Abstract
Insects constitute the most species-rich radiation of metazoa, a success that is due to the evolution of active flight. Unlike pterosaurs, birds and bats, the wings of insects did not evolve from legs1, but are novel structures that are attached to the body via a biomechanically complex hinge that transforms tiny, high-frequency oscillations of specialized power muscles into the sweeping back-and-forth motion of the wings2. The hinge consists of a system of tiny, hardened structures called sclerites that are interconnected to one another via flexible joints and regulated by the activity of specialized control muscles. Here we imaged the activity of these muscles in a fly using a genetically encoded calcium indicator, while simultaneously tracking the three-dimensional motion of the wings with high-speed cameras. Using machine learning, we created a convolutional neural network3 that accurately predicts wing motion from the activity of the steering muscles, and an encoder–decoder4 that predicts the role of the individual sclerites on wing motion. By replaying patterns of wing motion on a dynamically scaled robotic fly, we quantified the effects of steering muscle activity on aerodynamic forces. A physics-based simulation incorporating our hinge model generates flight manoeuvres that are remarkably similar to those of free-flying flies. This integrative, multi-disciplinary approach reveals the mechanical control logic of the insect wing hinge, arguably among the most sophisticated and evolutionarily important skeletal structures in the natural world.
Similar content being viewed by others
Main
Whether to forage, migrate, reproduce or avoid predators, aerial manoeuvrability is essential for flying insects. Most insects, including members of the species-rich orders Coleoptera (beetles), Hymenoptera (ants, bees and wasps) and Diptera (flies), actuate their wings using two morphologically and functionally distinct sets of muscles. Contractions of the large, indirect flight muscles (IFMs) are activated mechanically by stretch rather than by individual motor neuron spikes—a specialization that permits the production of elevated power at high wingbeat frequency5,6,7. There are two sets of IFMs: dorso-ventral muscles and the dorso-longitudinal muscles, arranged orthogonally to maintain a self-sustaining oscillation (Fig. 1a–c). The tiny deformations in the exoskeleton generated by the IFMs are mechanically amplified by the wing hinge8,9,10,11,12, an intricate joint containing an interconnected set of small, hard structures called sclerites embedded within more flexible exoskeleton and regulated by tiny control muscles13 (Fig. 1a,b). Here we used machine learning and physics-based simulations to gain insight into the underlying mechanics of the wing hinge of flies and its active regulation during flight.
Schematic drawings are based on a synthesis of prior work2,7,8,9,10,11,12,13,14,15,16,17,18,19 and our own morphological analysis (Supplementary Video 1). a, Contraction of the dorso-longitudinal muscles (DLMs) causes the scutellum (SCL) to move downward. PS, parascutal shelf; PWP, pleural wing process. b, Contraction of the dorso-ventral muscles (DVMs) cause the scutellum to move upward. These motions are transferred to the wing hinge via the parascutal shelf and SLA, with the wing’s fulcrum located at the pleural wing process. c, Side view of a flying fly indicating orientation in a and b. d, Schematic top view of the thorax, hinge and wing, illustrating wing veins (L1–L5) and scutum (SC). Dotted lines represent hypothesized flexure lines on the wing. e, Close-up of the wing hinge highlighting the parascutal shelf, SLA, ANP, PMNP, ax1, ax2, ax3, ax4, costal vein (CO), radial vein (RV), anal vein (AV) and the canoe. f, Layout of the steering muscles and sclerites in the thorax. g, Close-up of the sclerites and steering muscles. Tendons are marked in grey. h, Steering muscles grouped according to the sclerite to which they attached: basalare (b1, b2 and b3), first axillary (i1 and i2), third axillary (iii1, iii2 and iii3) and fourth axillary (iv1, iv2, iv3 and iv4) muscles.
Morphology of the dipteran wing hinge
The sclerites within the wing hinge are so small and move so rapidly that their mechanical operation during flight has not be accurately captured despite efforts using stroboscopic photography14, high-speed videography15 and X-ray tomography16. The current hypotheses for how the hinge works are inferred primarily from its static morphology in a variety of dipteran species2,9,10,12,16. Supplementary Video 1 illustrates the 3D arrangement of wing sclerites and their associated control muscles in Drosophila melanogaster. The consensus of prior studies suggests that the tiny deformations generated by contractions of the IFMs are transformed into wing motion via the actions of the anterior notal process (ANP) and posterior–medial notal process (PMNP) on the x-shaped first axillary sclerite (ax1) at the base of the wing (Fig. 1d–h). The ANP projects from the anterior end of the parascutal shelf and fits in the groove between the two anterior lobes of ax1. The PMNP sits at the anterior end of the large scutellar lever arm (SLA) in a notch between the two ventral lobes of ax1. Contractions of the dorso-longitudinal muscles during the downstroke are transmitted to the wing hinge via two main mechanisms: (1) a rotation of the scutellum along its junction with the notum that moves the SLA and PMNP upward; and (2) an elevation of the entire notum that raises the ANP at the lateral edge of the parascutal shelf (Fig. 1a). Contractions of the antagonistic dorso-ventral muscles cause the opposite effects during the upstroke (Fig. 1b). The more distal second axillary sclerite (ax2) possesses a dorsal flange that fits between the two dorsal protuberances of ax1, coupling the two sclerites during flight (Fig. 1g). Ax2 is a critical component of the hinge because it is rigidly attached to the radial vein (the structural spar of the anterior half of the wing) and has an anteriorly projecting ventral extension that fits into a notch in the pleural wing process, thus forming the fulcrum for wing motion. Because this fulcrum point is distal and anterior to ax1, upward motion of the SLA causes downward motion and pronation of the wing, whereas downward motion of the SLA causes upward motion and supination of the wing8.
In addition to the SLA, the anterior end of the scutellum is connected to another anteriorly projecting structure, the fourth axillary sclerite (ax4), providing a posterior mechanical pathway linking IFM strains to the wing base (Fig. 1a–e). Ax4 lies ventrolateral to the SLA and is attached by a long, previously unnamed sclerite (we term the ‘canoe’) to the third axillary sclerite (ax3). Ax3 contacts ax1, ax2 and the base of the anal vein (Fig. 1d,e). Because the anal vein serves as the structural spar of the posterior half of the wing, the linkage system consisting of the scutellum, ax4, canoe and ax3 could regulate wing camber and angle of attack10,17. In some prior studies, the canoe has been drawn as an extension of ax317,18, but the two structures are clearly distinct in our confocal reconstructions (Supplementary Video 1) and recent micro-computed tomography (μ-CT) data19.
In addition to the four axillary sclerites, another important hinge structure, the basalare, sits in the membranous episternal cleft just anterior to the wing14,16,20 (Fig. 1g,h). From the surface, the basalare looks like a tiny triangle, but its long internal invagination (apophysis) provides an attachment site for two of its three control muscles (Fig. 1h). During flight, the basalare is thought to oscillate passively due to the opening and closing of the episternal cleft. However, the magnitude of these oscillations and the mean position of the basalare are actively regulated by its three control muscles16,21, possibly influencing wing motion via a ligamentous connection to the radial vein.
Although the IFMs provide the power to oscillate the wings, they cannot regulate the rapid changes in kinematics required for flight manoeuvres or the sustained asymmetrical patterns of motion necessary to remain aloft with damaged wings22. In flies, that control is mediated by a set of 12 steering muscles that insert on 3 of the 4 axillary sclerites (ax1, ax3, ax4) and the basalare (Fig. 1f–h). The steering muscles are named after the sclerite on which they act (ax1: i1 and i2; ax3: iii1, iii2 and iii3; ax4: iv1, iv2, iv3 and iv4; basalare: b1, b2 and b3). Ax4 and its muscles are sometimes described using an alternative nomenclature (hg) after its German name hinter-Gelenkforsatz18. Each steering muscle is innervated by only a single excitatory motor neuron23,24, thus the entire flight motor system is remarkably sparse relative to comparably manoeuvrable vertebrates such as hummingbirds25.
Neural network model wing hinge
Machine learning can provide insight into many biological phenomena, for example, processing of visual information in mammalian cortex26. Although such approaches can accurately predict an output (for example, neuronal responses) from a set of inputs (for example, sensory stimuli), gaining mechanistic insight from the hidden features of a trained neural network is challenging. Our aim was to use machine learning to probe the musculoskeletal transformations that underlie the mechanical function of the hinge, using the activity of control muscles as an input, and the detailed 3D motion of the wings as an output. We constructed an experimental apparatus that captured the wing motion of tethered flies at 15,000 frames per second, while simultaneously measuring steering muscle activity using the Ca2+ indicator GCaMP7f27 (Fig. 2 and Extended Data Fig. 1). To capture a wide range of muscle activity, we collected a large set of sequences, each triggered whenever the activity of one of the steering muscles (chosen a priori before each trial) exceeded a user-defined threshold. To encourage a wide variety of flight behaviours, we displayed both open- and closed-loop visual stimuli on a cylindrical array of light-emitting diodes28 (LEDs) surrounding the fly. We recorded a total of 485 flight sequences from 82 flies. After excluding a subset of wingbeats from sequences when the fly either stopped flying or flew at an abnormally low wingbeat frequency, we obtained a final dataset of 72,219 wingbeats.
a, Three high-speed cameras with IR backlighting (not shown) capture a tethered fly from three orthogonal angles. An epifluorescence microscope projects blue light onto the left side of the fly’s thorax for imaging muscle activity using GCaMP7f via a dichroic mirror. Visual stimuli are provided by an array of orange LED panels surrounding the fly. A prism splits the top view between a high-speed camera and a kinefly camera (Methods) that tracks the stroke amplitude of the two wings for use in epochs of closed-loop visual display. b, Wing pose is determined from the high-speed image data using custom machine vision software. The wing state is subsequently described by four angles relative to the SRF, defined by performing principal component analysis on the motion of the left and right wing roots, with the x–y plane representing the mean stroke plane. Two angles: ϕ (stroke angle) and θ (deviation angle), describe the wing tip’s orientation in the SRF. c, Wing pitch angle (η) indicates the orientation of the leading edge relative to the z axis of the SRF. The deformation angle (ξ) represents the uniform rotation along three hinge lines that roughly align with the L3, L4 and L5 veins.
