Abstract
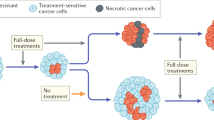
The emergence of drug-resistant cells, most of which have a mutated TP53 gene, prevents curative treatment in most advanced and common metastatic cancers of adults. Yet, a few, rarer malignancies, all of which are TP53 wild type, have high cure rates. In this Perspective, we discuss how common features of curable cancers offer insights into the evolutionary and developmental determinants of drug resistance. Acquired loss of TP53 protein function is the most common genetic change in cancer. This probably reflects positive selection in the context of strong ecosystem pressures including microenvironmental hypoxia. Loss of TP53’s functions results in multiple fitness benefits and enhanced evolvability of cancer cells. TP53-null cells survive apoptosis, and tolerate potent oncogenic signalling, DNA damage and genetic instability. In addition, critically, they provide an expanded pool of self-renewing, or stem, cells, the primary units of evolutionary selection in cancer, making subsequent adaptation to therapeutic challenge by drug resistance highly probable. The exceptional malignancies that are curable, including the common genetic subtype of childhood acute lymphoblastic leukaemia and testicular seminoma, differ from the common adult cancers in originating prenatally from embryonic or fetal cells that are developmentally primed for TP53-dependent apoptosis. Plus, they have other genetic and phenotypic features that enable dissemination without exposure to selective pressures for TP53 loss, retaining their intrinsic drug hypersensitivity.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Thun, M. J., DeLancey, J. O., Center, M. M., Jemal, A. & Ward, E. M. The global burden of cancer: priorities for prevention. Carcinogenesis 31, 100–110 (2010).
Nowell, P. C. The clonal evolution of tumor cell populations. Science 194, 23–28 (1976).
Greaves, M. & Maley, C. C. Clonal evolution in cancer. Nature 481, 306–313 (2012).
Gatenby, R. A. & Gillies, R. J. A microenvironmental model of carcinogenesis. Nat. Rev. Cancer 8, 56–61 (2008).
Anderson, A. R. A., Weaver, A. M., Cummings, P. T. & Quaranta, V. Tumor morphology and phenotypic evolution driven by selective pressure from the microenvironment. Cell 127, 905–915 (2006).
John, S. & Broggio, J. Stage 4 cancer survival data. Office for National Statistics, https://www.ons.gov.uk/peoplepopulationandcommunity/healthandsocialcare/conditionsanddiseases/bulletins/cancersurvivalinengland/latest (2019).
McGranahan, N. et al. Allele-specific HLA loss and immune escape in lung cancer evolution. Cell 171, 1259–1271.e11 (2017).
Huijben, S. et al. Aggressive chemotherapy and the selection of drug resistant pathogens. PLoS Pathog. 9, e1003578 (2013).
Galhardo, R. S., Hastings, P. J. & Rosenberg, S. M. Mutation as a stress response and the regulation of evolvability. Crit. Rev. Biochem. Mol. Biol. 42, 399–435 (2007).
Holohan, C., Van Schaeybroeck, S., Longley, D. B. & Johnston, P. G. Cancer drug resistance: an evolving paradigm. Nat. Rev. Cancer 13, 714–726 (2013).
Marine, J.-C., Dawson, S.-J. & Dawson, M. A. Non-genetic mechanisms of therapeutic resistance in cancer. Nat. Rev. Cancer 20, 743–756 (2020).
Kitano, H. Biological robustness. Nat. Rev. Genet. 5, 826–837 (2004).
Kreso, A. & Dick, J. E. Evolution of the cancer stem cell model. Cell Stem Cell 14, 275–291 (2014).
Greaves, M. Evolutionary determinants of cancer. Cancer Discov. 5, 806–820 (2015).
Shlush, L. I. et al. Tracing the origins of relapse in acute myeloid leukaemia to stem cells. Nature 547, 104–108 (2017).
Mani, S. A. et al. The epithelial-mesenchymal transition generates cells with properties of stem cells. Cell 133, 704–715 (2008).
