Abstract
Plants deploy receptor-like kinases and nucleotide-binding leucine-rich repeat receptors to confer host plant resistance (HPR) to herbivores1. These gene-for-gene interactions between insects and their hosts have been proposed for more than 50 years2. However, the molecular and cellular mechanisms that underlie HPR have been elusive, as the identity and sensing mechanisms of insect avirulence effectors have remained unknown. Here we identify an insect salivary protein perceived by a plant immune receptor. The BPH14-interacting salivary protein (BISP) from the brown planthopper (Nilaparvata lugens Stål) is secreted into rice (Oryza sativa) during feeding. In susceptible plants, BISP targets O. satvia RLCK185 (OsRLCK185; hereafter Os is used to denote O. satvia-related proteins or genes) to suppress basal defences. In resistant plants, the nucleotide-binding leucine-rich repeat receptor BPH14 directly binds BISP to activate HPR. Constitutive activation of Bph14-mediated immunity is detrimental to plant growth and productivity. The fine-tuning of Bph14-mediated HPR is achieved through direct binding of BISP and BPH14 to the selective autophagy cargo receptor OsNBR1, which delivers BISP to OsATG8 for degradation. Autophagy therefore controls BISP levels. In Bph14 plants, autophagy restores cellular homeostasis by downregulating HPR when feeding by brown planthoppers ceases. We identify an insect saliva protein sensed by a plant immune receptor and discover a three-way interaction system that offers opportunities for developing high-yield, insect-resistant crops.
Similar content being viewed by others
Main
During the millions of years of plant and insect co-evolution, herbivorous insects have developed diverse feeding strategies to retrieve nutrients from host plants. Such insects can cause up to 18% of annual losses of global crop yield3. Because plants are sessile and cannot escape continuous attack by pests, plants have evolved various defence strategies to combat herbivory. Unlike microbial pathogens, insects can move swiftly on or among plants as they feed, making it challenging to understand the mechanisms that underlie their interactions.
Herbivorous insects actively select their feeding sites and secrete saliva during feeding to facilitate nutrient acquisition from host plants1,4,5. Some salivary proteins induce host defences1,5. In the past decade, plant genes that encode nucleotide-binding leucine-rich repeat (NLR) receptors that confer resistance to piercing and sucking insects have been isolated through map-based cloning6,7,8,9,10,11,12. NLR receptors function as intracellular immune receptors that directly or indirectly detect cognate effectors13,14 and activate effector-triggered immunity (ETI)6,7,8,9,10, also known as HPR. The identities of the insect avirulence effectors that trigger ETI have not yet been discovered. Hence, how a host plant recognizes the insect and deploys and modulates resistance remains largely unknown.
The brown planthopper (BPH; N. lugens Stål) is the most destructive insect pest that threatens rice production globally15. Using its stylet mouthparts to probe and penetrate plant cells, it consumes phloem sap and eventually causes plant death and substantial yield losses16,17,18. BPHs migrate long distances, crossing national boundaries in search of better host plants15,16,17,18, which makes regional control of BPH challenging. For this reason, HPR has been crucial for BPH control. The first BPH-resistance gene isolated in rice, Bph14, encodes a coiled-coiled nucleotide-binding leucine-rich repeat (CC-NB-LRR) protein that forms a homomeric complex and interacts with WRKY transcription factors19. BPH14 enhances WRKY activity to confer resistance to BPH19.
Here we show that the BPH salivary protein BISP is directly recognized by BPH14. Notably, the perception of BISP not only triggers Bph14-mediated resistance but also activates NBR1-mediated selective autophagy. This results in the degradation of BISP to restrict hyperactivation of Bph14-regulated defences. We elucidate the molecular and cellular interactions that occur after BPH14 binds to the BISP effector to activate and modulate Bph14-mediated HPR.
BPH BISP interacts with rice BPH14
Bph14 has been widely used in rice breeding programmes, and many Bph14-resistant varieties have been released for rice production (Supplementary Table 1). Rice plants carrying Bph14 show little damage, whereas susceptible N14 plants die 7 days after BPH infestation (Fig. 1a and Extended Data Fig. 1a). Insect feeding and growth were inhibited on Bph14 plants, which resulted in lower weight gain and honeydew excretion than insects feeding on N14 plants expressing the susceptible BPH14 protein form N14 (Fig. 1b and Extended Data Fig. 1b). N14 shares 83% sequence identity with BPH14 (ref. 8). As Bph14 encodes a typical NLR receptor8,19, a BPH effector protein may be recognized by BPH14 to activate Bph14-mediated resistance.
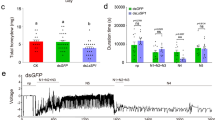
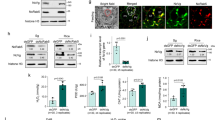
a, Phenotypes of rice plants carrying Bph14 and N14 after 7 days of BPH infestation. b, Weight gain and representative images of BPH insects feeding on Bph14 and N14 plants for 2 days (n = 20, biologically independent samples). The box limits indicate the 25th and 75th percentiles, the whiskers indicate the full range of the data and the centre line indicates the median. Individual data points are plotted. c–e, BISP interacted with BPH14 in Y2H (c), co-IP (d) and BiFC (e) assays. N14 and BISP(125–241) served as negative controls. DDO, SD/-Leu-Trp; QDO, SD/-Leu-Trp-His-Ade. bZIP63–CFP, nuclear marker. f, Quantification of relative YFP intensities in BiFC assays. Data are the mean ± s.d. (n = numbers of biologically independent cells). g, Amino acid sequence of BISP. The asterisk indicates the stop codon. Glycine residues are marked in red. The 13-amino acid and 4-amino acid tandem repeats are underlined in blue and black, respectively. h,i, Immunohistochemical localization of BISP in female BPH salivary glands using pre-immune rabbit serum (h) or anti-BISP antibodies (i). Red fluorescence (Cy3) and blue fluorescence show the localization of BISP and DAPI-stained nuclei, respectively. j, BISP was delivered into rice leaf sheaths during BPH infestation. Leaf sheath proteins were analysed using anti-BISP antibodies. Ponceau S staining served as the loading control. k, Immunohistochemical staining showing BISP in BPH-infested rice leaf sheaths. The non-infected (middle) and BPH-infested (bottom) sheaths were detected by anti-BISP antibodies and pre-immune rabbit serum, respectively, served as negative controls. In b and f, P values were derived by one-way analysis of variance (ANOVA). Experiments (a–e,h–k) were repeated at least three times, each giving similar results. The results of the other two repeats are presented in Supplementary Fig. 2. Scale bars, 5 μm (e), 25 μm (k), 100 μm (h,i) or 10 cm (a).
Planthoppers secrete salivary proteins into rice plants that may induce or suppress host defence responses during feeding1,4,5,16. Yeast two-hybrid (Y2H) screens identified BPH-secreted proteins (effectors) that interacted with BPH14. Of the 12 genes identified, one encoded the Bisp transcript that was abundant in the transcriptome of the BPH salivary gland (Supplementary Table 2). BISP interacted with BPH14, but not N14, based on repeated Y2H and co-immunoprecipitation (co-IP) experiments (Fig. 1c,d). Expression of BISP tagged with green fluorescent protein (BISP–GFP) in rice protoplasts showed that it localized to the nucleocytoplasm (Extended Data Fig. 1c,d), which is consistent with the subcellular localization of BPH14 (ref. 19). In addition, bimolecular fluorescence complementation (BiFC) confirmed the interaction between BISP and BPH14 and their nucleocytoplasmic co-localization in rice protoplasts (Fig. 1e,f and Extended Data Fig. 1e).
Bisp (LOC111051577) resides on chromosome 12 of BPH and encodes a 241-amino acid protein with an amino-terminal signal peptide (amino acids 1–25) and no transmembrane domain (Fig. 1g), which suggests that BISP is a secretory protein. Rich in glycine residues (32%), BISP has three 13-amino acid (GLADVGGLTNTVS) and 4-amino acid (CGGN/R) tandem repeats in its carboxy-terminal region (Fig. 1g). Quantitative PCR with reverse transcription (RT–qPCR) confirmed that Bisp is highly expressed in salivary glands and in whole female adults (Extended Data Fig. 1f,g). Immunohistochemical analysis using BISP antiserum revealed that BISP accumulated to higher levels in female salivary glands than in male salivary glands and guts (Fig. 1h,i and Extended Data Fig. 1h–m,p), a result that was consistent with Bisp RNA levels. Moreover, BISP signals were higher than the salivary protein NlSP1 (Extended Data Fig. 1n–p).
BPHs were allowed to feed on rice plants to determine whether BISP is secreted into rice leaf sheaths. Immunoblots detected BISP in leaf sheath protein extracts from BPH-infested but not non-infested plants (Fig. 1j). Immunolocalization assays verified the delivery of BISP into rice leaves. BISP was detected along the penetration path of stylets within the leaf sheaths (Fig. 1k). Immunogold electron microscopy further confirmed that BISP was secreted into rice tissues during BPH feeding (Extended Data Fig. 1q–t).
BISP suppresses plant defence
To determine whether BISP influenced the success of BPH on rice, we disrupted Bisp expression by microinjecting Bisp double-stranded RNAs (dsRNAs) into individual BPHs to induce RNA-mediated interference (RNAi). Bisp transcripts and proteins were significantly reduced in BPHs that received Bisp-RNAi compared with those that received GFP-RNAi (Extended Data Fig. 2a). Bisp-RNAi insects exhibited lower weight gain and honeydew excretion (Extended Data Fig. 2b,c). These insects also experienced significantly higher mortality when fed on the susceptible N14 rice plants than BPHs that received no injection (controls) or GFP-RNAi (Extended Data Fig. 2d). Therefore, Bisp plays a crucial role in the feeding and performance of BPHs on susceptible rice plants.
To determine the function of BISP in rice, we ectopically expressed BISP (without its signal peptide) in BPH-susceptible N14 rice (N14–Bisp) plants (Fig. 2a). The stature of N14–Bisp plants was similar to control N14 plants (Extended Data Fig. 3a,b). However, N14–Bisp plants were hypersensitive to BPHs, displayed more severe symptoms after BPH feeding and died more quickly than the BPH-infested N14 plants (Fig. 2b,c). Moreover, the weight gain and honeydew excretion of BPHs feeding on N14–Bisp plants were significantly higher than those feeding on N14 plants (Extended Data Fig. 3c,d). A two-host choice test showed that BPHs preferentially settled on N14–Bisp plants rather than N14 plants (Extended Data Fig. 3e). These results show that BISP increases the susceptibility of rice to BPHs.
a, Immunoblot detection of BISP in N14 and N14–Bisp transgenic rice lines. b,c, Phenotypes (b) and BPH resistance scores (c) of N14 and N14–Bisp plants after 4 days of BPH infestation. n = 36 plants examined over 3 independent experiments. d, Endogenous free SA levels in N14–Bisp and N14 plants. FW, fresh weight. Data are the mean ± s.d. (n = 3, biologically independent experiments). e, BISP interacted with OsRLCK185 and its kinase domain in a Y2H assay. OsRLCK185(1–85), OsRLCK185KD, OsRLCK185(354–491), OsRLCK185 deletion mutants; KD, kinase domain (amino acids 86–353); TDO, SD/-Leu-Trp-His with 1.5 mM 3-AT (3-amino-1,2,4-triazole). f, Co-IP showing the interaction between BISP and OsRLCK185. BISP(26–124) and OsRLCK159 served as negative controls. g, OsRLCK185 autophosphorylation activity was reduced by BISP. BISP(26–124)–GST–His proteins served as the negative control. Kinase activity was detected by autoradiography and input proteins shown after Coomassie Brilliant Blue staining (CBB). h,i, Phenotypes (h) and BPH-resistance scores (i) of Osrlck185 and WT plants after 4 days of BPH infestation. n = 45 plants examined over 3 independent experiments. In box plots in c and i, the box limits indicate the 25th and 75th percentiles, the whiskers indicate the full range of the data and the centre line indicates the median. Individual data points are plotted. In c,d and i, P values were derived by one-way ANOVA. The experiments (a,b,d–h) were repeated at least three times, each giving similar results. Scale bars, 10 cm (b,h).
We reasoned that the BISP effector was acting as a virulence factor that suppresses rice defence responses, which makes susceptible plants more vulnerable to BPHs. The endogenous levels of free salicylic acid (SA), a plant hormone vital for BPH resistance in rice10,11,19, were lower in N14–Bisp plants than in N14 plants (Fig. 2d). Similarly, the levels of SA biosynthesis and signalling genes (OsICS1 and OsNPR1) and downstream defence-related genes (OsPR1a, OsPR1b, OsPR5 and OsPR10) transcripts were all significantly lower in N14–Bisp plants than in N14 plants (Extended Data Fig. 3f).
Receptor-like kinases (RLKs) play crucial roles in plant defence against pathogens and herbivores1,20. We aimed to identify the target of BISP and to elucidate the molecular mechanism that underlies its ability to suppress defence responses in susceptible plants. To that end, we examined the interactions of RLKs, receptor-like cytoplasmic kinases (RLCKs) and MAP kinases with BISP in Y2H assays (Fig. 2e and Extended Data Fig. 4a–c). Among the eight kinases tested, only OsRLCK185 interacted with BISP. BISP interacted with the OsRLCK185 kinase domain (86–353 amino acids). The association of OsRLCK185 with BISP was confirmed by co-IP in rice protoplasts (Fig. 2f). As OsRLCK185 regulates plant immunity through autophosphorylation21,22, an in vitro phosphorylation assay and immunoblotting with phosphoserine/phosphothreonine-specific antibodies confirmed that OsRLCK185 was an active kinase with autophosphorylation activity (Extended Data Fig. 4d). Furthermore, OsRLCK185 autophosphorylation was attenuated by BISP (Fig. 2g and Extended Data Fig. 4d).
We used three OsRLCK185 knockout rice lines (Osrlck185-1–Osrlck185-3 plants) to further explore the roles of OsRLCK185 in rice defence against BPHs (Extended Data Fig. 4e). Osrlck185 lines were more susceptible to BPHs, as they were severely damaged compared with wild-type (WT) plants 4 days after BPH infestation (Fig. 2h,i). BPHs that fed on the Osrlck185 lines gained significantly more weight and excreted significantly more honeydew than those fed on WT plants (Extended Data Fig. 4f,g), which indicated that OsRLCK185 positively regulates basal immunity in BPH-susceptible plants.
