Abstract
The transmission of viruses from animal reservoirs to humans poses major threats to public health. Preparedness for future zoonotic outbreaks requires a fundamental understanding of how viruses of animal origin have adapted to binding to a cell surface component and/or receptor of the new host. Here we report on the specificities of human and animal viruses that engage with O-acetylated sialic acid, which include betacoronaviruses, toroviruses and influenza C and D viruses. Key to these studies was the development of a chemoenzymatic methodology that can provide almost any sialate-acetylation pattern. A collection of O-acetylated sialoglycans was printed as a microarray for the determination of receptor specificity. These studies showed host-specific patterns of receptor recognition and revealed that three distinct human respiratory viruses uniquely bind 9-O-acetylated α2,8-linked disialoside. Immunofluorescence and cell entry studies support that such a glycotope as part of a ganglioside is a functional receptor for human coronaviruses.

Similar content being viewed by others
Main
Eukaryotic cells are covered by a dense layer of glycans. These carbohydrates are structurally very diverse and expressed in tissue- and cell-specific manners. During evolution, glycan biosynthesis diverged and as a result structures of glycans differ considerably even between closely related species. These differences are, in part, driven by pathogenic pressure to hamper cross-species transmission1,2.
Numerous viruses initiate infection by binding to cell surface glycans of the host. The selectivities of viral receptor binding proteins for specific glycan structures critically determine host range, targeted tissues and cells, and pathogenesis3. Prototypic are influenza A viruses, which have an obligatory dependence for host glycans modified by a terminal sialic acid to infect cells4. The haemagglutinin (HA) of avian influenza A viruses preferentially binds sialic acid α2,3-linked to galactose (Gal), whereas human viruses recognize the α2,6-linked isomer. The influenza A viruses that circulates in humans are of avian origin and acquired an ability to use α2,6-linked sialosides through mutations in the receptor binding pocket of HA. This adaptation occurred because human upper airway tissue expresses high levels of α2,6-sialosides, whereas duck intestinal mucosa is rich in α2,3-sialosides.
For many other vertebrate viruses that employ sialoglycans as receptors, it has been difficult to determine receptor binding specificities5,6. This information is critical to establish to what extent differences in the sialoglycan repertoire between vertebrate species hamper cross-species transmission and how human viruses of zoonotic origin have overcome these barriers to establish infections in humans. Insight into receptor binding specificities is also critical for the development of surveillance, prevention and intervention strategies7,8.
Here we describe receptor binding specificities of a group of viruses that engage with O-acetylated variants of sialic acid, including several betacoronaviruses and toroviruses (subfamily Orthocoronavirinae and family Tobaniviridae, respectively), and influenza C and D viruses (family Orthomyxoviridae), and demonstrate host-specific patterns of receptor recognition that relate to both the O-acetylation pattern and sialic acid glycosidic linkage type. Our data revealed that viruses adapted to humans underwent convergent evolution to become highly selective for O-acetylated forms of α2,8-linked disialylated structures commonly found on glycolipids. Key to these discoveries was the implementation of a chemoenzymatic methodology that provided a previously inaccessible collection of O-acetylated sialoglycans.