To reconstruct the wing kinematics from high-speed images, we developed an automated tracking algorithm that used a trained convolutional neural network29 (CNN) and particle swarm optimization30 (Extended Data Fig. 2a and Supplementary Information). We specified wing pose relative to a fixed stroke plane reference frame (SRF) that approximated the mean stroke plane using three Tait–Bryan angles: stroke position (ϕ), deviation (θ) and wing pitch (η) (Fig. 2b). We also defined a fourth angle (ξ) that quantified wing camber (Fig. 2c). We parsed the kinematic traces into separate wingbeats, beginning and ending at dorsal stroke reversal. To reduce dimensionality, we fitted Legendre polynomials to all four kinematic angles, creating a vector of 80 coefficients (16 for ϕ, 20 for θ, 24 for η and 20 for ξ) that accurately captures the motion of one wing during each wingbeat.
We determined the Ca2+ signals originating from the 12 steering muscles using a real-time unmixing model27. The muscle activity traces were normalized over each trial duration, such that 0 and 1 correspond to −2 and +2 standard deviations, respectively (Fig. 3a). Owing to its binding kinetics31, GCaMP7f reports a low-pass filtered version of sarcoplasmic Ca2+ levels, which in addition to attenuating high frequencies, introduces a time delay between changes in ion concentration and fluorescence signals. Because of these effects, the muscle signals relevant for the kinematics of an individual wingbeat are spread out over several strokes, and the GCaMP7f data recorded after a particular wingbeat are better representative of the Ca2+ levels at the time the kinematic measurements were made. Thus, when training and evaluating our CNN, we associated the kinematics of each individual wingbeat with the Ca2+ signals recorded in the muscles during the subsequent nine wingbeats.
a, Example recording of normalized muscle fluorescence (grouped according to sclerite) and wingbeat frequency (freq). b, The CNN takes a 13 × 9 matrix as input, consisting of muscle activity and wingbeat frequency over a temporal window of 9 wingbeats. Two convolutional (conv) layers with SELU activations extract features from the input matrix. A fully connected (dense) layer with 1,024 neurons learns the relationships between the extracted features and an output layer of 80 neurons with linear activations. These 80 neurons correspond to 80 Legendre coefficients, encoding the four wing kinematic angles during the one wingbeat associated with each nine-wingbeat window of muscle activity. The wing kinematics are reconstructed by multiplying the Legendre coefficients with their corresponding basis functions (Xϕ, Xθ, Xη and Xξ). See Supplementary Information for more details regarding the CNN. c, Comparison between predicted (red) and tracked (black) traces of the four wing kinematic angles throughout the example recording. d, Training and validation error as a function of training epoch.
The details of the CNN and the metrics used to evaluate its accuracy are provided in Supplementary Information. We trained the network using 85% of our dataset, reserving 15% as a test set. For each wingbeat, the 13 × 9 matrix of muscle activity and wingbeat frequency provided the input to the network; the output consisted of a vector containing the 80 Legendre coefficients encoding the four wing pose angles (Fig. 3a,b). Even though the network was trained using a measure of muscle activity filtered by GCaMP7f kinetics, it nevertheless predicted the salient modulations in wing motion with remarkable accuracy (Fig. 3c and Extended Data Fig. 3). This accuracy was not guaranteed, given that prior biophysical experiments suggest that the properties of some steering muscles (most notably, b1 (ref. 20)) are regulated by changes in motor neuron firing phase, a physiological feature that is not detectable via Ca2+ imaging. This success of the network might imply that changes in firing phase are correlated with unknown features that are detectable by our imaging method, or that the prediction errors resulting from excluding firing phase are beyond the precision of our wing tracking. Nevertheless, the overall success of the network suggests that even a limited measure of steering muscle activity provides sufficient information to reconstruct the intricate pattern of wing motion.
A previous study found that steering muscles may be roughly categorized by their activity pattern into phasic muscles that fire in short bursts (for example, b2, i1, iii1 and iv1), and tonic muscles that exhibit more continuous patterns of activity (for example, b1, b3, i2, iii3 and iv4)27. We observed roughly the same pattern in our more limited dataset (Fig. 4a), which was deliberately enriched to capture sequences in which muscle activity and wing kinematics changed actively during the 1.1-s data sequences. To quantify the degree to which the activities of steering muscles were linked, we performed a correlation analysis on muscle activity, and found strong linear trends among some of the muscles (Extended Data Fig. 4 and Extended Data Table 1). We did not find any significant correlation between steering muscle activity and wingbeat frequency. This was not surprising, given that changes in wingbeat frequency are known to occur more slowly than other changes in the wing kinematics32, and the flight muscles implicated in frequency regulation (the IFMs33 and the two pleuro-sternal muscles34) were not recorded in this study.
a, Distribution of normalized muscle activity (ϵ) for all steering muscles. b, Muscle activity patterns generated in a series of virtual experiments performed on our CNN model of the wing hinge. Each of the twelve set of bars beneath each muscle indicates the normalized pattern of activity across all 12 muscles under two conditions: (1) baseline activity when no changes in wing motion were detected (inner grey bars); and (2) the patterns obtained by adjusting the activity of each muscle to its maximum value (outer coloured bars). The sets of grey bars are identical for each muscle, because the same baseline pattern is plotted for each case. When we changed the activity level of each individual muscle to ϵ = 1 (marked by an asterisk), we also adjusted the activity levels of all the other muscles according to the correlation patterns observed over our entire dataset (Extended Data Fig. 4) to ensure that the virtual manipulations were performed under conditions that approximated those under which the CNN was trained. Wingbeat frequency was maintained at a constant level for all patterns. c, CNN-predicted wing kinematics for the baseline and maximum activity patterns shown in b over a single wingbeat period (T). Coloured lines show kinematics resulting from maximum activity patterns of each muscle; baseline kinematics are shown in grey. The relative change in kinematics is plotted below.
Virtual manipulations using CNN model
We used our CNN model of the hinge to investigate the transformation between muscle activity and wing motion by performing a set of virtual manipulations, exploiting the network to execute experiments that would be difficult to perform on actual flies. First, we defined a baseline pattern of muscle activity that was derived from sequences without any noticeable changes in muscle fluorescence and wing motion (grey data, Fig. 4b,c). From this starting point, we could then manipulate the network inputs to systematically increase the activity of each individual steering muscle, along with proportional changes in the activity of the other muscles with which it is correlated (Extended Data Fig. 4). The kinematic consequences of increasing the normalized activity of each steering muscle to 1, representing near-maximal output, are plotted in the columns of Fig. 4c. From these data, it is possible to identify the correlations between muscle activation and changes in wing motion, as in an actual physiology experiment in which each steering muscle was recorded in isolation. The results of these virtual experiments are consistent with prior experimental data from the subset of steering muscles for which electrophysiological recordings have been possible. For example, activation of the b1 and b2 muscles is known to correlate with increases in stroke amplitude and deviation21,35. Although electrophysiological recordings from b3 have not been feasible owing to its small size, the antagonistic effects relative to b1 are consistent with the morphological arrangement of these two muscles. Similarly, the activity of i1 is known to correlate with decreases in stroke amplitude and deviation, whereas the activity of iii2 and iii3 are correlated with increases in these parameters35,36. The consistency of our model’s predictions with these prior observations provides confidence that the CNN converged on a solution that captures the salient features of wing hinge control.
Aerodynamic effects of steering muscles
The results of the virtual experiments using our CNN hinge model indicate that the 12 steering muscles act in a coordinated manner to regulate changes in wing motion. To examine the aerodynamic consequences of these changes, we used a dynamically scaled flapping robot37 (Extended Data Fig. 5). The motion of the leading edge of the robotic wing was controlled by a system of servo motors that specified the three wing angles (ϕ, θ and η). Unlike earlier versions that modelled the wing as a flat planform38, our new robotic wing consisted of four span-wise wedges connected by three hinge lines approximating the positions of wing veins L3, L4 and L5 (Figs. 1d and 2c). We could actuate the angles at each hinge line via servo motors, thus setting the fourth kinematic angle (ξ), which specifies camber (Extended Data Fig. 5a). A sensor at the base of the wing measured the aerodynamic forces and torques during flapping. Using prior estimates of wing mass39, we used the Newton–Euler equations to calculate the inertial forces generated by flapping and added them to the aerodynamic forces measured with the robot, thus obtaining a time history of total forces and torques generated by the wing for each wingbeat (Extended Data Fig. 5b,c). We then conducted this analysis for the 12 kinematic patterns predicted by our CNN for the maximum activity of each steering muscle (Fig. 4), creating a map linking muscle activity to flight forces (Extended Data Fig. 6).
