Abstract
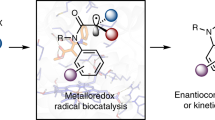
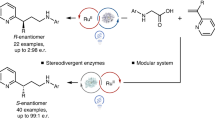
The formation of C–N bonds—of great importance to the pharmaceutical industry—can be facilitated enzymatically using nucleophilic and nitrene transfer mechanisms. However, neither natural nor engineered enzymes are known to generate and control nitrogen-centred radicals, which serve as valuable species for C–N bond formation. Here we use flavin-dependent ‘ene’-reductases with an exogenous photoredox catalyst to selectively generate amidyl radicals within the protein active site. These enzymes are engineered through directed evolution to catalyse 5-exo, 6-endo, 7-endo, 8-endo, and intermolecular hydroamination reactions with high levels of enantioselectivity. Mechanistic studies suggest that radical initiation occurs via an enzyme-gated mechanism, where the protein thermodynamically activates the substrate for reduction by the photocatalyst. Molecular dynamics studies indicate that the enzymes bind substrates using non-canonical binding interactions, which may serve as a handle to further manipulate reactivity. This approach demonstrates the versatility of these enzymes for controlling the reactivity of high-energy radical intermediates and highlights the opportunity for synergistic catalyst strategies to unlock previously inaccessible enzymatic functions.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
The data that support the findings in this study are available in this article and the Supplementary Information. Source data are provided with this paper.
References
Breuer, M. et al. Industrial methods for the production of optically active intermediates. Angew. Chem. Int. Ed. 43, 788–824 (2004).
Roughley, S. D. & Jordan, A. M. The medicinal chemist’s toolbox: an analysis of reactions used in the pursuit of drug candidates. J. Med. Chem. 54, 3451–3479 (2011).
Kerru, N., Gummidi, L., Maddila, S., Gangu, K. K. & Jonnalagadda, S. B. A review on recent advances in nitrogen-containing molecules and their biological applications. Molecules 25, 1909 (2020).
Slabu, I. S., Galman, J. L., Lloyd, R. C. & Turner, N. J. Discovery, engineering, and synthetic application of transaminase biocatalysts. ACS Catal. 7, 8263–8284 (2017).
Aleku, G. A. et al. A reductive aminase from Aspergillus oryzae. Nat. Chem. 9, 961–969 (2017).
Marshall, J. R. et al. Screening and characterization of a diverse panel of metagenomic imine reductases for biocatalytic reductive amination. Nat. Chem. 13, 140–148 (2021).
Parmeggiani, F., Weise, N. J., Ahmed, S. T. & Turner, N. J. Synthetic and therapeutic applications of ammonia-lyases and aminomutases. Chem. Rev. 118, 73–118 (2018).
Raj, H. et al. Engineering methylaspartate ammonia lyase for the asymmetric synthesis of unnatural amino acids. Nat. Chem. 4, 478–484 (2012).
Fu, H. et al. Chemoenzymatic asymmetric synthesis of the metallo-β-lactamase inhibitor aspergillomarasmine A and related aminocarboxylic acids. Nat. Chem. 1, 186–191 (2018).
Peatzold, J. & Bäckvall, J. E. Chemoenzymatic dynamic kinetic resolution of primary amines. J. Am. Chem. Soc. 127, 17620–17621 (2005).
Ghislieri, D. & Turner, N. J. Biocatalytic approaches to the synthesis of enantiomerically pure chiral amines. Top. Catal. 57, 284–300 (2014).
Athavale, S. V. et al. Biocatalytic, intermolecular C–H bond functionalization for the synthesis of enantioenriched amides. Angew. Chem. Int. Ed. 60, 24864–24869 (2021).
Farwell, C. C., Zhang, R. K., McIntosh, J. A., Hyster, T. K. & Arnold, F. H. Enantioselective enzyme-catalyzed aziridination enabled by active-site evolution of a cytochrome P450. ACS Cent. Sci. 1, 89–93 (2015).
Yang, Y. & Arnold, F. H. Navigating the unnatural reaction space: directed evolution of heme proteins for selective carbene and nitrene transfer. Acc. Chem. Res. 54, 1209–1224 (2021).
Zard, S. Z. Recent progress in the generation and use of nitrogen-centered radicals. Chem. Soc. Rev. 37, 1603–1618 (2008).
Kwon, K., Simons, R. T., Nandakumar, M. & Roizen, J. L. Strategies to generate nitrogen-centered radicals that may rely on photoredox catalysis: development in reaction methodology and applications in organic synthesis. Chem. Rev. 122, 2353–2428 (2022).
