Abstract
Endometrial carcinoma remains a public health concern with a growing incidence, particularly in younger women. Preserving fertility is a crucial consideration in the management of early-onset endometrioid endometrial carcinoma (EEEC), particularly in patients under 40 who maintain both reproductive desire and capacity. To illuminate the molecular characteristics of EEEC, we undertook a large-scale multi-omics study of 215 patients with endometrial carcinoma, including 81 with EEEC. We reveal an unexpected association between exposome-related mutational signature and EEEC, characterized by specific CTNNB1 and SIGLEC10 hotspot mutations and disruption of downstream pathways. Interestingly, SIGLEC10Q144K mutation in EEECs resulted in aberrant SIGLEC-10 protein expression and promoted progestin resistance by interacting with estrogen receptor alpha. We also identified potential protein biomarkers for progestin response in fertility-sparing treatment for EEEC. Collectively, our study establishes a proteogenomic resource of EEECs, uncovering the interactions between exposome and genomic susceptibilities that contribute to the development of primary prevention and early detection strategies for EEECs.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
Raw sequencing data generated in this study are deposited in the Genome Sequence Archive for Human (https://ngdc.cncb.ac.cn/gsa-human/) with accession number HRA003319. The MS proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE partner repository with the dataset identifier PXD046507. TCGA-UCEC and CPTAC-UCEC data were obtained from Cbioportal (https://www.cbioportal.org/) with the dataset identifier ucec_tcga_pan_can_atlas_2018 and ucec_cptac_2020. Panel sequencing of genomic data from the AACR GENIE Project was obtained from Synapse (https://www.synapse.org/) with the dataset identifier syn7222066. RNA-seq data from six public datasets (GSE1378, GSE1379, GSE6532, GSE9195, GSE12093 and GSE17705) of patients with breast cancer treated with endocrine therapy was obtained from Gene Expression Omnibus (GEO) repository. Source data are provided with this paper.
Code availability
Code required to reproduce the analyses in this paper is available through GitHub (https://github.com/Haz1y/EEEC_landscape) or Zenodo (https://doi.org/10.5281/zenodo.10756345)81.
References
Crosbie, E. J. et al. Endometrial cancer. Lancet 399, 1412–1428 (2022).
Sung, H. et al. Global Cancer Statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 71, 209–249 (2021).
Choi, J. et al. Distinct genomic landscapes in early-onset and late-onset endometrial. JCO Precis. Oncol. 6, e2100401 (2022).
Matsuo, K. et al. Ovarian conservation for young women with early-stage, low-grade endometrial cancer: a 2-step schema. Am. J. Obstet. Gynecol. 224, 574–584 (2021).
Jerzak, K. J., Duska, L. & MacKay, H. J. Endocrine therapy in endometrial cancer: an old dog with new tricks. Gynecol. Oncol. 153, 175–183 (2019).
Dou, Y. et al. Proteogenomic characterization of endometrial carcinoma. Cell 180, 729–748 (2020).
Cancer Genome Atlas Research Network et al.Integrated genomic characterization of endometrial carcinoma. Nature 497, 67–73 (2013).
Soumerai, T. E. et al. Clinical utility of prospective molecular characterization in advanced endometrial cancer. Clin. Cancer Res. 24, 5939–5947 (2018).
Wang, L.-E. et al. Roles of genetic variants in the PI3K and RAS/RAF pathways in susceptibility to endometrial cancer and clinical outcomes. J. Cancer Res. Clin. Oncol. 138, 377–385 (2012).
Liang, H. et al. Whole-exome sequencing combined with functional genomics reveals novel candidate driver cancer genes in endometrial cancer. Genome Res. 22, 2120–2129 (2012).
Kurnit, K. C. et al. CTNNB1 (β-catenin) mutation identifies low grade, early stage endometrial cancer patients at increased risk of recurrence. Mod. Pathol. 30, 1032–1041 (2017).
Westin, S. N. et al. PTEN loss is a context-dependent outcome determinant in obese and non-obese endometrioid endometrial cancer patients. Mol. Oncol. 9, 1694–1703 (2015).
Berg, A. et al. Molecular profiling of endometrial carcinoma precursor, primary and metastatic lesions suggests different targets for treatment in obese compared to non-obese patients. Oncotarget 6, 1327–1339 (2015).
