Abstract
The cyclic-oligonucleotide-based anti-phage signalling system (CBASS) is a type of innate prokaryotic immune system. Composed of a cyclic GMP–AMP synthase (cGAS) and CBASS-associated proteins, CBASS uses cyclic oligonucleotides to activate antiviral immunity. One major class of CBASS contains a homologue of eukaryotic ubiquitin-conjugating enzymes, which is either an E1–E2 fusion or a single E2. However, the functions of single E2s in CBASS remain elusive. Here, using biochemical, genetic, cryo-electron microscopy and mass spectrometry investigations, we discover that the E2 enzyme from Serratia marcescens regulates cGAS by imitating the ubiquitination cascade. This includes the processing of the cGAS C terminus, conjugation of cGAS to a cysteine residue, ligation of cGAS to a lysine residue, cleavage of the isopeptide bond and poly-cGASylation. The poly-cGASylation activates cGAS to produce cGAMP, which acts as an antiviral signal and leads to cell death. Thus, our findings reveal a unique regulatory role of E2 in CBASS.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Data supporting the findings of this study are available in this Article and its Supplementary Information. Genomes of S. marcescens were sequenced previously, and sequences are publicly available. GenBank accessions for the CBASS operons used in this study are found in Methods. Primers used in this study are provided in Supplementary Tables 3 and 4. The E. coli strain B/BL21-DE3 proteome database that was used in this study can be accessed at Uniprot (UP000002032). The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium (https://proteomecentral.proteomexchange.org) via the iProX partner repository56,57 with the dataset identifier PXD050178. For the cGAS–E2 complex, coordinates are available at the RCSB PDB (http://www.rcsb.org) under accession codes 8HSB and 8YJY, and electron microscopy maps are available at the Electron Microscopy Data Bank (EMDB; https://www.ebi.ac.uk/emdb/) under accession codes EMD-34992 and EMD-39353. Source data are provided with this paper.
Code availability
No custom code was used for data analyses.
References
Sun, L., Wu, J., Du, F., Chen, X. & Chen, Z. J. Cyclic GMP–AMP synthase is a cytosolic DNA sensor that activates the type I interferon pathway. Science 339, 786–791 (2013).
Wu, J. et al. Cyclic GMP–AMP is an endogenous second messenger in innate immune signaling by cytosolic DNA. Science 339, 826–830 (2013).
Davies, B. W., Bogard, R. W., Young, T. S. & Mekalanos, J. J. Coordinated regulation of accessory genetic elements produces cyclic di-nucleotides for V. cholerae virulence. Cell 149, 358–370 (2012).
Kranzusch, P. J., Lee, A. S., Berger, J. M. & Doudna, J. A. Structure of human cGAS reveals a conserved family of second-messenger enzymes in innate immunity. Cell Rep. 3, 1362–1368 (2013).
Whiteley, A. T. et al. Bacterial cGAS-like enzymes synthesize diverse nucleotide signals. Nature 567, 194–199 (2019).
Cohen, D. et al. Cyclic GMP–AMP signalling protects bacteria against viral infection. Nature 574, 691–695 (2019).
Morehouse, B. R. et al. STING cyclic dinucleotide sensing originated in bacteria. Nature 586, 429–433 (2020).
Ye, Q. et al. HORMA domain proteins and a Trip13-like ATPase regulate bacterial cGAS-like enzymes to mediate bacteriophage immunity. Mol. Cell 77, 709–722.e7 (2020).
Burroughs, A. M., Zhang, D., Schaffer, D. E., Iyer, L. M. & Aravind, L. Comparative genomic analyses reveal a vast, novel network of nucleotide-centric systems in biological conflicts, immunity and signaling. Nucleic Acids Res. 43, 10633–10654 (2015).
Millman, A., Melamed, S., Amitai, G. & Sorek, R. Diversity and classification of cyclic-oligonucleotide-based anti-phage signalling systems. Nat. Microbiol. 5, 1608–1615 (2020).
