Abstract
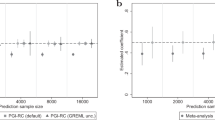
Polygenic indices (PGIs) are increasingly used to identify individuals at risk of developing disease and are advocated as screening tools for personalized medicine and education. Here we empirically assess rank concordance between PGIs created with different construction methods and discovery samples, focusing on cardiovascular disease and educational attainment. We find Spearman rank correlations between 0.17 and 0.93 for cardiovascular disease, and 0.40 and 0.83 for educational attainment, indicating highly unstable rankings across different PGIs for the same trait. Potential consequences for personalized medicine and gene–environment (G × E) interplay are illustrated using data from the UK Biobank. Simulations show how rank discordance mainly derives from a limited discovery sample size and reveal a tight link between the explained variance of a PGI and its ranking precision. We conclude that PGI-based ranking is highly dependent on PGI choice, such that current PGIs do not have the desired precision to be used routinely for personalized intervention.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Individual-level genotype and phenotype data are available by application via the UKB website (https://www.ukbiobank.ac.uk/enable-your-research/apply-for-access). The genome-wide summary statistics from 23andMe can be obtained by completing the 23andMe publication dataset access request form at https://research.23andme.com/dataset-access/. The genome wide summary statistics from CARDIoGRAM are available at http://www.cardiogramplusc4d.org/data-downloads/, file ‘CARDIoGRAMplusC4D 1000 Genomes-based GWAS - Additive’. The authors declare that the results supporting the findings of this study are available within the paper and its supplementary information files.
Code availability
Code for analyses and figures is available in a GitHub repository at https://github.com/DilnozaM/Rank-Concordance-of-PGI.
References
Visscher, P. M. et al. 10 Years of GWAS discovery: biology, function, and translation. Am. J. Hum. Genet. 101, 5–22 (2017).
Chabris, C. F., Lee, J. J., Cesarini, D., Benjamin, D. J. & Laibson, D. I. The fourth law of behavior genetics. Curr. Dir. Psychol. Sci. 24, 304–312 (2015).
Dudbridge, F. Power and predictive accuracy of polygenic risk scores. PLoS Genet 9, 1003348 (2013).
The International Schizophrenia Consortium. Common polygenic variation contributes to risk of schizophrenia and bipolar disorder. Nat. Lett. 460, 748–752 (2009).
Khera, A. V. et al. Genome-wide polygenic scores for common diseases identify individuals with risk equivalent to monogenic mutations. Nat. Genet. 50, 1219–1224 (2018).
Mega, J. L. et al. Genetic risk, coronary heart disease events, and the clinical benefit of statin therapy: an analysis of primary and secondary prevention trials. Lancet 385, 2264–2271 (2015).
Torkamani, A., Wineinger, N. E. & Topol, E. J. The personal and clinical utility of polygenic risk scores. Nat. Rev. Genet. 19, 581–590 (2018).
Kumar, A. et al. Whole-genome risk prediction of common diseases in human preimplantation embryos. Nat. Med. 28, 513–516 (2022).
Turley, P. et al. Problems with using polygenic scores to select embryos. N. Engl. J. Med. 385, 79–85 (2021).
Johnston, J. & Matthews, L. J. Polygenic embryo testing: understated ethics, unclear utility. Nat. Med. 28, 446–448 (2022).
Von Stumm, S. & Plomin, R. Using DNA to predict intelligence. Intelligence 86, 101530 (2021).
Shero, J. et al. The practical utility of genetic screening in school settings. NPJ Sci. Learn. 6, 1–10 (2021).
Biroli, P. et al. The economics and econometrics of gene-environment interplay. arXiv https://doi.org/10.48550/arXiv.2203.00729 (2022).
Pereira, R. D., van Kippersluis, H. & Rietveld, C. A. The interplay between maternal smoking and genes in offspring birth weight. J. Hum. Resour. https://doi.org/10.1101/2020.10.30.20222844 (2022).
