Abstract
Rare coding mutations cause ∼45% of congenital heart disease (CHD). Noncoding mutations that perturb cis-regulatory elements (CREs) likely contribute to the remaining cases, but their identification has been problematic. Using a lentiviral massively parallel reporter assay (lentiMPRA) in human induced pluripotent stem cell-derived cardiomyocytes (iPSC-CMs), we functionally evaluated 6,590 noncoding de novo variants (ncDNVs) prioritized from the whole-genome sequencing of 750 CHD trios. A total of 403 ncDNVs substantially affected cardiac CRE activity. A majority increased enhancer activity, often at regions with undetectable reference sequence activity. Of ten DNVs tested by introduction into their native genomic context, four altered the expression of neighboring genes and iPSC-CM transcriptional state. To prioritize future DNVs for functional testing, we used the MPRA data to develop a regression model, EpiCard. Analysis of an independent CHD cohort by EpiCard found enrichment of DNVs. Together, we developed a scalable system to measure the effect of ncDNVs on CRE activity and deployed it to systematically assess the contribution of ncDNVs to CHD.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
RNA-seq and MPRA next-generation sequencing data associated with this study have been deposited to Gene Expression Omnibus (GSE208283 and GSE210376). WGS data were reported previously6,7 and are available through dbGaP (phs001138.v4.p2, phs001194.v3.p2 and phs001735.v2.p1). Source data are provided with this paper.
Code availability
Custom code used in this study can be downloaded from Zenodo64 or GitHub:
(1) EpiCard https://github.com/pulab/CHD_DNVs;
(2) MPRA library design: https://github.com/pulab/CHD_DNVs/tree/main/MPRA-Enhancer/MPRA_library_designer-main
and (3) MPRA analysis: https://github.com/pulab/CHD_DNVs/tree/main/MPRA-Enhancer/CHD_MPRA_project
References
Van der Linde, D. et al. Birth prevalence of congenital heart disease worldwide: a systematic review and meta-analysis. J. Am. Coll. Cardiol. 58, 2241–2247 (2011).
Zaidi, S. et al. De novo mutations in histone-modifying genes in congenital heart disease. Nature 498, 220–223 (2013).
Homsy, J. et al. De novo mutations in congenital heart disease with neurodevelopmental and other congenital anomalies. Science 350, 1262–1266 (2015).
Jin, S. C. et al. Contribution of rare inherited and de novo variants in 2,871 congenital heart disease probands. Nat. Genet. 49, 1593–1601 (2017).
ENCODE Project Consortium An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Richter, F. et al. Genomic analyses implicate noncoding de novo variants in congenital heart disease. Nat. Genet. 52, 769–777 (2020).
Morton, S. U. et al. Genome-wide de novo variants in congenital heart disease are not associated with maternal diabetes or obesity. Circ. Genom. Precis. Med. 15, e003500 (2022).
Blow, M. J. et al. ChIP–seq identification of weakly conserved heart enhancers. Nat. Genet. 42, 806–810 (2010).
Huang, Y.-F., Gulko, B. & Siepel, A. Fast, scalable prediction of deleterious noncoding variants from functional and population genomic data. Nat. Genet. 49, 618–624 (2017).
Ernst, J. & Kellis, M. Chromatin-state discovery and genome annotation with ChromHMM. Nat. Protoc. 12, 2478–2492 (2017).
Hoffman, M. M., Buske, O. J., Wang, J., Weng, Z. & Bilmes, J. A. Unsupervised pattern discovery in human chromatin structure through genomic segmentation. Nat. Methods 9, 473–476 (2012).
Cooper, G. M. et al. Distribution and intensity of constraint in mammalian genomic sequence. Genome Res. 15, 901–913 (2005).
Inoue, F. & Ahituv, N. Decoding enhancers using massively parallel reporter assays. Genomics 106, 159–164 (2015).
Inoue, F. et al. A systematic comparison reveals substantial differences in chromosomal versus episomal encoding of enhancer activity. Genome Res. 27, 38–52 (2017).
Lian, X. et al. Directed cardiomyocyte differentiation from human pluripotent stem cells by modulating Wnt/β-catenin signaling under fully defined conditions. Nat. Protoc. 8, 162–175 (2013).