Simulating free flight manoeuvres
Using our aerodynamic data, we tested whether our CNN linking muscle activity to wing motion could generate free flight manoeuvres that resemble those executed by real flies. To do so, we used model predictive control40 (MPC) to determine the activity patterns within the array of 24 steering muscles (12 per side) that generate any arbitrary flight manoeuvre within a specified number of wingbeats. The details of our simulator are described in Supplementary Information (Fig. 5 and Extended Data Fig. 7). In brief, the state vector of the system (x) consists of 13 parameters specifying the linear and angular velocity of the body in the SRF and the body quaternion and position in an inertial reference frame. The control vector (u) consists of 24 parameters representing the left and right muscle activities. Our model also incorporates body drag, the inertial and aerodynamic damping of the wings41, and gravitational forces. Figure 5 shows a simulation of a stereotypical flight manoeuvre—a body saccade42—in which flies rapidly change direction32,43,44 in approximately 100 ms. The MPC simulation starts in forward flight (0.3 m s−1) and has to reach a goal state with a 90o yaw rotation within 10 wingbeats. Left and right steering muscle activities show relatively subtle changes during the manoeuvre, with the largest asymmetries in b1, b3 and iv4 activity (Fig. 5c). The saccade is achieved by a combination of roll, pitch and yaw rotations with only a slight deceleration in forward velocity (Fig. 5d). Using the time course of muscle activity, we could reconstruct the left and right wing motion patterns during the saccade using our CNN model (Fig. 5e). We provide example MPC simulations of other flight manoeuvres in Extended Data Fig. 7 and Supplementary Video 2. It is difficult to make precise one-to-one comparisons with the flight manoeuvres of real flies, because the most relevant free flight data were collected on a slightly larger species (Drosophila hydei) and even relatively stereotyped manoeuvres such as spontaneous body saccades vary extensively from event to event32,43. Also, to challenge our model, we deliberately ran our simulations such that the virtual flies had to complete their flight manoeuvres within ten wingbeats, whereas real flies are not necessarily operating under such a hard constraint. Nevertheless, free flight sequences animated in the same format as our simulations indicate that our model captures the salient features of real flies (Supplementary Videos 3–8). For example, both real and simulated flies change heading primarily by rolling their body to reorient their mean force vector, a feat that they accomplish via small changes in the wing deviation angle. It is noteworthy that even though our CCN network of the wing hinge was trained using muscle activity filtered by CGaMP7f kinetics, the model could nevertheless replicate the rapid manoeuvres of real flies, a result that is consistent with the physiology of the steering muscles. While GCaMP7f dynamics are slow relative to the time course of individual motor neuron spikes, the more relevant time scale is twitch duration, which is limited by the surface area of sarcoplasmic reticulum. Even tonic muscles such as b1 that exhibit changes in dynamic stiffness due to activation phase have twitch durations lasting several wingbeats20.
a, Top view of simulation after 30 ms, showing red and blue traces for the left and right wingtips, respectively. b, Orthographic view of wingtip traces of the simulation in a stationary body frame. c, Control inputs during the saccade, representing the activity of left and right steering muscles, grouped according to sclerite. d, Body state during the saccade, including linear (v) and angular (ω) velocities in the SRF, as well as body quaternion (q) and position (p) relative to an inertial frame. e, CNN-predicted wing kinematic angles derived from the steering muscle activity in c for left (red) and right (blue) wings. The baseline wingbeat (black) is plotted for comparison. Additional MPC simulation results can be found in Extended Data Figure 7 and Supplementary Video 2.
Latent variable analysis
In the simulated experiments summarized in Fig. 4, we deliberately incorporated the correlation patterns among muscles so that the results could be most accurately compared to prior work and were similar to the conditions under which we trained the network. However, this approach makes it difficult to isolate the effects of single muscles on wing motion. To gain more insight into the mechanical function of individual steering muscles, we constructed and trained an encoder–decoder4, with a network architecture that deliberately partitioned the input data from the steering muscles into four nodes according to the sclerites on which they act, with an additional node for wingbeat frequency (Extended Data Fig. 8). This narrow bottleneck of nodes projects muscle activity and frequency into a five-dimensional latent variable space, effectively performing a non-linear principal component analysis4. A subsequent layer of the network correlates each of these latent variables to changes in wing kinematics. After training this modified network on the dataset, we could then predict the changes in wing motion resulting from the activity of the four muscle groups (Fig. 6).
a, Muscle activity, ε, predicted by the decoder as a function of the sclerite latent parameters (see Extended Data Fig. 8). The z-scored latent variables were sampled in 9 steps (−3σ to +3σ), indicated by colour bars. b, Wing angles predicted by decoder as a function of latent state (coded by colour). c–f, Patterns of wing motion corresponding to extreme sclerite states (−3σ, blue; +3σ, red). Each figure plots instantaneous wing pitch and deformation at regular intervals superimposed over the wingtip traces. c, Cartoon illustrating proposed mechanism of action for the basalare, drawn for a left wing. The b1 and b2 rotate the basalare clockwise, exerting tension on a ligament attached to the radial vein, extending the wingbeat forward and advancing supination during ventral stroke reversal. The b3 rotates the basalare counter-clockwise and has the opposite effects. d, Cartoon illustrating proposed action of ax1 drawn for a left wing in cross section through the hinge. Contraction of i1 and i2 muscles pull ax1 downward, decreasing the ventral extent of the wingbeat owing to the inboard position of ax1 relative to ax2. e, Same view as in d, illustrating action of ax3. The iii2 and iii3 muscles pull ax3 downward, increasing the ventral extent of the wingbeat due to the outboard position of ax3 relative to ax2. f, Same view as in e, illustrating action of ax4. Ax4 is connected to ax3 and the anal vein via the canoe. The iv1 and iv2 muscles pull ax4 inward, leading to an increased angle of attack during the downstroke via the ax4-canoe-ax3-AV linkage. The iv3 and iv4 muscles pull ax4 outward, reducing the angle of attack during the downstroke.
Our analysis suggests that tension exerted on each sclerite modulates distinct aspects of wing motion (Fig. 6). For example, the b1 and b2 muscles act agonistically to increase stroke deviation and amplitude during the downstroke, as well as advancing the phase of wing rotation at the start of the upstroke. The increased tension on the ligament connecting the basalare to the radial vein caused by contraction of b1 and b2 could explain these results (Fig. 6b), with the b3 muscle having an antagonistic effect. Downward tension on ax1 caused by the activation of i1 and i2 decreases stroke amplitude and deviation at the start of the upstroke (Fig. 6d), with activation of iii2 and iii3 having almost precisely the opposite effect (Fig. 6e). The positions of the ax1 and ax3 muscle insertions relative to ax2 may provide a simple explanation for these antagonistic influences. Although the ax2 has no control muscles, this critical sclerite forms the hinge fulcrum via its joint with the pleural wing process. Whereas the muscle insertions on ax1 are medial to this fulcrum (Fig. 6d), those of ax3 are distal2,9,10,45 (Fig. 6e). Downward tension created by the i1 and i2 would tend to bias the mean stroke position of the wing upward (by pulling inboard of the fulcrum), tension created by the muscles of iii2 and iii3 would bias the wing downward (by pulling outboard of the fulcrum). The large iii1 muscle is rarely active during flight27 and is probably homologous with the ancestral retractor muscle that is used for folding the wing along the body axis45.
Our latent variable analysis suggests that the ax4 muscles exert a strong influence on wing motion (Fig. 6a,b,f). In particular, the large iv1 muscle decreases the magnitude of stroke amplitude and deviation, while increasing angle of attack during the downstroke, actions that are agonized by the activity of iv2. Both iv1 and iv2 insert at the same location on ax4 with a line of action that would pull the sclerite downward and inboard. The latent variable analysis suggests that iv3 has the opposite effect: increasing stroke amplitude and deviation while lowering angle of attack during the downstroke, actions that are mildly agonized by iv4. These results are roughly consistent with our virtual activation experiments using the CNN model (Fig. 4). A functional stratification of the ax4 muscles into two antagonistic groups has not been observed in previous physiological studies, although these muscles have proven difficult to record owing to their small size and tight clustering. However, prior morphological investigations have noted that these muscles insert via separate tendons that could cause different deflections of ax410,46. Our own morphological analysis (Figs. 1h and 6f and Supplementary Video 1) uncovered an intriguing feature of hinge morphology that further supports this functional stratification. In particular, the small, tonically active iv4 muscle does not insert on ax4 directly, but rather connects laterally to the tendon of iv3 by a conspicuously broad ligament that would act to pull the sclerite outboard, antagonistic to the actions of iv1 and iv2.