Studer, A. & Curran, D. P. Catalysis of radical reactions: a radical chemistry perspective. Angew. Chem. Int. Ed. 55, 58–102 (2016).
Horner, J. H., Musa, O. M., Bouvier, A. & Newcomb, M. Absolute kinetics of amidyl radical reactions. J. Am. Chem. Soc. 120, 7738–7748 (1998).
Rittle, J. & Green, M. T. Cytochrome P450 compound I: capture, characterization, and C–H bond activation kinetics. Science 330, 933–937 (2010).
Yokoyama, K. & Lilla, E. A. C–C bond forming radical SAM enzymes involved in the construction of carbon skeletons of cofactors and natural products. Nat. Prod. Rep. 35, 660–694 (2018).
Ye, Y., Fu, H. & Hyster, T. K. Activation modes in biocatalytic radical cyclization reactions. J. Ind. Microbiol. Biotechnol. 48, kuab021 (2021).
Toogood, H. S. & Scrutton, N. S. Discovery, characterization, engineering, and applications of ene-reductases for industrial biocatalysis. ACS Catal. 8, 3532–3549 (2018).
Biegasiewicz, K. F. et al. Photoexcitation of flavoenzymes enables a stereoselective radical cyclization. Science 364, 1166–1169 (2019).
Black, M. J. et al. Asymmetric redox-neutral radical cyclization catalysed by flavin-dependent ‘ene’-reductases. Nat. Chem. 12, 71–75 (2020).
Clayman, P. D. & Hyster, T. K. Photoenzymatic generation of unstabilized alkyl radicals: an asymmetric reductive cyclization. J. Am. Chem. Soc. 142, 15673–15677 (2020).
Fu, H. et al. Ground-state electron transfer as an initiation mechanism for biocatalytic C–C bond forming reactions. J. Am. Chem. Soc. 143, 9622–9629 (2021).
Page, C. G. et al. Quaternary charge-transfer complex enables photoenzymatic intermolecular hydroalkylation of olefins. J. Am. Chem. Soc. 143, 97–102 (2020).
Huang, X. et al. Photoenzymatic enantioselective intermolecular radical hydroalkylation. Nature 584, 69–74 (2020).
Grosheva, D. & Hyster, T. K. in Flavin-Based Catalysis: Principles and Applications (eds Cibulka, R. & Fraaije, M.) 291–313 (Wiley, 2021).
Davies, J., Morcillo, S. P., Douglas, J. J. & Leonori, D. Hydroxylamine derivatives as nitrogen-radical precursors in visible-light photochemistry. Chem. Eur. J. 24, 12154–12163 (2018).
Keum, H., Jung, H., Jeong, J., Kim, D. & Chang, S. Visible-light induced C(sp2)–H amidation with an aryl–alkyl σ-bond relocation via redox-neutral radical–polar crossover. Angew. Chem. Int. Ed. 60, 25235–25240 (2021).
Warren, J. J., Ener, M. E., Vlček, A., Winkler, J. R. & Gray, H. B. Electron hopping through proteins. Coord. Chem. Rev. 256, 2478–2487 (2012).
Sandoval, B. A. et al. Photoenzymatic reductions enabled by direct excitation of flavin-dependent ‘ene’-reductases. J. Am. Chem. Soc. 143, 1735–1739 (2021).
Biegasiewicz, K. F., Cooper, S. J., Emmanuel, M. A., Miller, D. C. & Hyster, T. K. Catalytic promiscuity enabled by photoredox catalysis in nicotinamide-dependent oxidoreductases. Nat. Chem. 10, 770–775 (2018).
Sandoval, B. A., Kurtoic, S. I., Chung, M. M., Biegasiewicz, K. F. & Hyster, T. K. Photoenzymatic catalysis enables radical-mediated ketone reduction in ene-reductases. Angew. Chem. Int. Ed. 58, 8714–8718 (2019).
Nakano, Y. et al. Photoenzymatic hydrogenation of heteroaromatic olefins using ‘ene’-reductases with photoredox catalysts. Angew. Chem. Int. Ed. 59, 10484–10488 (2020).
Prier, C. K., Rankic, D. A. & Macmillan, D. W. C. Visible light photoredox catalysis with transition metal complexes: applications in organic synthesis. Chem. Rev. 113, 5322–5363 (2013).