Barakat, T. S. et al. Functional dissection of the enhancer repertoire in human embryonic stem cells. Cell Stem Cell 23, 276–288 (2018).
Visel, A., Minovitsky, S., Dubchak, I. & Pennacchio, L. A. VISTA enhancer browser—a database of tissue-specific human enhancers. Nucleic Acids Res. 35, D88–D92 (2007).
Arnold, C. D. et al. Genome-wide quantitative enhancer activity maps identified by STARR-seq. Science 339, 1074–1077 (2013).
Tewhey, R. et al. Direct identification of hundreds of expression-modulating variants using a multiplexed reporter assay. Cell 165, 1519–1529 (2016).
Klein, J. C. et al. A systematic evaluation of the design and context dependencies of massively parallel reporter assays. Nat. Methods 17, 1083–1091 (2020).
Li, K. et al. Interrogation of enhancer function by enhancer-targeting CRISPR epigenetic editing. Nat. Commun. 11, 485 (2020).
Hilton, E. N. et al. Left-sided embryonic expression of the BCL-6 corepressor, BCOR, is required for vertebrate laterality determination. Hum. Mol. Genet. 16, 1773–1782 (2007).
Hamline, M. Y. et al. OFCD syndrome and extraembryonic defects are revealed by conditional mutation of the polycomb-group repressive complex 1.1 (PRC1.1) gene BCOR. Dev. Biol. 468, 110–132 (2020).
Wang, D. et al. Activation of cardiac gene expression by myocardin, a transcriptional cofactor for serum response factor. Cell 105, 851–862 (2001).
Huang, J. et al. Myocardin regulates BMP10 expression and is required for heart development. J. Clin. Invest. 122, 3678–3691 (2012).
Houweling, A. C. et al. Loss-of-function variants in myocardin cause congenital megabladder in humans and mice. J. Clin. Invest. 129, 5374–5380 (2019).
Santamaria, S. & de Groot, R. ADAMTS proteases in cardiovascular physiology and disease. Open Biol. 10, 200333 (2020).
Prins, B. P. et al. Exome-chip meta-analysis identifies novel loci associated with cardiac conduction, including ADAMTS6. Genome Biol. 19, 87 (2018).
Tian, E. et al. Galnt1 is required for normal heart valve development and cardiac function. PLoS ONE 10, e0115861 (2015).
Dykes, I. M. et al. HIC2 is a novel dosage-dependent regulator of cardiac development located within the distal 22q11 deletion syndrome region. Circ. Res. 115, 23–31 (2014).
Zhang, Q. et al. Multiplexed single-nucleus RNA sequencing using lipid-oligo barcodes. Curr. Protoc. 2, e579 (2022).
Wang, Z. et al. A non-canonical BCOR-PRC1.1 complex represses differentiation programs in human ESCs. Cell Stem Cell 22, 235–251 (2018).
Montefiori, L. E. et al. A promoter interaction map for cardiovascular disease genetics. eLife 7, e35788 (2018).
Avsec, Ž. et al. Effective gene expression prediction from sequence by integrating long-range interactions. Nat. Methods 18, 1196–1203 (2021).
Zhou, J. & Troyanskaya, O. G. Predicting effects of noncoding variants with deep learning-based sequence model. Nat. Methods 12, 931–934 (2015).
Shihab, H. A. et al. An integrative approach to predicting the functional effects of non-coding and coding sequence variation. Bioinformatics 31, 1536–1543 (2015).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Grant, C. E., Bailey, T. L. & Noble, W. S. FIMO: scanning for occurrences of a given motif. Bioinformatics 27, 1017–1018 (2011).
Hamad, S. et al. Generation of human induced pluripotent stem cell-derived cardiomyocytes in 2D monolayer and scalable 3D suspension bioreactor cultures with reduced batch-to-batch variations. Theranostics 9, 7222–7238 (2019).
Tohyama, S. et al. Distinct metabolic flow enables large-scale purification of mouse and human pluripotent stem cell-derived cardiomyocytes. Cell Stem Cell 12, 127–137 (2013).
