–1368 (2005).
Schubert, K. et al. The antituberculosis drug ethambutol selectively blocks apical growth in CMN group bacteria. MBio 8, e02213–16 (2017).
Kawai, Y. et al. A widespread family of bacterial cell wall assembly proteins. EMBO J. 30, 4931–4941 (2011).
Chan, Y. G., Frankel, M. B., Dengler, V., Schneewind, O. & Missiakas, D. Staphylococcus aureus mutants lacking the LytR-CpsA-Psr family of enzymes release cell wall teichoic acids into the extracellular medium. J. Bacteriol. 195, 4650–4659 (2013).
Baumgart, M., Schubert, K., Bramkamp, M. & Frunzke, J. Impact of LytR-CpsA-Psr proteins on cell wall biosynthesis in Corynebacterium glutamicum. J. Bacteriol. 198, 3045–3059 (2016).
Grzegorzewicz, A. E. et al. Assembling of the Mycobacterium tuberculosis cell wall core. J. Biol. Chem. 291, 18867–18879 (2016).
Harrison, J. et al. Lcp1 is a phosphotransferase responsible for ligating arabinogalactan to peptidoglycan in Mycobacterium tuberculosis. MBio 7, e00972–16 (2016).
Dautin, N. et al. Mycoloyltransferases: a large and major family of enzymes shaping the cell envelope of Corynebacteriales. Biochim. Biophys. Acta Gen. Subj. 1861, 3581–3592 (2017).
Touchette, M. H. & Seeliger, J. C. Transport of outer membrane lipids in mycobacteria. Biochim. Biophys. Acta Mol. Cell. Biol. Lipids 1862, 1340–1354 (2017).
Baumgart, M. et al. Construction of a prophage-free variant of Corynebacterium glutamicum ATCC 13032 for use as a platform strain for basic research and industrial biotechnology. Appl. Environ. Microbiol. 79, 6006–6015 (2013).
Okibe, N., Suzuki, N., Inui, M. & Yukawa, H. Efficient markerless gene replacement in Corynebacterium glutamicum using a new temperature-sensitive plasmid. J. Microbiol. Methods 85, 155–163 (2011).
Rodriguez-Rivera, F. P., Zhou, X., Theriot, J. A. & Bertozzi, C. R. Acute modulation of mycobacterial cell envelope biogenesis by front-line tuberculosis drugs. Angew. Chem. Int. Edn. Engl. 57, 5267–5272 (2018).
Besanceney-Webler, C. et al. Increasing the efficacy of bioorthogonal click reactions for bioconjugation: a comparative study. Angew. Chem. Int. Edn. Engl. 50, 8051–8056 (2011).
Paintdakhi, A. et al. Oufti: an integrated software package for high-accuracy, high-throughput quantitative microscopy analysis. Mol. Microbiol. 99, 767–777 (2016).
Acknowledgements
We thank B. Swarts (Central Michigan University) for providing O-alkTMM and J. Ngo (University of California, San Diego) for DBF-N3; J. Perrino, L. Joubert, and M. Footer (Stanford University) for assistance with the contrast-enhanced TEM sample preparation; A. Straight (Stanford University) for access to the microscope used for FRAP experiments; J. Frunzke (Institute of Bio- and Geosciences, IBG-1: Biotechnology, Forschungszentrum Jülich GmbH) for strain MB001; the Research Institute of Innovative Technology for the Earth in Kyoto Japan for plasmid pCRD206; M. Tsuchida (Open Imaging) for help with the laser ablation experiment; and E. Koslover (University of California, San Diego) for helpful discussions. X.Z. was supported by a Stanford University Interdisciplinary Graduate Fellowship. F.P.R.-R. was supported by a Ford Foundation Predoctoral Fellowship and a University of California Chancellor’s Fellowship. H.C.L. was supported by a Simons Foundation Life Science Research Foundation fellowship. J.C.B. was supported by an NIH Ruth Kirchstein National Research Service Award (F32GM116338). This work was supported by National Institutes of Health grants AI036929 (to J.A.T.), GM058867 (to C.R.B.), and AI051622 (to C.R.B.); Stanford Center for Systems Biology grant P50-GM107615 (to J.A.T.); and the Howard Hughes Medical Institute (to J.A.T. and C.R.B.) and an HHMI-Simons Faculty Scholar Award (to T.G.B.). The transmission electron microscope used in this project was supported in part by ARRA award 1S10RR026780-01 from the National Center for Research Resources (NCRR). The manuscript’s contents are solely the responsibility of the authors and do not necessarily represent the official views of the NCRR or the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
X.Z., F.P.R.-R., C.R.B., and J.A.T. conceived the project and designed experiments. X.Z. performed the experiments and analyzed data. F.P.R.-R. synthesized and characterized the fluorescent probes. H.C.L. and T.G.B. constructed bacterial strains. J.C.B. contributed expertise to the FRAP experiments and analysis. X.Z. and J.A.T. wrote the manuscript. All authors discussed the results and interpretation of the data, and read and commented on the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Text and Figures
Supplementary Table 1 and Supplementary Figures 1–14
Supplementary Video 1
Cell envelope assembly dynamics in C. glutamicum.
Supplementary Video 2
Confluency of mycomembrane in C. glutamicum.
Supplementary Video 3
Cell envelope assembly dynamics and delayed V snapping in ∆cgp_1735.
Supplementary Video 4
Perturbed cell envelope assembly and V snapping in EMB treated C. glutamicum
Rights and permissions
About this article
Cite this article
Zhou, X., Rodriguez-Rivera, F.P., Lim, H.C. et al. Sequential assembly of the septal cell envelope prior to V snapping in Corynebacterium glutamicum. Nat Chem Biol 15, 221–231 (2019). https://doi.org/10.1038/s41589-018-0206-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-018-0206-1
This article is cited by
-
Membrane manipulation by free fatty acids improves microbial plant polyphenol synthesis
Nature Communications (2023)
-
Eukaryotic-like gephyrin and cognate membrane receptor coordinate corynebacterial cell division and polar elongation
Nature Microbiology (2023)
-
Regulation of the cell division hydrolase RipC by the FtsEX system in Mycobacterium tuberculosis
Nature Communications (2023)
-
The mycobacterial cell envelope — a moving target
Nature Reviews Microbiology (2020)
-
Direct visualization of a molecular handshake that governs kin recognition and tissue formation in myxobacteria
Nature Communications (2019)