To further assess the function of BISP in BPH, we analysed the performance of BPHs treated with Bisp-RNAi or GFP-RNAi on Osrlck185-1 and WT plants. Bisp-RNAi insects showed significantly lower weight gain, honeydew excretion and survival rate than GFP-RNAi insects on Osrlck185-1 plants (Extended Data Fig. 2e–g). This result suggests that BISP has a specific function related to OsRLCK185 and may target other defence-related proteins in rice.
BISP activates BPH14 resistance
Pathogen effectors that are directly or indirectly recognized by NLR proteins trigger ETI13,14. Y2H and co-IP assays showed that BISP did not interact with either the CC or NB domain, but interacted with the LRR domain of BPH14 (Fig. 3a,b). Furthermore, the N terminus of BISP (amino acids 26–124) interacted with the LRR domain (Fig. 3c and Extended Data Fig. 5a). Biolayer interferometry (BLI) assays using the BPH14–LRR or N14–LRR domains and BISP purified from insect cells showed that BPH14–LRR bound to BISP with a dissociation constant (Kd) of 3.21 × 10−8 M (Fig. 3d) and N14–LRR binding to BISP was not detected (Extended Data Fig. 5b). BPH14–LRR also bound to BISP(26–124) with a Kd of 1.27 × 10−7 M (Extended Data Fig. 5c), but not bind to BISP(125–241) (Extended Data Fig. 5d). Binding competition experiments showed that BISP competed with BISP(26–124) for binding to BPH14–LRR (Fig. 3e). Microscale thermophoresis (MST) assays confirmed that BISP directly bound to BPH14 (Extended Data Fig. 5e,f).
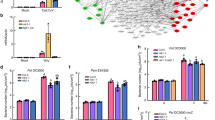
a,b, BISP interacted with the LRR domain of BPH14 in Y2H (a) and co-IP (b) assays. N14–LRR and BISP(125–241) served as negative controls. c, BISP(26–124) interacted with BPH14 and its LRR domain in a co-IP assay. N14–LRR served as a negative control. d, BLI analysis for binding kinetics between LRR (BPH14–LRR) and BISP. BLI response profile for BISP at different concentrations with sensor-immobilized LRR. Kd, equilibrium dissociation constant; Kon, association rate constant; Koff, dissociation rate constant. e, BLI analysis for competitive binding between BISP and BISP(26–124) with LRR. f, BISP increased the levels of BPH14 homomeric complex in a co-IP assay in rice protoplasts. BISP(125–241) and N14 served as negative controls. Numbers above the lanes indicate band intensity relative to co-precipitated BPH14–MYC, quantified using ImageJ. g, Co-expression of BPH14 and BISP increased WRKY72 levels. h,i, Protein (h) and relative transcript (i) levels of Wrky72 were increased in non-infested Bph14–Bisp lines compared with Bph14 and MH63 plants. j, Endogenous free SA levels in non-infested Bph14–Bisp and Bph14 plants. k,l, Phenotypes (k) and BPH-resistance scores (l) of Bph14–Bisp and Bph14 plants after BPH infestation for 14 days. n = 36 plants examined over 3 independent experiments. m, Weight gain of BPHs feeding on Bph14–Bisp and Bph14 plants for 48 h (n = 22, biologically independent samples). n,o, Photographs (n) and plant heights (o, n = 21 plants) of non-infested Bph14–Bisp and Bph14 plants at the 4-leaf stage. In box plots in l, m and o, the box limits indicate the 25th and 75th percentiles, the whiskers indicate the full range of the data and the centre line indicates the median. Individual data points are plotted. In g and h, numbers above/under the lanes indicate band intensity relative to actin (loading control) quantified using ImageJ. In h, i and j, data are the mean ± s.d. (n = 3, biologically independent experiments). In i, j, l, m and o, P values were derived by one-way ANOVA. Similar results were obtained from two (d,e) or three (a–c,f–h,k,m–o) independently replicated experiments. Scale bars, 10 cm (k,n).
BPH14 forms a homomeric complex and interacts with WRKY72 to increase WRKY72 accumulation, which in turn activates downstream defence signalling19. To examine the effects of BISP on the formation of BPH14 complexes and on WRKY72 protein levels, we co-expressed BPH14 and BISP in rice protoplasts. The levels of BPH14 self-association (Fig. 3f) and WRKY72 levels (Fig. 3g) were increased when BPH14 and BISP were co-expressed in rice protoplasts, which suggested that BISP stimulated the activity of BPH14.
To examine the activation of BPH14 by BISP, we constructed transgenic Bph14–Bisp rice lines in which Bisp was constitutively expressed in the Bph14 background (Extended Data Fig. 5g). WRKY72 protein and transcript levels were higher in three homozygous Bph14–Bisp plants relative to Bph14 plants (Fig. 3h,i). In addition, non-infested Bph14–Bisp plants had higher levels of free SA than Bph14 plants (Fig. 3j). Furthermore, the SA-responsive OsICS1 and OsNPR1 and four rice PR genes were significantly upregulated (Extended Data Fig. 5h). The intensity of Bph14-mediated resistance responses in the Bph14–Bisp plants positively correlated with the levels of BISP protein.
We evaluated BPH resistance in Bph14–Bisp plants. Whereas Bph14 plants showed little damage after 7 days of infestation (Fig. 1a), prolonged BPH feeding (14 days) caused substantial damage. By contrast, prolonged BPH feeding caused little damage to Bph14–Bisp plants (Fig. 3k,l), which suggested that ectopic expression of Bisp in Bph14 plants enhances resistance to BPHs. In addition, BPH adults preferred Bph14 over Bph14–Bisp plants in two-host choice assays and had poor performance on Bph14–Bisp plants based on BPH weight gain, honeydew excretion and survival (Fig. 3m and Extended Data Fig. 5i–k).
We further tested the performance of Bisp-RNAi and GFP-RNAi insects on Bph14 plants and susceptible control MH63 plants (Extended Data Fig. 2h–j). The GFP-RNAi and non-injected insects exhibited significantly decreased weight gain, honeydew excretion and survival on Bph14 plants compared with those on MH63 plants. By contrast, the Bisp-RNAi insects showed similar weight gain, honeydew excretion and survival rate when fed on Bph14 or MH63 plants. Moreover, the Bisp-RNAi insects exhibited more weight gain, honeydew excretion and a higher survival rate than GFP-RNAi insects on Bph14 plants. These results show that the knockdown of Bisp improved BPH performance on Bph14 plants.
Constitutive activation of immune responses to pathogens usually negatively affects plant growth23,24. We observed that ectopic expression of Bisp in the Bph14 background had significant fitness costs. Bph14–Bisp plants were smaller than Bph14 plants at both the seedling and heading stages (Fig. 3n,o and Extended Data Fig. 6a,b). Furthermore, the heading dates of Bph14–Bisp plants were advanced (Extended Data Fig. 6c). Finally, Bph14–Bisp plants exhibited poor agronomic traits, with significantly lower yields than Bph14 plants (Extended Data Fig. 6d–j). The degree of fitness costs in Bph14–Bisp plants positively correlated with the levels of BISP and the intensity of Bph14-mediated resistance responses (Fig. 3h–m and Extended Data Figs. 5g–k and 6). These results indicate that constitutively expressed BISP induces BPH14-mediated resistance in Bph14–Bisp plants, which has a substantial fitness cost in rice.
BPH14 mediates BISP turnover by autophagy
The fitness costs imposed by ectopic expression of BISP in Bph14 plants suggested that the activation of Bph14-mediated resistance should be tightly controlled in the natural habitats of rice. Several lines of evidence support this hypothesis. First, when BISP is expressed in rice protoplasts, BISP levels were lower when BISP was co-expressed with BPH14 (Fig. 4a), and BISP levels negatively correlated with BPH14 levels (Extended Data Fig. 7a). Indicative of the specificity of the BISP–BPH14 interaction, BISP levels were not reduced when BISP was co-expressed with N14 (Fig. 4a). These results indicate that BISP is degraded in a BPH14-dependent manner.
a, Effects of BPH14 and N14 on BISP levels in rice protoplasts. b, Effects of the 26S proteasome inhibitor MG132 on BISP levels in rice protoplasts. c, Effects of autophagy inhibitors on BISP protein levels in rice protoplasts. 3-MA, 3-methyladenine; CQ, chloroquine; E-64d, aloxistatin; LQ, leupeptin. DMSO was the solvent for all inhibitors. d, Transmission electron microscopy images of autophagic structures in the phloem of non-infested Bph14–Bisp plants. Insets show enlarged autophagosome image at higher magnification. Red arrows indicate the location of double-membrane autophagosomes. CC, companion cell; SE, sieve element cell; V, vacuole. Scale bars, 500 nm. e, Quantification of double-membraned autophagosomes. P values were derived by one-way ANOVA. Data are the mean ± s.e.m. (n = 5, biologically independent experiments with every 8 cells as a biological replicate). f, Immunoblot detection of BISP, OsATG8 and OsNBR1 in Bph14 and MH63 plants. g, Y2H assay of the interactions between BISP, BPH14, OsNBR1 and four OsATG8 proteins. h, Co-IP assays of interactions between OsNBR1 and BISP or BPH14 in rice protoplasts. i, Co-IP assay of interactions between OsNBR1 and four OsATG8 proteins in rice protoplasts. The OsNBR1 mutant N1 served as a negative control. j, Effects of OsNBR1 and BPH14 on BISP levels in rice protoplasts. N14 and N1 served as negative controls. k, BPH14 enhanced interactions between OsNBR1 and BISP in rice protoplasts. N14 served as a negative control. Numbers under the lanes indicate band intensity relative to co-precipitated OsNBR1–MYC, quantified using ImageJ. l, Immunoblot detection of BISP in protoplasts of MH63, Bph14 and OsNBR1 knockout (Osnbr1) plants. m, Immunoblot detection of BISP in MH63, Bph14 and Osnbr1 plants after BPH infestation. Numbers under the lanes (a–c,f,j,l,m) indicate protein abundance relative to that of actin (loading control), quantified using ImageJ. Experiments (a–d,f–m) were repeated three times and gave similar results.
The ubiquitin–proteasome system and autophagy are the main protein-degradation pathways in plant cells25. The ubiquitin–proteasome system is a key post-translational mechanism for controlling plant immune responses to pathogens26. To test whether the ubiquitin–proteasome system is involved in the modulation of BISP–BPH14 immune activation, the 26S proteasome inhibitor MG132 was added to protoplasts expressing BISP and BPH14. MG132 blocked the proteasome pathway degradation of WRKY72 (Extended Data Fig. 7b), but did not inhibit the degradation of BISP (Fig. 4b). By contrast, treatment with four autophagy inhibitors blocked the degradation of BISP (Fig. 4c and Extended Data Fig. 7c), which suggested that the autophagy pathway is involved in the degradation of BISP.
To further assess the role of autophagy in BISP turnover, we used cyan fluorescent protein-tagged ATG8F (CFP–ATG8F) as a marker27,28 to monitor autophagy (Extended Data Fig. 7d,e). When CFP–ATG8F was expressed with BISP or BPH14 individually in Nicotiana benthamiana leaves (hereafter Nb is used to denote N. benthamiana-related proteins or genes), few punctate CFP structures, corresponding to pre-autophagosomes and autophagosomes, were detected. However, when CFP–ATG8F was expressed with BISP and BPH14, the number of punctate CFP structures significantly increased, which indicated that BISP and BPH14 co-expression induced autophagy. This interaction was specific, as expressing CFP–ATG8F with BISP and N14 did not increase the number of autophagosomes. Concanamycin A (ConA) blocks autophagic flux28, and the numbers of autophagic bodies were substantially increased in cells that expressed both BISP and BPH14 and treated with ConA. When we silenced the core autophagy genes NbATG6, NbPI3K and NbATG7, the numbers of autophagic bodies were markedly decreased in the silenced leaves that expressed CFP–ATG8F with BISP–yellow fluorescent protein (YFP) and BPH14 (Extended Data Fig. 7f–h). Furthermore, using transmission electron microscopy, we observed more double-membraned autophagosome structures in the phloem companion cells of non-infested Bph14–Bisp plants than in Bph14 and MH63 plants (Fig. 4d,e). Accordingly, OsATG8 protein levels and OsATG8a, OsATG8b and OsATG8c transcript levels were significantly higher in non-infested Bph14–Bisp plants than in Bph14 and MH63 plants (Extended Data Fig. 7i,j). These observations indicate that BISP is degraded through autophagy in a BPH14-dependent manner.
As BPHs secrete saliva containing BISP into rice tissues when feeding (Fig. 1j,k and Extended Data Fig. 1q–t), we monitored BISP levels in infested BPH-susceptible (MH63) or BPH-resistant (Bph14) plants. In MH63 plants, BISP was first detected at 6 h and increased continuously for 72 h of BPH feeding (Fig. 4f). By contrast, BISP levels were lower in Bph14 plants, with levels sustained after 24 h of feeding (Fig. 4f). OsATG8 levels did not significantly change in BPH-infested susceptible plants. By contrast, in Bph14 plants, OsATG8 levels increased by 6 h after feeding and were maintained for 72 h (Fig. 4f). These results suggest that autophagy is activated in Bph14 plants but not in BPH-susceptible rice. These data correlated well with the upregulation of three OsATG8 genes in Bph14 plants compared with MH63 plants following BPH infestation (Extended Data Fig. 7k). ConA treatment substantially increased levels of ATG8 proteins in BPH-infested Bph14 plants (Extended Data Fig. 7l). These results indicate that BPH feeding activates autophagy and that BISP levels are tightly controlled in Bph14 plants.
OsNBR1-mediated autophagy controls BISP levels
Mechanisms of autophagy are conserved in yeast, plants and metazoans29. In rice and other plants, multiple ATG8 proteins mediate autophagy30,31,32. To clarify the molecular mechanisms that underlie the BPH14-dependent degradation of BISP, we examined the interactions of four OsATG8 proteins with BISP or BPH14 using Y2H assays. Neither BISP nor BPH14 interacted with the OsATG8 proteins (Fig. 4g and Extended Data Fig. 8a).
In addition to canonical autophagy, selective autophagy targets specific cargoes, such as intracellular pathogens and defective proteins, and plays pivotal roles in eukaryotic organisms33,34,35. NBR1 is a selective autophagy cargo receptor and interacts with ATG8 proteins to facilitate the autophagy-dependent degradation of target proteins33,34,35,36,37. We therefore examined the interaction of BISP or BPH14 with OsNBR1 in Y2H assays. Both BISP and BPH14 interacted with OsNBR1 (Fig. 4g and Extended Data Fig. 8a). Co-IP assays in rice protoplasts confirmed these interactions (Fig. 4h). Furthermore, OsNBR1 interacted with all four OsATG8 proteins in Y2H assays (Fig. 4g and Extended Data Fig. 8a) and in rice protoplasts (Fig. 4i), which suggested that OsNBR1 might mediate the degradation of BISP.