Yu, G., Wang, L.-G. & He, Q.-Y. ChIPseeker: an R/Bioconductor package for ChIP peak annotation, comparison and visualization. Bioinformatics 31, 2382–2383 (2015).
Hoang, T. T. et al. The Congenital Heart Disease Genetic Network Study: cohort description. PLoS ONE 13, e0191319 (2018).
Dickel, D. E. et al. Genome-wide compendium and functional assessment of in vivo heart enhancers. Nat. Commun. 7, 12923 (2016).
McLean, C. Y. et al. GREAT improves functional interpretation of cis-regulatory regions. Nat. Biotechnol. 28, 495–501 (2010).
Labun, K. et al. CHOPCHOP v3: expanding the CRISPR web toolbox beyond genome editing. Nucleic Acids Res. 47, W171–W174 (2019).
Mandegar, M. A. et al. CRISPR interference efficiently induces specific and reversible gene silencing in human iPSCs. Cell Stem Cell 18, 541–553 (2016).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 17, 10–12 (2011).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Vierstra, J. et al. Global reference mapping of human transcription factor footprints. Nature 583, 729–736 (2020).
Dubitzky, W., Wolkenhauer, O., Cho, K.-H. & Yokota, H. (eds.). Encyclopedia of Systems Biology, pp. 78 (Springer, 2013).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Anders, S., Pyl, P. T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
O’Leary, N. A. et al. Reference sequence (RefSeq) database at NCBI: current status, taxonomic expansion, and functional annotation. Nucleic Acids Res. 44, D733–D745 (2016).
Nadelmann, E. R. et al. Isolation of nuclei from mammalian cells and tissues for single-nucleus molecular profiling. Curr. Protoc. 1, e132 (2021).
Wolock, S. L., Lopez, R. & Klein, A. M. Scrublet: computational identification of cell doublets in single-cell transcriptomic data. Cell Syst. 8, 281–291 (2019).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587 (2021).
Phipson, B. et al. propeller: testing for differences in cell type proportions in single cell data. Bioinformatics 38, 4720–4726 (2022).
Yu, G., Wang, L.-G., Han, Y. & He, Q.-Y. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16, 284–287 (2012).
Alipanahi, B., Delong, A., Weirauch, M. T. & Frey, B. J. Predicting the sequence specificities of DNA- and RNA-binding proteins by deep learning. Nat. Biotechnol. 33, 831–838 (2015).
Kent, W. J. et al. The human genome browser at UCSC. Genome Res. 12, 996–1006 (2002).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Friedman, J., Hastie, T. & Tibshirani, R. Regularization paths for generalized linear models via coordinate descent. J. Stat. Softw. 33, 1–22 (2010).
Zhang, X., Morton, S. U., Seidman, J. G., Seidman, C. S. & Pu, W. T. Analysis code used to analyze ncDNVs in CHD. Zenodo https://zenodo.org/records/10294614 (2024).
Acknowledgements
We thank all patients and families who participated in this research. F.X. was supported by the AHA (20POST35200226). S.U.M. and J.G.S. were supported by NIH (R03 HL150412-01A1); S.U.M. was supported by NIH (1K08HL157653-01A1), an AHA Career Development Award, and the Boston Children’s Hospital Office of Faculty Development. W.T.P., C.E.S. and J.G.S. were supported by NIH (2U01HL098147 and U01 HL098166). C.E.S. and J.G.S. were supported by the Engineering Research Centers Program of the National Science Foundation (NSF Cooperative Agreement EEC-1647837). C.E.S. was supported by the Howard Hughes Medical Institute. The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
F.X., X.Z. and S.M. contributed equally to this work. F.X., W.T.P., J.G.S. and C.E.S. conceived and designed the study. F.X. performed the experiments and analyzed data. X.Z. and S.M. conducted bioinformatic analyses. X.Z. developed custom MPRA design and analysis software. S.W.K. and J.M.G. performed multiplexed snRNA-seq and associated analyses. F.X. and H.Z. performed EMSA and analyzed the data. Y.F., Y.C., N.M., P.B., J.C., X.L. P.Z. and T.W. generated plasmids, viruses and other necessary reagents and assisted with processing cells. S.M., J.H., F.R., Y.S. and B.G. analyzed WGS and annotated ncDNVs. F.X. and W.T.P. wrote the manuscript with contributions from the other authors. All authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Genetics thanks Stephanie Ware and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Establishment of the lentiMPRA platform to test cardiac enhancer activity in iPSC-CMs.