N-acetylneuraminic acid (Neu5Ac) and N-glycolylneuraminic acid (Neu5Gc) are the most common forms of sialic acid in vertebrates9. They occur in a variety of different glycosidic forms typically α2,3-linked to Gal, α2,6-linked to Gal and N-acetyl-galactosamine (GalNAc) or via an α2,8-linkage to another sialic acid (Fig. 1a). Further structural diversity arises from the substitution of the C-4, C-7, C-8 and/or C-9 hydroxyls by acetyl esters (Fig. 1b). Despite their presence in many different tissues10,11, O-acetylated sialic acids are Cinderella molecules12. There is limited knowledge of their biosynthesis, which precludes analysis through genetic manipulation. This class of glycans exhibits inherent chemical labilities and are prone to O-acetyl ester migration and hydrolysis, which complicates isolation and characterization. A lack of methods to prepare panels of O-acetylated sialosides is another major stumbling block, and reported synthetic approaches are largely limited to monosaccharide derivatives and C-9-modified sialosides (see Supplementary Fig. 1a–d for an overview of previously reported synthetic approaches)13,14,15,16,17,18,19,20,21,22,23. The synthetic inaccessibility of a complete series of O-acetylated sialosides stems from the stringent requirements to select protecting groups for a sialyl donor and acceptor to achieve high-yielding and stereoselective sialylations24. The selected protecting groups of the donor and acceptor also need to be compatible with the installation of different patterns of acetyl esters that have a propensity to migrate or hydrolyse even under mildly basic or acidic conditions, which results in a yet unmet synthetic challenge (Supplementary Fig. 1e).
a, Common backbone structures and glycosidic linkages. b, Naturally occurring partially O-acetylated variants of Neu5Ac9.
Results and discussion
A general approach to diverse O-acetylated sialyl oligosaccharides
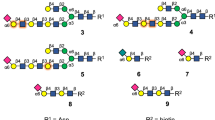
To develop a general method for the preparation of O-acetylated sialosides, we drew inspiration from the sialate-O-acetylesterase activities of viral haemagglutinin–esterases (HEs). These enzymes, which are expressed by various nido- and orthomyxoviruses, cleave acetyl esters of sialic acids with exquisite regioselectivity to facilitate a dynamic virion attachment to host cells. The bovine coronavirus (BCoV) HE is a 9-O-acetylesterase with a preference for 7,9-di-O-acetylsialosides10, whereas mouse hepatitis virus strain S (MHV-S) produces HEs specific for sialate-4-O-acetyl esters25. We envisaged that diversification of α2,3-, α2,6- and α2,8-linked sialosides 1a, 2a and 3a, respectively, which carried acetyl esters at C-4, C-7 and C-9 (Fig. 2a), by treatment with these enzymes combined with controlled acetyl ester migration from C-7 to C-9 (ref. 26) (Fig. 2b) could provide almost any sialate-acetylation pattern. The attraction of this approach is that from a common oligosaccharide precursor as many as seven sialate-O-acetylation patterns can be installed, thereby alleviating the need to select for each target the proper protecting groups, and establish the appropriate glycosylation conditions, which is almost an impossible task to achieve.
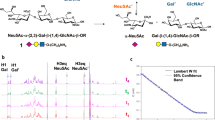
a, Chemically synthesized 4,7,9-tri-O-acetylated common precursors for HE-mediated regioselective de-O-acetylation. The biotinylated anomeric linker is denoted as R. b, Methods for chemoenzymatic remodelling of the common precursors. BCoV and MHV-S HEs regiospecifically cleave 9- and 4-O-acetyl groups, respectively. c, The glycosyl donor and acceptors for chemical assemblies of 1–3a. See Supplementary Fig. 2 for the preparation of glycosyl acceptors. Nap, 2-naphthyl.
The three common precursors 1a, 2a and 3a were prepared from universal donor 4 and glycosyl acceptors 5, 6 and 7, respectively (Fig. 2c). The 7,8-isopropylidene of 4 combined with the presence of a 4,5-carbamate ensured that glycosylations proceeded in a high yield with exquisite α-anomeric selectivity27. The latter was facilitated by using glycosyl acceptors 5, 6 and 7, which have highly reactive 3,4-, 4,6- and 7,8-diols that, in a regioselective manner, can be sialylated to provide α2,3-, α2,6- and α2,8-linked products, respectively. Oxidative acetal ring closure of the 2-naphthylmethyl ether of the glycosylation products protected the remaining alcohol, which allowed selective manipulation of the hydroxyls of the sialic acid moiety to install the required acetyl esters only at C-4, C-7 and C-9 (see Supplementary Fig. 1f for an analysis of the general strategy).