Lowey, B. et al. CBASS immunity uses CARF-related effectors to sense 3′–5′- and 2′–5′-linked cyclic oligonucleotide signals and protect bacteria from phage infection. Cell 182, 38–49.e17 (2020).
Kerscher, O., Felberbaum, R. & Hochstrasser, M. Modification of proteins by ubiquitin and ubiquitin-like proteins. Annu. Rev. Cell Dev. Biol. 22, 159–180 (2006).
Li, W. & Ye, Y. Polyubiquitin chains: functions, structures, and mechanisms. Cell. Mol. Life Sci. 65, 2397–2406 (2008).
Ledvina, H. E. et al. An E1–E2 fusion protein primes antiviral immune signalling in bacteria. Nature 616, 319–325 (2023).
Jenson, J. M., Li, T., Du, F., Ea, C. K. & Chen, Z. J. Ubiquitin-like conjugation by bacterial cGAS enhances anti-phage defence. Nature 616, 326–331 (2023).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Mirdita, M. & Schütze, K. ColabFold: making protein folding accessible to all. Nat. Methods 19, 679–682 (2022).
Holm, L., Laiho, A., Törönen, P. & Salgado, M. DALI shines a light on remote homologs: one hundred discoveries. Protein Sci. 32, e4519 (2023).
Dou, H., Buetow, L., Sibbet, G. J., Cameron, K. & Huang, D. T. BIRC7–E2 ubiquitin conjugate structure reveals the mechanism of ubiquitin transfer by a RING dimer. Nat. Struct. Mol. Biol. 19, 876–883 (2012).
Termathe, M. & Leidel, S. A. The Uba4 domain interplay is mediated via a thioester that is critical for tRNA thiolation through Urm1 thiocarboxylation. Nucleic Acids Res. 46, 5171–5181 (2018).
Hendrickx, A. P., Budzik, J. M., Oh, S. Y. & Schneewind, O. Architects at the bacterial surface—sortases and the assembly of pili with isopeptide bonds. Nat. Rev. Microbiol. 9, 166–176 (2011).
Jacobitz, A. W., Kattke, M. D., Wereszczynski, J. & Clubb, R. T. Sortase transpeptidases: structural biology and catalytic mechanism. Adv. Protein Chem. Struct. Biol. 109, 223–264 (2017).
Lowther, J. et al. The importance of pH in regulating the function of the Fasciola hepatica cathepsin L1 cysteine protease. PLoS Negl. Trop. Dis. 3, e369 (2009).
Zhao, L. et al. Structural analysis of asparaginyl endopeptidase reveals the activation mechanism and a reversible intermediate maturation stage. Cell Res. 24, 344–358 (2014).
Verma, S., Dixit, R. & Pandey, K. C. Cysteine proteases: modes of activation and future prospects as pharmacological targets. Front. Pharmacol. 7, 107 (2016).
Elsässer, B. et al. Distinct roles of catalytic cysteine and histidine in the protease and ligase mechanisms of human legumain as revealed by DFT-based QM/MM simulations. ACS Catal. 7, 5585–5593 (2017).
Feliciangeli, S. F. et al. Identification of a pH sensor in the furin propeptide that regulates enzyme activation. J. Biol. Chem. 281, 16108–16116 (2006).
Williamson, D. M., Elferich, J., Ramakrishnan, P., Thomas, G. & Shinde, U. The mechanism by which a propeptide-encoded pH sensor regulates spatiotemporal activation of furin. J. Biol. Chem. 288, 19154–19165 (2013).
Vernet, T. et al. Processing of the papain precursor. The ionization state of a conserved amino acid motif within the Pro region participates in the regulation of intramolecular processing. J. Biol. Chem. 270, 10838–10846 (1995).
Fuentes-Prior, P. & Salvesen, G. S. The protein structures that shape caspase activity, specificity, activation and inhibition. Biochem. J. 384, 201–232 (2004).
