Abstract
Hepatocellular carcinoma (HCC) is the most common form of primary adult liver cancer. After nearly a decade with sorafenib as the only approved treatment, multiple new agents have demonstrated efficacy in clinical trials, including the targeted therapies regorafenib, lenvatinib and cabozantinib, the anti-angiogenic antibody ramucirumab, and the immune checkpoint inhibitors nivolumab and pembrolizumab. Although these agents offer new promise to patients with HCC, the optimal choice and sequence of therapies remains unknown and without established biomarkers, and many patients do not respond to treatment. The advances and the decreasing costs of molecular measurement technologies enable profiling of HCC molecular features (such as genome, transcriptome, proteome and metabolome) at different levels, including bulk tissues, animal models and single cells. The release of such data sets to the public enhances the ability to search for information from these legacy studies and provides the opportunity to leverage them to understand HCC mechanisms, rationally develop new therapeutics and identify candidate biomarkers of treatment response. Here, we provide a comprehensive review of public data sets related to HCC and discuss how emerging artificial intelligence methods can be applied to identify new targets and drugs as well as to guide therapeutic choices for improved HCC treatment.
Key points
-
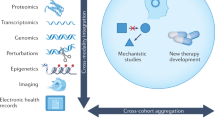
The past few years have witnessed the generation of big omics data across multiple modalities in hepatocellular carcinoma (HCC) — from primary to metastatic cancer, from bulk tissues to single cells and from patients to preclinical models.
-
Big data brings new hope but also new challenges in translating data points to therapeutics.
-
Multiple new targeted therapies have shown efficacy in HCC, yet the optimal choice and sequence of therapies for individual patients is unknown, without established clinical biomarkers of response or resistance.
-
A systems approach that aims to target a list of disease molecular features, such as gene expression signatures, can be used to complement the conventional target-based approach.
-
Big data analysis, including pan-cancer studies, might help quantify biological differences between preclinical models and patients, further guiding translational research, which is especially critical for understudied cancers such as HCC.
-
Emerging artificial intelligence methods, including deep learning, could empower big data in HCC therapeutic discovery and identification of predictive biomarkers.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Change history
11 March 2020
A Correction to this paper has been published: https://doi.org/10.1038/s41575-020-0288-6
References
Bray, F. et al. Global Cancer Statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 68, 394–424 (2018).
Global Burden of Disease Liver Cancer Collaboration et al. The burden of primary liver cancer and underlying etiologies from 1990 to 2015 at the global, regional, and national level: results from the global burden of disease study 2015. JAMA Oncol. 3, 1683–1691 (2017).
Ryerson, A. B. et al. Annual report to the nation on the status of cancer, 1975-2012, featuring the increasing incidence of liver cancer. Cancer 122, 1312–1337 (2016).
American Cancer Society. Key statistics about liver cancer. American Cancer Society https://www.cancer.org/cancer/liver-cancer/about/what-is-key-statistics.html (2019).
Singal, A. G. & El-Serag, H. B. Hepatocellular carcinoma from epidemiology to prevention: translating knowledge into practice. Clin. Gastroenterol. Hepatol. 13, 2140–2151 (2015).
de Lope, C. R., Tremosini, S., Forner, A., Reig, M. & Bruix, J. Management of HCC. J. Hepatol. 56 (Suppl. 1), S75–S87 (2012).
Mazzaferro, V. et al. Liver transplantation for the treatment of small hepatocellular carcinomas in patients with cirrhosis. N. Engl. J. Med. 334, 693–699 (1996).
Llovet, J. M. et al. Arterial embolisation or chemoembolisation versus symptomatic treatment in patients with unresectable hepatocellular carcinoma: a randomised controlled trial. Lancet 359, 1734–1739 (2002).
Llovet, J. M. et al. Sorafenib in advanced hepatocellular carcinoma. N. Engl. J. Med. 359, 378–390 (2008).
Cheng, A. L. et al. Efficacy and safety of sorafenib in patients in the Asia-Pacific region with advanced hepatocellular carcinoma: a phase III randomised, double-blind, placebo-controlled trial. Lancet. Oncol. 10, 25–34 (2009).