Thus, a trimethylsilyl triflate (TMSOTf)-mediated glycosylation of 4 with 5 gave a trisaccharide that was immediately subjected to wet trifluoroacetic acid (TFA) to remove the isopropylidene acetal to provide 8 in a yield of 79% over two steps as only the α-anomer (Fig. 3a). 2,3-dichloro-5,6-dicyano-p-benzoquinone (DDQ)-mediated oxidative acetal ring closure of 8 in anhydrous dichloromethane (DCM) gave 9, which was converted into 10 by a one-pot procedure that involved hydrolysis of the methyl ester and 4,5-carbamate of the sialic acid moiety followed by selective acetylation of the resulting amine by reaction with acetyl chloride. Next, the azido moieties of 10 were reduced by the Staudinger reaction to give diamine 11, which was converted into 12 by selective amine manipulations. The free amine and the C-4, C-7 and C-8 hydroxyls of 12 were acetylated using acetic anhydride in pyridine to give 13. Hydrogenation of the latter compound resulted in the removal of the benzyl ethers and reduction of the azido moiety into an amine. The target trisaccharide 1a was obtained in a high overall yield by controlled migration of the acetyl ester at C-8 to C-9 using a CsF–acetic acid buffer in dimethylsulfoxide (DMSO) followed by biotinylation of the amine of the anomeric spacer.
a, Chemical synthesis of 1a. See Supplementary Fig. 4 for details on the regioselective C-8-to-C-9 acetyl ester migration performed on deprotected oligosaccharides in DMSO. b, Modifications of 1a and 2a using HE-catalysed O-acetyl removal and base-mediated O-acetyl migration. Enzymatic reactions went to full conversion. Representative yields in parentheses were determined post-HPLC purification. 7-O-acetyl remained stable (for sialoforms 1b, 2b, 1d, 2d, 1f and 2f) during the course of reaction and purification. See Supplementary Figs. 5–8 and Supplementary Table 2 for the structural confirmation. c, Chemical synthesis and enzymatic modification of 3a using HEs. MHV-S HE removed 4-O-acetyl groups of both the non-reducing and reducing end Neu5Ac residues. Lc, 6-aminocaproic acid linker; NBI, 5-norbornene-2,3-dicarboxylic acid imido; Su, succinimido.
Exposure of 1a to MHV-S or BCoV HE in ammonium formate buffer provided the 4,7- and 7,9-di-O-acetylated derivatives 1b and 1d, respectively (Fig. 3b). Although bovines do not express C-4 acetylated sialosides, BCoV HE readily converted 1a into 1b. Exposure of the product of MHV-S (1d) with BCoV HE gave a 7-O-acetylated derivative 1f. Treatment of 1b and 1f with aqueous ammonium bicarbonate (100 mM) resulted in an efficient migration of the C-7 acetyl ester to C-9, which provided 1c and 1g, respectively. Finally, subjecting 1c to BCoV gave the 4-O-acetyl derivative 1e. Compound 2a (see Supplementary Fig. 3 for the chemical synthesis) could be diversified into the mono- and di-O-acetylated derivatives 2b–2e by using MHV-S and BCoV HE combined with controlled O-acetyl ester migrations.
The preparation of 2,8-linked disialosides is particularly challenging due to the low reactivity of the C-8 hydroxyl of a sialic acid acceptor28. Coupling of 4 with 729 using TMSOTf as the activator gave the corresponding 2,8-linked disialoside in high yields, which could easily be converted into 3a (Fig. 3c) and then diversified by MHV-S and BCoV HE treatment and a controlled acetyl ester migration to provide 3d, 3f and 3g. We found that the 4-O-acetyl ester of the terminal and the internal sialoside of 3a can readily be cleaved by MHV-S HE and thereby provide access to biologically highly relevant glycans9.