Mahdi, H., Schlick, C. J., Kowk, L.-L., Moslemi-Kebria, M. & Michener, C. Endometrial cancer in Asian and American Indian/Alaskan native women: tumor characteristics, treatment and outcome compared to non-Hispanic white women. Gynecol. Oncol. 132, 443–449 (2014).
Rassen, J. A. et al. One-to-many propensity score matching in cohort studies. Pharmacoepidemiol. Drug Saf. 21, 69–80 (2012).
Lin, H. et al. Stanniocalcin 1 is a phagocytosis checkpoint driving tumor immune resistance. Cancer Cell 39, 480–493 (2021).
Yan, Y., Ham, B.-K., Chong, Y. H., Yeh, S.-D. & Lucas, W. J. A plant small RNA-binding protein 1 family mediates cell-to-cell trafficking of RNAi signals. Mol. Plant 13, 321–335 (2020).
Gautam, S. K. et al. MUCIN-4 (MUC4) is a novel tumor antigen in pancreatic cancer immunotherapy. Semin. Immunol. 47, 101391 (2020).
Gao, Q. et al. Integrated proteogenomic characterization of HBV-related hepatocellular carcinoma. Cell 179, 561–577 (2019).
Machin, P. et al. Microsatellite instability and immunostaining for MSH-2 and MLH-1 in cutaneous and internal tumors from patients with the Muir–Torre syndrome. J. Cutan. Pathol. 29, 415–420 (2002).
Pugh, T. J. et al. AACR project GENIE: 100,000 cases and beyond. Cancer Discov. 12, 2044–2057 (2022).
Pearlman, R. et al. Prevalence and spectrum of germline cancer susceptibility gene mutations among patients with early-onset colorectal cancer. JAMA Oncol. 3, 464–471 (2017).
Gerhauser, C. et al. Molecular evolution of early-onset prostate cancer identifies molecular risk markers and clinical trajectories. Cancer Cell 34, 996–1011 (2018).
Rahman, N. Realizing the promise of cancer predisposition genes. Nature 505, 302–308 (2014).
Latham, A. et al. Microsatellite instability is associated with the presence of Lynch syndrome pan-cancer. J Clin. Oncol. 37, 286–295 (2019).
Srinivasan, P. et al. The context-specific role of germline pathogenicity in tumorigenesis. Nat. Genet. 53, 1577–1585 (2021).
Yehia, L., Keel, E. & Eng, C. The clinical spectrum of PTEN mutations. Annu. Rev. Med. 71, 103–116 (2020).
Tan, M.-H. et al. Lifetime cancer risks in individuals with germline PTEN mutations. Clin. Cancer Res. 18, 400–407 (2012).
Ten Broeke, S. W. et al. Lynch syndrome caused by germline PMS2 mutations: delineating the cancer risk. J. Clin. Oncol. 33, 319–325 (2015).
De Jonge, M. M. et al. Germline BRCA-associated endometrial carcinoma is a distinct clinicopathologic entity. Clin. Cancer Res. 25, 7517–7526 (2019).
Alexandrov, L. B. et al. Signatures of mutational processes in human cancer. Nature 500, 415–421 (2013).
Alexandrov, L. B. et al. The repertoire of mutational signatures in human cancer. Nature 578, 94–101 (2020).
Nik-Zainal, S. et al. Mutational processes molding the genomes of 21 breast cancers. Cell 149, 979–993 (2012).
Li, H.-D. et al. Polymerase-mediated ultramutagenesis in mice produces diverse cancers with high mutational load. J. Clin. Invest. 128, 4179–4191 (2018).
Meier, B. et al. Mutational signatures of DNA mismatch repair deficiency in C. elegans and human cancers. Genome Res. 28, 666–675 (2018).
Kucab, J. E. et al. A compendium of mutational signatures of environmental agents. Cell 177, 821–836 (2019).
Touat, M. et al. Mechanisms and therapeutic implications of hypermutation in gliomas. Nature 580, 517–523 (2020).
Crisafulli, G. et al. Temozolomide treatment alters mismatch repair and boosts mutational burden in tumor and blood of colorectal cancer patients. Cancer Discov. 12, 1656–1675 (2022).
Smith, R. B. et al. The relationship between MX [3-chloro-4-(dichloromethyl)-5-hydroxy-2(5H)-furanone], routinely monitored trihalomethanes, and other characteristics in drinking water in a long-term survey. Environ. Sci. Technol. 49, 6485–6493 (2015).