The roles of OsNBR1 in the autophagic degradation of BISP were examined by immunoblotting. The expression of BISP with OsNBR1 in rice protoplasts did not influence BISP levels. By contrast, the expression of BISP, OsNBR1 and BPH14 reduced the level of BISP (Fig. 4j), which suggested that OsNBR1 facilitated BPH14-dependent degradation of BISP. We used CFP–ATG8F to determine the effects of OsNBR1 on the numbers of autophagic vesicles. The expression of OsNBR1 with BISP and BPH14 induced more punctuate CFP–ATG8F structures in N. benthamiana leaves than expression of only BISP and BPH14 (Extended Data Fig. 8b,c). Additionally, OsNBR1 protein and transcript levels were significantly higher in non-infested Bph14–Bisp plants than in Bph14 and MH63 plants (Extended Data Fig. 8d,e). Following BPH infestation, OsNBR1 protein and RNA levels were increased in Bph14 plants, but not in BPH-susceptible MH63 plants (Fig. 4f and Extended Data Fig. 8f). Moreover, ConA treatment significantly increased OsNBR1 levels in Bph14 plants following BPH infestation (Extended Data Fig. 7l). Y2H and co-IP assays showed that the CC and LRR domains of BPH14 interacted with OsNBR1 (Extended Data Fig. 9a,b). Co-IP assays also showed that BPH14 enhanced the interaction of OsNBR1 with BISP (Fig. 4k).
NBR1 homopolymerizes through its PB1 domain, binds to ubiquitin through its UBA domain and interacts with ATG8 through a conserved LIR motif to promote selective autophagy37. OsNBR1 contains these interaction domains, as well as conserved ZZ-type zinc finger and BRCA1 domains. To determine which OsNBR1 domain (or domains) participates in the degradation of BISP, we generated four truncated forms (N1 to N4) and three site-directed mutants (N5 to N7) of OsNBR1 (Extended Data Fig. 9c). Co-expression assays showed that none of mutants were able to degrade BISP (Extended Data Fig. 9d). Whereas the Osnbr1 N5 mutant lost the ability to form homopolymers (Extended Data Fig. 9h), the other Osnbr1 mutants retained the ability to form homopolymers. However, these mutants failed to interact with BPH14, BISP or OsATG8C (Extended Data Fig. 9e–i). These results suggest that OsNBR1 is involved in BISP degradation through autophagy and that domains outside the UBA in OsNBR1 are important for interacting with its target BISP.
We generated OsNBR1 knockout mutants in the Bph14 background (Bph14–Osnbr1) to further analyse the roles of OsNBR1 in the degradation of BISP (Extended Data Fig. 9j). OsNBR1 was undetectable in the Bph14–Osnbr1 mutants (Extended Data Fig. 9k). We then expressed BISP and BPH14 in Bph14–Osnbr1, Bph14 and MH63 protoplasts. Consistent with a role for OsNBR1 in BISP turnover, BISP levels were higher in Bph14–Osnbr1 protoplasts than in Bph14 and MH63 protoplasts (Fig. 4l). BISP was more abundant in infested Bph14–Osnbr1 plants than in Bph14 plants, but less abundant than in MH63 plants (Fig. 4m). These results indicate that OsNBR1-mediated autophagic degradation controls BISP levels.
BISP degradation attenuates resistance
On susceptible plants, BPHs consume phloem sap for prolonged times17. By contrast, on BPH-resistant plants, various resistance factors prevent sustained phloem ingestion11,12,16,17,38. BPHs have a higher frequency of stylet probing, penetrating numerous sites in an attempt to find an acceptable feeding site17,39.
To provide insights into the duration of Bph14-mediated immunity, we assessed the longevity of BISP and the immune status of rice plants after BPH feeding had ceased. To this end, BPH-resistant Bph14 plants and BPH-susceptible MH63 plants were infested with BPHs for 24 h. At this time, insects were removed, and BISP, OsATG8, OsNBR1 and WRKY72 levels were monitored for 48 h. In MH63 plants, BISP levels were similar at 0 and 36 h after BPH removal. By contrast, in Bph14 plants, BISP levels decreased markedly at 6 h and were barely detected at 36 h (Fig. 5a). Correlating well with the decreased BISP levels, the levels of OsATG8 and OsNBR1 proteins and their transcripts were more abundant in Bph14 plants than in MH63 plants (Fig. 5a and Extended Data Fig. 10a). These results suggest that after BPH feeding and secretion of saliva have ceased, BISP is stable for prolonged times in susceptible rice but is rapidly eliminated through autophagy in Bph14 plants.
a, Immunoblot detection of BISP, OsATG8, OsNBR1, and WRKY72 in Bph14 and MH63 plants after cessation of BPH feeding. b, Relative Wrky72 transcript levels in Bph14 and MH63 plants after cessation of BPH feeding. c, Co-IP detection of levels of BPH14 homomeric complex formation. N1 served as a negative control. Numbers above the lanes indicate band intensity relative to co-precipitated BPH14–MYC, quantified using ImageJ. d, Immunoblot detection of BISP and WRKY72 in Bph14 and Bph14–Osnbr1 plants after cessation of BPH feeding. e, Relative Wrky72 transcript levels in Bph14 and Bph14–Osnbr1 plants after cessation of BPH feeding. f,g, Photographs (f) and resistance scores (g) of Bph14 and Bph14–Osnbr1 plants after 14 days of BPH infestation. Scale bar, 10 cm. The box limits indicate the 25th and 75th percentiles, the whiskers indicate the full range of the data and the centre line indicates the median. Individual data points are plotted (n = 42 plants examined over 3 independent experiments). h, A working model for BISP-induced immunity regulation. In BPH-susceptible rice, BISP interacts with OsRLCK185 and suppresses its phosphorylation activity, thereby inhibiting basal defence and promoting BPH feeding. In rice carrying the BPH resistance gene Bph14, BISP binds directly to the LRR domain of BPH14 and activates HPR. Continuous activation of HPR is detrimental to plant growth and reproduction. Therefore, following activation of BPH14-mediated immunity, BISP–BPH14 binds to OsNBR1, which mediates the autophagic degradation of BISP. The degradation of BISP leads to the termination of BPH14-mediated immunity, which restores cellular homeostasis and prevents the fitness costs associated with prolonged activation of BPH14-mediated immunity. The timing of dissociation of the BISP–BPH14 complex after association with OsNBR1 and the fate of BPH14 are yet to be determined. Numbers under lanes (a,d) indicate protein abundance relative to that of actin (loading control, a,c,d), quantified using ImageJ. In b, c, e and g, P values were derived by one-way ANOVA. In b and e, data are the mean ± s.d. (n = 3, biologically independent experiments). Experiments (a,c,d,f) were repeated three times, each giving similar results. PRRs, pattern recognition receptors.
Moreover, the degradation of BISP was accompanied by a mitigation of the signal outputs of BPH14-activated resistance (Fig. 5a,b and Extended Data Fig. 10b). For example, WRKY72 protein and transcript levels rapidly reduced (within 12 h) in Bph14 plants and were slower in susceptible plants after BPH feeding had ceased (Fig. 5a,b). In addition, OsICS1 and OsNPR1 transcripts showed a similar regulation pattern (Extended Data Fig. 10b). The rapid dissipation of BPH14-regulated immunity based on BISP depletion was supported by the dynamics of formation of the BPH14 homomeric complex, which was reduced when BPH14 was expressed with BISP and OsNBR1 to induce autophagy (Fig. 5c). These results suggest that autophagy contributes to the elimination of BISP and the attenuation of resistance in Bph14 plants after BPH feeding is terminated.
Further support for this theory was garnered from examining the fate of BISP and defence-signalling components in Bph14–Osnbr1 plants and WT Bph14 plants after BPH feeding was terminated. BISP levels were significantly higher in Bph14–Osnbr1 plants than in Bph14 plants at 0, 24 and 48 h after removing the insects (Fig. 5d), which confirmed that OsNBR1 deficiency slowed the degradation of BISP. WRKY72 protein and transcript levels, as well as Osics1 and Osnpr1 transcripts, were significantly higher at 0–48 h in the Bph14–Osnbr1 plants than the Bph14 plants (Fig. 5d,e and Extended Data Fig. 10c). This result suggests that the time span of BPH14-activated signalling is extended in Bph14–Osnbr1 plants.
Finally, we examined the impact of OsNBR1 on rice resistance to BPHs. Whereas Bph14 plants were damaged by prolonged BPH infestation (14 days), Bph14–Osnbr1 plants had limited symptoms and displayed a higher level of BPH resistance (Fig. 5f,g). BPH adults preferred Bph14 plants to Bph14–Osnbr1 plants in a two-host choice test. On the basis of honeydew excretion, weight gain and survival, BPHs had poorer performance when feeding on Bph14–Osnbr1 plants (Extended Data Fig. 10d–g). These results indicate that OsNBR1 deficiency enhances BPH resistance in Bph14 plants.
Discussion
HPR conferred by NLR proteins is widely used for controlling insect pests40. Here we provide molecular and cellular insights into the long-standing question of how plant NLR receptors perceive insect-feeding signals to deploy and modulate ETI resistance mediated by these proteins. Notably, we discovered that the tripartite interactions of BISP, its cognate NLR receptor BPH14 and the selective-autophagy receptor NBR1 enable a self-regulated resistance mechanism to BPHs (Fig. 5h).
In this tripartite interaction system, the BISP effector is secreted into rice leaf sheaths to bind to one of its targets, OsRLCK185. By interfering with OsRLCK185 autophosphorylation, BISP suppresses rice immune responses to promote BPH feeding and success. However, in resistant rice, the NLR receptor BPH14 directly binds to BISP. Following recognition, resistance is activated, which stops BPHs from ingesting phloem sap8,17,19. Constitutive activation of BISP-triggered resistance in Bph14 plants is detrimental to plant fitness, reducing plant stature and yields. Therefore, mechanisms to temper the resistance response must be deployed to promote plant vitality. The importance of the ubiquitin–proteasome system in effector and NLR protein turnover in pathogen–plant ETI is known26,41. Here we revealed that the governor of Bph14-mediated resistance is OsNBR1-regulated autophagy, which controls BISP turnover. Both BISP and BPH14 physically interact with OsNBR1. In turn, OsNBR1 interacts with OsATG8 to promote the degradation of BISP and, thereby, fine-tune the level of Bph14-mediated resistance. Whether the BISP–BPH14 complex dissociates before BISP turnover by autophagy or whether BPH14 is also degraded by autophagy is an essential topic of research in the future. Finally, although autophagy can be induced by transient reprogramming in response to plant hormones42, the role of SA, which is induced in the BISP–BPH14 interaction, remains to be elucidated.
The BISP–BPH14–OsNBR1 resistance response adapts to changes in BPH feeding pressure. BPHs terminate feeding when a host becomes undesirable, which can induce individual insects or entire BPH populations to migrate long distances in search of better host plants15,16,17,18. When BPHs feed on Bph14 plants, BISP triggers resistance, which makes Bph14 plants unpalatable and forces the insects to stop feeding. With the cessation of feeding and the departure of insects from these sites, the activated autophagy pathway rapidly degrades BISP to restore cellular homeostasis. Immune systems resume their ‘off’ status, thereby allowing rice to reallocate resources to growth and reproduction. The newly described BISP–BPH14–OsNBR1 tripartite interaction system enables rice to resist BPHs without compromising yield performance in the natural habitats43. By leveraging the autoregulatory mechanisms used in Bph14 gene-for-gene resistance in new ways, we have the ability to create a new generation of resistant rice that can feed the increasing world population in a sustainable manner and, significantly, to control insect pests without relying on insecticide sprays.
Methods
Plant materials and growth conditions
Nipponbare (designated as N14) is a model O. sativa japonica rice variety that contains the N14-susceptible allele of Bph14. N14 was used in early experiments to establish the principles of BISP–BPH14 interactions. The RI35 line (designated as Bph14) is a recombinant inbred line that contains the BPH resistance gene Bph14 (ref. 8). Minghui 63 (MH63) is the BPH-susceptible parent of RI35 (ref. 44) and is a model variety for O. sativa indica rice breeding and genomics. MH63 and BPH14 were used for most studies to elucidate BISP–BPH14–NBR1 interactions owing to the agronomic importance of MH63. The following transgenic lines were developed in this study: N14–Bisp (pUBI::Bisp-Myc, N14 recipient), Bph14–Bisp (pUBI::Bisp–Myc, Bph14 recipient), Osrlck185 (OsRLCK185–Cas9, ZH11 recipient), Bph14–Osnbr1 (OsNBR1–Cas9, Bph14 recipient). The Osrlck185 mutants were generated in the ZH11 background45; ZH11 is a readily transformable O. sativa japonica rice that BPH-susceptible genotype.
Insect materials and growth conditions
The BPH insects were maintained on susceptible cultivar Taichung Native 1 at 28 ± 2 °C with 60–80% relative humidity and a photoperiod of 16 h of light–8 h of dark. RNAi of BPH by dsRNA injection was developed as part of this study: GFP-RNAi (microinjected with dsGFP) and Bisp-RNAi (microinjected with dsBisp).
BPH bioassays on rice plants
To evaluate BPH resistance of transgenic and WT rice plants, sets of around 15 or 20 seeds from the same plant were sown in a plastic cup (10 cm in diameter, 20 cm in height). At the three-leaf stage, all plants were placed under a large gauze cover, and each seedling was infested with ten second- or third-instar BPH nymphs. When all of the susceptible plants had died (scored as 9), each seedling of the other cultivars or lines was given a score of 0, 1, 3, 5, 7, or 9 according to the degree of damage, as previously described8,46,47, and the BPH resistance score was calculated. At least three replicates were used for each cultivar or line.
To measure the weight gain and honeydew excretion of BPHs, a newly emerged short-winged female adult and a Parafilm sachet were weighed using a 1/100,000 Shimadzu AUW120D electronic balance11,12. The insect was placed in the pre-weighed sachet, which was then attached to the leaf sheath near the lower part of a rice plant. After 48 h, the insect and Parafilm sachet were weighed again. The difference between the first and second measurements of the weight of the insect was recorded as the BPH weight gain; the difference between the first and second measurements of the weight of the sachet was recorded as amount of honeydew excreted. At least ten replicates were used for each cultivar or line.