Li, X. et al. Cyclic GMP–AMP synthase is activated by double-stranded DNA-induced oligomerization. Immunity 39, 1019–1031 (2013).
Yu, L. & Liu, P. Cytosolic DNA sensing by cGAS: regulation, function, and human diseases. Signal Transduct. Target. Ther. 6, 170 (2021).
Duncan-Lowey, B., McNamara-Bordewick, N. K., Tal, N., Sorek, R. & Kranzusch, P. J. Effector-mediated membrane disruption controls cell death in CBASS antiphage defense. Mol. Cell 81, 5039–5051.e5035 (2021).
David, Y., Ziv, T., Admon, A. & Navon, A. The E2 ubiquitin-conjugating enzymes direct polyubiquitination to preferred lysines. J. Biol. Chem. 285, 8595–8604 (2010).
Stewart, M. D., Ritterhoff, T., Klevit, R. E. & Brzovic, P. S. E2 enzymes: more than just middle men. Cell Res. 26, 423–440 (2016).
Burroughs, A. M., Jaffee, M., Iyer, L. M. & Aravind, L. Anatomy of the E2 ligase fold: implications for enzymology and evolution of ubiquitin/Ub-like protein conjugation. J. Struct. Biol. 162, 205–218 (2008).
Pearce, M. J., Mintseris, J., Ferreyra, J., Gygi, S. P. & Darwin, K. H. Ubiquitin-like protein involved in the proteasome pathway of Mycobacterium tuberculosis. Science 322, 1104–1107 (2008).
Burns, K. E. et al. “Depupylation” of prokaryotic ubiquitin-like protein from mycobacterial proteasome substrates. Mol. Cell 39, 821–827 (2010).
Iyer, L. M., Burroughs, A. M. & Aravind, L. Unraveling the biochemistry and provenance of pupylation: a prokaryotic analog of ubiquitination. Biol. Direct 3, 45 (2008).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Bepler, T. et al. Positive-unlabeled convolutional neural networks for particle picking in cryo-electron micrographs. Nat. Methods 16, 1153–1160 (2019).
Zivanov, J., Nakane, T. & Scheres, S. H. W. A Bayesian approach to beam-induced motion correction in cryo-EM single-particle analysis. IUCrJ 6, 5–17 (2019).
Punjani, A., Zhang, H. & Fleet, D. J. Non-uniform refinement: adaptive regularization improves single-particle cryo-EM reconstruction. Nat. Methods 17, 1214–1221 (2020).
Punjani, A. & Fleet, D. J. 3D variability analysis: resolving continuous flexibility and discrete heterogeneity from single particle cryo-EM. J. Struct. Biol. 213, 107702 (2021).
Sanchez-Garcia, R. et al. DeepEMhancer: a deep learning solution for cryo-EM volume post-processing. Commun. Biol. 4, 874 (2021).
Pettersen, E. F. et al. UCSF ChimeraX: structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D. 66, 486–501 (2010).
Wang Ray, Y.-R. et al. Automated structure refinement of macromolecular assemblies from cryo-EM maps using Rosetta. eLife 5, e17219 (2016).
Liebschner, D., Afonine, P. V., Baker, M. L., Bunkóczi, G. & Adams, P. D. Macromolecular structure determination using X-rays, neutrons and electrons: recent developments in Phenix. Acta Crystallogr. D. 75, 861–877 (2019).
Leman, J. K. et al. Macromolecular modeling and design in Rosetta: recent methods and frameworks. Nat. Methods 17, 665–680 (2020).
Hu, M., Liu, Y., Yu, K. & Liu, X. Decreasing the amount of trypsin in in-gel digestion leads to diminished chemical noise and improved protein identifications. J. Proteomics 109, 16–25 (2014).
Mazzocco, A., Waddell, T. E., Lingohr, E. & Johnson, R. P. Enumeration of bacteriophages using the small drop plaque assay system. Methods Mol. Biol. 501, 81–85 (2009).