Harden, J., Jewell, A., Donaldson, F. P. & Nyman, M. C. Benzidine transformation processes in natural sediments. Environ. Toxicol. Chem. 25, 1969–1974 (2006).
Guo, K.-F. et al. Acute administration of methyleugenol impairs hippocampus-dependent contextual fear memory and increases anxiety-like behavior in mice. J. Agric. Food Chem. 68, 7490–7497 (2020).
Lee, I., Zhang, G., Mesaros, C. & Penning, T. M. Estrogen receptor dependent and independent roles of benzo[a]pyrene in Ishikawa cells. J. Endocrinol. 247, 139–151 (2020).
Woolston, A. et al. Mutational signatures impact the evolution of anti-EGFR antibody resistance in colorectal cancer. Nat. Ecol. Evol. 5, 1024–1032 (2021).
Akbani, R. et al. A pan-cancer proteomic perspective on the cancer genome atlas. Nat. Commun. 5, 3887 (2014).
Berger, A. C. et al. A comprehensive pan-cancer molecular study of gynecologic and breast cancers. Cancer Cell 33, 690–705 (2018).
Ding, L. et al. Perspective on oncogenic processes at the end of the beginning of cancer genomics. Cell 173, 305–320 (2018).
Hoadley, K. A. et al. Multiplatform analysis of 12 cancer types reveals molecular classification within and across tissues of origin. Cell 158, 929–944 (2014).
Szklarczyk, D. et al. STRING v11: protein–protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 47, D607–D613 (2019).
Senkal, C. E. et al. Ceramide is metabolized to acylceramide and stored in lipid droplets. Cell Metab. 25, 686–697 (2017).
Liberzon, A. et al. The molecular signatures database hallmark gene set collection. Cell Syst. 1, 417–425 (2015).
Garg, K. & Soslow, R. A. Endometrial carcinoma in women aged 40 years and younger. Arch. Pathol. Lab. Med. 138, 335–342 (2014).
De Heer, E. C., Jalving, M. & Harris, A. L. HIFs, angiogenesis, and metabolism: elusive enemies in breast cancer. J. Clin. Invest. 130, 5074–5087 (2020).
Rosario, S. R. et al. Pan-cancer analysis of transcriptional metabolic dysregulation using The Cancer Genome Atlas. Nat. Commun. 9, 5330 (2018).
Kim, N. H. et al. Snail reprograms glucose metabolism by repressing phosphofructokinase PFKP allowing cancer cell survival under metabolic stress. Nat. Commun. 8, 14374 (2017).
Liu, J. et al. Hypoxia induced ferritin light chain (FTL) promoted epithelia mesenchymal transition and chemoresistance of glioma. J. Exp. Clin. Cancer Res. 39, 137 (2020).
Lachance, J. A. et al. The effect of age on clinical/pathologic features, surgical morbidity, and outcome in patients with endometrial cancer. Gynecol. Oncol. 101, 470–475 (2006).
Obermair, A. et al. Fertility-sparing treatment in early endometrial cancer: current state and future strategies. Obstet. Gynecol. Sci. 63, 417–431 (2020).
Nitecki, R., Woodard, T. & Rauh-Hain, J. A. Fertility-sparing treatment for early-stage cervical, ovarian, and endometrial malignancies. Obstet. Gynecol. 136, 1157–1169 (2020).
Derbyshire, A. E., Ryan, N. & Crosbie, E. J. Biomarkers needed to predict progestin response in endometrial cancer. BJOG 124, 1584 (2017).
Hanker, A. B., Sudhan, D. R. & Arteaga, C. L. Overcoming endocrine resistance in breast cancer. Cancer Cell 37, 496–513 (2020).
Chang, M. T. et al. Identifying recurrent mutations in cancer reveals widespread lineage diversity and mutational specificity. Nat. Biotechnol. 34, 155–163 (2016).
Hoyos, D. et al. Fundamental immune-oncogenicity trade-offs define driver mutation fitness. Nature 606, 172–179 (2022).
Elez, E. et al. RNF43 mutations predict response to anti-BRAF/EGFR combinatory therapies in BRAFV600E metastatic colorectal cancer. Nat. Med. 28, 2162–2170 (2022).
Barkal, A. A. et al. CD24 signalling through macrophage Siglec-10 is a target for cancer immunotherapy. Nature 572, 392–396 (2019).