The results demonstrate that, regardless of glycosidic linkage type and O-acetylation pattern, MHV-S and BCoV HEs efficiently remove C-4 and C-9 acetyl esters, respectively. Furthermore, we found that the viral esterases are highly enzymatically active, can be stored for prolonged period of times and tolerate a wide range of pH values, which makes them attractive tools for chemoenzymatic synthesis. All the compounds were fully characterized by two-dimensional NMR experiments, which confirmed the homogeneity and a proper pattern of O-acetylation. The latter was evident from a large downfield shift of α-proton peaks (Supplementary Table 2), which in heteronuclear multiple-bond correlation experiments showed correlation with carbonyl carbons (Supplementary Fig. 6–8).
Viral receptor analysis using glycan microarray technology
The comprehensive library of O-acetylated sialosides allowed a systematic examination of the receptor specificities of an extensive range of viruses known to use this class of glycans for host-cell attachment and entry (see Supplementary Table 1 for the specifications). Among these were three established human viruses, namely coronaviruses (CoVs) OC43 and HKU1 (genus Betacoronavirus, subgenus Embecovirus) and influenza C virus (ICV). Infections with these viruses can cause mild upper respiratory illnesses in healthy adults, but can lead to serious complications in newborns, the elderly and immunocompromised individuals30. In addition, receptor specificities of a wide range of related animal viruses were investigated, which included BCoV, the presumed ancestor of OC43, and influenza D virus (IDV), which is related to ICV. We also examined receptor specificities of toroviruses from swine and cattle that are of veterinary importance and can cause respiratory and gastrointestinal illnesses. The CoVs attach to O-acetylated sialosides via their spike protein for virion uptake. They also express a homodimeric HE, composed of a sialic acid binding and esterase domain, which is important for the destruction of decoy receptors and virion release. Toroviruses encode a gene homologous to HE, whereas ICV and IDV express a multifunctional haemagglutinin-esterase-fusion protein (HEF), in which the haemagglutinin domain is responsible for receptor binding, the esterase region for receptor destruction and the fusion domain for merging the viral envelope with the host membrane.
The receptor binding (S1A) domains of the spike proteins and the enzymatically inactive forms of the ectodomains of HEs and HEFs were expressed in HEK293T cells as Fc fusion proteins. The O-acetylated sialosides were printed on streptavidin-coated glass slides to give a microarray that was exposed to different concentrations of the Fc fusion proteins and binding was detected by an anti-Fc antibody tagged with AlexaFluor-647 (the full structures of the compounds on the microarray are shown in Supplementary Fig. 9). Quality control was performed to ensure that no acetyl migration had occurred during printing and slide storage (see Supplementary Fig. 10 for the liquid chromatography–mass spectroscopy analysis of 1f in the printing solution and Fig. 11 for the stability assessment on-slide).
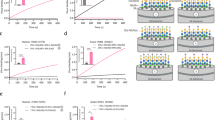
Spike proteins of the human CoVs OC43 and HKU1 showed a strong preference for α2,8-linked disialic acid modified by 9-O-acetyl ester (Fig. 4a). HKU1, which entered the human population centuries earlier than OC4331, exhibited the highest selectivity for this glycoform. Both virolectins also bound disialic acid with an additional C-7 acetyl ester, but with a much lower responsiveness. The HEF of ICV/Johannesburg, a human orthomyxovirus, targets the same structure, but does not tolerate an acetyl ester at C-7.