Chakraborty, S., Mizusaki, H. & Kenney, L. J. A FRET-based DNA biosensor tracks OmpR-dependent acidification of Salmonella during macrophage infection. PLoS Biol. 13, e1002116 (2015).
Burgstaller, S. et al. pH-Lemon, a fluorescent protein-based pH reporter for acidic compartments. ACS Sens. 4, 883–891 (2019).
Ma, J. et al. iProX: an integrated proteome resource. Nucleic Acids Res. 47, D1211–d1217 (2019).
Chen, T. & Ma, J. iProX in 2021: connecting proteomics data sharing with big data. Nucleic Acids Res. 50, D1522–d1527 (2022).
Acknowledgements
We would like to express our special gratitude to L. Liu of Tsinghua University for his invaluable guidance in the field of protein chemistry. We thank E.-D. Wang, W. Yan and H. Hu for their advice, P. Xu for providing materials, X. Li for technical support and all lab members for helpful discussions. We thank D. Li, X. Li and Y. Zeng of the Core Facility of Wuhan University for their assistance with cryo-EM grid screening and data collection. This work was supported by the National Natural Science Foundation of China (grants 32150009 and 31870165 to B.Z., 22174003 and 21974002 to X. Liu, and 31900032 to F.H.), the National Key Research and Development Program of China (2022YFA0912200 to L.W. and 2022YFA1304500 to X. Liu) and the Fund from Science, Technology and Innovation Commission of Shenzhen Municipality (grant JCYJ20210324115811032 to B.Z.).
Author information
Authors and Affiliations
Contributions
F.H. and B.Z. conceived the project. Y.Y., J.X., W.X., F.H., D.W., X. Liu, L.W. and B.Z. designed the experiments. Y.Y., J.X., W.X. and D.W. performed the experiments. J.X. and L.W. determined and analysed structures. Y.Y., F.H., W.X., G.K.O., C.V.R., Hao Wu, D.W., X. Liu, L.W. and B.Z. analysed the data and wrote the paper. All authors discussed the results and contributed to the final paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Microbiology thanks Karl-Peter Hopfner and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 In vitro characterization of the enzymatic activity of cGAS.
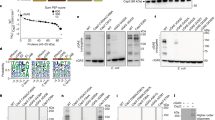
a, HPLC analysis of nucleotide second messenger synthesis by cGAS using various nucleotide substrates. cGAS synthesizes a product only in the presence of both ATP and GTP. b, cGAS synthesis activity is dependent on Mn2+. c, Optimal Mn2+ concentration for cGAS. The products synthesized at various Mn2+ concentrations were quantified by HPLC. Data are mean ± s.d. for n = 3 independent replicates. d, Comparison of the activities of cGAS and its D84A/D86A mutant. e, cGAS product was degraded by nuclease P1 but not CIP, as analyzed by HPLC. f, Comparison of the retention time of possible cGAMP variants (3′,3′-cGAMP, 2′,3′-cGAMP, and 3′,2′-cGAMP) and the cGAS product. g, MS/MS fragmentation spectra of the cGAS product (top) and the standard 3′,2′-cGAMP (bottom), and the chemical structure of 3′,2′-cGAMP. h, Michaelis-Menten kinetics plot of ATP and GTP for cGAS. Km, kcat and kcat/Km for ATP are 393.0 ± 15.3 μM, 2.4 ± 0.3 min−1 and 5.9 × 10-3, respectively. For GTP, Km, kcat and kcat/Km are 94.2 ± 11.1 μM, 2.6 ± 0.2 min−1 and 2.8 × 10-2, respectively. Data are mean ± s.d. for n = 3 independent replicates. All data are representative of three independent experiments.