a. Strategy for pilot experiment to test lentiviral reporter assay in iPSC-CMs. b. Flow cytometry analysis of cTNT+ iPSC-CMs at differentiation day 12. Cells were gated with SSC and FSC to exclude debris and doublets. Flow cytometry plots displayed a biomodal distribution between fluorescent and non-fluorescent cells. Gates determining the percent of fluorescent cells were drawn at the local minimum between these distributions. c. Activities of PSC-specific enhancer (OCT4 PE) and cardiac enhancers (VISTA enhancer browser hs2330 and hs1670) in iPSCs and iPSC-CMs. Representative images from 4 independent experiments. Scale bar, 100 μm. d. Strategy for pilot experiment to measure enhancer activity by Amplicon-seq. e. Enhancer activities of PSC enhancers (Enh1–4) and cardiac enhancers (Enh 5–19). Activity of the empty vector (EV) was set 1. Enhancer activity was normalized to EV. Data are represented as mean ± SEM of 4 independent experiments (2-sided unpaired t test).
Extended Data Fig. 2 Assessment of human cardiac enhancer activity with hiPSC-CMs and lentiSTARR-seq.
a. Minimal read coverage of designed regions in DNA replicates. Red line shows minimum coverage for inclusion in analysis (FPM ≥ 20). b. Pearson correlation of MPRA activity between biological replicates at D17 and D24. There was excellent correlation both within group and across time points. c. Summary of MPRA results. Plot at the bottom shows a vertical line for each tested region with the indicated annotation. Enrichment score indicates enrichment of a set of regions of interest toward the ends of the ranked list of all regions. Enrichment p-value was determined by 1-sided permutation test (see Methods) with Bonferroni correction. Active enhancers were those enriched in RNA compared to DNA (DESeq2 Padj < 0.05). d. Violin plot with the log2(RNA/DNA) results of all candidates, active candidates, inactive candidates and negative controls. Kruskal-Wallis test p-values vs. neg control are shown. Center, box and whiskers indicate median, 25th and 75th percentiles and value closest to 25th percentile minus or 75th percentile plus 1.5 times the interquartile range. e. Twenty-four candidate cardiac enhancers of known cardiovascular disease genes with a range of MPRA enhancer activity were individually cloned into the lentiMPRA vector, in which a minimal promoter drives GFP expression. Red color indicates enhancers that were classified as active by MPRA. GFP expression was evaluated by epifluorescent imaging. Representative images from 4 independent experiments. Scale bar, 100 µm.
Extended Data Fig. 3 Functional dissection of active cardiac enhancers by tiling deletion mutagenesis.
a. Coverage of designed regions. Red line shows minimum coverage for inclusion in analysis (FPM ≥ 20). 97.6% of regions had coverage ≥20 FPM. b. Summary of activity of regions in the mutagenesis MPRA. Line plot at the bottom shows a vertical line for each tested region with the indicated annotation. Enrichment score indicates how the indicated annotations are distributed across the regions, ranked by activity. Enrichment p-value with Bonferroni correction was calculated using a 1-sided permutation test (see Methods). Active enhancers had barcodes that were overrepresented in RNA compared to DNA (DESeq Padj < 0.05). c. Validation of effects of mutations on transcription factor binding. Transcription factor binding was evaluated by electrophoretic mobility shift assay. The indicated wild-type and mutant oligonucleotide pairs were incubated with transcription factors with predicted altered motifs and analyzed by gel electrophoresis. Results are representative of at least 2 independent experiments.