a, Column graphs of viral receptor selectivities. See Supplementary Fig. 9 for the chemical structures of the compounds. Additional results with different concentrations of proteins are presented in Supplementary Fig. 12. Columns show the background-subtracted average relative fluorescence unit (RFU) values of four replicates. Error bars indicate the s.d. of the RFUs. b, Heat-map presentations of viral receptor selectivities. In the heat map, the signal intensities are normalized with the highest value in each protein defined as 1.0 and shown with a colour gradient. Concentrations of Fc-tagged proteins presented are: BCoV S1A 0.3 μg ml–1, OC43 S1A 0.3 μg ml–1, HKU1 S1A 30 μg ml–1, ICV HEF 0.3 μg ml–1, IDV HEF 0.3 μg ml–1, porcine torovirus (PToV) HE 3 μg ml–1, BToV HE 3 μg ml–1, BCoV HE 3 μg ml–1, ECoV HE 3 μg ml–1, RbCoV HE 3 μg ml–1, CRCoV HE 3 μg ml–1, MHV-S HE 3 μg ml–1, ECoV S1A 3 μg ml–1, RbCoV S1A 3 μg ml–1 and CRCoV S1A 10 μg ml–1. Representative surface dissociation constants (Kd,surf) were obtained for the binding of HKU1 S1A with compounds 3g (9-O-Ac, 60 nM) and 3d (7,9-di-O-Ac, 85 nM). See supplementary Fig. 13 for binding curves. Sialoforms 1b and 2b (4,7-di-O-Ac Neu5Ac) were not included in the library because they have not been documented to be naturally occurring.
The binding selectivities of the spike proteins from animal CoVs differ from those of humans. In the case of BCoV, the presumable ancestor of OC43, the spike is rather promiscuous and binds several 9-mono- and 7,9-di-O-acetylated sialosides preferentially linked in an α2,3-fashion. Equine, canine respiratory and rabbit CoVs (ECoV, CRCoV and RbCoV, respectively) also encode spikes that recognize C-9 acetylated sialosides, but again with the preferences for specific glycosidic forms distinctive from those of the human viruses. Similarly, IDV/Oklahoma, an orthomyxovirus of swine and distantly related to the human pathogen ICV32, prefers α2,3- and 2,6-sialosides.
Surprisingly, the HEs of the animal CoVs display ligand fine specificities different to that of those corresponding spike protein (Fig. 4b). For example, the spikes of ECoV, RbCoV and CRCoV preferentially bind 2,3-linked sialosides modified by a 9-O-monoacetyl ester, whereas their HEs recognize di- or tri-O-acetylated structures. The MHV-S spike and HE represent an extreme case of engaging with different structures, and bind a proteinaceous receptor (CEACAM-1a)33 and 4-O-acetylated sialosides, respectively. The lectin domain of the HEs probably evolved to recognize O-acetylated glycans rich in the mucus layer of the host. It contributes to virion attachment, but also regulates the sialate-O-acetylesterase activity of HE to destroy decoy receptors to facilitate virion migration through the mucus and the release of viral progeny34,35. As a main function, the spike protein has to bind to a cell surface receptor to promote virion uptake followed by membrane fusion and ultimately release of the nucleocapsid into the cytoplasm36. Owing to promiscuity, a spike may also have affinity for decoy receptors in the mucus, and for BCoV and OC43 it has been shown that the HE and spike function as a two-component system for dynamic receptor interactions35.
The binding selectivities of the HEs for specific O-acetylation forms correlate well with the sialic acid repertoire of the mucus of the respective hosts. Cattle produce ample quantities of 7,9-di-O-acetylated sialosides37, which is preferred by BCoV and bovine torovirus (BToV). Equine, murine and rabbits harbour 4-O-acetylated sialic acids across multiple tissues38, which is in agreement with the specificities of the HEs from ECoV, MHV-S and RbCoV, respectively. The loss of HE lectin function in the human coronaviruses OC43 and HKU1 resulted in a substantial reduction in sialate-O-acetylesterase activity towards multivalently displayed 9-O-acetylated sialosides39, possibly as an adaptation to low receptor densities in the human respiratory mucus40.
Identification of functional receptor for human coronaviruses
Although reversible virion attachment to sialosides is key during transit through the mucus, a more stable association at the epithelial cell surface is required for entry. The array studies showed that the spikes of the human viruses OC43 and HKU1 recognize 9-O-acetylated α2,8-linked disialosides. These observations prompted us to examine whether spike-mediated binding to such a glycotope can facilitate infection.