Abramovitz, A. S. & Massey, V. Interaction of phenols with old yellow enzyme. Physical evidence for charge-transfer complexes. J. Biol. Chem. 251, 5327–5336 (1976).
Marshall, S. J., Krause, D., Blencowe, D. K. & White, G. F. Characterization of glycerol trinitrate reductase (NerA) and the catalytic role of active-site residues. J. Bacteriol. 186, 1802–1810 (2004).
Steward, R. C. & Massey, V. Potentiometric studies of native and flavin-substituted old yellow enzyme. J. Biol. Chem. 260, 13639–13647 (1985).
Reetz, M. T. & Carballeira, J. D. Iterative saturation mutagenesis (ISM) for rapid directed evolution of functional enzymes. Nat. Protoc. 2, 891–903 (2007).
Nicholls, B. et al. Engineering a non-natural photoenzyme for improved photon efficiency. Angew. Chem. Int. Ed. https://doi.org/10.1002/anie.202113842 (2022).
Bougioukou, D. J., Kille, S., Taglieber, A. & Reetz, M. T. Directed evolution of an enantioselective enoate-reductase: testing the utility of iterative saturation mutagenesis. Adv. Synth. Catal. 351, 3287–3305 (2009).
Sim, B. A., Griller, D. & Wayner, D. D. M. Reduction potentials for substituted benzyl radicals: pKa values for the corresponding toluenes. J. Am. Chem. Soc. 111, 754–755 (1989).
Sorigué, D. et al. Mechanism and dynamics of fatty acid photodecarboxylase. Science https://doi.org/10.1126/science.abd5687 (2021).
Weinberg, D. R. et al. Proton-coupled electron transfer. Chem. Rev. 112, 4016–4093 (2012).
Kato, Y., Nagao, R. & Noguchi, T. Redox potential of the terminal quinone electron acceptor QB in photosystem II reveals the mechanism of electron transfer regulation. Proc. Natl Acad. Sci. USA 113, 620–625 (2016).
Acknowledgements
The research reported here was supported by the National Institutes of Health National Institute of General Medical Sciences (R01 GM127703). Transient absorption spectroscopy studies were supported by the Division of Chemical Sciences, Geosciences, and Biosciences, Office of Basic Energy Sciences of the US Department of Energy (DOE) through grant DE-SC0019370. We thank the Princeton Catalysis Initiative and Merck Sharp & Dohme LLC, a subsidiary of Merck & Co., Inc., Rahway, NJ, USA, for financial support. This work made use of the Cornell University NMR Facility, which is supported, in part, by the NSF though MRI Award CHE-1531632. D.G.O. acknowledges support from the Postgraduate Scholarships Doctoral Program of the Natural Sciences and Engineering Research Council of Canada. We thank K. Meihaus, P. Clayman, K. Biegasiewicz and Y. Liu for assistance in preparing this manuscript, and T. Qiao for assistance with density functional theory calculations.
Author information
Authors and Affiliations
Contributions
T.K.H. conceived and directed the project. Y.Y. and J.C. performed and analysed the experiments. D.G.O. performed the transient absorption spectroscopy experiments. D.G.O. and G.D.S. analysed and interpreted the spectroscopy results. D.V. performed the MD simulations. D.V. and C.K.P. analysed and interpreted the MD simulations. All authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Chemistry thanks Fabio Parmeggiani and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Parts I–XVI (Figs. 1–32, Tables 1–24, NMR spectra and HPLC traces).
Source data
Source Data Fig. 2
Statistical source data for Fig. 2c.
Source Data Fig. 3
Statistical source data for Fig. 3a,c.
Source Data Fig. 3
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ye, Y., Cao, J., Oblinsky, D.G. et al. Using enzymes to tame nitrogen-centred radicals for enantioselective hydroamination. Nat. Chem. 15, 206–212 (2023). https://doi.org/10.1038/s41557-022-01083-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41557-022-01083-z
This article is cited by
-
An evolved artificial radical cyclase enables the construction of bicyclic terpenoid scaffolds via an H-atom transfer pathway
Nature Chemistry (2024)
-
Stereoselective amino acid synthesis by photobiocatalytic oxidative coupling
Nature (2024)
-
Enantioselective alkene hydroalkylation overcoming heteroatom constraints via cobalt catalysis
Nature Synthesis (2024)
-
Remote stereocontrol with azaarenes via enzymatic hydrogen atom transfer
Nature Chemistry (2024)
-
Enantioselective decarboxylative alkylation using synergistic photoenzymatic catalysis
Nature Catalysis (2023)