Barcellos, S. H., Carvalho, L. S. & Turley, P. Education can reduce health differences related to genetic risk of obesity. Proc. Natl Acad. Sci. USA 115, E9765–E9772 (2018).
Slob, E. A. W. & Rietveld, C. A. Genetic predispositions moderate the effectiveness of tobacco excise taxes. PLoS ONE 16, e0259210 (2021).
Wand, H. et al. Improving reporting standards for polygenic scores in risk prediction studies. Nature 591, 211–219 (2021).
Lambert, S. A. et al. The polygenic score catalog as an open database for reproducibility and systematic evaluation. Nat. Genet. 53, 420–425 (2021).
Becker, J. et al. Resource profile and user guide of the Polygenic Index Repository. Nat. Hum. Behav. 5, 1744–1758 (2021).
Schultz, L. M. et al. Stability of polygenic scores across discovery genome-wide association studies. HGG Advances. 3, 100091 (2022)
Mills, M. C., Barban, N. & Tropf, F. C. An Introduction to Statistical Genetic Data Analysis (Cambridge MIT Press, 2020).
Aragam, K. G. et al. Limitations of contemporary guidelines for managing patients at high genetic risk of coronary artery disease. J. Am. Coll. Cardiol. 75, 2769–2780 (2020).
Inouye, M. et al. Genomic risk prediction of coronary artery disease in 480,000 adults: implications for primary prevention. J. Am. Coll. Cardiol. 72, 1883–1893 (2018).
Bycroft, C. et al. The UK Biobank resource with deep phenotyping and genomic data. Nature 562, 203–209 (2018).
Nikpay, M. et al. A comprehensive 1,000 genomes-based genome-wide association meta-analysis of coronary artery disease. Nat. Genet. 47, 1121–1130 (2015).
Lee, J. J. et al. Gene discovery and polygenic prediction from a genome-wide association study of educational attainment in 1.1 million individuals. Nat. Genet. 50, 1112–1121 (2018).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Chang, C. C. et al. Second-generation PLINK: rising to the challenge of larger and richer datasets. Gigascience 4, 1–16 (2015).
Vilhjálmsson, B. J. et al. Modeling linkage disequilibrium increases accuracy of polygenic risk scores. Am. J. Hum. Genet. 97, 576–592 (2015).
Ni, G. et al. A comparison of ten polygenic score methods for psychiatric disorders applied across multiple cohorts. Biol. Psychiatry. 90, 611–620 (2021).
Conley, D., Laidley, T. M., Boardman, J. D. & Domingue, B. W. Changing polygenic penetrance on phenotypes in the 20th century among adults in the US population. Sci. Rep. 6, 6–10 (2016).
Yengo, L., Yang, J. & Visscher, P. M. Expectation of the intercept from bivariate LD Score regression in the presence of population stratification. Preprint at bioRxiv https://doi.org/10.1101/310565 (2018).
Adhyaru, B. B. & Jacobson, T. A. Safety and efficacy of statin therapy. Nat. Rev. Cardiol. 15, 757–769 (2018).
Grundy, S. M. et al. 2018 AHA/ACC/AACVPR/AAPA/ABC/ACPM/ADA/AGS/APhA/ASPC/NLA/PCNA guideline on the management of blood cholesterol: executive summary. J. Am. Coll. Cardiol. 73, 3168–3209 (2019).
Goff, D. C. et al. 2013 ACC/AHA guideline on the assessment of cardiovascular risk: a report of the American College of Cardiology/American Heart Association Task Force on practice guidelines. J. Am. Coll. Cardiol. 63, 2935–2959 (2014).
Van Kippersluis, H. et al. Overcoming attenuation bias in regressions using polygenic indices: a comparison of approaches. Preprint at bioRxiv https://doi.org/10.1101/2021.04.09.439157 (2022).