The role of passive wing flexion
Whereas our the latent variable analysis predicts a large influence of ax4 muscles on stroke amplitude and deviation, prior anatomical analysis suggest that this sclerite’s primary effect would be to regulate angle of attack10. According to this hypothesis, iv1 and iv2 muscles cause an inward movement of ax4 that depresses the anal vein of the wing during the downstroke via its linkage to ax3, thereby increasing angle of attack and reducing the lift-to-drag ratio (Fig. 6f). We speculate that this active control of angle of attack and the resulting effects on flight forces might lead to passive changes in stroke amplitude and deviation via a flexure joint at the base of the wing. Evidence for just such a flexure line has been proposed for other fly species47 and is evident in high-speed imaging data collected during free flight in D. hydei, which show that the wing blade extends beyond the maximum angular limit of the wing base during stroke reversal, with the deformation concentrated at a narrow point (Extended Data Fig. 9a). Using confocal microscopy, we detected a strong autofluorescence signal with 405 nm excitation (Extended Data Figure 9b), consistent with the presence of the elastic protein resilin along a chord-wise line at the base of the wing48. Resilin is a rubber-like protein first discovered by Weis-Fogh in the wing hinge of locusts and flight muscle tendons of dragonflies49. This chord-wise flexure line could explain how alterations in angle of attack caused by activity of the ax4 muscles could generate the large measured changes in stroke amplitude and deviation, and might also have a role in the storage and recovery of inertial energy via elastic storage50,51,52.
Conclusions
In this study, we used machine learning to construct a CNN model of the fly’s wing hinge based on the simultaneous measurement of steering muscle activity and wing kinematics. There are several reasons suggesting that our CNN model provides an accurate representation of the biomechanical processes that transform steering muscle activity into wing motion. First, when we used the network to predict the pattern of wing motion generated by the maximal activity of each muscle, the model’s output was consistent with all prior experiments in which muscle activity has been measured directly with electrodes (Fig. 4). Second, when we embedded the CNN hinge model into a state-space simulation of different canonical flight manoeuvres, the results resemble the known behaviour of freely flying flies32,43,53,54 (Fig. 5). Third, when we constructed an encoder–decoder in which the steering muscle activity was parsed into nodes representing the muscle groups inserting on each of the four wing sclerites (Fig. 6), the model’s predictions were consistent with known features of hinge morphology and differences in the insertion patterns of control muscles. Collectively, our results generate a suite of specific hypotheses (Fig. 6c–f) that can be tested via a combination of additional experiments and a detailed, physics-based model of the hinge. Further, with the recent availability of a connectome for the ventral nerve cord of Drosophila55,56, we expect that these results will help provide a deeper understanding of flight control circuitry that underlies the recruitment of steering muscles during flight manoeuvres.
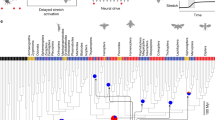
Until recently, the morphologically complex wing hinge of flies and other neopterous insects was considered a derived trait that allows the wing to fold along the body axis when not in use. The hinge found in extant paleopterous insects (dragonflies, damselflies and mayflies), which is actuated by direct flight muscles and does not permit folding, was considered representative of the ancestral condition57. However, a new phylogeny derived from a comprehensive genetic analysis of the Polyneoptera provides a compelling reinterpretation58, in which the unfoldable wing of the paleopterous orders is more parsimoniously interpreted as a radical simplification of the ancestral condition. According to this hypothesis, many features of the odonates—including the inability to fold their wings—are secondary specializations of their large size, perching habit, and predatory lifestyle59,60,61. Thus, despite the fact that Drosophila is a crown taxon, the analysis of its wing hinge could provide general insight into biomechanical innovations underling the evolution of insect flight.
Methods
Flies
We generated flies expressing GCaMP7f in steering muscles by crossing w[1118]; + ;P{y[+t7.7] w[+mC]=R22H05-Gal4}attP2 and +[HCS];P{20XUAS-IVS-GCaMP7f}attP40;+. All experiments were conducted on the three-day-old female offspring. We anaesthetized flies on a 4 °C cold plate to immobilize them and removed the anterior two pairs of legs at the coxa. The flies were then attached to a tungsten wire at the notum using UV-curing glue (Bondic) and placed into our experimental setup after a 10-min recovery period.
Imaging and constructing 3D model of the wing hinge
Our reconstruction of cuticle morphology using confocal microscopy was based on a previously published method62. Flies were anaesthetized with acetone and briefly washed with 70% ethanol. We placed the animals in PBS with 2% paraformaldehyde and 0.1% Triton X-100 and before carefully removing their abdomen. After overnight fixation at 4 °C, we isolated the thoraces and bleached the preparations for 24 h in 20% hydrogen peroxide. The samples were then embedded in 7% agarose and cut into 0.3 mm sagittal sections using a vibratome (Leica VT1000s). The slices were incubated overnight at 37 °C in PBS, 0.1% Triton X-100, and 0.2 mg ml−1 of trypsin to remove soft tissues. We then gradually dehydrated the preparations in a glycerol series (2% to 80%), followed by ethanol series (20% to 100%), before mounting them between two cover slips in methyl salicylate for imaging. Serial optical sections were obtained on a laser confocal microscope (Zeiss 980) at 1 µm intervals using a LD-LCI 25×, 0.8 NA objective, or at 0.3 µm intervals, using a Plan-Apochromat 40×, 0.8 NA objective. We detected the green autofluorescence characteristic of hard, sclerotized cuticle by exciting the tissue with 488 nm light. We extracted 3D meshes from the confocal stacks using a viewer plugin for FIJI (http://fiji.sc/) and imported the data into Blender (http://www.blender.org.). The resulting image data from the 0.3-mm sections were then processed to segment individual structures (for example, sclerites, muscles and apodemes) and reconstruct the continuous 3D morphology of the thorax in the vicinity of the wing hinge. The results were used to create the 2D cartoons in Fig. 1, and the animation in Supplementary Video 1. The same tissue preparation protocol was used to capture the image of the wing base in Extended Data Figure 9b, using a Zeiss 880 microscope with a Plan-Apochromat 40×, 0.8 NA objective. We collected autofluorescence signals at 405, 488 and 568 nm excitation for evidence of dityrosine bonds consistent with the presence of resilin.
High-speed camera recordings
We used three synchronized high-speed cameras (SA5, Photron) to image the flies from orthogonal views. The cameras recorded continuously at a rate of 15,000 frames per second (fps) with an electronic shutter speed of 33.3 μs, telecentric lenses (0.5X PlatinumTL, Edmund Optics) and collimated infrared (850 nm) backlights (M850LP1 LED and SM2P collimation lens, Thor Labs). The 3D calibration of the high-speed cameras was achieved using the direct linear transformation (DLT) method (http://www.kwon3d.com/theory/dlt/dlt.html). The recorded images had a resolution of 256 × 256 pixels with 8-bit depth. To optimize data storage, we divided the camera memory buffers into 8 partitions, each accommodating one 1.1-s sequence of 16,376 frames. Once all partitions were full, the data were transferred to a hard drive. During each experiment, we also operated a low-speed tracking system called kinefly63, which tracked the stroke amplitude of the left and right wings so that we could implement visual closed-loop of the cylindrical array of LEDs28 surrounding the fly.
Real-time calcium imaging
We used a previously described technique27 to visualize muscle activity using GCaMP7f in the fly by utilizing single photon excitation through the cuticle. A blue LED (M470L3, Thor Labs) served as the excitation light source, which was directed through a 480/40 nm excitation filter (Chroma) and focused onto the fly using a 4× lens (CFI Plan Fluor, Nikon). The resulting fluorescent light passed through a 535/50 nm emission filter (Chroma) and was captured by a machine vision camera (BF3-U3-04S2M-CS, FLIR). To synchronize the imaging process, we utilized an optoelectronic wingbeat analyser64 to strobe the blue excitation LED for 1 ms at the dorsal stroke reversal of each wingbeat. The fluorescence camera was strobed at half the wingbeat frequency (~100 fps), resulting in each frame representing the sum of two consecutive illuminations. To synchronize the high-speed and fluorescence image streams, we employed a microcontroller (Teensy 3.2) that received synchronization pulses from the Photron cameras and ventral stroke reversal signals from the wingbeat analyser, and coordinated the strobe signals to the blue excitation LED and the machine vision camera. The fluorescence images, along with a high-speed pulse count serving as a timestamp, were saved by our data acquisition software implemented in Robotic Operating System (ROS; https://www.ros.org/) using Python scripts. An unmixing algorithm, described elsewhere27, was applied to extract the Ca2+ signal from the overlapping muscles. To initialize an experiment, a GUI was used to orient and scale the 3D muscle model to the image using an affine transformation. The resulting fluorescence image and data vector capturing the activity of the steering muscles were saved in an hdf5 file. The muscle activity vector was stored in a rolling buffer within ROS, covering approximately 30 s of data.
Our objective was to trigger the data sequences during rapid changes in muscle activity. To achieve this, a separate ROS node monitored the activity of a predefined muscle of interest throughout the experiment. At the start of each trial, we selected a muscle and an activity threshold for the trigger level. During the experiment, the trigger node calculated the gradient of muscle activity over 3 frames (equivalent to 6 wingbeats) and normalized this signal by dividing it by the standard deviation over a 30-s rolling buffer. Whenever the absolute value of the gradient exceeded the activity threshold, indicating a significant increase or decrease in muscle activity, the high-speed cameras were triggered in centre mode, such that we saved 8,188 frames before the trigger event and 8,188 frames after. A 30-s refractory period after each trigger event ensured there was no overlap between subsequent high-speed sequences. In total, we recorded data from 82 flies, resulting in 485 high-speed videos (Extended Data Table 2). We aimed to record sequences triggered on all 12 muscles from a minimum of 5 flies each. However, the iii1 muscle exhibited sporadic activity, primarily during flight starts and stops, whereas iii2 displayed more gradual changes in activity compared to other muscles. As a result, we were only able to capture data from one fly for iii1 and four flies for iii2.