Kudo, M. et al. Lenvatinib versus sorafenib in first-line treatment of patients with unresectable hepatocellular carcinoma: a randomised phase 3 non-inferiority trial. Lancet 391, 1163–1173 (2018).
Bruix, J. et al. Regorafenib for patients with hepatocellular carcinoma who progressed on sorafenib treatment (RESORCE): a randomised, double-blind, placebo-controlled, phase 3 trial. Lancet 389, 56–66 (2017).
Abou-Alfa, G. K. et al. Cabozantinib in patients with advanced and progressing hepatocellular carcinoma. N. Engl. J. Med. 379, 54–63 (2018).
Zhu, A. X. et al. Ramucirumab after sorafenib in patients with advanced hepatocellular carcinoma and increased α-fetoprotein concentrations (REACH-2): a randomised, double-blind, placebo-controlled, phase 3 trial. Lancet Oncol. 20, 282–296 (2019).
El-Khoueiry, A. B. et al. Nivolumab in patients with advanced hepatocellular carcinoma (CheckMate 040): an open-label, non-comparative, phase 1/2 dose escalation and expansion trial. Lancet 389, 2492–2502 (2017).
Zhu, A. X. et al. Pembrolizumab in patients with advanced hepatocellular carcinoma previously treated with sorafenib (KEYNOTE-224): a non-randomised, open-label phase 2 trial. Lancet Oncol. 19, 940–952 (2018).
Okusaka, T. & Ikeda, M. Immunotherapy for hepatocellular carcinoma: current status and future perspectives. ESMO Open. 3 (Suppl. 1), e000455 (2018).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT03434379 (2019).
Chen, B. & Butte, A. J. Leveraging big data to transform target selection and drug discovery. Clin. Pharmacol. Ther. 99, 285–297 (2016).
Wooden, B., Goossens, N., Hoshida, Y. & Friedman, S. L. Using big data to discover diagnostics and therapeutics for gastrointestinal and liver diseases. Gastroenterology 152, 53–67.e3 (2017).
Llovet, J. M., Montal, R., Sia, D. & Finn, R. S. Molecular therapies and precision medicine for hepatocellular carcinoma. Nat. Rev. Clin. Oncol. 15, 599–616 (2018).
Cancer Genome Atlas Research Network. Comprehensive and integrative genomic characterization of hepatocellular carcinoma. Cell 169, 1327–1341.e23 (2017).
Lin, C.-P., Liu, C.-R., Lee, C.-N., Chan, T.-S. & Liu, H. E. Targeting c-Myc as a novel approach for hepatocellular carcinoma. World J. Hepatol. 2, 16–20 (2010).
Belmar, J. & Fesik, S. W. Small molecule Mcl-1 inhibitors for the treatment of cancer. Pharmacol. Ther. 145, 76–84 (2015).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT02508467 (2019).
Stein, S. et al. Safety and clinical activity of 1L atezolizumab + bevacizumab in a phase Ib study in hepatocellular carcinoma (HCC). J. Clin. Oncol. 36 (Suppl. 15), 4074 (2018).
Schulze, K. et al. Exome sequencing of hepatocellular carcinomas identifies new mutational signatures and potential therapeutic targets. Nat. Genet. 47, 505–511 (2015).
Ahn, S. M. et al. Genomic portrait of resectable hepatocellular carcinomas: implications of RB1 and FGF19 aberrations for patient stratification. Hepatology 60, 1972–1982 (2014).
Fujimoto, A. et al. Whole-genome mutational landscape and characterization of noncoding and structural mutations in liver cancer. Nat. Genet. 48, 500–509 (2016).
Guichard, C. et al. Integrated analysis of somatic mutations and focal copy-number changes identifies key genes and pathways in hepatocellular carcinoma. Nat. Genet. 44, 694–698 (2012).
Totoki, Y. et al. Trans-ancestry mutational landscape of hepatocellular carcinoma genomes. Nat. Genet. 46, 1267–1273 (2014).
Chaudhary, K. et al. Multimodal meta-analysis of 1,494 hepatocellular carcinoma samples reveals significant impact of consensus driver genes on phenotypes. Clin. Cancer Res. 25, 463–472 (2019).