Humans express six α2,8-sialyltransferases (ST8Sia1–ST8Sia6). ST8Sia2, 3 and 4 are involved in the biosynthesis of α2,8-linked linear homopolymers of sialic acid, which are mainly found in the brain, where they play a role in the developmental plasticity of the nervous system41,42. These sialosides have also been detected in secondary lymphatic tissues, where they are involved in the recruitment of immune cells43, and in cancer cells where they play a role in metastasis44. Polysialic acids of mammals are not known to be modified by O-acetyl esters, and have not been observed on epithelial cells, and thus it is unlikely that they function as a receptor for the respiratory viruses.
ST8Sia1 (GD3 synthase), ST8Sia5 (GT3 synthase) and ST8Sia6 transfer a single sialic acid residue to NeuAcα2-3Gal moieties, and thereby form α2,8-linked disialosides. The first two enzymes modify gangliosides, whereas the latter one acts on O-linked glycans45,46,47. A recent study demonstrated that the human respiratory tract expresses α2,8-linked disialylated glycolipids, and in particular the ganglioside GD3 (NeuAcα2,8NeuAcα2,3Galβ1,4Glcβ1-Cer) was abundantly present48. This ganglioside is one of a few glycoconjugates in humans known to be modified by a C-9 acetyl ester (9-Ac-GD3).
At the plasma membrane, gangliosides cluster in microdomains, where they can engage in multivalent interactions with receptor binding proteins, which results in high avidity binding, receptor clustering and endocytosis49. Indeed, they are known to promote cellular uptake of bacterial toxins50 and viruses, which include ICV51. In the case of CoVs, the attachment of virions to gangliosides via the 20 nm spike proteins is expected to keep the 8 nm HE proteins at some distance from the cell surface, and thereby protect these receptors from HE-mediated destruction and prevent virion elution39. Thus, we asked whether gangliosides, such as 9-Ac-GD3, may function as a receptor for OC43 and HKU1.
The biosynthesis of 9-Ac-GD3 involves the concerted action of N-acetylneuraminate 9-O-acetyltransferase (CASD1), which catalyses the addition of an acetyl ester to C-9 of cytidine-monophosphate-linked sialic acid to give CMP-Neu5,9Ac2 and ST8Sia1, which can transfer this sugar nucleotide to GM352. HEK293T cells naturally express CASD1, and by transfecting these cells with ST8Sia1, produce 9-Ac-GD3. Thus, we examined by immunofluorescence the binding of OC43 and HKU1 S1A-Fc fusion proteins to cells transfected by ST8Sia1 (Fig. 5a). OC43 S1A-Fc bound to mock-transfected cells, in accordance with the susceptibility of HEK293T-cells to OC43-1967/USA infection39, but binding was strongly increased on ST8Sia1 overexpression. HKU1 can only be propagated in epithelial cell cultures and select cell lines53, and not in HEK293T cells. As anticipated, HKU1 S1A-Fc did not detectably bind to mock-transfected HEK293T cells. However, it strongly bound to ST8Sia1-transfected cells, which demonstrates that the latter enzyme is involved in the biosynthesis of the receptor for this virus. Substitution of the critical S1A receptor binding site residue Trp90 by Ala abolished binding54, which shows that an intact receptor binding site is required for binding. Treatment of transfected cells with BCoV HE strongly reduced the binding, which confirms a dependence of the O-acetylation of sialic acid for binding. Finally, a strong reduction in binding was also observed when transfected cells were pretreated with an inhibitor of glycolipid biosynthesis (Supplementary Fig. 14)55, which supports that S1A-Fc binds to 9-O-acetylated disialogangliosides.