Automated wing pose reconstruction
We developed an automated tracking system to extract body and wing pose, called Flynet, which consists of two steps: (1) a trained CNN that predicts pose vectors of the body and wing; and (2) a particle swarm optimization (PSO) step that refines the CNN prediction via 3D model fitting. A GUI was created to load images, scale the 3D fly model, annotate frames, and run the automated tracking algorithm. The 3D fly model includes the head, thorax, abdomen, left wing, and right wing, each with a pose vector comprising a quaternion, q, and position, p. The CNN, built using Tensorflow (https://www.tensorflow.org/) and Keras (https://keras.io/) in Python 3, consists of a convolutional block to extract image features in each camera view, followed by a fully connected layer of 1,024 neurons with SELU activation functions65 and a final layer of 37 neurons with linear activations corresponding to the pose vectors of the five 3D model components (Extended Data Fig. 2a). Further details regarding the training and operation of the automated wing tracker are provided in Supplementary Information.
We defined the SRF by conducting a principal component analysis on the left and right wing root traces; the first, second, and third principal components of the root traces determined the y, x and z axes of the SRF, respectively. Once the SRF was defined, we used three Tait–Bryan angles to specify the orientation of the leading edge of the wing: stroke angle ϕ, deviation angle θ, and wing pitch angle η. Additionally, we defined a fourth angle (ξ) to describe chord-wise wing deformation (that is, camber) along three flexure lines roughly corresponding to wing veins L3, L4 and L5 (Fig. 1a). We assumed equal division of the total deformation angle among these three span-wise flexure lines (Extended Data Fig. 2b).
Following the tracking process with Flynet, we applied an extended Kalman filter (EKF)66 to achieve temporal smoothing of body and wing pose. Each pose vector was independently filtered, incorporating the first and second order temporal derivatives to ensure smooth motion. The system matrix utilized the temporal derivative of quaternions and a Maclaurin series of position and wing deformation. To achieve optimal smoothing with no phase shift, we performed both a forward pass of the EKF and a backward pass using a Rauch–Tung–Striebel smoother. The Flynet GUI enables users to adjust the system covariance, allowing control over the degree of filtering for each pose parameter. To facilitate comparison between wingbeats of different durations, we normalized the wingbeat period to 1 and computed the wingbeat frequency f for each wingbeat, starting and ending with dorsal stroke reversal. To accurately quantify the complex time history of wing motion, we fit Legendre polynomials to the 4 wing kinematic angles for each wingbeat as described more fully in Supplementary Information.
Outlier rejection, normalization and training of the CNN
From the initial dataset of 83,056 wingbeats, we removed unrealistic flight conditions by excluding wingbeats with frequencies outside the range of 150 to 250 Hz and outliers exceeding biologically plausible angle limits for each wing kinematic angle ([−120o ≤ ϕ ≤ 120o], [−60o ≤ θ ≤ 60o], [−150o ≤ η ≤ 150o], [−90o ≤ ξ ≤ 90o]). These two criteria eliminated 13% of the data, resulting in a final dataset of 72,219 wingbeats. Prior to training the CNN, we normalized the muscle activity dataset by dividing the GCaMP7f traces by their respective standard deviations over the entire experiment duration such that 0 corresponds to −2σ and 1 to +2σ, and normalized wingbeat frequency by mapping 150 Hz to 0 and 250 Hz to 1. These normalized values formed a 13 × 9 input matrix representing muscle activity and wingbeat frequency over 9 wingbeats. The output data consisted of the 80 Legendre coefficients for the left wing kinematic angles, normalized to range between −1 and 1 by dividing by π. The CNN employed a sliding window of 9 wingbeats as input and predicted the Legendre coefficients for the first wingbeat of the window. Because the muscle fluorescence was recorded at half the wingbeat frequency, we interpolated the data to predict a value for each wingbeat. We allocated the first 30 wingbeats from each high-speed video sequence for validation, while the remaining wingbeats were used for training. The training and validation datasets, consisting of 61,351 and 10,868 wingbeats respectively, were randomly shuffled. The CNN was trained for 1000 epochs using a batch size of 100, a learning rate of 10−4, and a learning rate decay of 10−7 per epoch, with the mse serving as the loss function. Gaussian noise with a standard deviation of 0.05 was added to the network input to enhance noise robustness. After shuffling the dataset following each epoch, the training and validation losses reached mse values of 10−3.33 and 10−3.07, respectively (Fig. 3d). To evaluate the accuracy, the trained CNN was used to predict wing motion for each validation sequence based on the muscle activity data. The predicted coefficient vector was multiplied by π and separated into coefficients for the four wing kinematic angles. Multiplication of the Legendre coefficient vectors by the Legendre polynomial bases recovered the four wing kinematic angles during each wingbeat. Figure 3 and Extended Data Fig. 3 illustrate examples of different sequences of wing motion predicted from muscle activity and wingbeat frequency.
Muscle activity correlation analysis
We conducted a linear correlation analysis on the muscle activity in our dataset. For each steering muscle, we selected samples with an average gradient larger than 0.005 over the 9-wingbeat window of muscle fluorescence, a criterion based on the sharp rise and slow decay of the GCaMP7f fluorescence kernel. Scatterplots illustrating the correlations between steering muscle fluorescence and wingbeat frequency for the gradient-selected wingbeats are presented in Extended Data Fig. 4. By employing linear models with RANSAC fitting67, we determined the slopes and intercepts for all correlations. Excluding the wingbeat frequency, muscle activity can be assumed to reside on a 12-dimensional plane. To ensure that all linear correlations in muscle activity share a common origin, we defined a baseline muscle activity pattern. This involved identifying sequences in the dataset with no significant changes in muscle activity or wing motion and setting the wingbeat frequency to 200 Hz. In Extended Data Fig. 5, the intercepts of the linear model were adjusted such that the baseline muscle activity pattern aligns with the 12-dimensional plane (Extended Data Table 1).
Virtual experiments
To conduct virtual experiments on the wing hinge (Fig. 4), we utilized the measured muscle activity correlation (Extended Data Fig. 4 and Extended Data Table 1) in conjunction with the trained CNN. In order to obtain realistic inputs to the CNN, we used a linear model to find steering muscle activity patterns that were within the manifold of the dataset that was used to train the CNN. First, we determined the baseline wing kinematics through the CNN’s prediction of the wing motion corresponding to the baseline pattern of muscle activity repeated over 9 wingbeats. To find the maximum muscle activity patterns for each muscle, we traversed the 12-dimensional surface from the baseline muscle activity pattern to a point where the specific muscle reached an activity value of 1.
Robotic wing experiments
The aerodynamic forces generated by the baseline and maximum muscle activity patterns predicted by the CNN were assessed using a dynamically scaled robotic wing consisting of a stepper motor (IMS M-2231-3) controlling ϕ through two gears with a 1:3 ratio, and two servo motors (HiTec D951TW) controlling θ and η. A six-degree-of-freedom sensor (ATI Nano 17) captured forces and torques along three orthogonal axes at the wing’s base. A microcontroller (Teensy 3.2), operating at 100 Hz, updated stepper and servo positions and collected sensor data (Extended Data Fig. 5a). The wing itself consisted of four acrylic panels and three micro-servos (HiTec HS-7115TH) at the base, collectively controlling the fourth wing kinematic angle, ξ. Each trial involved repeating the programmed motion pattern for seven wingbeats. The commands for the first and last wingbeats of each sequence were multiplied by the first and second quarter, respectively, of a sin2(t/T) function, ensuring that the wing began and ended at the home position (ϕ = 0, θ = 0, η = 0, ξ = 0). To reach steady-state conditions for wake effects, the measurements from the first three wingbeats were discarded68, and we calculated median force and torque values over wingbeats 4, 5 and 6. In a separate experiment, gravitational and buoyancy forces acting on the wing were measured by playing back the wing motion at a 0.2× speed, enabling subtraction of the gravity measurement from the corresponding 1× speed experiment to isolate aerodynamic forces and torques. As described in Supplementary Information, the values for total forces and torques that we report (Ftotal, Ttotal) are the sum of aerodynamic components measured using the dynamically scaled wing and inertial components calculated using the Newton–Euler equations.
MPC simulations
To simulate free flight manoeuvres, we integrated a non-linear, discrete-time, state-space model of a flying fly, described in the Supplementary Information, into a MPC69 loop using the do-mpc Python package40 (Fig. 5, Extended Data Fig. 7 and Supplementary Video 2). Each MPC simulation involved defining a start state (xinit), a goal state (xgoal), and a time period for the manoeuvre execution. The optimization process computed the trajectory that minimizes the mse between the current state and the goal state, aiming to reach the goal state in the shortest possible time. The MPC controller utilized dynamic programming to optimize the state-space trajectory over a finite horizon, considering the objective cost function and non-linear constraints. In our simulations, the finite horizon was set to 10 wingbeats, ensuring continuous trajectory computation toward the goal state. The control inputs were the activity patterns of the left and right steering muscles. We restricted the muscle activity patterns based on the correlation analysis shown in Extended Data Fig. 4. Specifically, we limited the steering muscle activity to lie within 10% of the normalized 12-dimensional muscle activity plane for both wings. The muscle activity of each muscle was also bounded between −0.2 and 1.2. We assigned a penalization weight of 1 for each muscle, with the exception of muscles b2 and iii1, which were assigned penalization weights of 2 and 10, respectively. These higher weights were chosen because muscles b2 and iii1 are so seldom active27. In a similar fashion, the importance of the different components of the objective function was adjusted by setting weights. For instance, for symmetrical manoeuvres such as forward acceleration, ascent and descent, we assigned higher weights to state variables that broke symmetry.