Iizuka, N. et al. Differential gene expression in distinct virologic types of hepatocellular carcinoma: association with liver cirrhosis. Oncogene 22, 3007–3014 (2003).
Chen, X. et al. Gene expression patterns in human liver cancers. Mol. Biol. Cell 13, 1929–1939 (2002).
Zhu, Z. W. et al. Enhanced glypican-3 expression differentiates the majority of hepatocellular carcinomas from benign hepatic disorders. Gut 48, 558–564 (2001).
Jia, H.-L. et al. Gene expression profiling reveals potential biomarkers of human hepatocellular carcinoma. Clin. Cancer Res. 13, 1133–1139 (2007).
GTEx Consortium. The genotype-tissue expression (GTEx) project. Nat. Genet. 45, 580–585 (2013).
Li, J. et al. TCPA: a resource for cancer functional proteomics data. Nat. Methods 10, 1046–1047 (2013).
Tsai, T.-H. et al. LC-MS/MS based serum proteomics for identification of candidate biomarkers for hepatocellular carcinoma. Proteomics 15, 2369–2381 (2015).
Jiang, Y. et al. Proteomics identifies new therapeutic targets of early-stage hepatocellular carcinoma. Nature 567, 257–261 (2019).
Huang, Q. et al. Metabolic characterization of hepatocellular carcinoma using nontargeted tissue metabolomics. Cancer Res. 73, 4992–5002 (2013).
Di Poto, C. et al. Metabolomic characterization of hepatocellular carcinoma in patients with liver cirrhosis for biomarker discovery. Cancer Epidemiol. Biomarkers Prev. 26, 675–683 (2017).
Chen, T. et al. Serum and urine metabolite profiling reveals potential biomarkers of human hepatocellular carcinoma. Mol. Cell. Proteomics 10, M110.004945 (2011).
Pfister, S. X. & Ashworth, A. Marked for death: targeting epigenetic changes in cancer. Nat. Rev. Drug Discov. 16, 241–263 (2017).
Revill, K. et al. Genome-wide methylation analysis and epigenetic unmasking identify tumor suppressor genes in hepatocellular carcinoma. Gastroenterology 145, 1424–1435.e1-25 (2013).
Robinson, D. R. et al. Integrative clinical genomics of metastatic cancer. Nature 548, 297–303 (2017).
Zheng, C. et al. Landscape of infiltrating T cells in liver cancer revealed by single-cell sequencing. Cell 169, 1342–1356.e16 (2017).
MacParland, S. A. et al. Single cell RNA sequencing of human liver reveals distinct intrahepatic macrophage populations. Nat. Commun. 9, 4383 (2018).
Zheng, H. et al. Single-cell analysis reveals cancer stem cell heterogeneity in hepatocellular carcinoma. Hepatology 68, 127–140 (2018).
Hou, Y. et al. Single-cell triple omics sequencing reveals genetic, epigenetic, and transcriptomic heterogeneity in hepatocellular carcinomas. Cell Res. 26, 304–319 (2016).
Han, X. et al. Mapping the mouse cell atlas by Microwell-seq. Cell 172, 1091–1107.e17 (2018).
Tabula Muris Consortium et al. Single-cell transcriptomics of 20 mouse organs creates a Tabula Muris. Nature 562, 367–372 (2018).
Regev, A. et al. The human cell atlas. eLife 6, e27041 (2017).
Barretina, J. et al. The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature 483, 603–607 (2012).
Ghandi, M. et al. Next-generation characterization of the Cancer Cell Line Encyclopedia. Nature 569, 503–508 (2019).
Klijn, C. et al. A comprehensive transcriptional portrait of human cancer cell lines. Nat. Biotechnol. 33, 306–312 (2015).
Yang, W. et al. Genomics of drug sensitivity in cancer (GDSC): a resource for therapeutic biomarker discovery in cancer cells. Nucleic Acids Res. 41, D955–D961 (2013).
Li, J. et al. Characterization of human cancer cell lines by reverse-phase protein arrays. Cancer Cell 31, 225–239 (2017).
Qiu, Z. et al. A pharmacogenomic landscape in human liver cancers. Cancer Cell 36, 179–193.e11 (2019).