Witte, J. S., Visscher, P. M. & Wray, N. R. The contribution of genetic variants to disease depends on the ruler. Nat. Rev. Genet. 5, 765–776 (2014).
De Vlaming, R. Genetic-nuture and assortative-mating-effects simulator. GitHub https://github.com/devlaming/gnames (2022).
Bulik-Sullivan, B. et al. LD Score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat. Genet. 47, 291–295 (2015).
De Vlaming, R. et al. Meta-GWAS Accuracy and Power (MetaGAP) calculator shows that hiding heritability is partially due to imperfect genetic correlations across studies. PLoS Genet 13, e1006495 (2017).
Choi, S. W., Mak, T. S.-H. & O’Reilly, P. F. Tutorial: a guide to performing polygenic risk score analyses. Nat. Protoc. 15, 2759–2772 (2020).
Ware, E. B. et al. Heterogeneity in polygenic scores for common human traits. Preprint at bioRxiv https://doi.org/10.1101/106062 (2017).
Pain, O. et al. Evaluation of polygenic prediction methodology within a reference-standardized framework. PLoS Genet. 17, 1–22 (2021).
Clifton, L., Collister, J. A., Liu, X., Littlejohn, T. J. & Hunter, D. J. Assessing agreement between different polygenic risk scores in the UK Biobank. Sci. Rep. 12, 12812 (2022).
Sun, J. et al. Translating polygenic risk scores for clinical use by estimating the confidence bounds of risk prediction. Nat. Commun. 12, 1–9 (2021).
Ding, Y. et al. Large uncertainty in individual polygenic risk score estimation impacts PRS-based risk stratification. Nat. Genet. 54, 30–39 (2022).
Glasziou, P. P., Irwig, L., Heritier, S., Simes, R. J. & Tonkin, A. Monitoring cholesterol levels: measurement error or true change? Ann. Intern. Med. 148, 656–661 (2008).
Jiang, L. et al. A resource-efficient tool for mixed model association analysis of large-scale data. Nat. Genet. 51, 1749–1755 (2019).
Acknowledgements
UK Biobank has obtained ethical approval from the National Research Ethics Committee (11/NW/0382). This research has been conducted using the UK Biobank Resource under application number 41382. The authors gratefully acknowledge funding from NORFACE through the Dynamic of Inequality across the Life Course (DIAL) programme (GEIGHEI 462-16-100). Research reported in this publication was also supported by the National Institute on Aging of the National Institutes of Health under Award R56AG058726. S.F.W.M. gratefully acknowledges funding from the European Union’s Horizon 2020 research and innovation programme under the Marie Skłodowska-Curie grant agreement (GENIO 101019584). C.A.R. and S.v.H. gratefully acknowledge funding from the European Research Council (GEPSI 946647; DONNI 851725). We are grateful for A. Okbay and employees and research participants of the 23andMe, Inc. cohort for sharing GWAS summary statistics for EA, and we thank P. Biroli, T. Galama, E. Slob and R. de Vlaming for insightful comments. This work made use of the Dutch national e-infrastructure with the support of the SURF Cooperative using grant EINF-1107. The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
All authors designed and oversaw the study. S.F.W.M. and D.M. conducted the GWAS in UKB and the meta-analyses with other GWAS summary statistics, constructed the PGIs and prepared the illustrative applications. R.D.P. performed the G × E analyses. H.v.K. and C.A.R. conducted the simulations. C.A.R. and S.v.H. assisted with the empirical analyses. All authors contributed to preparing and critically reviewing the manuscript and the supplementary file.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Human Behaviour thanks Sam Trejo and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary methods, results, references, Tables 1–5 and Fig. 1.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Muslimova, D., Dias Pereira, R., von Hinke, S. et al. Rank concordance of polygenic indices. Nat Hum Behav 7, 802–811 (2023). https://doi.org/10.1038/s41562-023-01544-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41562-023-01544-6