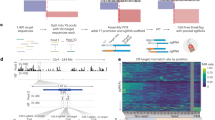
Extended Data Fig. 4 CHD MPRA library characterization.
a. The CHD MPRA library included 6590 REF-ALT pairs. After pooled library synthesis of barcoded oligos, the oligos were PCR amplified and cloned into lentivirus genome backbone. A minimal promoter (miniP)-GFP cassette was then inserted into the cloned oligo library. b. Summary of activity of CHD MPRA library. Plot on bottom indicates the occurrence of the indicated annotation with a vertical line. Enrichment score represents enrichment of the indicated set of annotations at either end of the list of all regions, ranked by activity. Enrichment p-value was determined by 1-sided permutation test, with Bonferroni correction. Active enhancers had barcodes overrepresented in RNA compared to DNA (DESeq2 Padj < 0.05). c. Pearson correlation (PCC) between regions shared between the Mutagenesis MPRA and the CHD MPRA. The same genomic sequences had different barcodes in the two assays. d. Validation of the effect of variants on transcription factor binding. EMSA assay was used to test the binding of SRF or TBX20 to REF or ALT variant sequences. For the GLB1L3 CRE, ALT disrupted the SRF motif and reduced SRF binding in the EMSA assay. For the PIP4K2A CRE, ALT generated a TBX20 motif and increased TBX20 binding in the EMSA assay. Representative of three independent experiments. Two-tailed t-test. n = 3 per group. Graph shows mean ± SD.
Extended Data Fig. 5 Genomic loci of CHD-associated ncDNVs.
a–d. WashU Epigenome Browser views of loci containing 4 ncDNVs. Promoter capture Hi-C and RNA-seq in iPSCs and iPSC-CMs from ref. 33, PMID 29988018. Genes dysregulated by DNVs are indicated in red. Green lines highlight 171 bp REF region with DNV in the center. e–h. Sanger sequencing traces of genome edited iPSC lines.
Extended Data Fig. 6 Characterization of iPSC-CMs with knockin of CHD gene-associated noncoding DNVs.
a. BCOR downregulation in SMAD2 Het and KO iPSC-CMs. Gene expression was measured by RNA-seq. One-way ANOVA with Dunnett’s multiple comparison test versus WT. n = 3. b. Effect of ncDNVs on binding of transcription factors to CREs near CHD genes. 39 bp duplexes centered on ncDNVs neighboring 4 CHD genes were synthesized. Binding of purified, recombinant proteins to the REF or ALT sequence was measured by electrophoretic mobility shift assay (EMSA). SMAD2 and HIC2 bound CREs near BCOR and ACVRL1 more strongly for REF compared to ALT. In contrast, SRF and TBX20 bound CREs near ADAMTS6 and MYOCD more strongly for ALT compared to REF. Note lower free probe in MYOCD-ALT compared to REF. Results are representative of at least three independent experiments. Quantification of TBX20 EMSA: mean ± SD; n = 3; two-sided t-test. Graphs in a and b show mean ± SD.
Extended Data Fig. 7 snRNA-seq characterization of the impact of four ncDNVs that impact MPRA activity on iPSC differentiation to iPSC-CMs.
a. Expression of cardiac marker genes. Most nuclei contained cardiomyocyte marker genes. b. Two independent iPSC clones per ncDNV (ACVRL1, ADAMTS6, MYOCD) or knockin pools (BCOR) were separately differentiated into iPSC-CMs and then analyzed by multiplexed snRNA-seq. After clustering, UMAP plots of individual cells are shown separately for each independent differentiation. c–e. Pseudo-bulk differential gene expression analysis. The number of differentially expressed genes for each independent replicate vs. wild type was analyzed from snRNA-seq data. Differentially expressed genes for the two replicates showed excellent overlap (c). Gene ontology terms enriched in differentially expressed genes shared between biological replicates for ACVRL1 ncDNV KI lines (d) or ADAMTS6 ncDNV KI lines (e). BH-corrected hypergeometric p-values. f. CHD genes differentially expressed in iPSC-CMs containing indicated ncDNV knockins compared to wild-type (WT). The selected CHD genes were mouse or human CHD genes (see Supplementary Data 5) that overlapped with genes differentially expressed in both replicates of any of the four introduced ncDNVs. BH-corrected P values were reported by Seurat FindMarkers function. g. Comparison of genes upregulated in BCOR ncDNV KI pool iPSC-CMs compared to BCOR genome occupancy in H1 hESCs (GSE104690). One-sided permutation test (10000 permutations).
Extended Data Fig. 8 snRNA-seq characterization of the impact of five ncDNVs that did not alter MPRA activity in iPSC-CMs.