Huttlin, E. L. et al. Dual proteome-scale networks reveal cell-specific remodeling of the human interactome. Cell 184, 3022–3040 (2021).
Wu, D.-P. et al. Cx43 deficiency confers EMT-mediated tamoxifen resistance to breast cancer via c-Src/PI3K/Akt pathway. Int. J. Biol. Sci. 17, 2380–2398 (2021).
Cornel, K. M. C. et al. Overexpression of 17β-hydroxysteroid dehydrogenase type 1 increases the exposure of endometrial cancer to 17β-estradiol. J. Clin. Endocrinol. Metab. 97, E591–E601 (2012).
Konings, G. F. et al. Blocking 17β-hydroxysteroid dehydrogenase type 1 in endometrial cancer: a potential novel endocrine therapeutic approach. J. Pathol. 244, 203–214 (2018).
Wright, R. H. G. et al. ADP-ribose–derived nuclear ATP synthesis by NUDIX5 is required for chromatin remodeling. Science 352, 1221–1225 (2016).
Page, B. D. G. et al. Targeted NUDT5 inhibitors block hormone signaling in breast cancer cells. Nat. Commun. 9, 250 (2018).
Van Roosmalen, W. et al. Tumor cell migration screen identifies SRPK1 as breast cancer metastasis determinant. J. Clin. Invest. 125, 1648–1664 (2015).
Liu, P.-H. et al. Association of obesity with risk of early-onset colorectal cancer among women. JAMA Oncol. 5, 37–44 (2019).
Lynch, H. T., Watson, P., Conway, T., Fitzsimmons, M. L. & Lynch, J. Breast cancer family history as a risk factor for early onset breast cancer. Breast Cancer Res. Treat. 11, 263–267 (1988).
Safdar, N. S. et al. Genomic determinants of early recurrences in low-stage, low-grade endometrioid endometrial carcinoma. J. Natl Cancer Inst. 114, 1545–1548 (2022).
Ugai, T. et al. Is early-onset cancer an emerging global epidemic? Current evidence and future implications. Nat. Rev. Clin. Oncol. 19, 656–673 (2022).
Esposito, G. et al. Diabetes risk reduction diet and endometrial cancer risk. Nutrients 13, 2630 (2021).
Friedenreich, C. M., Ryder-Burbidge, C. & McNeil, J. Physical activity, obesity and sedentary behavior in cancer etiology: epidemiologic evidence and biologic mechanisms. Mol. Oncol. 15, 790–800 (2021).
Westin, S. N. et al. Prospective phase II trial of levonorgestrel intrauterine device: nonsurgical approach for complex atypical hyperplasia and early-stage endometrial cancer. Am. J. Obstet. Gynecol. 224, 1–15 (2021).
Chatila, W. K. Genomic and transcriptomic determinants of response to neoadjuvant therapy in rectal cancer. Nat. Med. 28, 26 (2022).
Xu, Y. et al. Endometrium-derived mesenchymal stem cells suppress progression of endometrial cancer via the DKK1-Wnt/β-catenin signaling pathway. Stem Cell Res. Ther. 14, 159 (2023).
tjmu-hz. Haz1y/EEEC_landscape: update-20240302-NatGen. Zenodo https://doi.org/10.5281/zenodo.10756345 (2024).
Xiao, W. et al. Toward best practice in cancer mutation detection with whole-genome and whole-exome sequencing. Nat. Biotechnol. 39, 1141–1150 (2021).
Acknowledgements
C.S. and G.C. acknowledge grants from the National Key R&D Program of China (2022YFC2704300 and 2022YFC2704301). X.C. acknowledges grants from the National Key R&D Program of China (2022YFC2704300 and 2022YFC2704305) and the Shanghai Technical Innovation Program (20Z1190070). The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
C.S. and Z. Hu conceived the study idea. J.L. and W. Liu collected samples. Z. Hu conducted the multi-omic profiling, with the help of J.F., C.F. and J.P. Y.N., Q.W. and C.W. interpreted the pathological slides. J.L. and Z. Wu performed experimental validations, with the help of W. Li, X.Y., Q.L., S.C., W. Liu, Y.D., W.W., F.L., X.K., Z.X., X.Z. and T.Q. C.S. and Z. Hu wrote the manuscript with input from all the authors. G.B.M., D.M., Z.W., J.W., J.J. and X.W. provided expertise and feedback. C.S., X.C., G.C., K.L., Q.G. and W.D. supervised the project and provided valuable critical discussion. All authors reviewed and approved the manuscript before publication.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Genetics thanks Russell Broaddus and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 The workflow of the TJFD-EC proteogenomic study.