Latent variable analysis
As described more fully in the Supplementary Information, we employed an encoder–decoder architecture for latent variable analysis4, in which the latent variables predicted both muscle activity and wing motion (Extended Data Figure 9a). The encoder split the input data into five streams, each processed by a separate CNN, to project onto an individual latent variable. The muscle activity decoder reconstructed the input matrix using a fully connected dense layer with hyperbolic tangent (tanh) activation and a deconvolutional layer with SELU activation. To focus on the effects of sclerite state on wing kinematics, we introduced a back-propagation stop between the latent space and the muscle activity decoder. Instead of relying on the muscle activity reconstruction error, we used the wing kinematics decoder’s reconstruction mse to update the encoder weights. The wing kinematics decoder utilized a dense layer of 1,024 neurons with SELU activation to predict the 80 Legendre coefficients representing wing motion during a wingbeat. After training the latent network for 1,000 epochs with mse as the error function, the training error for wing kinematic reconstruction was mse = 6.0 × 10−4, and the validation error was mse = 7.9 × 10−4. The changes in wing motion predicted by variation in the latent variables associated with each wing sclerite (Fig. 6b) were roughly consistent with the results of our virtual muscle activation experiments (Fig. 4c). Besides sclerite functionality, the latent variable analysis also predicts the effect of wingbeat frequency on wing motion. Our analysis predicted a decrease in stroke amplitude and downstroke-to-upstroke ratio for increasing frequency, consistent with a study on the power requirements of forward flight39. In Extended Data Fig. 8d, we present four aerodynamic parameters: absolute angle of attack, wingtip speed, and non-dimensional lift and drag in the SRF, where the quasi-steady model was used to compute lift and drag forces. The computed lift and drag values were lower than the measurements from the dynamically scaled model wing, a discrepancy that might be due to the omission of wing deformation in the quasi-steady calculations.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The data required to perform the analyses in this paper and reconstruct all the data figure are available in the following files: main_muscle_and_wing_data.h5, flynet_data.zip, robofly_data.zip, which are available from the Caltech Data website: https://doi.org/10.22002/aypcy-ck464. main_muscle_and_wing_data.h5 contains the time series of muscle activity and wing kinematics used to train the muscle-to-wing motion CNN and the encoder–decoder used in the latent variable analysis. flynet_data.zip contains a series of data files for training and running Flynet: (1) camera/calibration/cam_calib.txt (example camera calibration data); (2) movies/session_01_12_2020_10_22 (folder containing example movies); (3) labels.h5 and valid_labels.h5 (data for training); and (4) weights_24_03_2022_09_43_14.h5 (example weights). robofly_data.zip contains the MATLAB data files with force and torque data acquired using the dynamically scaled robotic fly.
Code availability
The code required to perform the analyses in this paper and reconstruct all the data figures are available at https://github.com/FlyRanch/mscode-melis-siwanowicz-dickinson. The software is organized into seven submodules: flynet, flynet-kalman, flynet-optimizer, latent-analysis, mpc-simulations, robofly and wing-hinge-cnn. The installation instructions, system requirements and dependency information are given separately in their respective folders. flynet is a neural network and GUI application that requires the dataset flynet_data.zip, and may be used to create Extended Data Fig. 2. An example demonstrating how to train the network can be found in the examples sub-directory and is called train_flynet.py. flynet-kalman is a Kalman filter Python extension used by Flynet. flynet-optimizer is a particle swarm optimization extension module used by Flynet. latent-analysis is a Python library and Jupyter notebook for performing latent variable analysis that requires the dataset main_muscle_and_wing_data.h5, and may be used to create Fig. 6 and Extended Data Fig. 8. mpc-simulations is a Python library and Jupyter notebook for MPC simulations, and may be used to create Fig. 5 and Extended Data Fig. 7. robofly is a Python library and Jupyter notebook for extracting force and torque data from the robotic fly experiments and plotting forces superimposed on 3D wing kinematics. It requires dataset robofly_data.zip, and may be used to create Extended Data Figs. 5 and 6. wing-hinge-cnn is a Python library and Jupyter notebook for creating the muscle-to-wing motion CNN. It requires main_muscle_and_wing_data.h5, and may be used to create Figs. 3 and 4 and Extended Data Fig. 3. An example demonstrating how to train the network can be found in the examples sub-directory as is called train_wing_hinge_cnn.py. The files containing the raw videos of the muscle Ca2+ images and high-speed videos of wing motion are too large to be hosted on a publicly accessible website. Example high-speed videos are provided in the folder movies/session_01_12_2020_10_22 mentioned in Data availability. Additional sequences are available upon request by contacting the corresponding author.
References
Grimaldi, D. & Engel, M. S. Evolution of the Insects (Cambridge Univ. Press, 2005).
Deora, T., Gundiah, N. & Sane, S. P. Mechanics of the thorax in flies. J. Exp. Biol. 220, 1382–1395 (2017).
Gu, J. et al. Recent advances in convolutional neural networks. Pattern Recognit. 77, 354–377 (2018).
Kramer, M. A. Nonlinear principal component analysis using autoassociative neural networks. AlChE J. 37, 233–243 (1991).
Pringle, J. W. S. The excitation and contraction of the flight muscles of insects. J. Physiol. 108, 226–232 (1949).
Josephson, R. K., Malamud, J. G. & Stokes, D. R. Asynchronous muscle: a primer. J. Exp. Biol. 203, 2713–2722 (2000).
Gau, J. et al. Bridging two insect flight modes in evolution, physiology and robophysics. Nature 622, 767–774 (2023).
Boettiger, E. G. & Furshpan, E. The mechanics of flight movements in diptera. Biol. Bull. 102, 200–211 (1952).
Pringle, J. W. S. Insect Flight (Cambridge Univ. Press, 1957).
Miyan, J. A. & Ewing, A. W. How Diptera move their wings: a re-examination of the wing base articulation and muscle systems concerned with flight. Phil. Trans. R. Soc. B 311, 271–302 (1985).
Wisser, A. Wing beat of Calliphora erythrocephala: turning axis and gearbox of the wing base (Insecta, Diptera). Zoomorph. 107, 359–369 (1988).
Ennos, R. A. A comparative study of the flight mechanism of diptera. J. Exp. Biol. 127, 355–372 (1987).
Dickinson, M. H. & Tu, M. S. The function of dipteran flight muscle. Comp. Biochem. Physiol. A 116, 223–238 (1997).
Nalbach, G. The gear change mechanism of the blowfly (Calliphora erythrocephala) in tethered flight. J. Comp. Physiol. A 165, 321–331 (1989).
Walker, S. M., Thomas, A. L. R. & Taylor, G. K. Operation of the alula as an indicator of gear change in hoverflies. J. R. Soc. Inter. 9, 1194–1207 (2011).
Walker, S. M. et al. In vivo time-resolved microtomography reveals the mechanics of the blowfly flight motor. PLoS Biol. 12, e1001823 (2014).
Wisser, A. & Nachtigall, W. Functional-morphological investigations on the flight muscles and their insertion points in the blowfly Calliphora erythrocephala (Insecta, Diptera). Zoomorph. 104, 188–195 (1984).
Heide, G. Funktion der nicht-fibrillaren Flugmuskeln von Calliphora. I. Lage Insertionsstellen und Innervierungsmuster der Muskeln. Zool. Jahrb., Abt. allg. Zool. Physiol. Tiere 76, 87–98 (1971).
Fabian, B., Schneeberg, K. & Beutel, R. G. Comparative thoracic anatomy of the wild type and wingless (wg1cn1) mutant of Drosophila melanogaster (Diptera). Arth. Struct. Dev. 45, 611–636 (2016).
Tu, M. & Dickinson, M. Modulation of negative work output from a steering muscle of the blowfly Calliphora vicina. J. Exp. Biol. 192, 207–224 (1994).
Tu, M. S. & Dickinson, M. H. The control of wing kinematics by two steering muscles of the blowfly (Calliphora vicina). J. Comp. Physiol. A 178, 813–830 (1996).
Muijres, F. T., Iwasaki, N. A., Elzinga, M. J., Melis, J. M. & Dickinson, M. H. Flies compensate for unilateral wing damage through modular adjustments of wing and body kinematics. Interface Focus 7, 20160103 (2017).
O’Sullivan, A. et al. Multifunctional wing motor control of song and flight. Curr. Biol. 28, 2705–2717.e4 (2018).
Azevedo, A. et al. Tools for comprehensive reconstruction and analysis of Drosophila motor circuits. Preprint at BioRxiv https://doi.org/10.1101/2022.12.15.520299 (2022).
Donovan, E. R. et al. Muscle activation patterns and motoranatomy of Anna’s hummingbirds Calypte anna and zebra finches Taeniopygia guttata. Physiol. Biochem. Zool. 86, 27–46 (2013).
Bashivan, P., Kar, K. & DiCarlo, J. J. Neural population control via deep image synthesis. Science 364, eaav9436 (2019).
Lindsay, T., Sustar, A. & Dickinson, M. The function and organization of the motor system controlling flight maneuvers in flies. Curr. Biol. 27, 345–358 (2017).