a, Immunofluorescent staining of mock-transfected or ST8Sia1-transfected HEK293T cells using wild-type or W90A-mutant S1A-mouse-Fc from OC43 or HKU1. Cells pretreated with the sialate-9-O‐acetyl esterase (9-O-SOAE), BCoV HE, were used as the receptor-depleted control. W, tryptophan; A, alanine. See Supplementary Fig. 14 for cell treatment with the glycolipid biosynthesis inhibitor. Scale bars, 50 μm. This panel shows representative micrographs from three repeats. b, Infectivity assays using OC43 and HKU1 spike-pseudotyped virions. Wild-type or CASD1-knockout (ΔCASD1) HEK293T cells were mock-transfected or ST8Sia1-transfected before infection. VSV particles that encoded a luciferase gene were pseudotyped with a wild-type or W90A-mutant spike from OC43 or HKU1. Infectivity was quantified with relative light units by measuring the signal produced from reaction of luciferase with the substrate. Relative infectivity is obtained by normalizing the relative light units (averages of three technical replicates) against the highest value in OC43 or HKU1. The data points correspond to relative infectivity from three biologically independent experiments. Columns show the mean relative infectivity. Error bars represent the s.d. (n = 3). P values (two-tailed) calculated by Welch’s unequal variances t-test are indicated in the graphs. c, Immunohistochemical staining of nasal (left) and lung (right) epithelia using wild-type (wt) or W90A-mutated OC43 or HKU1 S1A fused with mouse Fc. Scale bars, 200 μm. Shown in this panel are representative tissue micrographs. The tissue staining experiments were repeated twice.
To determine whether 9-O-acetylated disialogangliosides support cell entry, infection assays were performed with vesicular stomatitis virus (VSV) particles pseudotyped with coronavirus spikes. Mock-transfected HEK293T cells were susceptible to infection with VSV pseudotyped with an OC43 spike, but the infection increased significantly on ST8Sia1 overexpression (Fig. 5b). HEK293T cells were not detectably infected by particles pseudotyped with a HKU1 spike, but became susceptible on overexpression of ST8Sia1. VSV particles, pseudotyped with spike proteins defective of receptor binding sites, were non-infectious, and infection by particles that bore wildtype spikes was prohibited in CASD1-knockout HEK293T cells transfected with ST8Sia1.
The microarray and cell-based studies indicate that the spikes of OC43 and HKU1 can bind to gangliosides that contain α2,8-disialosyl, such as 9-Ac-GD3, via their S1A in a 9-O-acetyl-dependent fashion, and thereby facilitate cell entry. Future studies will focus on extending the chemoenzymatic methodology to the preparation of intact ganglioside glycans for direct binding studies to investigate the importance of the underlying glycan for recognition by the human coronaviruses.
Finally, we examined the distribution of O-acetylated disialoside receptors in the human respiratory tract using OC43 and HKU1 S1A fused with mouse Fc (Fig. 5c). The two wild-type S1A proteins stained ciliated pseudostratified columnar epithelial cells of the human nasal cavity and lung tissue. No binding was observed for a receptor-binding-site-deficient S1A, which indicates that the staining is dependent on the interaction with sialate-9-O-acetyl. The results demonstrate that 9-O-acetylated disialoside glycotopes are expressed in the human upper and lower respiratory tracts, with the former being the major site for viral replication in natural infections.
Conclusions
CoVs that emerge from zoonotic introduction pose threats to public health, as poignantly illustrated by the current SARS-CoV-2 pandemic56. Four CoVs of zoonotic origin have become endemic in the human population, which include the alphacoronaviruses NL63 and 229E and the betacoronaviruses OC43 and HKU1, which cause up to 30% of mild respiratory tract infections, but can also lead to severe disease and fatality in young children, the elderly and immunocompromised individuals. Future zoonotic transmissions of CoVs are likely given that there is large animal reservoir and their ability to cross species barriers. Preparedness for future outbreaks will require a fundamental understanding how these viruses adapt to replication in humans, which includes insight into the human-associated patterns of receptor preference. Here, we describe a platform to examine receptor requirements of viruses that engage with O-acetylated sialosides, which include Embecoviruses. Our findings highlight that the human CoVs OC43 and HKU1, and human orthomyxovirus ICV, preferentially bind to 9-O-acetylated α2,8-linked sialosides. Immunofluorescence and cell-entry studies indicate that such a glycotope as part of the glycolipids is employed for viral entry. Preferential usage of a common receptor type by three distinct human respiratory viruses is indicative of a convergent evolution by adapting to the sialoglycome of the human respiratory tract. This knowledge will facilitate predicting future cross-species transmissions and will be important for the implementation of prevention and intervention strategies. In addition to host–virus interactions, O-acetylated sialosides have also been implicated in innate and adaptive immunity37, cancer57 and bacterial foraging and pathogenesis58,59. The glycan array described here will find utility to examine at a molecular level how O-acetylated sialosides modulate physiological and disease processes.