For the two-host choice tests, a transgenic plant and a WT plant were grown in a plastic cup as described elsewhere10,11,12. At the four-leaf stage, the cup was covered with gauze, and 20 or 30 second- or third-instar nymphs were released into the cup (the exact number depended on the BPH resistance of the WT plant). The number of nymphs that settled on each plant was recorded at 6, 12, 24, 48, 72, 96, and 120 h after release (the moment of release was considered time 0). Ten cups were analysed for each cultivar or line examined.
To study BPH nymph survival on rice, second- or third-stage BPH nymphs were released (ten insects per plant), and the cups were covered with a light-transmitting mesh. The nymphs on each plant were counted 8 or 9 days after release. Survival rates were calculated as the number of surviving nymphs divided by the total number of nymphs released at the start of the experiment. Ten cups were analysed for each cultivar or line.
Plasmid construction
Routine molecular cloning techniques were used to prepare the constructs. The plasmids and primers used in this work are listed in Supplementary Table 3. All of the resulting recombinant vectors were sequenced.
To construct the plasmid for Y2H screening, the full-length Bph14 sequence was amplified from Bph14 leaf sheath cDNA and cloned into the Xma I site of the pGBKT7 vector, which produced the BPH14–BD vector.
To construct the plasmids for the Y2H assays, the coding sequences of Bisp (ORF sequence without its 1–25 amino acid residue signal peptide; amino acids 26–241), Bisp26–124(amino acids 26–124) and Bisp125–241 (amino acids 125–241) were amplified and cloned into the Xma I site of the pGADT7 vector to produce BISP–AD, BISP(26–124)–AD and BISP(125–241)–AD, respectively. The coding sequences of the CC, NB and LRR domains and full-length Bph14 were amplified from the BPH14-BD vector and cloned into the Xma I sites of the pGADT7 and pGBKT7 vector to produce CC–AD, NB–AD, LRR–AD, BPH14–AD, CC–BD, NB–BD, and LRR–BD, respectively. The coding sequence of N14 (Bph14 allele in the rice variety Nipponbare) was amplified from N14 leaf sheath cDNA and cloned into the Xma I site of the pGBKT7 vector to produce N14–BD.
Eight kinases that were differentially phosphorylated during BPH-infested and non-infested plants were chosen for this study. The coding sequences for the OsRLCK185, OsRLCK1851–85 (amino acids 1–85), OsRLCK185KD (kinase domain, amino acids 86–353), OsRLCK185354–491 (amino acids 354–491), OsMAPKKKε, OsMKK6, OsMPK20, OsMPK16, OsCDPK14, OsCDPK20, and OsRLCK259 fragments were amplified from N14 leaf sheath cDNA and cloned into the pGBKT7 vector to produce OsRLCK185–BD, OsRLCK185(1–85)–BD, OsRLCK185KD–BD, OsRLCK185(354–491)–BD, OsMAPKKKε–BD, OsMKK6–BD, OsMPK20–BD, OsMPK16–BD, OsCDPK14–BD, OsCDPK20–BD, and OsRLCK259–BD, respectively. The coding sequences of OsNBR1, OsATG8a, OsATG8b, OsATG8c, and OsATG8e were amplified from Bph14 leaf sheath cDNA and cloned into pGBKT7 and pGADT7 to produce OsNBR1–BD, OsATG8A–BD, OsATG8B–BD, OsATG8C–BD, and OsATG8E–BD, respectively.
Constructs used for rice protoplast transfection were generated using pCXUN-4×HA and pCXUN-4×Myc19. The coding sequences of the CC, NB and LRR domains, Bph14, N14, the N14 LRR domain (N14–LRR), Wrky72, Bisp (ORF sequence without its signal peptide), Bisp26–124, Bisp125–241, OsRLCK185, OsRLCK259, OsNBR1, OsNBR1 mutant N1, OsATG8a, OsATG8b, OsATG8c, and OsATG8e were amplified and cloned into the pCXUN-4×Myc vector to produce CC–MYC, NB–MYC, LRR–MYC, BPH14–MYC, N14–MYC, N14–LRR–MYC, WRKY72–MYC, BISP–MYC, BISP(26–124)–MYC, BISP(125–241)–MYC, OsRLCK185–MYC, OsRLCK259–MYC, N1–MYC, OsNBR1–MYC, MYC–OsATG8A, MYC–OsATG8B, MYC–OsATG8C, and MYC–OsATG8E, respectively. Meanwhile, the fragments for the GFP, CC, NB, LRR domains, Bph14, N14, the N14 LRR domain (N14–LRR), Bisp, Bisp26–124, Bisp125–241, OsRLCK185, and OsNBR1 were amplified and cloned into the pCXUN-4×HA vector to produce GFP–HA, CC–HA, NB–HA, LRR–HA, BPH14–HA, N14–HA, N14–LRR–HA, BISP–HA, BISP(26–124)–HA, BISP(125–241)–HA, OsRLCK185–HA, and OsNBR1–HA, respectively. Additionally, we generated a series of Osnbr1 fragments, including the isolated N1 (amino acids 1–755), N2 (amino acids 756–832), N3 (amino acids 1–782), N4 (amino acids 1–755 and 798–832), and mutant N5 (K13A), N6 (WL788/791AA) and N7 (K13A and WL788/791AA) fragments and cloned them into the pCXUN-4×HA vector to produce N1–HA, N2–HA, N3–HA, N4–HA, N5–HA, N6–HA, and N7–HA, respectively.
To analyse the subcellular localization of BISP, the coding sequence of Bisp (ORF sequence without its signal peptide) was cloned downstream of the maize (Zea mays) Ubiquitin 1 promoter, in-frame with GFP in the binary vector pCAMBIA1300, which produced the BISP–GFP construct.
For BiFC analysis, the coding sequences of Bph14, N14, Bisp (ORF sequence without its signal peptide), and Bisp125–241 were cloned in-frame into the Xma I sites of the pUSYNE and pUSYCE vectors19, which produced the constructs BISP–YN, BISP(125–241)–YN, BPH14–YC, and N14–YC, respectively.
The Escherichia coli recombinant protein vectors used for the expression and purification of proteins in the phosphorylation activity assays, designated pET-MBP-His and pET-GST-His, were created by adding a C-terminal MBP or GST tag, respectively, to the pET-28a expression vector (EMD Biosciences, Novagen). The coding sequences of Bisp (ORF sequence without its signal peptide), Bisp26–124 and OsRLCK185 were amplified and cloned into the pET-MBP-His and pET-GST-His vector, which produced OsRLCK185–MBP–His, BISP(26–124)–GST–His and BISP–GST–His constructs, respectively.
For the N. benthamiana leaf agroinfiltration experiments, the coding sequences of Bph14, N14, OsNBR1, and N1 (an OsNBR1 mutant) were cloned into the pEarleyGate 203 vector48 to produce BPH14-203, N14-203, OsNBR1-203, and N1-203, respectively. The coding sequence of Bisp (ORF sequence without its signal peptide) was cloned into the pEarleyGate 201 and pEarleyGate 101 vector48 to produce BISP-201 and BISP-101 (BISP–YFP), respectively.
To generate the CRISPR–Cas9 construct, the target sites for CRISPR–Cas9 were designed using the web-based tool CRISPR-P (v.2.0; http://crispr.hzau.edu.cn/CRISPR2/). The CRISPR–Cas9 binary construct was prepared as previously described49, which produced the construct OsNBR1–Cas9.
Y2H screening
The Matchmaker Gold Yeast Two-Hybrid System (Clontech, 630489) was used to screen for BPH14-interacting proteins. Y2H Gold cells carrying BPH14–BD were mixed with 1 ml of a BPH salivary gland cDNA library and incubated overnight before plating on TDO (SD/-Leu-Trp-His)-selective medium. Candidate clones growing on TDO medium were confirmed following the manufacturer’s protocol (Clontech, 630489). The BPH cDNA library was constructed using the Make Your Own “Mate & Plate” Library System (Clontech, 630490) with a simple and highly efficient protocol. Total RNA was isolated from BPH salivary glands using TRIzol reagent (Takara, 9109), and cDNA was synthesized from the isolated mRNA using SMART cDNA synthesis technology (Clontech, 634926).
For the Y2H assay of protein interactions, the yeast strain AH109 was transformed with the indicated bait and prey plasmids according to the manufacturer’s instructions. Co-transformants were simultaneously plated on selection medium containing DDO (SD/-Leu-Trp), TDO (SD/-Leu-Trp-His) with the appropriate concentration of 3-amino-1,2,4-triazole (3-AT) or QDO (SD/-Leu-Trp-His-Ade) and incubated at 30 °C.
Cloning and sequencing of Bisp cDNA
The cDNA sequences of Bisp were obtained from BPH salivary gland transcriptomes5. To obtain the full-length counterparts of the truncated sequences in the transcriptomes, 5′- and 3′-RACE amplification was performed using a SMARTer RACE cDNA Amplification kit (Clontech, 634923) following the manufacturer’s instructions. Bisp-specific primers were obtained based on sequencing data. The RACE products were amplified, and the purified products were ligated into the pMD18-T vector (Takara, 6011) and sequenced.
Protein extraction, immunoblot analysis and in vivo co-IP assays
Proteins were extracted from the leaf sheaths of rice seedlings at the four-leaf stage. The leaf sheaths were wiped with a cotton ball soaked in alcohol to remove honeydew and ground in liquid nitrogen. Total protein was extracted from the leaf tissue in rice protein extraction buffer (50 mM Tris-HCl pH 7.5, 150 mM NaCl, 10% glycerol, 0.1% NP-40, plant protease inhibitor cocktail (Roche)). Equal amounts of total protein were analysed by SDS–PAGE and detected by immunoblotting using anti-actin (Abbkine, ABL1050; 1:3,000), anti-BISP (1:500), anti-OsWRKY72 (Beijing Protein Innovation, AbP80456-A-SE; 1:500) and anti-AtNBR1 (Agrisera, AS194281; 1:1,000) antibodies. For OsATG8 analysis, equal amounts of proteins were subjected to SDS–PAGE with 6 M urea50 and detected by immunoblotting using anti-AtATG8A antibodies (Abcam, ab77003; 1:1,000). The anti-BISP and anti-NISP1 antibodies were prepared by expressing BISP and NISP1 (cloned into pET28a) in E. coli strain BL21 (DE3), respectively. The expressed recombinant proteins were collected and injected into two rabbits (DIA·AA Biotech). Ponceau S solution (Sigma-Aldrich, P7170) was used to stain the PVDF membrane and was used as a loading control. The intensities of protein signals were quantified using ImageJ.
Protein samples from rice protoplasts were prepared in rice protein extraction buffer (100 mM Tris-HCl pH 7.5, 1 mM EDTA, 5 mM MgCl2, 0.5% (w/v) Triton X-100, with a plant protease inhibitor cocktail (Roche)). Total soluble proteins were extracted from rice protoplast samples, each in 100 µl of rice protoplast protein extraction buffer. Next, 10 µl of the extract was separated by SDS–PAGE and subjected to immunoblotting using anti-HA (MBL, M180-3; 1:1,000), anti-MYC (MBL, M192-3; 1:1,000) or anti-actin (Abbkine, ABL1050; 1:3,000) antibodies.
For the co-IP assays, rice protoplasts were incubated at 28 °C for 14–20 h after transfection. Total protein extracts were collected from the protoplasts in the rice protein extraction buffer as described above. The supernatants were incubated with 10 μl Protein G Agarose beads (Millipore, 16–266) and 1 μl anti-HA (MBL, M180-3) or 1 μl anti-MYC (MBL, M192-3) antibodies at 4 °C for 5 h, and detected using anti-HA or anti-HA mAb-HRP-DirecT (MBL, M180-7, clone: TANA2; 1:1,000), anti-actin and anti-MYC or anti-MYC mAb-HRP-DirecT (MBL, M192-7, clone: My3, 1:1,000) antibodies, respectively. The intensities of protein signals were quantified using ImageJ.
For the yeast protein immunoblots, total yeast protein was extracted as previously described51. In brief, single yeast colonies were resuspended in 4 ml YPDA liquid medium and grown to OD600 = 0.6 at 30 °C. The pelleted yeast cells were resuspended in 1 ml distilled water, combined with 100 μl 0.2 M NaOH, incubated for 5 min at room temperature and pelleted and resuspended in 50 μl loading buffer. The proteins were boiled for 10 min at 95 °C and detected by immunoblotting with anti-HA or anti-MYC antibodies. Ponceau S solution was used to stain the PVDF membrane for loading control.
Rice protoplast isolation and transfection
Rice protoplasts were prepared from etiolated seedlings and transfected as previously described11,19. About 3–10 μg of plasmid DNA was used to transfect around 2 × 105 protoplasts using the PEG-mediated method. The fluorescent or epitope-tagged proteins were detected at 14–20 h after transfection.
Subcellular localization and BiFC analysis
For subcellular localization, the BISP–GFP plasmid and the nuclear marker bZIP63–RFP11,19 were transformed into rice protoplasts. Following incubation at 28 °C for 14–20 h, the protoplasts were imaged using a confocal microscope (Leica, DMi8).
For BiFC analysis, protoplasts were transfected with the indicated expression vectors. Following incubation at 28 °C for 14–20 h, the protoplasts were imaged by confocal microscopy. BiFC fluorescent signals were quantified using previously described methods52. The intensity of YFP and CFP signals in individual protoplasts was determined using ImageJ software. The average YFP/CFP intensity ratio (per cent) of the whole cell was measured, and the means were calculated from five to ten independent cells. The fluorescent or epitope-tagged proteins were detected with anti-GFP (Roche, 11814460001; 1:1,000), anti-HA or anti-MYC antibodies.
RNA isolation, real-time and semi-quantitative RT–PCR analysis
Total RNA was isolated from rice leaf sheaths, BPHs or N. benthamiana leaves using TRIzol reagent (Takara, 9109). First-strand cDNA was synthesized using a PrimeScript RT Reagent kit with gDNA Eraser (Takara, RR047A) following the manufacturer’s instructions.
Real-time RT–qPCR was carried out using a CFX96 Real-Time system (Bio-Rad) with SYBR Green Supermix (Bio-Rad, 172–5274) following the manufacturer’s instructions. For RT–qPCR in rice, three housekeeping genes, Osubiquitin, Ostbp and Oshsp53, were used as reference genes for calibration of real-time RT–PCR data. For RT–qPCR in BPH, NlRPS18, Nlubiquitin and NlACTB (which encodes β-actin) were used as reference genes. The sequences of the primers used are listed in Supplementary Table 3.