Broutier, L. et al. Human primary liver cancer-derived organoid cultures for disease modeling and drug screening. Nat. Med. 23, 1424–1435 (2017).
Chen, X. & Calvisi, D. F. Hydrodynamic transfection for generation of novel mouse models for liver cancer research. Am. J. Pathol. 184, 912–923 (2014).
Ruiz de Galarreta, M. et al. β-catenin activation promotes immune escape and resistance to anti-PD-1 therapy in hepatocellular carcinoma. Cancer Discov. 9, 1124–1141 (2019).
Joshi, J. J. et al. H3B-6527 is a potent and selective inhibitor of FGFR4 in FGF19-driven hepatocellular carcinoma. Cancer Res. 77, 6999–7013 (2017).
Huynh, H. et al. Infigratinib mediates vascular normalization, impairs metastasis, and improves chemotherapy in hepatocellular carcinoma. Hepatology 69, 943–958 (2019).
Lee, J.-S. et al. Application of comparative functional genomics to identify best-fit mouse models to study human cancer. Nat. Genet. 36, 1306–1311 (2004).
Conte, N. et al. PDX Finder: a portal for patient-derived tumor xenograft model discovery. Nucleic Acids Res. 47, D1073–D1079 (2019).
Su, W.-H. et al. OncoDB.HCC: an integrated oncogenomic database of hepatocellular carcinoma revealed aberrant cancer target genes and loci. Nucleic Acids Res. 35, D727–D731 (2007).
He, S. et al. PDXliver: a database of liver cancer patient derived xenograft mouse models. BMC Cancer 18, 550 (2018).
Cocco, E., Scaltriti, M. & Drilon, A. NTRK fusion-positive cancers and TRK inhibitor therapy. Nat. Rev. Clin. Oncol. 15, 731–747 (2018).
National Cancer Institute. Targeted Cancer Therapies Fact Sheet. NCI https://www.cancer.gov/about-cancer/treatment/types/targeted-therapies/targeted-therapies-fact-sheet (2019).
Vilchez, V., Turcios, L., Marti, F. & Gedaly, R. Targeting Wnt/β-catenin pathway in hepatocellular carcinoma treatment. World J. Gastroenterol. 22, 823–832 (2016).
Meek, D. W. Regulation of the p53 response and its relationship to cancer. Biochem. J. 469, 325–346 (2015).
Toledo, F. & Wahl, G. M. MDM2 and MDM4: p53 regulators as targets in anticancer therapy. Int. J. Biochem. Cell Biol. 39, 1476–1482 (2007).
Ruden, M. & Puri, N. Novel anticancer therapeutics targeting telomerase. Cancer Treat. Rev. 39, 444–456 (2013).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT02989857 (2019).
Rubio-Perez, C. et al. In silico prescription of anticancer drugs to cohorts of 28 tumor types reveals targeting opportunities. Cancer Cell 27, 382–396 (2015).
Lin, D.-C. et al. Genomic and epigenomic heterogeneity of hepatocellular carcinoma. Cancer Res. 77, 2255–2265 (2017).
Thillai, K., Ross, P. & Sarker, D. Molecularly targeted therapy for advanced hepatocellular carcinoma - a drug development crisis? World J. Gastrointest. Oncol. 8, 173–185 (2016).
Rimassa, L. et al. Tivantinib for second-line treatment of MET-high, advanced hepatocellular carcinoma (METIV-HCC): a final analysis of a phase 3, randomised, placebo-controlled study. Lancet Oncol. 19, 682–693 (2018).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT02528643 (2019).
Liu, M. et al. Integrative epigenetic analysis reveals therapeutic targets to the DNA methyltransferase inhibitor guadecitabine (SGI-110) in hepatocellular carcinoma. Hepatology 68, 1412–1428 (2018).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT01752933 (2019).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT03257761 (2019).
Dudley, J. T. et al. Computational repositioning of the anticonvulsant topiramate for inflammatory bowel disease. Sci. Transl. Med. 3, 96ra76 (2011).