Extended Data Fig. 2 Predicted structure of the cGAS-E2 complex.
a, SDS-PAGE analysis of the in vitro assembly of the covalent cGAS-E2 protein. Two proteins were mixed at a molar ratio of 1:1 and incubated at 4 °C for the indicated periods (cGAS+E2). Since both cGAS and E2 were His-tagged during the in vitro reconstruction, the molecular weight of cGAS-E2 protein was slightly higher than that obtained in vivo from His-tagged cGAS and non-tagged E2. Data are representative of three independent experiments. b,c, Chromatography traces and SDS-PAGE analysis of purified cGAS-E2 complex. Data are representative of three independent experiments. d, Representative cryo-EM micrograph of cGAS-E2 complex. Data are representative of three independent experiments. e, Ribbon diagram of the top ranked structure prediction of cGAS-E2 complex colored by the per-residue LDDT scores. f, Ribbon diagram of the top ranked structure prediction of cGAS-E2 complex with cGAS and E2 colored in green and orange, respectively.
Extended Data Fig. 3 Cryo-EM workflows and the quality of reconstructed cryo-EM maps.
a,b, Workflow of 3D reconstruction of the initial cGAS-E2 dataset (a) and the UltrAuFoil cGAS-E2 dataset (b). c, Directional FSC of the cGAS-E2 cryo-EM density map. d, Orientation distributions of cGAS-E2 3D reconstruction. e, Local resolutions of the cGAS-E2 cryo-EM map. f, Superimposed predicted and experimental structures of cGAS-E2.
Extended Data Fig. 4 Sequence alignment of cGAS and E2 homologs.
a, Sequence alignment of the C-terminal residues of cGAS and CD-NTase homologs from other E2-CBASSs. A conserved C-terminal glycine is marked by a red triangle. See also Supplementary Fig. 1. b, Sequence alignment of E2 and its orthologs shows two conserved cysteine residues (marked by red triangles). See also Supplementary Fig. 2.
Extended Data Fig. 5 Characterization of the bond formed between cGAS Y405 and E2 K159.
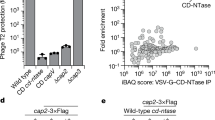
a, Magnified view of the E2 active site in cGAS-E2 complex showing continuous density between Y405 of cGAS and K159 of E2. b, Extracted ion chromatograms (left panel) and corresponding mass spectra of precursor ions (right panel) of the control peptide HPRPSEVKNER and the modified peptide HPRPSEVKNER bearing an NTFY remnant at K159 (red). MS signals of the control peptide were only detected in E2 but not cGAS or cGAS-E2 samples. In contrast, MS signals of the modified peptide were only detected in cGAS-E2 but not E2 or cGAS samples. All data are representative of three independent experiments.
Extended Data Fig. 6 Comparison of CD-NTase and E2 homologs.
a, Sequence logos for CD-NTase (cGAS) from type II CBASSs. Left and right show sequence logos for the C-terminal 10 residues of CD-NTase from E1E2/JAB-CBASSs and E2-CBASSs, respectively. Data are depicted as bits and signified by the height of each residue. Different colors indicate the chemical properties of amino acids. See also Supplementary Fig. 1, 3. b, Sequence alignment of E2 homologs from E2-CBASSs and E1E2/JAB-CBASSs show that they share low homology. The two crucial histidine residues (marked by red triangles) are conservative in E2 from E2-CBASSs but not E1E2/JAB-CBASSs. See also Supplementary Fig. 2, 4, 5. c, Sequence logos for the active site sequences of Cap2 (E2) from E2-CBASSs. Top left and right show 20 amino acids upstream and downstream of the active site cysteine, and 5 amino acids upstream and downstream of the residues corresponding to H152, respectively. The positions corresponding to H87, C101, and H152 are marked by red triangles. Bottom, Sequence logo for the active site sequences of E2 domain of Cap2 (E1E2) from E1E2/JAB-CBASSs. 20 amino acids upstream and downstream of the active site cysteine are shown. The active site cysteine is marked by a red triangle. Data are depicted as bits and signified by the height of each residue. Different colors indicate the chemical properties of different amino acids. See also Supplementary Figs. 2, 4. d, Superimposed structures of E2 (orange) and human E2 (gray, PDB ID 4AUQ) show that no similar catalytic site is found in human E2.