Shibue, T. & Weinberg, R. A. EMT, CSCs, and drug resistance: the mechanistic link and clinical implications. Nat. Rev. Clin. Oncol. 14, 611–629 (2017).
Greaves, M. Cancer stem cells as ‘units of selection’. Evol. Appl 6, 102–108 (2013).
Fong, C. Y. et al. BET inhibitor resistance emerges from leukaemia stem cells. Nature 525, 538–542 (2015).
Chaidos, A. et al. Clinical drug resistance linked to interconvertible phenotypic and functional states of tumor-propagating cells in multiple myeloma. Blood 121, 318–328 (2013).
Chen, J. et al. A restricted cell population propagates glioblastoma growth after chemotherapy. Nature 488, 522–526 (2012).
Levine, A. J. & Oren, M. The first 30 years of p53: growing ever more complex. Nat. Rev. Cancer 9, 749–758 (2009).
Kandoth, C. et al. Mutational landscape and significance across 12 major cancer types. Nature 502, 333–339 (2013).
Papaemmanuil, E. et al. Genomic classification and prognosis in acute myeloid leukemia. N. Engl. J. Med. 374, 2209–2221 (2016).
Berlanga, P. et al. The European MAPPYACTS Trial: precision medicine program in pediatric and adolescent patients with recurrent malignancies. Cancer Discov. 12, 1266–1281 (2022).
Stracquadanio, G. et al. The importance of p53 pathway genetics in inherited and somatic cancer genomes. Nat. Rev. Cancer 16, 251–265 (2016).
Brown, C. J., Lain, S., Verma, C. S., Fersht, A. R. & Lane, D. P. Awakening guardian angels: drugging the p53 pathway. Nat. Rev. Cancer 9, 862–873 (2009).
Amadou, A., Achatz, M. I. W. & Hainaut, P. Revisiting tumor patterns and penetrance in germline TP53 mutation carriers: temporal phases of Li–Fraumeni syndrome. Curr. Opin. Oncol. 30, 23–29 (2018).
Abegglen, L. M. et al. Potential mechanisms for cancer resistance in elephants and comparative cellular response to DNA damage in humans. J. Am. Med. Assoc. 314, 1850–1860 (2015).
Lu, W.-J., Amatruda, J. F. & Abrams, J. M. p53 ancestry: gazing through an evolutionary lens. Nat. Rev. Cancer 9, 758–762 (2009).
Bieging, K. T., Mello, S. S. & Attardi, L. D. Unravelling mechanisms of p53-mediated tumour suppression. Nat. Rev. Cancer 14, 359–370 (2014).
Vousden, K. H. Activation of the p53 tumor suppressor protein. Biochim. Biophys. Acta 1602, 47–59 (2002).
Levine, A. J. p53, the cellular gatekeeper for growth and division. Cell 88, 323–331 (1997).
Muller, P. A. J. & Vousden, K. H. p53 mutations in cancer. Nat. Cell Biol. 15, 2–8 (2013).
Losos, J. Improbable Destinies: How Predictable is Evolution? (Penguin, 2018).
Wong, T. N. et al. Role of TP53 mutations in the origin and evolution of therapy-related acute myeloid leukaemia. Nature 518, 552–555 (2015).
Bristow, R. G. & Hill, R. P. Hypoxia and metabolism. Hypoxia, DNA repair and genetic instability. Nat. Rev. Cancer 8, 180–192 (2008).
Fang, J. S., Gillies, R. D. & Gatenby, R. A. Adaptation to hypoxia and acidosis in carcinogenesis and tumor progression. Semin Cancer Biol. 18, 330–337 (2008).
Gatenby, R. A. & Gillies, R. J. Why do cancers have high aerobic glycolysis? Nat. Rev. Cancer 4, 891–899 (2004).
Pan, Y., Oprysko, P. R., Asham, A. M., Koch, C. J. & Simon, M. C. p53 cannot be induced by hypoxia alone but responds to the hypoxic microenvironment. Oncogene 23, 4975–4983 (2004).