The TJFD-EC study of 229 patients, including 14 AEHs and 215 EECs, was used for WES and proteomics analysis.
Extended Data Fig. 2 Quality assessments for WES and MS data.
(a) The body mass index (BMI) between EEECs and LEECs (all FIGO stages were included). The p value was calculated by two-sided Mann–Whitney U test. Data are represented as boxplots with the middle line indicating the median, while the upper and lower hinges indicate the 25% and 75% quartiles, respectively. (b, c) The WES quality control results for mapping quality, insert size and coverage. Notes: all of these results fall within the prescribed guidelines of ‘Toward best practice in cancer mutation detection with whole-genome and whole-exome sequencing’, published in Nature Biotechnology82. (d) The distribution of GIV scores in our WES samples. Notes: according to the thresholds recommended in the literature in Nature Biotechnology82, samples with a GIV score exceeding 1.5 are deemed to exhibit severe DNA damage. Specifically, only 4.2% of our samples surpass the GIV score threshold of 1.5. (e) The relative proportion of [C>T] transition in TCGA, CPTAC and TJFD datasets. Notes: we did not observe a significant enrichment of C>T mutations (FFPE-induced errors often exhibit a bias toward C>T transitions) in our dataset but even slightly lower trend of C>T mutations compared to TCGA study (p = 0.01). The p value was calculated by two-sided Mann–Whitney U test. Data are represented as boxplots with the middle line indicating the median, while the upper and lower hinges indicate the 25% and 75% quartiles, respectively. (f) Longitudinal quality control of mass spectrometry using tryptic digest of HeLa cells every three days. Correlation matrix of all measured HeLa samples based on Pearson’s correlation. (g) Histograms showing the distribution of peptides per protein (top panel) and the mass of proteins (bottom panel). (h) Principal component analysis (PCA) of proteomic data. Red triangles: tumors; blue dots: normal adjacent tissues. (i) The Venn diagram comparing the detected proteins in the CPTAC-EC study and our own TJFD study. (j) The pathway enrichment analysis on the 4189 unique proteins identified in the CPTAC dataset. Notes: only the TNF-α pathway in the hallmark genesets displayed significant enrichment (q < 0.1). This suggests that our comparative proteomic analysis with FFPE accurately reflects the biological characteristics of the samples, except for proteins involved in the activity of the TNF-α pathway.
Extended Data Fig. 3 Genomic characterization of TJFD and comparison with TCGA dataset.
(a) To eliminate the selection bias of patients in TJFD and TCGA databases caused by baseline demographic and clinical factors, propensity score matching was used to balance the two datasets. Baseline clinicopathological characteristics including FIGO stage, grade and TCGA subtype were fit into a multivariate logistic model, and the nearest neighbor algorithm and one-to-one match were set in the logistic model. R package ‘MatchIt’ was used for the calculation algorithm. (b, c) Clinical characteristics (b) and mutation profiles (c) between two datasets after propensity score matching (PSM). (d) Forest plot of differentially mutated genes between the two datasets before (left) and after PSM (right). Data are represented as a forest plot of ORs with 95% confidence intervals represented as circles and error bars. OR: odds ratio. The p values were calculated by two-sided chi-square test or Fisher’s exact test as appropriate.
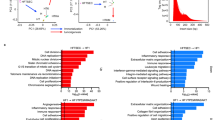
Extended Data Fig. 4 Molecular characterization of early-onset EEC in the whole TJFD dataset.
To further investigate and validate the molecular characterization of early-onset EEC, the whole TJFD dataset (of all FIGO stages) was included. (a–d) Comparison of mutation landscape (a), tumor mutation burden (b), proportion of TCGA subtype (c) and frequently mutated genes (d) between age groups of the whole dataset. The p values were calculated by two-sided Mann–Whitney U test and two-sided Fisher’s exact test. For b, data are represented as boxplots with the middle line indicating the median, while the upper and lower hinges indicate the 25% and 75% quartiles, respectively. (e) To further depict the distribution difference of CTNNB1 mutation of early-onset EEC, the AACR GENIE dataset21 of 2401 endometrial carcinomas was included in analysis. The CTNNB1 mutations in late-onset tumors were distributed across the gene, while the early-onset tumors showed remarkable enrichment in the region of exon 3. (f) Correlation between the probability of specific somatic mutation and age-at-diagnosis in TJFD dataset. (g) The correlation in TCGA and CPTAC datasets was also performed for further validation. For f and g, logistics regression analysis was performed, and black dots represent the distribution of mutation status of specific genes in each patient alongside their respective ages.