Extended Data Fig. 7 Characterization of the thioester bond in cGAS-E2.
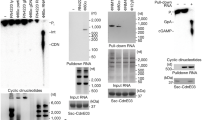
a, Extracted ion chromatograms (upper panel) and corresponding mass spectra of precursor ions (lower panel) of the modified peptide LCLFLPR bearing an NTFY remnant at C101 (red). MS signals of the modified peptide were only detected in cGAS-E2 samples without DTT treatment but not samples of E2, cGAS or cGAS-E2 treated with DTT. b, SDS-PAGE analysis showed that the thioester bond (-DTT) and isopeptide bond (+DTT) in cGAS-E2 are interconvertible at various pH. c, SDS-PAGE analysis of the in vitro processing of cGAS-MBP by E2, E2 (C101A), or E2 (H87A). All data are representative of three independent experiments.
Extended Data Fig. 8 Characterization of the isopeptide bond in poly-cGAS.
a, Extracted ion chromatograms (left panel) and corresponding mass spectra of precursor ions (right panel) of the control peptide EQGSLHLDKK and the modified peptide EQGSLHLDKK bearing an NTFY remnant at K381 (red). b, Extracted ion chromatograms (left panel) and corresponding mass spectra of precursor ions (right panel) of the control peptide TGGLIATGLAGTAAQAGVPKNTFYGE and the modified peptide TGGLIATGLAGTAAQAGVPKNTFYGE bearing an NTFY remnant at K401 (red). In both cases, MS signals of the modified peptides were only detected in cGAS-cGAS samples but not cGAS alone. c, SDS-PAGE analysis of cGAS mutants (indicated on above gel) co-purified with E2 K159A. All data are representative of three independent experiments.
Extended Data Fig. 9 Regulation of cGAS activity by E2-mediated poly-cGASylation.
a, HPLC analysis and quantification of the 3′,2′-cGAMP synthesized by cGAS and variants as shown in Fig. 5a, with concentrations of ATP, GPT, and Mn being 250 uM, 250 uM, and 2.5 mM, respectively. b, HPLC analysis and quantification of the 3′,2′-cGAMP synthesized by cGAS and variants as shown in Fig. 5b, with the concentration of ATP, GTP, or Mn2+ in the reaction reduced by half compared to those in a. All data are representative of three independent experiments.
Extended Data Fig. 10 Mechanisms underlying poly-cGAS formation.
a, 2D class averages of cGAS-E2 dimer (left) and the modeled structure based on the 2D averages (right). b, Intracellular pH change of E. coli BL21 during T4 phage infection. pH was determined using the intracellular pH probe BCECF-AM (red) or the FRET-based protein sensor (blue), respectively. Data are mean ± s.d. for n = 3 biological replicates.
Supplementary information
Supplementary Information
Supplementary Tables 1–4 and Figs. 1–5.
Source data
Source Data Fig. 1
Unprocessed gels.
Source Data Fig. 2
Unprocessed gels and statistical source data.
Source Data Fig. 3
Unprocessed gels and statistical source data.
Source Data Fig. 4
Unprocessed gels and statistical source data.
Source Data Fig. 5
Unprocessed western blots and gels, and statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Unprocessed gels and statistical source data.
Source Data Extended Data Fig. 7
Unprocessed gels.
Source Data Extended Data Fig. 8
Unprocessed gels.
Source Data Extended Data Fig. 10
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yan, Y., Xiao, J., Huang, F. et al. Phage defence system CBASS is regulated by a prokaryotic E2 enzyme that imitates the ubiquitin pathway. Nat Microbiol (2024). https://doi.org/10.1038/s41564-024-01684-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41564-024-01684-z