De Palma, M., Biziato, D. & Petrova, T. V. Microenvironmental regulation of tumour angiogenesis. Nat. Rev. Cancer 17, 457–474 (2017).
Lugano, R., Ramachandran, M. & Dimberg, A. Tumor angiogenesis: causes, consequences, challenges and opportunities. Cell. Mol. Life Sci. 77, 1745–1770 (2020).
Lopci, E. et al. PET radiopharmaceuticals for imaging of tumor hypoxia: a review of the evidence. Am. J. Nucl. Med Mol. Imaging 4, 365–384 (2014).
O’Connor, J. P. B., Robinson, S. P. & Waterton, J. C. Imaging tumour hypoxia with oxygen-enhanced MRI and BOLD MRI. Br. J. Radiol. 92, 20180642 (2019).
Bhandari, V. et al. Molecular landmarks of tumor hypoxia across cancer types. Nat. Genet. 51, 308–318 (2019).
Walsh, J. C. et al. The clinical importance of assessing tumor hypoxia: relationship of tumor hypoxia to prognosis and therapeutic opportunities. Antioxid. Redox Signal 21, 1516–1554 (2014).
Zschaeck, S. et al. Individual patient data meta-analysis of FMISO and FAZA hypoxia PET scans from head and neck cancer patients undergoing definitive radio-chemotherapy. Radiother. Oncol. 149, 189–196 (2020).
Höckel, M. et al. Association between tumor hypoxia and malignant progression in advanced cancer of the uterine cervix. Cancer Res. 56, 4509–4515 (1996).
Bhandari, V. et al. Divergent mutational processes distinguish hypoxic and normoxic tumours. Nat. Commun. 11, 737 (2020).
Graham, N. A. et al. Recurrent patterns of DNA copy number alterations in tumors reflect metabolic selection pressures. Mol. Syst. Biol. 13, 914 (2017).
Bartesaghi, S. et al. Inhibition of oxidative metabolism leads to p53 genetic inactivation and transformation in neural stem cells. Proc. Natl Acad. Sci. USA 112, 1059–1064 (2015).
Graeber, T. G. et al. Hypoxia-mediated selection of cells with diminished apoptotic potential in solid tumours. Nature 379, 88–91 (1996).
Zhang, Y., Yan, W. & Chen, X. Mutant p53 disrupts MCF-10A cell polarity in three-dimensional culture via epithelial-to-mesenchymal transitions. J. Biol. Chem. 286, 16218–16228 (2011).
Greaves, M. Does everyone develop covert cancer? Nat. Rev. Cancer 14, 209–210 (2014).
Meslin, F., Thiery, J., Richon, C., Jalil, A. & Chouaib, S. Granzyme B-induced cell death involves induction of p53 tumor suppressor gene and its activation in tumor target cells. J. Biol. Chem. 282, 32991–32999 (2007).
Shouval, R. et al. Impact of TP53 genomic alterations in large B-cell lymphoma treated with CD19-chimeric antigen receptor T-cell therapy. J. Clin. Oncol. 40, 369–381 (2022).
Sarkisian, C. J. et al. Dose-dependent oncogene-induced senescence in vivo and its evasion during mammary tumorigenesis. Nat. Cell Biol. 9, 493–505 (2007).
Junttila, M. R. & Evan, G. I. p53—a Jack of all trades but master of none. Nat. Rev. Cancer 9, 821–829 (2009).
Halazonetis, T. D., Gorgoulis, V. G. & Bartek, J. An oncogene-induced DNA damage model for cancer development. Science 319, 1352–1355 (2008).
Baslan, T. et al. Ordered and deterministic cancer genome evolution after p53 loss. Nature 608, 795–802 (2022).
de Bruin, E. C. et al. Spatial and temporal diversity in genomic instability processes defines lung cancer evolution. Science 346, 251–256 (2014).
Nik-Zainal, S. et al. The life history of 21 breast cancers. Cell 149, 994–1007 (2012).