Jahchan, N. S. et al. A drug repositioning approach identifies tricyclic antidepressants as inhibitors of small cell lung cancer and other neuroendocrine tumors. Cancer Discov. 3, 1364–1377 (2013).
Brum, A. M. et al. Connectivity Map-based discovery of parbendazole reveals targetable human osteogenic pathway. Proc. Natl Acad. Sci. USA 112, 12711–12716 (2015).
Lamb, J. et al. The Connectivity Map: using gene-expression signatures to connect small molecules, genes, and disease. Science 313, 1929–1935 (2006).
Pessetto, Z. Y. et al. In silico and in vitro drug screening identifies new therapeutic approaches for Ewing sarcoma. Oncotarget 8, 4079–4095 (2017).
Chen, B. et al. Computational discovery of niclosamide ethanolamine, a repurposed drug candidate that reduces growth of hepatocellular carcinoma cells in vitro and in mice by inhibiting cell division cycle 37 signaling. Gastroenterology 152, 2022–2036 (2017).
Chen, B. et al. Reversal of cancer gene expression correlates with drug efficacy and reveals therapeutic targets. Nat. Commun. 8, 16022 (2017).
Caicedo, J. C., Singh, S. & Carpenter, A. E. Applications in image-based profiling of perturbations. Curr. Opin. Biotechnol. 39, 134–142 (2016).
Duffy, A. G. et al. Tremelimumab in combination with ablation in patients with advanced hepatocellular carcinoma. J. Hepatol. 66, 545–551 (2017).
Robert, C. et al. Pembrolizumab versus ipilimumab in advanced melanoma. N. Engl. J. Med. 372, 2521–2532 (2015).
Borghaei, H. et al. Nivolumab versus docetaxel in advanced nonsquamous non-small-cell lung cancer. N. Engl. J. Med. 373, 1627–1639 (2015).
Thorsson, V. et al. The immune landscape of cancer. Immunity 48, 812–830.e14 (2018).
Varn, F. S., Wang, Y., Mullins, D. W., Fiering, S. & Cheng, C. Systematic pan-cancer analysis reveals immune cell interactions in the tumor microenvironment. Cancer Res. 77, 1271–1282 (2017).
Rooney, M. S., Shukla, S. A., Wu, C. J., Getz, G. & Hacohen, N. Molecular and genetic properties of tumors associated with local immune cytolytic activity. Cell 160, 48–61 (2015).
Sia, D. et al. Identification of an immune-specific class of hepatocellular carcinoma, based on molecular features. Gastroenterology 153, 812–826 (2017).
Rohr-Udilova, N. et al. Deviations of the immune cell landscape between healthy liver and hepatocellular carcinoma. Sci. Rep. 8, 6220 (2018).
Grinberg-Bleyer, Y. et al. NF-κB c-Rel is crucial for the regulatory T cell immune checkpoint in cancer. Cell 170, 1096–1108.e13 (2017).
Lee, J.-S. et al. Classification and prediction of survival in hepatocellular carcinoma by gene expression profiling. Hepatology 40, 667–676 (2004).
Hoshida, Y. et al. Integrative transcriptome analysis reveals common molecular subclasses of human hepatocellular carcinoma. Cancer Res. 69, 7385–7392 (2009).
Liu, G., Dong, C. & Liu, L. Integrated multiple “-omics” data reveal subtypes of hepatocellular carcinoma. PLOS ONE 11, e0165457 (2016).
Chaudhary, K., Poirion, O. B., Lu, L. & Garmire, L. X. Deep learning-based multi-omics integration robustly predicts survival in liver cancer. Clin. Cancer Res. 24, 1248–1259 (2018).
Zucman-Rossi, J., Villanueva, A., Nault, J.-C. & Llovet, J. M. Genetic landscape and biomarkers of hepatocellular carcinoma. Gastroenterology 149, 1226–1239.e4 (2015).
Iorio, F. et al. A landscape of pharmacogenomic interactions in cancer. Cell 166, 740–754 (2016).
Seashore-Ludlow, B. et al. Harnessing connectivity in a large-scale small-molecule sensitivity dataset. Cancer Discov. 5, 1210–1223 (2015).
Basu, A. et al. An interactive resource to identify cancer genetic and lineage dependencies targeted by small molecules. Cell 154, 1151–1161 (2013).