Reiser, M. B. & Dickinson, M. H. A modular display system for insect behavioral neuroscience. J. Neurosci. Meth. 167, 127–139 (2008).
Albawi, S., Mohammed, T. A. & Al-Zawi, S. Understanding of a convolutional neural network. In 2017 International Conference on Engineering and Technology (ICET) 1–6 https://doi.org/10.1109/ICEngTechnol.2017.8308186 (2017).
Kennedy, J. & Eberhart, R. Particle swarm optimization. In Proc. ICNN’95—International Conference on Neural Networks Vol. 4, 1942–1948 (1995).
Dana, H. et al. High-performance calcium sensors for imaging activity in neuronal populations and microcompartments. Nat. Methods 16, 649–657 (2019).
Muijres, F. T., Elzinga, M. J., Melis, J. M. & Dickinson, M. H. Flies evade looming targets by executing rapid visually directed banked turns. Science 344, 172–177 (2014).
Gordon, S. & Dickinson, M. H. Role of calcium in the regulation of mechanical power in insect flight. Proc. Natl Acad. Sci. USA 103, 4311–4315 (2006).
Nachtigall, W. & Wilson, D. M. Neuro-muscular control of dipteran flight. J. Exp. Biol. 47, 77–97 (1967).
Heide, G. & Götz, K. G. Optomotor control of course and altitude in Drosophila melanogaster is correlated with distinct activities of at least three pairs of flight steering muscles. J. Exp. Biol. 199, 1711–1726 (1996).
Balint, C. N. & Dickinson, M. H. The correlation between wing kinematics and steering muscle activity in the blowfly Calliphora vicina. J. Exp. Biol. 204, 4213–4226 (2001).
Elzinga, M. J., Dickson, W. B. & Dickinson, M. H. The influence of sensory delay on the yaw dynamics of a flapping insect. J. R. Soc. Interface 9, 1685–1696 (2012).
Dickinson, M. H., Lehmann, F.-O. & Sane, S. P. Wing rotation and the aerodynamic basis of insect flight. Science 284, 1954–1960 (1999).
Lehmann, F. O. & Dickinson, M. H. The changes in power requirements and muscle efficiency during elevated force production in the fruit fly Drosophila melanogaster. J. Exp. Biol. 200, 1133–1143 (1997).
Lucia, S., Tătulea-Codrean, A., Schoppmeyer, C. & Engell, S. Rapid development of modular and sustainable nonlinear model predictive control solutions. Control Eng. Pract. 60, 51–62 (2017).
Cheng, B., Fry, S. N., Huang, Q. & Deng, X. Aerodynamic damping during rapid flight maneuvers in the fruit fly Drosophila. J. Exp. Biol. 213, 602–612 (2010).
Collett, T. S. & Land, M. F. Visual control of flight behaviour in the hoverfly, Syritta pipiens L. J. Comp. Physiol. 99, 1–66 (1975).
Muijres, F. T., Elzinga, M. J., Iwasaki, N. A. & Dickinson, M. H. Body saccades of Drosophila consist of stereotyped banked turns. J. Exp. Biol. 218, 864–875 (2015).
Syme, D. A. & Josephson, R. K. How to build fast muscles: synchronous and asynchronous designs. Integr. Comp. Biol. 42, 762–770 (2002).
Snodgrass, R. E. Principles of Insect Morphology (Cornell Univ. Press, 2018).
Williams, C. M. & Williams, M. V. The flight muscles of Drosophila repleta. J. Morphol. 72, 589–599 (1943).
Wootton, R. The geometry and mechanics of insect wing deformations in flight: a modelling approach. Insects 11, 446 (2020).
Lerch, S. et al. Resilin matrix distribution, variability and function in Drosophila. BMC Biol. 18, 195 (2020).
Weis-Fogh, T. A rubber-like protein in insect cuticle. J. Exp. Biol. 37, 889–907 (1960).
Weis-Fogh, T. Energetics of hovering flight in hummingbirds and in Drosophila. J. Exp. Biol. 56, 79–104 (1972).
Ellington, C. P. The aerodynamics of hovering insect flight. VI. Lift and power requirements. Phil. Trans. R. Soc. B 305, 145–181 (1984).
Alexander, R. M. & Bennet-Clark, H. C. Storage of elastic strain energy in muscle and other tissues. Nature 265, 114–117 (1977).
Mronz, M. & Lehmann, F.-O. The free-flight response of Drosophila to motion of the visual environment. J. Exp. Biol. 211, 2026–2045 (2008).
Ristroph, L., Bergou, A. J., Guckenheimer, J., Wang, Z. J. & Cohen, I. Paddling mode of forward flight in insects. Phys. Rev. Lett. 106, 178103 (2011).
Takemura, S. et al. A connectome of the male Drosophila ventral nerve cord. Preprint at bioRxiv https://doi.org/10.1101/2023.06.05.543757 (2023).
Cheong, H. S. J. et al. Transforming descending input into behavior: The organization of premotor circuits in the Drosophila male adult nerve cord connectome. Preprint at BioRxiv https://doi.org/10.1101/2023.06.07.543976 (2023).
Martynov, A. B. Über zwei Grundtypen der Flügel bei den Insecten und ihre Evolution. Z. Morph. Ökol. Tiere 4, 465–501 (1925).
Wipfler, B. et al. Evolutionary history of Polyneoptera and its implications for our understanding of early winged insects. Proc. Natl Acad. Sci. USA 116, 3024–3029 (2019).
Hasenfuss, I. The evolutionary pathway to insect flight—a tentative reconstruction. Arthr. System. Phylog. 66, 19–35 (2008).
Willkommen, J. & Hörnschemeyer, T. The homology of wing base sclerites and flight muscles in Ephemeroptera and Neoptera and the morphology of the pterothorax of Habroleptoides confusa (Insecta: Ephemeroptera: Leptophlebiidae). Arthro. Struc. Develop. 36, 253–269 (2007).
Willmann, R. in Arthropod Relationships (eds Fortey, R. A. & Thomas, R. H.) 269–279 (Springer, 1998); https://doi.org/10.1007/978-94-011-4904-4_20.
Shao, L. et al. A neural circuit encoding the experience of copulation in female Drosophila. Neuron 102, 1025–1036.e6 (2019).
Suver, M. P., Huda, A., Iwasaki, N., Safarik, S. & Dickinson, M. H. An array of descending visual interneurons encoding self-motion in Drosophila. J. Neurosci. 36, 11768–11780 (2016).
Götz, K. G. Course-control, metabolism and wing interference during ultralong tethered flight in Drosophila melanogaster. J. Exp. Biol. 128, 35–46 (1987).
Klambauer, G., Unterthiner, T., Mayr, A. & Hochreiter, S. in Advances in Neural Information Processing Systems Vol. 30 (Curran Associates, 2017).
Grewal, M. S. & Andrews, A. P. Kalman Filtering: Theory and Practice with MATLAB (John Wiley & Sons, 2014).
Fischler, M. A. & Bolles, R. C. Random sample consensus: a paradigm for model fitting with applications to image analysis and automated cartography. Commun. ACM 24, 381–395 (1981).
Birch, J. M. & Dickinson, M. H. The influence of wing–wake interactions on the production of aerodynamic forces in flapping flight. J. Exp. Biol. 206, 2257–2272 (2003).
Kouvaritakis, B. & Cannon, M. Model Predictive Control: Classical, Robust and Stochastic (Springer, 2016).
Acknowledgements
The authors thank W. Dickson for extensive expertise in instrumentation, programming, data analysis, formatting all the data and code for public repositories, and creating the animations of free flight data in Supplementary Videos 3–8; T. Lindsay for assistance in the design of the epifluorescence microscope and data acquisition software used for muscle imaging; A. Erickson for helpful comments on the manuscript and Supplementary Information; A. Huda for assistance in the construction of genetic lines; J. Omoto for collecting confocal images of wings to visualize resilin using autofluorescence; J. Tuthill and T. Azevedo for a tomographic dataset of the Drosophila wing hinge that was collected at the European Synchrotron Radiation Facility in Grenoble, France; S. Whitehead for analysis of this tomography data to provide a preliminary reconstruction of the hinge sclerites, and for critical feedback on the manuscript text and data presentation; and B. Fabian and R. G. Beutel for providing μ-CT data from their publication on the morphology of the adult fly body. The research reported in this publication was supported by the National Institute of Neurological Disorders and Stroke of the NIH (U19NS104655). I.S. was supported through the AniBody Project Team at HHMI’s Janelia Research Campus for this work.
Author information
Authors and Affiliations
Contributions
J.M.M. collected all the data presented in the manuscript and developed the software for data analysis. J.M.M. and M.H.D. collaborated on planning the experiments, preparing figures, and writing the manuscript. I.S. collected the high-resolution morphological images of the Drosophila thorax and created Supplementary Video 1.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks the anonymous reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Automated setup for simultaneous recording of muscle fluorescence and wing motion.
a, Illustration of experimental apparatus, created using Solidworks (www.solidworks.com). High-speed cameras, equipped with 0.5X telecentric lenses and collimated IR back-lighting capture synchronized frames of the fly from three orthogonal angles at a rate of 15,000 frames per second. An epi-fluorescence microscope with a muscle imaging camera records GCaMP7f fluorescence in the left steering muscles at approximately 100 frames per second, utilizing a strobing mechanism triggered every other wingbeat. A blue LED provides a brief, 1 ms illumination of the fly’s thorax during dorsal stroke reversal. A camera operating at 30 fps captures a top view of the fly for the kinefly wing tracker. b, Image of the flight arena featuring the components of the setup: LED panorama, IR diode and wingbeat analyzer for triggering the muscle camera and blue LED, prism for splitting the top view between the high-speed camera and kinefly camera, IR backlight, 4X lens of the epi-fluorescence microscope, and a tethered fly illuminated by the blue LED.