McPherson, A. et al. Divergent modes of clonal spread and intraperitoneal mixing in high-grade serous ovarian cancer. Nat. Genet. 48, 758–767 (2016).
Liu, Y. et al. p53 regulates hematopoietic stem cell quiescence. Cell Stem Cell 4, 37–48 (2009).
Cicalese, A. et al. The tumor suppressor p53 regulates polarity of self-renewing divisions in mammary stem cells. Cell 138, 1083–1095 (2009).
Zhao, T. & Xu, Y. p53 and stem cells: new developments and new concerns. Trends Cell Biol. 20, 170–175 (2010).
Mizuno, H., Spike, B. T., Wahl, G. M. & Levine, A. J. Inactivation of p53 in breast cancers correlates with stem cell transcriptional signatures. Proc. Natl Acad. Sci. USA 107, 22745–22750 (2010).
Chen, S. et al. Mutant p53 drives clonal hematopoiesis through modulating epigenetic pathway. Nat. Commun. 10, 5649 (2019).
Mohyeldin, A., Garzón-Muvdi, T. & Quiñones-Hinojosa, A. Oxygen in stem cell biology: a critical component of the stem cell niche. Cell Stem Cell 7, 150–161 (2010).
Smith, B. A. et al. A human adult stem cell signature marks aggressive variants across epithelial cancers. Cell Rep. 24, 3353–3366.e5 (2018).
Pece, S. et al. Biological and molecular heterogeneity of breast cancers correlates with their cancer stem cell content. Cell 140, 62–73 (2010).
Merlos-Suárez, A. et al. The intestinal stem cell signature identifies colorectal cancer stem cells and predicts disease relapse. Cell Stem Cell 8, 511–524 (2011).
Eppert, K. et al. Stem cell gene expression programs influence clinical outcome in human leukemia. Nat. Med. 17, 1086–1093 (2011).
Shah, S. P. et al. The clonal and mutational evolution spectrum of primary triple-negative breast cancers. Nature 486, 395–399 (2012).
Samanta, D., Gilkes, D. M., Chaturvedi, P., Xiang, L. & Semenza, G. L. Hypoxia-inducible factors are required for chemotherapy resistance of breast cancer stem cells. Proc. Natl Acad. Sci. USA 111, E5429–E5438 (2014).
Payne, J. L. & Wagner, A. The causes of evolvability and their evolution. Nat. Rev. Genet. 20, 24–38 (2019).
Gutekunst, M. et al. p53 hypersensitivity is the predominant mechanism of the unique responsiveness of testicular germ cell tumor (TGCT) cells to cisplatin. PLoS ONE 6, e19198 (2011).
Visvader, J. E. Cells of origin in cancer. Nature 469, 314–322 (2011).
Greaves, M. A causal mechanism for childhood acute lymphoblastic leukaemia. Nat. Rev. Cancer 18, 471–484 (2018).
Marshall, G. M. et al. The prenatal origins of cancer. Nat. Rev. Cancer 14, 277–289 (2014).
Gudkov, A. V. & Komarova, E. A. The role of p53 in determining sensitivity to radiotherapy. Nat. Rev. Cancer 3, 117–129 (2003).
Sarosiek, K. A. et al. Developmental regulation of mitochondrial apoptosis by c-Myc governs age- and tissue-specific sensitivity to cancer therapeutics. Cancer Cell 31, 142–156 (2017).
Vousden, K. H. & Lane, D. P. p53 in health and disease. Nat. Rev. Mol. Cell Biol. 8, 275–283 (2007).
Zhang, G. et al. p53 pathway is involved in cell competition during mouse embryogenesis. Proc. Natl Acad. Sci. USA 114, 498–503 (2017).
Oosterhuis, J. W. & Looijenga, L. H. J. Human germ cell tumours from a developmental perspective. Nat. Rev. Cancer 19, 522–537 (2019).
Runyan, C. et al. Steel factor controls midline cell death of primordial germ cells and is essential for their normal proliferation and migration. Development 133, 4861–4869 (2006).