Zhu, A. X. et al. REACH-2: A randomized, double-blind, placebo-controlled phase 3 study of ramucirumab versus placebo as second-line treatment in patients with advanced hepatocellular carcinoma (HCC) and elevated baseline α-fetoprotein (AFP) following first-line sorafenib. J. Clin. Oncol. 36 (Suppl. 15), 4003 (2018).
Hoshida, Y. et al. Molecular classification and novel targets in hepatocellular carcinoma: recent advancements. Semin. Liver Dis. 30, 35–51 (2010).
Yoshihara, K. et al. Inferring tumour purity and stromal and immune cell admixture from expression data. Nat. Commun. 4, 2612 (2013).
Chen, B., Sirota, M., Fan-Minogue, H., Hadley, D. & Butte, A. J. Relating hepatocellular carcinoma tumor samples and cell lines using gene expression data in translational research. BMC Med. Genomics 8 (Suppl. 2), S5 (2015).
Uhlen, M. et al. A pathology atlas of the human cancer transcriptome. Science 357, eaan2507 (2017).
Hirschfield, H. et al. In vitro modeling of hepatocellular carcinoma molecular subtypes for anti-cancer drug assessment. Exp. Mol. Med. 50, e419 (2018).
Chen, H., Engkvist, O., Wang, Y., Olivecrona, M. & Blaschke, T. The rise of deep learning in drug discovery. Drug Discov. Today 23, 1241–1250 (2018).
Mamoshina, P., Vieira, A., Putin, E. & Zhavoronkov, A. Applications of deep learning in biomedicine. Mol. Pharmaceutics 13, 1445–1454 (2016).
Ma, J., Sheridan, R. P., Liaw, A., Dahl, G. E. & Svetnik, V. Deep neural nets as a method for quantitative structure–activity relationships. J. Chem. Inf. Model. 55, 263–274 (2015).
Coley, C. W., Barzilay, R., Green, W. H., Jaakkola, T. S. & Jensen, K. F. Convolutional embedding of attributed molecular graphs for physical property prediction. J. Chem. Inf. Model. 57, 1757–1772 (2017).
Lusci, A., Pollastri, G. & Baldi, P. Deep architectures and deep learning in chemoinformatics: the prediction of aqueous solubility for drug-like molecules. J. Chem. Inf. Model. 53, 1563–1575 (2013).
Aliper, A. et al. Deep learning applications for predicting pharmacological properties of drugs and drug repurposing using transcriptomic data. Mol. Pharm. 13, 2524–2530 (2016).
Kadurin, A., Nikolenko, S., Khrabrov, K., Aliper, A. & Zhavoronkov, A. druGAN: an advanced generative adversarial autoencoder model for de novo generation of new molecules with desired molecular properties in silico. Mol. Pharmaceutics 14, 3098–3104 (2017).
Merkwirth, C. & Lengauer, T. Automatic generation of complementary descriptors with molecular graph networks. J. Chem. Inf. Model. 45, 1159–1168 (2005).
Liu, B. et al. Retrosynthetic reaction prediction using neural sequence-to-sequence models. ACS Cent. Sci. 3, 1103–1113 (2017).
Alakwaa, F. M., Chaudhary, K. & Garmire, L. X. Deep learning accurately predicts estrogen receptor status in breast cancer metabolomics data. J. Proteome Res. 17, 337–347 (2018).
Christiansen, E. M. et al. In silico labeling: predicting fluorescent labels in unlabeled images. Cell 173, 792–803.e19 (2018).
Zeng, W. Z. D., Glicksberg, B. S., Li, Y. & Chen, B. Selecting precise reference normal tissue samples for cancer research using a deep learning approach. BMC Med. Genomics 12 (Suppl. 1), 21 (2019).
Yala, A., Lehman, C., Schuster, T., Portnoi, T. & Barzilay, R. A deep learning mammography-based model for improved breast cancer risk prediction. Radiology 292, 60–66 (2019).
Topol, E. J. High-performance medicine: the convergence of human and artificial intelligence. Nat. Med. 25, 44–56 (2019).