For the semi-quantitative RT–PCR assays, first-strand cDNA (5 μl) was diluted to 50 μl final volume with TE buffer, and 1 μl of the dilution was used as template for PCR amplification using gene-specific primers. The internal standard NbACTB was used. The number of cycles used for amplification with each primer pair was adjusted to ensure that the amplification was in the linear range. The PCR products were sequenced to ensure that they were derived from the targeted genes. The sequences of the primers used are listed in Supplementary Table 3.
Immunohistochemistry
For immunolocalization of BISP in rice leaf-sheath sections, N14 plants at the two-leaf stage were infested with ten fourth- or fifth-instar nymphs for 2 days or served as non-infested controls. The outer leaf sheaths were collected, fixed, dehydrated, and embedded in paraffin (Paraplast Plus, Sigma-Aldrich, P3683) as previously described11,17. The sections were incubated in dimethylbenzene and an alcohol gradient to remove the paraffin and to gradually rehydrate. The sections were incubated with anti-BISP antibodies or pre-immune rabbit serum (prepared as described above, 1:100) at 4 °C in the dark overnight. The next day, the sections were washed 3 times (15 min each time) with PBST (PBS with 0.1% (v/v) Triton X-100), blocked with 5% bovine serum albumin in PBST at room temperature for 1 h, washed 3 times at 15-min intervals with PBST, and incubated with the secondary Cy3-conjugated goat anti-rabbit antibodies (Jackson ImmunoResearch, 111-165-003, 1:500) at 4 °C overnight. The sections were washed extensively with PBST at 15-min intervals and imaged under a confocal microscope (Leica, DMi8).
For immunolocalization in BPH salivary glands and guts, female or male insects were anaesthetized with carbon dioxide (CO2). The salivary glands and guts were dissected in 0.65% NaCl, washed 3 times in PBST and fixed in 4% (v/v) formaldehyde in PBST at 4 °C overnight. The next day, the organs were extensively washed with PBST, incubated with anti-BISP antibodies, pre-immune serum or anti-NlSP1 antibodies54 (1:100) at 4 °C in the dark overnight, and treated as described above. Nuclei were stained with 100 nM 4,6-diamidino-2-phenylindole (DAPI) at room temperature in the dark for 5 min. The salivary glands and guts were extensively washed with PBST and placed on a glass slide. Photographs were taken under a confocal microscope (Leica, DMi8). For the quantification of Cy3, the Cy3 and DAPI fluorescence signals in individual cells were quantified using ImageJ. The average Cy3/DAPI intensity ratio of each cell was measured, and mean ratios were calculated from five to ten independent cells.
RNAi and bioassay of BPHs after dsRNA injection
A 506-bp fragment of Bisp was amplified with the primers listed in Supplementary Table 3, including the T7 promoter sequence in the forward primer. After cloning the Bisp fragment into the pMD18-T vector (Takara, 6011), dsRNA was synthesized from PCR-generated DNA templates using a MEGAscript T7 Transcription kit (Ambion, AM1354). BPH nymphs in their third, fourth or fifth stage were injected with Bisp-RNAi or GFP-RNAi using a Nanoliter 2010 injector (World Precision Instruments) as previously described5. The efficiency of gene silencing was determined by RT–qPCR at 1–10 days after dsRNA treatment.
After the injected BPHs were placed and reared on rice plants for 12 h, the survival rate, weight gain and honeydew excretion of GFP-RNAi BPHs, Bisp-RNAi BPHs and non-injected control insects were measured as described above. Five replicates were set up for each treatment to measure BPH survival rates, and 20 replicates were performed for each treatment group to measure weight gain and honeydew excretion.
Plant transformation
To constitutively express Bisp in rice plants, the BISP–MYC plasmids were transformed into N14 or Bph14 plants through Agrobacterium-mediated transformation8. To knockout OsNBR1 by genome editing, the CRISPR–Cas9 plasmids OsNBR1–Cas9 were introduced into Agrobacterium tumefaciens strain EHA105 and transformed into Bph14 as described elsewhere8. Genomic DNA was extracted from these transformants, and PCR amplification was performed using primer pairs flanking the designed target site. The PCR products were sequenced to determine whether the gene editing was successful.
Quantification of free SA
Endogenous free SA levels were determined in rice leaf sheaths. Fifteen leaf sheaths from each plant or line were ground to a fine powder in liquid nitrogen as a replicate. Three biological replicates of each frozen sample were prepared. Each sample (100 mg) was added to 750 μl precooled extraction buffer (methanol:water:acetic acid, 80:19:1, v/v/v) supplemented with 3 µg naphthaleneacetic acid (Sigma-Aldrich, 35745) as an internal standard and shaken at 4 °C overnight. After centrifugation, the supernatant was filtered through a 0.22-μm nylon membrane and dried under a nitrogen flow at room temperature for at least 4 h. Finally, 200 μl methanol was added to dissolve the extracts. Endogenous free SA measurements were performed as described elsewhere55.
Recombinant protein expression and purification in E. coli and BISP antibody specificity analysis
The OsRLCK185–MBP–His, BISP–GST–His and BISP(26–124)–GST–His constructs were expressed in E. coli BL21 (DE3) cells. GST-tagged or MBP-tagged recombinant proteins were purified using Glutathione Sepharose beads (GE Healthcare, 17075601) and Amylose Resin (New England Biolabs, E8022S), respectively, according to the manufacturer’s instructions. Protein concentrations were determined by BCA protein assay (Beyotime, P0010S).
For the BISP antibody specificity analysis, the purified BISP–GST–His protein and total protein from whole insect and a twofold dilution series were separated on SDS–PAGE gels. BISP–GST–His was detected in immunoblots using anti-BISP and anti-His (GenScript, A00186; 1:2,000) antibodies, respectively. Ponceau S solution was used to stain the PVDF membrane and used as the total BPH protein loading control.
In vitro phosphorylation assays
To test the autophosphorylation of OsRLCK185, 1 µg purified recombinant OsRLCK185–MBP–His, BISP(26–124)–GST–His or BISP–GST–His was incubated in 25 µl kinase reaction buffer containing 50 mM HEPES (pH 7.5), 10 mM MgCl2, 1 mM DTT, 1 mM ATP, and 2.5 µCi [γ-32P]ATP at 30 °C for 10–30 min. The kinase reaction was subsequently analysed using a 10% SDS–PAGE gel for autoradiography. A similar gel was subjected to Coomassie Brilliant Blue staining as a control.
To test the phosphorylation of OsRLCK185 at the serine and threonine residues, phosphorylated OsRLCK185 was detected by immunoblotting with anti-phosphoserine/phosphothreonine antibodies (Millipore, 05-368, 1:1000) and anti-His antibody (GenScript, A00186).
Recombinant protein expression and purification in insect cells and BLI analyses
LRR, N14–LRR, BISP, BISP(26–124), and BISP(125–241) were cloned into pFastBac 1 with an N-terminal 6×His-SUMO tag and expressed in SF9 insect cells (Thermo Fisher Scientific, 11496015) at 27 °C. SF9 cells (500 ml) cultured in Sf-900 II SFM medium (Invitrogen, 10902088) were infected with 25 ml recombinant baculovirus and were cultured for 4 days at 27 °C. Cells were collected and re-suspended in PBS solution (2 mM KH2PO4, 8 mM Na2HPO4, 136 mM NaCl, and 2.6 mM KCl, pH 7.4). After sonication and centrifugation, recombinant LRR, N14–LRR, BISP, BISP(26–124), and BISP(125–241) proteins in the supernatant were purified with Ni-NTA beads (Novagen,70666-4) and then were dialysed in PBS solution.
The binding kinetics and affinities of LRR and N14–LRR with BISP, BISP(26–124) or BISP(125–241) was determined using an Octet RED96 system (FortéBio) as previously described56,57. Purified LRR and N14–LRR were biotinylated with NHS-biotin (Sangon Biotech, C100212) according to the manufacturer’s instructions. All streptavidin-coated biosensors (FortéBio) were hydrated in PBS buffer (2 mM KH2PO4, 8 mM Na2HPO4, 136 mM NaCl, 2.6 mM KCl, 0.05% Tween 20, and 0.5% bovine serum albumin, pH 7.4) for 10 min. Biotinylated LRR and N14–LRR were diluted in PBS buffer to a final concentration of 20 μg ml−1 and immobilized onto a streptavidin-coated biosensor. Biosensors with immobilized LRR and N14–LRR were diluted in binding buffer containing different concentrations of BISP, BISP(26–124) or BISP(125–241) for the binding kinetics analysis (association step). Subsequently, the streptavidin-coated biosensor was returned to PBS buffer for 180 s (dissociate step). PBS buffer was used in all BLI experiments to reduce nonspecific binding to the biosensors except for experiments in the presence of BISP, BISP(26–124) or BISP(125–241). The experiments included the following steps at 25 °C: (1) loading (60 s); (2) baseline (120 s); (3) immobilization of proteins onto sensors (60 s); (4) association (180 s); and (5) disassociation (180 s). The data were analysed, and sensor-grams were step-corrected and reference-corrected. Global fitting of the kinetic rates was performed, then the equilibrium dissociation constant (Kd), association (Kon) and dissociation (Koff) rate constants and their standard errors were analysed using FortéBio data analysis software (v.1.1.0.16, FortéBio). The assays were repeated two times for each affinity measurement.
For the competition assay, an Octet RED96 system (FortéBio) was used as previously described58. Biotinylated LRR was diluted in PBS buffer to a final concentration of 20 μg ml−1 and immobilized onto a streptavidin-coated biosensor. The association of BISP(26–124) or PBS was measured for 360 s at 25 °C. Next, the competitors (BISP, PBS or BISP(26–124)) were added to the BISP(26–124)-associated or PBS-associated samples. The experiments included the following steps at 25 °C: (1) loading (60 s); (2) baseline (120 s); (3) immobilization of protein onto sensors (60 s); (4) baseline (240 s); (5) association with BISP(26–124) or PBS (360 s); (6) disassociation (240 s); (7) association with the competitors BISP, PBS or BISP(26–124) (120 s); and (8) dissociation (180 s). The data were analysed using FortéBio data analysis software (v.1.1.0.16, FortéBio). The assays were repeated two times for each measurement.
MST assay
The MST assay was performed as previously described59,60. The affinity of purified LRR or N14–LRR with BISP, BISP(26–124) or BISP(125–241) was measured using a Monolith NT.115 (Nanotemper Technologies). Proteins were fluorescently labelled using a Monolith Protein Labelling kit (RED-NHS 2nd Generation; Nanotemper Technologies, MO-L011) according to the manufacturer’s protocol, and the labelled protein used for each assay was about 20 nM. A solution of unlabelled BISP, BISP(26–124) or BISP(125–241) protein was diluted for an appropriate serial concentration gradient. The samples were loaded into capillaries (Nanotemper Technologies, MO-K022) after incubation at room temperature for 5 min. Measurements were performed in 20 μl of PBS buffer containing 0.05% Tween 20. The assays were repeated three times for each affinity measurement. Data analyses were performed using Nanotemper Analysis software provided by the manufacturer.
For the competition assays between LRR with BISP and BISP(26–124) by MST, 50 nM LRR proteins and 200 nM BISP(26–124) were combined in PBS buffer with 0.05% Tween 20 and incubated for 10 min to enable protein interactions. A series of different concentrations of BISP was then added to the LRR–BISP(26–124) mixture before measurement. The assays were repeated three times for each measurement. The dissociation constant (Ki) from the MST assays was calculated using the Ki Finder software from the Nanotemper website (https://nanotemper.my.site.com/explore/s/article/Can-competition-assays-be-performed-with-MST).
Inhibition of protein degradation in rice protoplasts
After PEG-mediated transformation, the rice protoplasts were placed in 1 ml W5 buffer containing 10 μM 3-MA (3-methyladenine, Selleck, S2767), 100 μM chloroquine (MCE, HY-17589A), 100 μM leupeptin (MCE, HY-18234A), or 20 μM E-64d (Aloxistatin, Sigma-Aldrich, E8640) to inhibit autophagy or 50 μM MG132 (Selleck, S2619) to inhibit the 26S proteasome. The treated protoplasts were incubated at 28 °C for 14–20 h before protein extraction. Anti-actin (Abbkine, ABL1050) was used as a reference to quantify total protein levels. Rice WRKY72 is degraded by the 26S proteasome pathway19 and was used as a positive control to ensure that the proteasome inhibitor MG132 was active. The rice HD1 protein is degraded in the dark by the autophagy pathway61, and OsNBR1 was used as a positive control to assure autophagy inhibitors were active. The intensity of protein signals in immunoblots was quantified by ImageJ.
Live-cell imaging of autophagic structures and virus-induced gene silencing in N. benthamiana leaves
To observe autophagic structures in N. benthamiana leaves by confocal microscopy, A. tumefaciens strain GV3101 cells harbouring CFP–NbATG8F (designed as CFP–ATG8F)27 and the constructed expression vectors, BISP-201, BPH14-203, N14-203, OsNBR1-203, and N1-203 were infiltrated into N. benthamiana leaves for 60 h. The samples were then infiltrated with or without the autophagy inhibitor ConA (first made in DMSO at 0.5 M, then 1:1,000 added) for 10 h before being examined under a confocal microscope (Leica, DMi8).
For virus-induced gene silencing (VIGS) assays, pTRV1 and pTRV2 or its derivatives were introduced into Agrobacterium strain GV3101. VIGS assays were performed as previously described62. In brief, the Agrobacterium cultures with pTRV1 and pTRV2 or its derivatives were mixed at a 1:1 ratio and were infiltrated into the leaves of six-leaf stage N. benthamiana plants. Silenced phenotypes appeared in the upper leaves about 10 days after infiltration. A. tumefaciens strain GV3101 cells harbouring CFP–NbATG8F and the constructed expression vectors BISP-101 (BISP–YFP) and BPH14-203 were infiltrated into the N. benthamiana upper leaves.