Braun, R. E. Every sperm is sacred—or is it? Nat. Genet. 18, 202–204 (1998).
Masters, J. R. W. & Köberle, B. Curing metastatic cancer: lessons from testicular germ-cell tumours. Nat. Rev. Cancer 3, 517–525 (2003).
Awuah, S. G., Riddell, I. A. & Lippard, S. J. Repair shielding of platinum–DNA lesions in testicular germ cell tumors by high-mobility group box protein 4 imparts cisplatin hypersensitivity. Proc. Natl Acad. Sci. USA 114, 950–955 (2017).
Usanova, S. et al. Cisplatin sensitivity of testis tumour cells is due to deficiency in interstrand-crosslink repair and low ERCC1-XPF expression. Mol. Cancer 9, 248 (2010).
Greaves, M. F. & Wiemels, J. Origins of chromosome translocations in childhood leukaemia. Nat. Rev. Cancer 3, 639–649 (2003).
Böiers, C. et al. A human IPS model implicates embryonic B-myeloid fate restriction as developmental susceptibility to B acute lymphoblastic leukemia-associated ETV6-RUNX1. Dev. Cell 44, 362–377.e7 (2018).
Kersey, J. H. Fifty years of studies of the biology and therapy of childhood leukemia. Blood 90, 4243–4251 (1997).
Eswaran, J. et al. The pre-B-cell receptor checkpoint in acute lymphoblastic leukaemia. Leukemia 29, 1623–1631 (2015).
Greaves, M. & Hughes, W. Cancer cell transmission via the placenta. Evol. Med. Public Health 2018, 106–115 (2018).
Seckl, M. J., Sebire, N. J. & Berkowitz, R. S. Gestational trophoblastic disease. Lancet 376, 717–729 (2010).
Shih, I.-M. & Kuo, K.-T. Power of the eternal youth: nanog expression in the gestational choriocarcinoma. Am. J. Pathol. 173, 911–914 (2008).
Stovall, T. G., Ling, F. W., Gray, L. A., Carson, S. A. & Buster, J. E. Methotrexate treatment of unruptured ectopic pregnancy: a report of 100 cases. Obstet. Gynecol. 77, 749–753 (1991).
Coorens, T. H. H. et al. Embryonal precursors of Wilms tumor. Science 366, 1247–1251 (2019).
Spreafico, F. et al. Wilms tumour. Nat. Rev. Dis. Prim. 7, 75 (2021).
Savage, P. Chemotherapy curability in leukemia, lymphoma, germ cell tumors and gestational malignancies: a reflection of the unique physiology of their cells of origin. Front. Genet 11, 426 (2020).
Perez-Atayde, A. R. et al. Spectrum of tumor angiogenesis in the bone marrow of children with acute lymphoblastic leukemia. Am. J. Pathol. 150, 815–821 (1997).
Donato, D. P., Johnson, M. T., Yang, X. J. & Zynger, D. L. Expression of carbonic anhydrase IX in genitourinary and adrenal tumours. Histopathology 59, 1229–1239 (2011).
Huang, D. Y. & Sidhu, P. S. Focal testicular lesions: colour Doppler ultrasound, contrast-enhanced ultrasound and tissue elastography as adjuvants to the diagnosis. Br. J. Radiol. 85, S41–S53 (2012).
Kalavska, K. et al. Prognostic value of intratumoral carbonic anhydrase IX expression in testicular germ cell tumors. Oncol. Lett. 13, 2177–2185 (2017).
Alexandrov, L. B. et al. Signatures of mutational processes in human cancer. Nature 500, 415–421 (2013).
The ICGC/TCGA Pan-Cancer Analysis of Whole Genomes Consortium Pan-cancer analysis of whole genomes. Nature 578, 82–93 (2020).
Brady, S. W. et al. The genomic landscape of pediatric acute lymphoblastic leukemia. Nat. Genet. 54, 1376–1389 (2022).