Extended Data Fig. 5 Mutation signature associated with endogenous and environmental mutagens.
To reveal the association of de novo mutational signatures with the endogenous and exogenous mutagens, cosine similarity analysis against mutation signatures defined in COSMIC31, 32 and environmental agents36 was performed. (a) Comparison of de novo mutational signatures identified in this study with previously published COSMIC31 and SBS32 mutational signatures. (b) The mutational sig6 showed high similarity to COSMIC signatures about tobacco, aflatoxin, ROS, aromatic amines, ROS-like methyl eugenol and HR/NER deficiency. Moreover, de novo Sig6 was highly associated with environmental mutagens. (c) Comparison of the relative proportion of de novo mutation signatures between early-onset and late-onset EECs. The p values were calculated by two-sided t-test. Data are represented as boxplots with the middle line indicating the median, while the upper and lower hinges indicate the 25% and 75% quartiles, respectively. (d) Spearman’s correlation between the exposome-related mutation sig6 burden and age-at-diagnosis, and the green shadow represents 95% confidence interval. The p value from Spearman correlation with permutation test.
Extended Data Fig. 6 3D-Sig-Explorer uncovers exposure-molecular-phenotype links.
(a) Schematic overview of the 3D-Sig-Explorer model. The principle and computation details are described in Methods. (b) The mutational signatures contribution to LEEC-related genes.
Extended Data Fig. 7 The impact of SIGLEC10 on hormone treatment resistance in vitro and in vivo.
(a) Sanger sequencing validation of p.Q144K (or named as c.430C>A) mutation of SIGLEC10. (b) BioPlex database reveals protein interaction of SIGLEC10 and ESR1. (c) Validation of Siglec-10 overexpressing in cell line by western blot. (d) Cell proliferation of SIGLEC10Q144K, SIGLEC10wt, negative control (NC) and parental cells (CON), which were treated with progesterone (PRG, 25 μM) for 72 hours. The p values were calculated by the ANOVA test followed by Tukey’s honestly significant difference (HSD) post hoc test. (e, f) Representative images and quantification (n = 3 per group) of 3D tumor spheroids formation in negative control, SIGLEC10wt and SIGLEC10Q144K overexpressing RL95-2 and MCF-7 cells after treatment with PRG (20 μM) and medroxyprogesterone acetate (MPA, 10 μM) for 7 days. Scale bar, 20 μm. The p values were calculated by the ANOVA test followed by Tukey’s honestly significant difference (HSD) post hoc test. (g) Validation of Siglec-10 protein overexpressing in PDO model by western blot. (h) Multiplex immunofluorescence (mIF) images of Siglec-10, ER and PR in negative control, SIGLEC10wt and SIGLEC10Q144K overexpressing PDO models. Scale bar, 100 μm. (i) Tumor growth curves of subcutaneous EC models derived from negative control (top) and SIGLEC10Q144K overexpressing Ishikawa (bottom) after treatment with MPA (300 mg/kg/d, i.g.) for 21 days (n = 5 per group). The p values were calculated by two-sided t-tests. (j) Kaplan–Meier curves according to SRPK1 expression in GSE12093 and GSE17705 were shown. The cutoff for high and low level of SRPK1 was 1201.3 and 10.26426, respectively. For d and f, data represent the mean ± SD from three independent replicates. For i, data represent the mean ± SEM from five independent replicates. i.g.: oral gavage.
Supplementary information
Supplementary Information
Supplementary Methods.
Supplementary Tables
Supplementary Tables 1–5.
Source data
Source Data Fig. 7 and Extended Data Fig. 7
Unprocessed western blots.
Source Data Figs. 7 and 8
Statistical source data of the IHC quantification.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Hu, Z., Wu, Z., Liu, W. et al. Proteogenomic insights into early-onset endometrioid endometrial carcinoma: predictors for fertility-sparing therapy response. Nat Genet 56, 637–651 (2024). https://doi.org/10.1038/s41588-024-01703-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-024-01703-z