Transmission electron microscopy
For immunogold electron microscopy, N14 leaf sheaths infested with BPH for 24 h were cut into 0.1-mm pieces, fixed in precooled 3% paraformaldehyde with 0.1% glutaraldehyde (in 100 mM PBS, pH 7.4) and immediately placed under a vacuum until the samples were completely submerged in the liquid. The samples were transferred to fresh 0.1% glutaraldehyde and incubated at 4 °C overnight. The next day, the samples were washed 3 times with PBS (pH 7.4) for 30 min. The samples were cleared by successive washes with 30% and 50% ethanol for 60 min each at 4 °C, then 70%, 80%, 85%, 90%, and 95% ethanol for 30 min each at −20 °C, and finally 100% ethanol 3 times for 60 min each at −20 °C. After ethanol washes, the samples were embedded in Lowicry K4M resin stepwise, according to the manufacturer’s instructions. Ultrathin sections (70 nm) of the embedded tissues were cut with a diamond knife using an Ultracut E Ultramicrotome (Reichart-Jung) and collected on 3-mm copper (mesh) grids. Immunolabelling of leaf sections was done using standard procedures as previously described59 with anti-BISP antibodies or pre-immune serum at 100 µg ml–1 and anti-rabbit IgG gold-coupled secondary antibodies (Sigma-Aldrich, G7277- 4ML) at 1:50 dilution. The sections were then stained with 2% uranyl acetate (in 70% ethanol) for 15 min, washed in distilled water 3 times, stained with lead citrate for 12 min, washed in NaOH buffer, washed in distilled water 3 times, and examined under a JEM-1230 electron microscope.
For transmission electron microscopy observations, 2-week-old Bph14–Bisp transgenic lines, Bph14 and MH63 leaf sheaths were cut into 0.1-mm pieces, fixed in precooled 4% paraformaldehyde with 2% glutaraldehyde (in 100 mM PBS, pH 7.4) and immediately placed under a vacuum until the samples were completely submerged in the liquid. The samples were transferred to fresh 2% glutaraldehyde and incubated at 4 °C overnight. The next day, the samples were washed 5 times with PBS (pH 7.4) for 20 min and submerged in a 1% solution of osmium tetroxide at room temperature for 1–2 h until they turned completely black because of oxidation. The samples were washed 5 times with PBS at 20-min intervals. The samples were cleared by successive washes with 15%, 30%, 50%, and 70% ethanol for 30 min each, then 80%, 85%, 90%, and 95% ethanol for 20 min each, and finally 100% ethanol twice for 45 min each. After ethanol washing, the samples were embedded in Spurr resin stepwise as instructed by the manufacturer. Ultrathin sections (70 nm) of the embedded tissues were cut with a diamond knife using an Ultracut E Ultramicrotome (Reichart-Jung), collected on 3-mm copper (mesh) grids, stained with 2% uranyl acetate (in 70% ethanol) for 10 min, washed in distilled water 3 times, stained with lead citrate for 15 min, washed in NaOH buffer, and examined under a JEM-1230 electron microscope. To quantify autophagosomes, the double-membrane autophagosomes in every eight transmission electron microscope images were counted and served as one biological replicate; each genotype had five replicates.
BPH infestation
Seeds were sown in 10-cm diameter plastic cups and grown in a plant growth chamber (CONVIRON PGC2000) as described elsewhere11. To analyse the relationship between BISP levels and autophagy, 4-leaf rice plants were infested with second- or third-instar BPH nymphs (10 per plant) at the selected time points (6, 12, 24, 48, and 72 h before the end of the experiment). Control groups were maintained in parallel but without BPH infestation. At the end of the treatments, leaf sheaths were wiped with cotton wool balls soaked in alcohol to remove honeydew, leaf sheaths were excised, frozen, and ground to a fine powder in liquid nitrogen, and proteins isolated. Immunoblotting was performed following the procedures described above in the section ‘Protein extraction, immunoblotting and in vivo co-IP assays’. All treatments were repeated three times.
To measure autophagy and BISP levels following BPH infestation, 4-leaf rice plants were infested with second- or third-instar BPH nymphs (10 per plant) for 24 h. The insects were then removed, and the plants were allowed to grow under standard conditions (described above). At the selected time points after BPH removal, honeydew was removed from leaf sheaths, leaves were excised, frozen, and ground to a fine powder in liquid nitrogen, and proteins isolated. Immunoblotting was performed following the procedures described above. All treatments were repeated three times.
For the autophagy inhibitor ConA treatment, the BPH-infested and non-infested rice plants were treated with or without ConA (10 μM) for 12 h before samples were collected. Leaf sheath total protein extraction and immunoblotting were performed according to the procedures described above. All treatments were repeated three times.
Agronomic performance test
To investigate the agronomic performance of Bph14 and Bph14–Bisp plants, plants were grown in a field in Wuhan, China, under routine management. At harvest, the middle plants in the central row of each plot were assessed for the following nine agronomic traits: plant height, sword leaf length, sword leaf width, number of panicles per plant, number of filled grains per panicle, number of filled grains per plant, percentage of filled grain, 1,000-grain weight, and grain yield per plant63.
Statistical analysis
No statistical methods were used to predetermine the sample sizes. The experiments were not randomized, and investigators were not blinded to allocation during the experiments and outcome assessment. Quantification analyses of protein abundance were conducted using ImageJ software. All values are presented as the mean ± s.d. or mean ± s.e.m. as indicated. Data points are plotted on the graphs, and the number of samples is indicated in the corresponding figure legends. In all cases, sampling for replicates and between time points was performed by collecting material from new plants. All statistical analyses were performed by one-way analysis of variance with MS Excel and GraphPad Prism (v.8.0.1, GraphPad Software). Detailed information about statistical analysis values for all experiments is provided in Supplementary Table 4.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data are available within this article and its Supplementary Information. Original gel blots are shown in Supplementary Fig. 1. Original data points in graphs are shown in the source data files. Statistical analyses of this study are provided in Supplementary Table 4. The sequences of Bisp have been deposited and made publicly available in GenBank with accession number MH885414. Source data are provided with this paper.
References
Kaloshian, I. & Walling, L. Hemipteran and dipteran pests: effectors and plant host immune regulators. J. Integr. Plant Biol. 58, 350–361 (2016).
Hatchett, J. H. & Gallun, R. L. Genetics of the ability of the Hessian fly, Mayetiola destructor, to survive on wheats having different genes for resistance. Ann. Entomol. Soc. Am. 63, 1400–1407 (1970).
Oerke, E. C. Crop losses to pests. J. Agric. Sci. 144, 31–43 (2006).
Jiang, Y., Zhang, C., Chen, R. & He, S. Challenging battles of plants with phloem-feeding insects and prokaryotic pathogens. Proc. Natl Acad. Sci. USA 116, 23390–23397 (2019).
Shangguan, X. et al. A mucin-like protein of planthopper is required for feeding and induces immunity response in plants. Plant Physiol. 176, 552–565 (2018).
Rossi, M. et al. The nematode resistance gene Mi of tomato confers resistance against the potato aphid. Proc. Natl Acad. Sci. USA 95, 9750–9754 (1998).
Dogimont, C., Chovelon, V., Pauquet, J., Boualem, A. & Bendahmane, A. The Vat locus encodes for a CC-NBS-LRR protein that confers resistance to Aphis gossypii infestation and A. gossypii-mediated virus resistance. Plant J. 80, 993–1004 (2014).
Du, B. et al. Identification and characterization of Bph14, a gene conferring resistance to brown planthopper in rice. Proc. Natl Acad. Sci. USA 106, 22163–22168 (2009).
Tamura, Y. et al. Map-based cloning and characterization of a brown planthopper resistance gene BPH26 from Oryza sativa L. ssp. indica cultivar ADR52. Sci. Rep. 4, 5872 (2014).
Zhao, Y. et al. Allelic diversity in an NLR gene BPH9 enables rice to combat planthopper variation. Proc. Natl Acad. Sci. USA 113, 12850–12855 (2016).
Guo, J. et al. Bph6 encodes an exocyst-localized protein and confers broad resistance to planthoppers in rice. Nat. Genet. 50, 297–306 (2018).
Shi, S. et al. Bph30 confers resistance to brown planthopper by fortifying sclerenchyma in rice leaf sheaths. Mol. Plant 14, 1714–1732 (2021).
Jones, D. G. & Dangl, J. L. The plant immune system. Nature 444, 323–329 (2006).
Cui, H., Tsuda, K. & Parker, J. E. Effector-triggered immunity: from pathogen perception to robust defense. Annu. Rev. Plant Biol. 66, 487–511 (2015).
Dyck, V. & Thomas, B. in Brown Planthopper: Threat to Rice Production in Asia (ed. Brady, N. C.) 3–17 (International Rice Research Institute, 1979).
Sogawa, K. The rice brown planthopper: feeding physiology and host plant interactions. Annu. Rev. Entomol. 27, 49–73 (1982).
Hao, P. et al. Herbivore-induced callose deposition on the sieve plates of rice: an important mechanism for host resistance. Plant Physiol. 146, 1810–1820 (2008).
Liu, F. et al. Ultrabithorax is a key regulator for the dimorphism of wings, a main cause for the outbreak of planthoppers in rice. Nat. Sci. Rev. 7, 1181–1189 (2020).
Hu, L. et al. The coiled-coil and nucleotide binding domains of Brown Planthopper Resistance14 function in signaling and resistance against planthopper in rice. Plant Cell 29, 3157–3185 (2017).
Tang, D., Wang, G. & Zhou, J. Receptor kinases in plant–pathogen interactions: more than pattern recognition. Plant Cell 29, 618–637 (2017).
Yamaguchi, K. et al. A receptor-like cytoplasmic kinase targeted by a plant pathogen effector is directly phosphorylated by the chitin receptor and mediates rice immunity. Cell Host Microbe 13, 347–357 (2013).
Wang, C. et al. OsCERK1-mediated chitin perception and immune signaling requires receptor-like cytoplasmic kinase 185 to activate an MAPK cascade in rice. Mol. Plant 10, 619–633 (2017).
Nelson, R., Wiesner-Hanks, T., Wisser, R. & Balint-Kurti, P. Navigating complexity to breed disease-resistant crops. Nat. Rev. Genet. 19, 21–33 (2018).
Xu, G. et al. uORF-mediated translation allows engineered plant disease resistance without fitness costs. Nature 545, 491–494 (2017).
Dikic, I. Proteasomal and autophagic degradation systems. Annu. Rev. Biochem. 86, 193–224 (2017).
Wersch, S., Tian, L., Hoy, R. & Li, X. Plant NLRs: the whistleblowers of plant immunity. Plant Commun. 1, 100016 (2020).
Han, S. et al. Cytoplastic glyceraldehyde-3-phosphate dehydrogenases interact with ATG3 to negatively regulate autophagy and immunity in Nicotiana benthamiana. Plant Cell 27, 1316–1331 (2015).
Klionsky, D. et al. Guidelines for the use and interpretation of assays for monitoring autophagy (4th edition). Autophagy 17, 1–382 (2021).
Dikic, I. & Elazar, Z. Mechanism and medical implications of mammalian autophagy. Nat. Rev. Mol. Cell Biol. 19, 349–364 (2018).
Kellner, R., Concepcion, J., Maqbool, A., Kamoun, S. & Dagdas, Y. ATG8 expansion: a driver of selective autophagy diversification? Trends Plant Sci. 22, 204–214 (2017).
Yu, J., Zhen, X., Li, X., Li, N. & Xu, F. Increased autophagy of rice can increase yield and nitrogen use efficiency (NUE). Front. Plant Sci. 10, 584 (2019).
Brillada, C. et al. Exocyst subunit Exo70B2 is linked to immune signaling and autophagy. Plant Cell 33, 404–419 (2021).
Zhang, Y. & Chen, Z. Broad and complex roles of NBR1-mediated selective autophagy in plant stress responses. Cells 9, 2562 (2020).
Hofius, D., Li, L., Hafrén, A. & Coll, N. Autophagy as an emerging arena for plant–pathogen interactions. Curr. Opin. Plant Biol. 38, 117–123 (2017).
Thirumalaikumar, V. et al. Selective autophagy regulates heat stress memory in Arabidopsis by NBR1-mediated targeting of HSP90 and ROF1. Autophagy 17, 2184–2199 (2021).
Hafrén, A. et al. Selective autophagy limits cauliflower mosaic virus infection by NBR1-mediated targeting of viral capsid protein and particles. Proc. Natl Acad. Sci. USA 114, E2026–E2035 (2017).
Svenning, S., Lamark, T., Krause, K. & Johansen, T. Plant NBR1 is a selective autophagy substrate and a functional hybrid of the mammalian autophagic adapters NBR1 and p62/SQSTM1. Autophagy 7, 993–1010 (2011).
Will, T. & Bel, A. Physical and chemical interactions between aphids and plants. J. Exp. Bot. 57, 729–737 (2006).
Ghaffar, M., Pritchard, J. & Ford-Lioyd, B. Brown planthopper (N. lugens Stål) feeding behaviour on rice germplasm as an indicator of resistance. PLoS ONE 6, e22137 (2011).
Michel, A. & Harris, M. Editorial overview: why modern research justifies the re-emergence of host–plant resistance as a focus for pest management. Curr. Opin. Insect Sci. 45, iii–v (2021).
Wang, J. et al. Two VOZ transcription factors link an E3 ligase and an NLR immune receptor to modulate immunity in rice. Mol. Plant 14, 253–266 (2021).
Rodriguez, E. et al. Autophagy mediates temporary reprogramming and dedifferentiation in plant somatic cells. EMBO J. 39, e103315 (2020).
Jiang, H. et al. Evaluation and breeding application of six brown planthopper resistance genes in rice maintainer line Jin 23B. Rice 11, 22 (2018).
Shi, Z. Y. et al. Construction of genomic library of a BPH-resistant rice line with binary vector and physical map of Qbp1 locus. Plant Sci. 165, 879–885 (2003).
Lu, Y. et al. Genome-wide targeted mutagenesis in rice using the CRISPR/Cas9 system. Mol. Plant 10, 1242–1245 (2017).
Huang, Z. et al. Identification and mapping of two brown planthopper resistance genes in rice. Theor. Appl. Genet. 102, 929–934 (2001).
Heinrichs, E., Medrano, F. & Rapusas, H. Genetic Evaluation for Insect Resistance in Rice (International Rice Research Institute, 1985).
Earley, K. et al. Gateway-compatible vectors for plant functional genomics and proteomics. Plant J. 45, 616–629 (2006).
Ma, X. et al. A robust CRISPR/Cas9 system for convenient, high-efficiency multiplex genome editing in monocot and dicot plants. Mol. Plant 8, 1274–1284 (2015).
Hofius, D. et al. Autophagic components contribute to hypersensitive cell death in Arabidopsis. Cell 137, 773–783 (2009).