Five ncDNVs that did not affect MPRA activity (MPRA-NC) and were knocked into WTC-11 iPSCs. a,b. Two independent knockin clones of ARMC4, DDX11, DTNA or PDE2A ncDNV, a SOX9 ncDNV knockin clone, a BCOR ncDNV knockin pool (positive control) and WTC-11 (two independent replicates) were differentiated into iPSC-CMs. On day 10, nuclei were analyzed by multiplexed snRNA-seq. Clustering identified 4 cell states (a) that express iPSC-CM markers (b). c. The distribution of iPSC-CMs among the 4 cell states was reproducible in biological replicate samples. d. Analysis of iPSC-CM state distribution by genotype. BCOR significantly expanded cluster 1 compared to WT (ANOVA with Dunnett’s test versus WT for each iPSC-CM state). The ncDNVs that did not affect MPRA activity had no significant effect on iPSC-CM state distribution.
Extended Data Fig. 9 Characterization of EpiCard scores.
a. Comparison of EpiCard, HeartENN and Enformer scores by MPRA region activity. Two-sided t-test. b. Correlation between EpiCard, HeartENN and Enformer scores expressed as Pearson coefficient (p-value) across 3745 ncDNVs with scores available. c,d. Comparison of functional scores for ncDNVs in an independent CHD cohort and non-CHD cohort, compared by 2-sided t-test with nominal p-values reported. c. All ncDNVs meeting prioritization criteria (see Fig. 3a). Right, subset of prioritized ncDNVs near HHE genes. ncDNVs (n = 6211 CHD and 10224 non-CHD). d. Subset of ncDNVs near HHE genes (n = 3120 CHD and 5195 non-CHD). DNVs. Center, box and whiskers indicate median, 25th and 75th percentiles and value closest to 25th percentile minus or 75th percentile plus 1.5 times the interquartile range.
Extended Data Fig. 10 Schematic of enrichment score calculation.
Given a ranked list L and a specific group of regions R that is a subset of L, the enrichment score at position i (ESi) is the difference between the cumulative probability of membership in R compared to L.
Supplementary information
Supplementary Tables
Supplementary Table 1: Candidate regions used to establish the lentiMPRA platform. VISTA regions are active cardiac enhancers from the VISTA enhancer browser (https://enhancer.lbl.gov/). Supplementary Table 2: Twenty-four candidate cardiac enhancers with a range of activities in iPSC-CM MPRA were validated individually using GFP reporter assays. FACS FC, GFP mean fluorescence intensity FC compared to EV. Supplementary Table 3: DNVs detected by WGS of CHD trios. Supplementary Table 4: Summary of ten CHD ncDNVs that altered MPRA activity tested by knockin at the endogenous locus. Knockin of four ncDNVs significantly affected the expression of neighboring CHD-associated genes. ns, not significant; -, not tested. Supplementary Table 5: Oligonucleotides used in this study. The HDR templates were the same sequence as the 171 nt ALT sequence used in the MPRA. ALT variant is indicated in red. sgRNAs were selected to overlap the ALT variant. Supplementary Table 6: Correlation of MPRA activity with human fetal heart chromatin features. Supplementary Table 7: List of inputs used for the LASSO model. Files to load into the LASSO model can be obtained at https://github.com/pulab/CHD_DNVs
Supplementary Data 1
Cardiac enhancer MPRA design and results.
Supplementary Data 2
Cardiac enhancer mutagenesis.
Supplementary Data 3
CHD DNV MPRA.
Supplementary Data 4
Epicard scores.
Supplementary Data 5
Prioritized CHD genes.
Supplementary Data 6
Nonredundant motif database and expressed transcription factors in iPSC-CMs.
Supplementary Data 7
ncDNVs in CHD and non-CHD probands of the validation cohort.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data and unprocessed EMSA gels.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data and unprocessed EMSA gels.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 9
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Xiao, F., Zhang, X., Morton, S.U. et al. Functional dissection of human cardiac enhancers and noncoding de novo variants in congenital heart disease. Nat Genet 56, 420–430 (2024). https://doi.org/10.1038/s41588-024-01669-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-024-01669-y