Arisdakessian, C., Poirion, O., Yunits, B., Zhu, X. & Garmire, L. DeepImpute: an accurate, fast and scalable deep neural network method to impute single-cell RNA-seq data. Genome Biol. 20, 211 (2018).
Eraslan, G., Simon, L. M., Mircea, M., Mueller, N. S. & Theis, F. J. Single-cell RNA-seq denoising using a deep count autoencoder. Nat. Commun. 10, 390 (2019).
Hoadley, K. A. et al. Cell-of-origin patterns dominate the molecular classification of 10,000 tumors from 33 types of cancer. Cell 173, 291–304.e6 (2018).
Yu, L.-X. & Schwabe, R. F. The gut microbiome and liver cancer: mechanisms and clinical translation. Nat. Rev. Gastroenterol. Hepatol. 14, 527–539 (2017).
Gao, J. et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci. Signal. 6, pl1 (2013).
Acknowledgements
This research was supported by grants K01ES025434 awarded by the National Institute of Environmental Health Sciences through funds provided by the trans-NIH Big Data to Knowledge (BD2K) initiative (www.bd2k.nih.gov), P20 COBRE GM103457 awarded by the NIH National Institute of General Medical Sciences, R01 LM012373 awarded by the National Library of Medicine, R01 HD084633 awarded by the National Institute of Child Health and Human Development, and Hawaii Community Foundation Medical Research Grant 14ADVC-64566 to L.X.G; the CJ Huang Foundation, HM Lui Foundation, and TS Kwok Liver Research Foundation to M.S.C; and R21 TR001743 and K01 ES028047 and the MSU Global Impact Initiative to B.C.
Author information
Authors and Affiliations
Contributions
All authors contributed equally to the article.
Corresponding author
Ethics declarations
Competing interests
R.K.K. declares the following competing interests: research funding and/or supply of study drug to institution for conduct of clinical trials from Adaptimmune, Agios, AstraZeneca, Bayer, Bristol–Myers Squibb, Eli Lilly and Co, EMD Serono, Exelixis, Merck, Novartis, Partner Therapeutics, QED, Taiho; funding (to individual) for Independent Data Monitoring Committee membership by Genentech/Roche; Steering Committee/Advisory Board memberships (funding to institution) by Agios, AstraZeneca, Bristol–Myers Squibb; Steering Committee (without compensation): Exelixis. The other authors declare no competing interests.
Additional information
Peer review information
Nature Reviews Gastrenterology & Hepatology thanks J. Lee, D. Sia and C.-M. Wong for their contribution to the peer review of this work.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Broad GDAC: https://gdac.broadinstitute.org/
Cancer Cell Line Encyclopedia: https://portals.broadinstitute.org/ccle
Cancer Therapeutics Response Portal: http://portals.broadinstitute.org/ctrp.v2.1/
cBioPortal: http://www.cbioportal.org/
CHNPP Data Portal LIVER: http://liver.cnhpp.ncpsb.org/auth/login
dbGaP: https://www.ncbi.nlm.nih.gov/gap/
EGA portal: http://www.ebi.ac.uk/ega/
Gene Expression Omnibus: https://www.ncbi.nlm.nih.gov/geo/
Genomics of Drug Sensitivity in Cancer Project: https://www.cancerrxgene.org/
GTEx Portal: https://www.gtexportal.org/
Human Proteome Map: http://humanproteomemap.org/
Library of Integrated Network-based Cellular Signatures: http://www.lincscloud.org/
Liver Cancer Model Repository: http://www.picb.ac.cn/limore/
NCI Genomic Data Commons portal: https://portal.gdc.cancer.gov/
PDXLiver: http://www.picb.ac.cn/PDXliver/
Project Achilles: https://depmap.org/portal/achilles/
The Human Protein Atlas: http://www.proteinatlas.org/
Rights and permissions
About this article
Cite this article
Chen, B., Garmire, L., Calvisi, D.F. et al. Harnessing big ‘omics’ data and AI for drug discovery in hepatocellular carcinoma. Nat Rev Gastroenterol Hepatol 17, 238–251 (2020). https://doi.org/10.1038/s41575-019-0240-9
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41575-019-0240-9