Kushnirov, V. Rapid and reliable protein extraction from yeast. Yeast 16, 857–860 (2000).
Kudal, J. & Bock, R. Lighting the way to protein–protein interactions: recommendations on best practices for bimolecular fluorescence complementation analyses. Plant Cell 28, 1002–1008 (2016).
Hu, J. et al. The Bphi008a gene interacts with the ethylene pathway and transcriptionally regulates MAPK genes in the response of rice to brown planthopper feeding. Plant Physiol. 156, 856–872 (2011).
Huang, J. et al. Salivary Protein 1 of brown planthopper is required for survival and induces immunity response in plants. Front. Plant Sci. 11, 571280 (2020).
Liu, H., Li, X., Xiao, J. & Wang, S. A convenient method for simultaneous quantification of multiple phytohormones and metabolites: application in study of rice–bacterium interaction. Plant Methods 8, 2 (2012).
Cao, L. et al. Design of protein-binding proteins from the target structure alone. Nature 605, 551–560 (2022).
Hong, Q. et al. Molecular basis of receptor binding and antibody neutralization of omicron. Nature 604, 546–552 (2022).
Alanine, D. et al. Human antibodies that slow erythrocyte invasion potentiate malaria-neutralizing antibodies. Cell 178, 216–288 (2019).
Friml, J. et al. ABP1-TMK auxin perception for global phosphorylation and auxin canalization. Nature 609, 575–581 (2022).
Mamou, G. et al. Peptidoglycan maturation controls outer membrane protein assembly. Nature 606, 953–959 (2022).
Hu, Z. et al. Autophagy targets Hd1 for vacuolar degradation to regulate rice flowering. Mol. Plant 15, 1137–1156 (2022).
Liu, Y. et al. Autophagy regulates programmed cell death during the plant innate immune response. Cell 121, 567–577 (2005).
You, A. Q. et al. Identification of QTLs across recombinant inbred lines and testcross populations for traits of agronomic importance in rice. Genetics 172, 1287–1300 (2006).
Acknowledgements
We thank Y. Liu (Tsinghua University) for providing plasmids CFP–NbATG8F, pTRV1, pTRV2, pTRV2-NbPI3K, pTRV2-NbATG6 and pTRV2-NbATG7; Y. Liu (South China Agricultural University) for providing plasmid pYLCRISPR/Cas9-MH; S. Zhang (Huazhong Agricultural University) for helpful suggestions for the BLI assays; and J. L. Dangl, the University of North Carolina, for critical reading and insightful discussions. This work was supported by grants from the National Natural Science Foundation of China (31630063, 31901518 and 31230060), the Major Program of Guangdong Basic and Applied Research, China (2019B030302006), Program of Department of Science and Technology of Hubei Province (2022EJD011), the National Key Research and Development Program of China (2016YFD0100903 and 2016YFD0100600), the National Program on Research & Development of Transgenic Plants (2016ZX08009-003-001), the National Postdoctoral Program for Innovative Talents (BX20180230) and the China Postdoctoral Science Foundation (2018M640729).
Author information
Authors and Affiliations
Contributions
G.H. conceived and supervised the project. J.G., H.W., W.G. and G.H. designed the experiments. J.G., H.W. and W.G. carried out most of the experiments, including the Y2H assays, cloning, co-IP, BiFC, subcellular localization, autophagy analysis, BLI, MST, phenotype assays, microscopy, RNA analysis and western blotting. Q.G., J.W. and J.S. performed transmission electron microscopy, SA measurement and phosphorylation assays. J.Y., S.S., X.S., S.J. and B.L. performed cloning, dsRNA injection, BPH performance analysis and RNA analysis. Y.P., M.G., X.Z. and D.W. performed cloning, western blotting, protoplast isolation and immunogold microscopy. J.Z. and C.X. performed VIGS and microscopy. J.H., W.R. and C.Z. maintained the BPH population, performed phenotype and immunohistochemistry. B.D., R.C. and L.Z generated material used in this study and performed agronomic traits. J.G., H.W. and W.G. prepared the figures. Y.Z., Q.Z. and L.L.W. provided theoretical contributions to the project. J.G., H.W., W.G., Y.Z., Q.Z., L.L.W. and G.H. analysed data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks Thorsten Nurnberger and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Host-plant resistance evaluation, BISP subcellular localization in rice protoplasts, Bisp gene and BISP protein expression profile in BPH, and BISP delivery into rice leaf sheath.
a, BPH-resistance scores of Bph14 and N14 plants after BPH infestation. Bph14 and N14 plants express the resistant NLR (BPH14) and the susceptible allele variant (N14), respectively. Lower scores indicate higher resistance to the insects. Data were collected 7 days after BPH infestation (n = 36 plants examined over 3 independent experiments). P values were derived by one-way ANOVA. b, Honeydew excretion of BPHs fed on Bph14 and N14 plants for 2 days. P values were derived by one-way ANOVA (n = 20, biologically independent plants). c, BISP-GFP had a nucleocytoplasmic localization in rice protoplasts. bZIP63-RFP was used as a nuclear marker. The confocal images were taken 14–20 h after transformation. Scale bar, 5 μm. d, Immunoblot detection of full-length BISP-GFP expression in rice protoplasts. BISP-GFP was detected by anti-GFP antibody. e, Immunoblot detection of the expressed proteins in BiFC assays. BISP-YN, BISP125–241-YN(BISP(125–241)-YN), and BPH14-YC or N14-YC were co-expressed in rice protoplasts and the total proteins were detected using anti-Myc, anti-HA or anti-ACTIN antibody. f, Relative levels of Bisp RNA in different BPH tissues. SG, salivary glands; FB, fat body. Data are mean ± s.d. (n = 3, biologically independent experiments). g, Relative level of Bisp RNAs during different developmental stages. 1st to 5th, are first to fifth instars; Female and Male are adults. Data are mean ± s.d. (n = 3, biologically independent experiments). h, (Upper) Determination of the anti-BISP specificity using purified BISP-GST-His. The purified BISP-GST-His protein was serially diluted and detected by anti-BISP or anti-His antisera. (Bottom) Correlation between the anti-BISP labeled signal and anti-His labeled signals. The densities of anti-BISP and anti-His labeled signals were quantified using ImageJ with the undiluted sample set as 1. i, (Upper) Determination of the anti-BISP specificity using a dilution series of total mixed adults BPH proteins. BISP was detected using the anti-BISP antiserum. Ponceau staining (Ponceau S) is a loading control. (Bottom) Correlation between the anti-BISP labeled and Ponceau S signals. The density of anti-BISP labeled and Ponceau S signals were quantified using ImageJ with the undiluted sample set as 1. j,k, Immunohistochemical localization of BISP in male BPH salivary glands. Anti-BISP (j) or pre-immune rabbit serum (k) was used as the primary antiserum. Red fluorescence (Cy3) shows the localization of BISP and blue fluorescence shows the location of DAPI-stained nuclei. Scale bars, 100 μm. l,m, Immunohistochemical localization of BISP in mixed adults BPH guts. Anti-BISP (l) or pre-immune rabbit serum (m) was used as the primary antiserum. Red fluorescence (Cy3) shows the localization of BISP and blue fluorescence shows the location of DAPI-stained nuclei. Scale bars, 100 μm. n,o, Immunohistochemical localization of NlSP1 in female BPH salivary glands. Either anti-NlSP1 (n) or pre-immune rabbit serum (o) was used as the primary antiserum. Red fluorescence (Cy3) shows the localization of NlSP1 and blue fluorescence shows the location of DAPI-stained nuclei. Scale bars, 100 μm. p, Quantification of Cy3 intensities. The Cy3 fluorescence in samples using anti-BISP, anti-NlSP1 and pre-immune serum was normalized relative to DAPI fluorescence (blue) intensity. P values were derived by one-way ANOVA. Data are mean ± s.d. (n = 5, biologically independent experiments). q-t, Immunogold electron microscopy analysis of the distribution of BISP in the rice leaf sheaths during BPH feeding. q,s, TEM image of the BPH salivary sheaths in the rice phloem tissue. Either anti-BISP (q) or pre-immune rabbit serum (s) was used as the primary antiserum. Scale bars, 20 μm. r,t, TEM image of the boxed area in (q) and (s). Red arrows indicate the anti-BISP-labeled immunogold particles (dark spots). Scale bars, 200 nm. SH, salivary sheaths; P, phloem cells. In box plots in a,b, the box limits indicate the 25th and 75th percentiles, the whiskers indicate the full range of the data, and the center line indicates the median. Individual data points are plotted. Experiments were independently repeated two (q–t) or three (b–e, h–o) times, each giving similar results.
Extended Data Fig. 2 BISP played crucial roles in the feeding and performance of BPH on rice plants.
a, Effects of dsRNA injection on Bisp transcript levels in BPHs. Bisp-Ri (Bisp-RNAi) insects were assayed daily for 10 days. GFP-Ri (GFP-RNAi) and CK insects were assayed on day 3. Data are mean ± s.d. (n = 3, biologically independent experiments). The inset shows a representative immunoblot demonstrating the relative abundance of BISP in GFP-Ri BPHs and Bisp-Ri BPHs at 3 days after injection. Total BPH proteins were extracted from injected or control insects and immunoblotted. BISP was detected with anti-BISP antibodies and Ponceau S staining was used as the loading control. b,c,d, Weight gain (b), honeydew excretion (c) and survival rates (d) of dsRNA-injected and non-injected BPHs fed on N14 plants. e,f,g, Weight gain (e), honeydew excretion (f) and survival rates (g) of dsRNA-injected and non-injected BPHs fed on the Osrlck185−-1 (Osrlck185-1) and WT plants. h,i,j, Weight gain (h), honeydew excretion (i) and survival rates (j) of dsRNA-injected BPHs fed on the Bph14 and MH63 plants. In a–j, BPHs were microinjected with Bisp dsRNA (Bisp-Ri) or GFP dsRNA (GFP-Ri) and non-injected BPHs (CK) served as a control. For b–j, P values were derived by one-way ANOVA. In b,c,e,f,h,i, data are mean ± s.d. (n = 20, biologically independent plants); in d,g,j, data are mean ± s.e.m. (n = 5, biologically independent experiments). In box plots in b,c,e,f,h,i, the box limits indicate the 25th and 75th percentiles, the whiskers indicate the full range of the data, and the center line indicates the median. Individual data points are plotted. In e–j, different letters indicate significant differences (P < 0.05, one-way ANOVA); exact P values are provided in Supplementary Table 4. All the experiments were repeated three times, each giving similar results.
Extended Data Fig. 3 Effects of Bisp on BPH resistance in N14-Bisp transgenic lines.
a, Photographs of the independent N14-Bisp homozygous T2 transgenic lines (N14-Bisp-3 and N14-Bisp-11) and N14 plants at the 3-leaf stage without BPH feeding. Scale bar, 10 cm. b, Plant height of N14-Bisp lines and N14 at the 3-leaf stage. P values were derived by one-way ANOVA (n = 30, biologically independent samples). c, d, Weight gain (c) and honeydew excretion (d) of BPHs reared on the N14-Bisp transgenic and N14 plants for 2 d. P values were derived by one-way ANOVA (n = 10, biologically independent samples). e, Results of the two-host choice assay examining the settling of BPHs on the N14-Bisp transgenic and N14 plants. P values were derived by one-way ANOVA. Data are means ± s.d. (for N14-3 and N14-Bisp-11, n = 7 and 10, respectively, biologically independent experiments). f, Transcript levels analysis of SA biosynthesis- and signaling-related genes OsICS1 and OsNPR1 and defense-related genes OsPR1a, OsPR1b, OsPR5 and OsPR10 in two N14-Bisp transgenic lines and N14 plants. P values were derived by one-way ANOVA. Data are mean ± s.d. (n = 3, biologically independent experiments). In box plots in b, c, d, the box limits indicate the 25th and 75th percentiles, the whiskers indicate the full range of the data, and the center line indicates the median. Individual data points are plotted. All the experiments were repeated three times, each giving similar results.
Extended Data Fig. 4 Identification of OsRLCK185 and its role in plant defense.
a, Y2H screen for BISP-interacting kinases. Yeast strain AH109 was co-transformed with the indicated constructs and grown on DDO (SD/-Leu-Trp) and TDO (SD/-Leu-Trp-His) selective medium containing 1.5 mM 3-amino-1,2,4-triazole (3-AT). b, Immunoblot detection of proteins expressed in yeast using anti-HA and anti-Myc (anti-MYC) antibodies, respectively. Ponceau S staining was used as the loading control. Asterisks indicate specific bands detected by immunoblotting analysis. c, Structure of OsRLCK185. The orange box indicates the kinase domain. d, BISP but not BISP26–124 (BISP(26–124)) reduced OsRLCK185 autophosphorylation activity at Ser and Thr residues in vitro. OsRLCK185 proteins were immunoblotted and phosphorylated OsRLCK185 was detected with anti-phosphoserine/ phosphothreonine antibodies. The input proteins were immunoblotted and detected with anti-His antibodies. e, Verification of three independent OsRLCK185 knockout lines (Osrlck185, Osrlck185−-1, −2, and −3) by PCR-based sequencing. f,g, Weight gain (f) and honeydew excretion (g) of BPHs on three Osrlck185− lines and WT plants for 2 d. P values were derived by one-way ANOVA (n = 30, biologically independent samples). In box plots, the box limits indicate the 25th and 75th percentiles, the whiskers indicate the full range of the data, and the centre line indicates the median. Individual data points are plotted. Experiments were repeated three times (a,b,d,f,g) or two times (e), each giving similar results.