Extended Data Fig. 2 Flynet workflow and definitions of wing kinematic angles.
a, The Flynet algorithm takes three synchronized frames as input. Each frame undergoes CNN processing, resulting in a 256-element feature vector extracted from the image. These three feature vectors are concatenated and analyzed by a fully connected (dense) layer with Scaled Exponential Linear Unit (SELU) activation, consisting of 1024 neurons. The output of the neural network is the predicted state (37 elements) of the five model components represented by a quaternion (q), translation vector (p), and wing deformation angle (ξ). Subsequently, the state vector is refined using 3D model fitting and particle swarm optimization (PSO). Normally distributed noise is added to the predicted state, forming the initial state for 16 particles. During the 3D model fitting, the particles traverse the state-space, maximizing the overlap between binary body and wing masks of the segmented frames (Ib) and the binary masks of the 3D model projected onto the camera views (Ip). The cost function (Ib ∆Ip)/(Ib∪Ip) is evaluated iteratively for a randomly selected 3D model component. The PSO algorithm tracks the personal best cost encountered by each particle and the overall lowest cost (global best). After 300 iterations, the refined state is determined by selecting the global best for each 3D model component. See Supplementary Information for more details. b, Training and validation error of the Flynet CNN as a function of training epoch.
Extended Data Fig. 3 CNN-predicted wing motion for example flight sequences.
a, The top five traces show activity of the steering muscles in the four sclerite groups as well as wingbeat frequency during a full, 1.1 second recording. The bottom four traces indicate comparison between the tracked (black) and CNN-predicted (red) wing kinematic angles throughout the sequence. Expanded plots of a 100-ms sequence (0.5 to 0.6 seconds) are plotted on the right. b, c, d. Same as but for a different flight sequences.
Extended Data Fig. 4 Correlation analysis of steering muscle fluorescence and wingbeat frequency.
Linear models (colored lines) fitted to wingbeats in the entire dataset of 72,219 wingbeats from 82 flies. Gray dots represent the normalized baseline muscle activity level, while colored dots represent the normalized maximum muscle activity level. The correlation coefficients associated with these plots are provided in Extended Data Table 1. For more detail on regression methods, see Supplementary Information.
Extended Data Fig. 5 Aerodynamic force measurements and inertial force calculations.
a, Dynamically scaled flapping fly wing model immersed in mineral oil. b, Non-dimensional forces and torques in the strokeplane reference frame (SRF) for the baseline wingbeat. The four traces in each panel correspond to the total (black: Ftotal, Ttotal), aerodynamic (blue: Faero, Taero), inertial components due to acceleration (green: Facc, Tacc), and inertial components due to angular velocity (red: Fangvel; Tangvel). See Supplementary Information for more details. c, Representation of total forces during the baseline wingbeat, viewed from the front, left, and top. Gray trace represents the wing trajectory; cyan arrows represent instantaneous total force on the wing. At the wing joint, three arrows depict the total mean force, half the body weight, and half the estimated body drag.
Extended Data Fig. 6 Aerodynamic and inertial forces for maximum muscle activity wingbeats.
Figures depict the CNN-predicted wing motion for maximum muscle activity patterns, viewed from the front, left, and top. Instantaneous vectors depicting the sum of aerodynamic and inertial forces are shown in cyan. The wingbeat-averaged force vector is indicated by the color corresponding to the specific steering muscle set to maximum activity. Note that the scaling for the wingbeat-averaged forces differs from that for the instantaneous forces. The black gravitational force and blue body drag force are plotted as in Extended Data Fig. 5c.
Extended Data Fig. 7 Simulation of free flight maneuvers using the state-space system and Model Predictive Control.
a, Schematic of the state-space system and MPC loop, including system matrix (A), control matrix (B), the state vector (x), temporal derivative (\(\dot{x}\)) left and right steering muscle activity (uL, uR), initial state (xinit) and goal state (xgoal). b, Forward flight simulation with wingtip traces in red and blue. c, Wing motion during forward flight simulation plotted in stationary body frame. d, Backward flight simulation. e, Wing motion during backward flight simulation plotted in stationary body frame. f, Left and right steering muscle activity during the forward flight manoeuvre. g, State vector during forward flight manoeuvre. h, Steering muscle activity for the backward flight manoeuvre. i, State vector for the backward flight maneuver. j, CNN-predicted left (red) and right (blue) wing kinematics for the forward flight manoeuvre. Note that because this is a bilaterally symmetric flight manoeuvre, the model generates left and right wing kinematics that are identical. The left wing kinematics are displayed underneath the right kinematics,and thus cannot be seen. A baseline wingbeat is shown to emphasize the relative changes in wing motion. k, CNN-predicted wing motion for the backward flight manoeuvre.
Extended Data Fig. 8 Latent variable analysis reveals sclerite function using an encoder–decoder.
a, The network architecture consists of an encoder (red), muscle activity decoder (green), and wing kinematics decoder (blue). The encoder splits the input data into five streams corresponding to different muscle groups and frequency. Feature extraction is performed using convolutional and fully connected layers with SELU activation. Each stream is projected onto a single latent variable. In the muscle activity decoder, the latent variables are transformed back into the input data. A backpropagation stop prevents weight adjustments in the encoder based on the muscle activity reconstruction. The wing kinematics decoder predicts the Legendre coefficients of wing motion using the latent variables. See Supplementary Information for more details. b, Predicted muscle activity (replotted from Fig. 6) and normalized wingbeat frequency as a function of each latent parameter varied within the range of −3σ to +3σ. Color bar indicates the latent variable value in panels (c) and (e). c, Predicted wing motion by the wing kinematics decoder for the five latent parameters. d, Absolute angle-of-attack (|α|), wingtip velocity (utip) in mm s−1, non-dimensional lift (L mg−1), and non-dimensional drag (D mg−1). The non-dimensional lift and drag were computed using a quasi-steady model as described in Supplementary Information.
Extended Data Fig. 9 Flexible wing root facilitates elastic storage during wingbeat and allows wing to passively respond to changes in lift and drag throughout stroke.
a, Top view of ventral stroke reversal in free flight. Red circles mark the estimated position of the wing hinge, dotted lines indicate the expected position of the wing if a chord-wise flexure line was not present. Images are reproduced from previously publish data32 from Drosophila hydei. b, Composite confocal image of the wing base on Drosophila melanogaster, indicating a bright blue band of auto-fluorescence consistent with the presence of resilin and existence of a chord-wise flexure line (dashed arrows). The image shown is characteristic of the 4 wings (from 4 individual female flies) that we processed for confocal microscopy.
Supplementary information
Supplementary Information
This file contains Supplementary Information, including Supplementary Figs. 1–4.
Supplementary Video 1
The left side of a Drosophila thorax, annotated to illustrate the arrangement of wing sclerites and associated musculature. The colour scheme used for the sclerites and muscles are consistent with Fig. 1.
Supplementary Video 2
Animations of seven simulated flight maneuvers shown in world and body frames (forward acceleration, backward acceleration, upward acceleration, downward acceleration, left saccade, right saccade, and sideways flight) generated using the CNN model of the wing hinge and state-space model operating with a MPC loop (see Fig. 5 and Extended Data Fig. 7).
Supplementary Video 3
A previously published free flight maneuver of D. hydei (Mujires et al., 2014), animated in the same format as that used to depict the simulated flight maneuvers in Supplementary Video 2. The sequence provides examples of slow and fast saccades and forward acceleration.
Supplementary Video 4
A previously published free flight maneuver of D. hydei (Mujires et al., 2014), animated in the same format as that used to depict the simulated flight maneuvers in Supplementary Video 2. The sequence provides examples of slow ascent, slow descent, and fast ascent.
Supplementary Video 5
A previously published free flight maneuver of D. hydei (Mujires et al., 2014), animated in the same format as that used to depict the simulated flight maneuvers in Supplementary Video 2. The sequence provides examples slow ascent with sideslip and a saccade.
Supplementary Video 6
A previously published free flight maneuver of D. hydei (Mujires et al., 2014), animated in the same format as that used to depict the simulated flight maneuvers in Supplementary Video 2. The sequence provides an example of a very fast saccade.
Supplementary Video 7
A previously published free flight maneuver of D. hydei (Mujires et al., 2014), animated in the same format as that used to depict the simulated flight maneuvers in Supplementary Video 2. The sequence provides an example of backwards flight.
Supplementary Video 8
A previously published free flight maneuver of D. hydei (Mujires et al., 2014), animated in the same format as that used to depict the simulated flight maneuvers in Supplementary Video 2. The sequence provides examples of forward flight with ascent and descent.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Melis, J.M., Siwanowicz, I. & Dickinson, M.H. Machine learning reveals the control mechanics of an insect wing hinge. Nature 628, 795–803 (2024). https://doi.org/10.1038/s41586-024-07293-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-024-07293-4
This article is cited by
-
An exploration of how the insect-wing hinge functions
Nature (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.