Moorman, A. V. New and emerging prognostic and predictive genetic biomarkers in B-cell precursor acute lymphoblastic leukemia. Haematologica 101, 407–416 (2016).
Ford, A. M., Colman, S. & Greaves, M. Covert pre-leukaemic clones in healthy co-twins of patients with childhood acute lymphoblastic leukaemia. Leukemia https://doi.org/10.1038/s41375-022-01756-1 (2022).
Mori, H. et al. Chromosome translocations and covert leukemic clones are generated during normal fetal development. Proc. Natl Acad. Sci. USA 99, 8242–8247 (2002).
Papaemmanuil, E. et al. RAG-mediated recombination is the predominant driver of oncogenic rearrangement in ETV6-RUNX1 acute lymphoblastic leukemia. Nat. Genet. 46, 116–125 (2014).
Anderson, K. et al. Genetic variegation of clonal architecture and propagating cells in leukaemia. Nature 469, 356–361 (2011).
Paulsson, K. et al. The genomic landscape of high hyperdiploid childhood acute lymphoblastic leukemia. Nat. Genet. 47, 672–676 (2015).
Meyer, L. H. et al. Early relapse in ALL is identified by time to leukemia in NOD/SCID mice and is characterized by a gene signature involving survival pathways. Cancer Cell 19, 206–217 (2011).
Rehe, K. et al. Acute B lymphoblastic leukaemia-propagating cells are present at high frequency in diverse lymphoblast populations. EMBO Mol. Med. 5, 38–51 (2013).
Savage, P. et al. A case of intraplacental gestational choriocarcinoma; characterised by the methylation pattern of the early placenta and an absence of driver mutations. BMC Cancer 19, 744 (2019).
Litchfield, K. et al. Whole-exome sequencing reveals the mutational spectrum of testicular germ cell tumours. Nat. Commun. 6, 5973 (2015).
Taylor-Weiner, A. et al. Genomic evolution and chemoresistance in germ-cell tumours. Nature 540, 114–118 (2016).
Al-Lazikani, B., Banerji, U. & Workman, P. Combinatorial drug therapy for cancer in the post-genomic era. Nat. Biotechnol. 30, 679–692 (2012).
Zhang, J., Cunningham, J., Brown, J. & Gatenby, R. Evolution-based mathematical models significantly prolong response to abiraterone in metastatic castrate-resistant prostate cancer and identify strategies to further improve outcomes. eLife 11, e76284 (2022).
Acar, A. et al. Exploiting evolutionary steering to induce collateral drug sensitivity in cancer. Nat. Commun. 11, 1923 (2020).
Acknowledgements
This Perspective is dedicated to the memory of Dr Donald Pinkel (1926–2022) pioneer of curative chemotherapy for children with leukaemia (Supplementary Information and Supplementary Fig. 2). The authors are grateful for funding received from the Wood family—in memory of Artemis. We gratefully acknowledge the FAZA images courtesy of D Vriens, Section of Nuclear Medicine, Department of Radiology, Leiden University Medical Centre (LUMC), Leiden, the Netherlands and Holland Proton Therapy Centre (HollandPTC), Delft, the Netherlands and the bone marrow images with microvessels courtesy of S. E. Sallan, Dana-Farber Cancer Institute and Boston Children’s Hospital, Boston, USA. We also thank A. Moorman for kindly providing Supplementary Fig. 1 (previously unpublished data). We are grateful to J. Vormoor, A. Roy, T. Graham and A. Borkhardt for helpful comments.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Ecology & Evolution thanks Subhayan Chattopadhyay, Wei Gu and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1 and 2, Text/Information on childhood ALL curability and on TP53, self-renewal and the cure of APML, Table 1 and References.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Mansur, M.B., deSouza, N.M., Natrajan, R. et al. Evolutionary determinants of curability in cancer. Nat Ecol Evol 7, 1761–1770 (2023). https://doi.org/10.1038/s41559-023-02159-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-023-02159-w
This article is cited by
-
Convergent TP53 loss and evolvability in cancer
BMC Ecology and Evolution (2023)