Extended Data Fig. 5 BISP triggered Bph14-mediated BPH resistance in Bph14 plants.
a, Y2H assays between the full-length BPH14, the BPH14 LRR domain (LRR) and the N14 LRR domain (N14-LRR) and the BISP deletion mutants BISP26–124 (BISP(26–124)) and BISP125–241(BISP(125–241)). DDO, SD/-Leu-Trp; TDO, SD/-Leu-Trp-His with 2.5 mM 3-AT. b, Biolayer interferometry (BLI) determination of the binding kinetics between N14-LRR and BISP. BLI response profile of BISP at different concentrations with the sensor-immobilized N14-LRR. c, BLI analysis for the binding kinetics between the BPH14 LRR and BISP26–124. BLI response profile of BISP26–124 at different concentrations with the sensor-immobilized LRR. Kd, equilibrium dissociation constant; Kon, association rate constant; Koff, dissociation rate constant. d, BLI analysis for the binding kinetics between the BPH14 LRR and BISP125–241. BLI response profile of BISP125–241 at different concentrations with the sensor-immobilized LRR. e, MST analysis of binding affinity between fluorescently labeled LRR, N14-LRR and BISP, BISP26–124 and BISP125–241. The Kd values show the binding between LRR and BISP, BISP26–124. ΔFnorm is the normalized fluorescence minus the relative fluorescence of the unbound state. Data are mean ± s.d. (n = 3, biologically independent experiments). f, MST analysis of the competitive binding between BISP and BISP26–124 with LRR. ΔFnorm is the normalized fluorescence minus the relative fluorescence of the unbound state. Data are mean ± s.d. (n = 3, biologically independent experiments). g, BISP protein levels in the independent Bph14-Bisp homozygous T2 transgenic lines and Bph14 control plants. Total proteins were immunoblotted and detected with anti-Myc and anti-ACTIN antibodies, respectively. h, Relative transcript levels of Bph14-activated SA biosynthesis- and signaling-related genes OsICS1 and OsNPR1, and defense-related genes OsPR1a, OsPR1b, OsPR5 and OsPR10 in the Bph14-Bisp transgenic lines and WT Bph14 plants. P values were derived by one-way ANOVA. Data are mean ± s.d. (n = 3, biologically independent experiments). i, Honeydew excretion by BPHs reared on the Bph14-Bisp transgenic lines and WT Bph14 plants for 48 h. P values were derived by one-way ANOVA (n = 20, biologically independent plants). In box plots, the box limits indicate the 25th and 75th percentiles, the whiskers indicate the full range of the data, and the center line indicates the median. Individual data points are plotted. j, Survival rate of BPHs feeding on the Bph14-Bisp transgenic lines and WT Bph14 plants for 9 days. P values were derived by one-way ANOVA. Data are mean ± s.e.m. (n = 5, biologically independent experiments). k, Two-host choice test of BPHs on the Bph14-Bisp transgenic plants and WT Bph14 plants. P values were derived by one-way ANOVA. Data are mean ± s.e.m. (n = 8, biologically independent experiments). All the experiments were repeated two (b–d) or three (a,g,i–k) times, each giving similar results.
Extended Data Fig. 6 Agronomic traits of the Bph14-Bisp transgenic lines and WT Bph14 plants.
a, Phenotypes of the Bph14-Bisp transgenic lines and Bph14 plants at the heading stage. Scale bar, 30 cm. b, c, Plant height (b) and heading date (c) of the Bph14-Bisp transgenic lines and Bph14 plants at the heading stage. P values were derived by one-way ANOVA. For b, n = 18, biologically independent samples. For c, n = 30, biologically independent samples. d–j, Agronomic traits of the Bph14-Bisp transgenic lines and WT Bph14 plants at the mature stage. (d), Flag leaf length. (e), Flag leaf width. (f), The number of panicles per plant. (g), The number of filled grains per plant. (h), Percentage of filled grains. (i), 1,000-grain weight. (j), Grain yield per plant. One-way ANOVA (n = 18, biologically independent samples). In box plots in b–j, the box limits indicate the 25th and 75th percentiles, the whiskers indicate the full range of the data, and the center line indicates the median. Individual data points are plotted.
Extended Data Fig. 7 The autophagic degradation of BISP was promoted by BPH14.
a, Dosage effect of BPH14 on BISP levels in rice protoplasts. BISP-MYC was expressed with or without BPH14-MYC in rice protoplasts. Total proteins were immunoblotted and detected with anti-Myc and anti-ACTIN antibodies, respectively. b, Immunoblots showing that proteasome degradation of WRKY72-HA was blocked by MG132 treatment. Total protoplast proteins were immunoblotted and detected with anti-HA and anti-ACTIN antibodies, respectively. c, Immunoblots showing that the autophagy degradation of Hd1-HA was blocked by treatments with autophagy inhibitors. Total protoplast proteins were immunoblotted and detected with anti-HA and anti-ACTIN antibodies, respectively. d, (Upper) Confocal images of CFP-ATG8f-labeled autophagic puncta structures in the absence (left row) or presence (right row) of ConA. Autophagosomes in cells co-expressing CFP-ATG8f with BISP, N14, or BPH14 (individually) and CFP-ATG8f and BISP with N14 or BPH14 after agroinfiltration of N. benthamiana leaves are displayed. Red arrows indicate CFP-ATG8f-labeled autophagic puncta. (Bottom) Insets show enlarged autophagosome image at higher magnification. Scale bars, 25 μm. e, Average number of CFP-NbATG8-labeled autophagic puncta in the presence or absence ConA. P values were derived by one-way ANOVA. Data are mean ± s.d. (n = 6, biologically independent experiments). f,g,h, Silencing of NbATG6, NbPI3K or NbATG7 suppressed the production of CFP-ATG8f-labeled autophagic puncta. (f), Confocal images of CFP-ATG8f-labeled autophagic puncta structures co-localized with BISP-YFP. CFP-ATG8f, BISP-YFP and BPH14 were transiently co-expressed in the silenced leaves (VIGS-NbATG6, VIGS-NbPI3K and VIGS-NbATG7) or the non-silenced control leaves (VIGS-EV). Red arrows indicate CFP-ATG8f-labeled autophagic puncta co-localized with BISP-YFP. (Bottom) Insets show enlarged autophagosome image at higher magnification. Scale bars, 25 μm. (g), The transcript levels of NbATG6, NbPI3K and NbATG7 by semi-quantitative RT-PCR. The transcript levels of NbACTB were used as a loading control. (h), Average number of CFP-NbATG8-labeled autophagic puncta in the silenced and control leaves. P values were derived by one-way ANOVA. Data are mean ± s.d. (n = 6, biologically independent experiments). i, Immunoblot detection of OsATG8 in the non-infested Bph14-Bisp transgenic lines, Bph14 and MH63 plants. Total proteins were immunoblotted and detected with anti-AtATG8A and anti-ACTIN antibodies, respectively. j, Relative transcript levels of OsATG8s (OsATG8a, OsATG8b, OsATG8c) in the non-infested Bph14-Bisp transgenic lines, Bph14 and MH63 plants. P values were derived by one-way ANOVA. Data are mean ± s.d. (n = 3, biologically independent experiments). k, Relative transcript levels of OsATG8s (OsATG8a, OsATG8b, OsATG8c) in Bph14 and MH63 plants after BPH feeding for different durations. P values were derived by one-way ANOVA. Data are mean ± s.d. (n = 3, biologically independent experiments). l, Immunoblot detection of OsATG8 and OsNBR1 in the non-infested and BPH-infested MH63 and Bph14 plants in the absence or presence of ConA treatment. Leaf sheaths total proteins were immunoblotted and detected with anti-AtATG8A, anti-AtNBR1 and anti-ACTIN antibodies, respectively. Numbers under the lanes (a,c,i,l) indicate protein abundance relative to that of ACTIN, quantified by ImageJ. Experiments (a–d,f,g,i,l) were repeated three times, each giving similar results.
Extended Data Fig. 8 OsNBR1 played a role in the autophagic degradation of BISP by BPH14.
a, Immunoblot detection of proteins expressed in yeast using anti-HA and anti-Myc antibodies, respectively. Ponceau S staining was used as the loading control. Asterisks indicate specific bands detected by immunoblotting analysis. b, Confocal images of CFP-NbATG8f-labeled autophagic puncta structures in the absence (left) or presence (right) of ConA. The photos showing cells co-expressing CFP-NbATG8f and BISP with N14 and OsNBR1, BPH14, BPH14 and OsNBR1, or BPH14 and OsNBR1 mutant N1 after agroinfiltration of N. benthamiana leaves. Red arrows indicate CFP-NbATG8f-labeled autophagic puncta. (Bottom Row) Insets show enlarged autophagosome image at higher magnification. EV, Empty Vector. Scale bars, 25 μm. c, Average number of CFP-NbATG8-labeled autophagic puncta in the absence or presence of ConA. Data are mean ± s.d. (n = 10, biologically independent samples). Different letters indicate significant differences (P < 0.05, one-way ANOVA); exact P values are provided in Supplementary Table 4. d, Immunoblot detection of OsNBR1 levels in the non-infested Bph14-Bisp transgenic lines, Bph14 and MH63 plants. Total proteins were immunoblotted and detected with anti-AtNBR1 and anti-ACTIN antibodies, respectively. Numbers under the lanes indicate protein abundance relative to that of ACTIN, quantified by ImageJ. e, Relative OsNBR1 transcript levels in the non-infested Bph14-Bisp transgenic lines, Bph14 and MH63 plants. P values were derived by one-way ANOVA. Data are means ± s.d. (n = 3, biologically independent experiments). f, Relative OsNBR1 transcript levels in Bph14 and MH63 plants after BPH infestation. P values were derived by one-way ANOVA. Data are mean ± s.d. (n = 3, biologically independent experiments). Experiments (a,b,d) were repeated at least three times, each giving similar results.
Extended Data Fig. 9 The PB1 domain, UBA domain and LIR motif of OsNBR1 played essential roles in BPH14-promoted autophagic degradation of BISP.
a, Y2H assay of the interactions between OsNBR1 and the CC, NB, and LRR domains of BPH14. DDO, SD/-Leu-Trp; TDO, SD/-Leu-Trp-His with 1 mM 3-AT. b, Co-IP assay of the interactions between OsNBR1 and the CC, NB, and LRR domains of BPH14 in rice protoplasts. OsNBR1-MYC and CC-HA, NB-HA or LRR-HA were co-expressed in rice protoplasts, and used in the co-IP assay with anti-Myc agarose; the input and immunoprecipitated proteins were detected with anti-HA, anti-Myc or anti-ACTIN antibodies. c, Schematic illustration of OsNBR1 and the N1-N7 mutant constructs used in this study. d, OsNBR1 mutants were unable to degrade BISP in rice protoplasts. BISP-MYC was co-expressed with BPH14-MYC and the full-length OsNBR1 (FL-HA) or its HA-tagged mutants (N1-N7-HA) in rice protoplasts. The levels of BISP-MYC protein were detected with anti-Myc antibody. e–h, Co-IP assays of the interactions between the OsNBR1 mutants and BISP (e), BPH14 (f), OsATG8c (g) and OsNBR1 (h) in rice protoplasts. BISP-MYC, BPH14-MYC, MYC- OsATG8c, or OsNBR1-MYC was co-expressed with OsNBR1(FL-HA) and its HA-tagged mutants (N1-N7-HA) in rice protoplasts. Proteins interacting with OsNBR1-HA or its mutants were identified in a co-IP assay with anti-MYC agarose; the input and immunoprecipitated proteins were detected with anti-HA, anti-Myc or anti-ACTIN antibodies. i, Summary of interactions between the OsNBR1 mutants and BISP, BPH14, OsATG8c, and OsNBR1, respectively. j, Verification of the mutations in the independent homozygous OsNBR1 knockout (Bph14-Osnbr1, Osnbr1−) lines (Bph14-Osnbr1−-1 and Bph14-Osnbr1−-2) by PCR-based sequencing. k, Immunoblot detection of OsNBR1 proteins in Bph14 and two OsNBR1 knockout lines (Bph14-Osnbr1−-1 and Bph14-Osnbr1−-2). Total proteins were immunoblotted and detected with anti-AtNBR1 and anti-ACTIN antibodies, respectively. In c-i, FL, full-length of OsNBR1. N1-N7, the seven OsNBR1 mutants. Experiments were repeated three times (a,b,d–h,k) or two times (j) with similar results.
Extended Data Fig. 10 Effects of BPH infestation cessation in BPH14 activation and termination and the role OsNBR1 played in BPH resistance.
a, Relative transcript levels of OsATG8s and OsNBR1 in Bph14 and MH63 plants upon the cessation of BPH feeding. b, Relative transcript levels of Bph14-mediated resistance signaling genes, OsICS1 and OsNPR1, in Bph14 and MH63 plants upon the cessation of BPH feeding. c, Relative transcript levels of Bph14-mediated resistance signaling genes, OsICS1 and OsNPR1, in the Bph14-Osnbr1− (Bph14-Osnbr1) lines (Bph14-Osnbr1−-1 and Bph14-Osnbr1−-2) and Bph14 plants upon the cessation of BPH feeding. d, BPH weight gain on the Bph14-Osnbr1− lines (Bph14-Osnbr1−-1 and Bph14-Osnbr1−-2) and Bph14 plants after 48 h of feeding. e, Honeydew excretion by BPHs reared on the Bph14-Osnbr1− lines and Bph14 plants after 48 h of feeding. f, Two-host choice test of BPHs feeding on the Bph14-Osnbr1− lines and Bph14 plants. g, Survival rates of BPHs feeding on the Bph14-Osnbr1− lines and Bph14 plants for 8 days. In a,b,c, P values were derived by one-way ANOVA. Data are mean ± s.d. (n = 3, biologically independent experiments). In d,e, P values were derived by one-way ANOVA (n = 18, biologically independent samples). In f,g,P values were derived by one-way ANOVA. Data are mean ± s.e.m. (n = 10, biologically independent samples). In box plots in d,e, the box limits indicate the 25th and 75th percentiles, the whiskers indicate the full range of the data, and the center line indicates the median. Individual data points are plotted.
Supplementary information
Supplementary Fig. 1
Uncropped blots and gel images.
Supplementary Fig. 2
The other two repeats of Fig. 1a–e,h–k.
Supplementary Table 1
New rice varieties introgressed with Bph14 and authorized for release into production.
Supplementary Table 2
BPH14-interacting proteins from BPH identified in a Y2H assay.
Supplementary Table 3
List of primers used in this study.
Supplementary Table 4
Statistical summary—a summary of all statistical analysis is provided. Source data for all of the statistical analysis can be found in the source data files.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Guo, J., Wang, H., Guan, W. et al. A tripartite rheostat controls self-regulated host plant resistance to insects. Nature 618, 799–807 (2023). https://doi.org/10.1038/s41586-023-06197-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-023-06197-z
This article is cited by
-
Advances in molecular interactions between rice and insect herbivores
Crop Health (2024)
-
Callose deposition regulates differences in cotton aphid resistance among six watermelon varieties
Journal of Pest Science (2024)
-
Autophagy receptor OsNBR1 modulates salt stress tolerance in rice
Plant Cell Reports (2024)
-
Genome-wide identification of nucleotide-binding domain leucine-rich repeat (NLR) genes and their association with green peach aphid (Myzus persicae) resistance in peach
BMC Plant Biology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.