Methods
General procedure for sialylation and isopropylidene removal
A suspension of the diol acceptors 5, 6 or 7 and the sialyl donor 4 (1.2 equiv. for 5 and 6, and 2.0 equiv. for 7) and 4 Å molecular sieves in anhydrous DCM (50 mM based on acceptors) was placed under an atmosphere of argon and then stirred at room temperature for 1 h. The mixture was cooled (−78 °C) and TMSOTf (1.2 equiv. for 5 and 6, and 2.0 equiv. for 7) was added dropwise. The reaction mixture was stirred at −78 °C for 5–10 min after which it was quenched with triethylamine. The mixture was filtered through Celite, diluted with DCM and washed with saturated NaHCO3. The organic phase was dried (Na2SO4) and concentrated in vacuo. The residue was dissolved in DCM, to which a few drops of water was added. TFA (10–15% v/v) was added, and the biphasic reaction mixture was stirred vigorously at room temperature for 2 h. The mixture was directly poured into cold saturated NaHCO3. The organic phase was collected when no more CO2 was generated, dried (Na2SO4), filtered and the filtrate concentrated in vacuo. The reside was purified by silica gel column (toluene:ethyl acetate 3:1–2:1 for 4 + 5 and 4 + 6, and toluene:acetonitrile 4:1–3:1 for 4 + 7).
General procedure for hydrogenation, regioselective acetyl migration and biotin conjugation
In a double-necked flask, the starting material (13, 16 or S15) was dissolved in THF/water that contained acetic acid (5 equiv. to starting materials). The flask was filled with hydrogen followed by the addition of palladium hydroxide (Degussa type). On completion of the reaction, the catalyst was filtered off over Celite and the filtrate concentrated in vacuo and further freeze-dried. The residue was dissolved in DMSO-d6 in the presence of 6 equiv. acetic acid-d4 and 3 equiv. caesium fluoride. The mixture was heated (80 °C) for 14–20 h, after which a proton NMR spectrum was recorded, which showed a shift of Sia H-8 from ~5 ppm to ~4 ppm, which confirmed completion of the reaction. The mixture was lyophilized and the residue directly dissolved in DMSO to which N-hydroxysuccinimde-activated biotin or biotin-Lc was added. On completion of the reaction, as indicated by electrospray ionization mass spectroscopy, the solution was freeze-dried, the resultant residue passed over a Biogel p-2 column and then further purified with reversed-phase HPLC (C18, 5%–40% acetonitrile gradient in water, 50 min, for 1a and 2a) or hydrophilic interaction chromatography HPLC (95%–65% acetonitrile gradient in water, 60 min, for 3a).
HE-mediated de-O-acetylation
The precursor 1a, 2a or 3a was dissolved in 50 mM ammonium formate buffer (pH 6.4) to achieve 5 mM concentration. BCoV and/or MHV-S HEs were added to achieve a concentration of 10 μg ml–1. The progress of the reaction was monitored by electrospray ionization mass spectroscopy (negative mode). On completion, the reaction mixture was lyophilized and purified with reversed-phase HPLC (C18, 5%–40% acetonitrile gradient in water, 50 min, for the 1 and 2 series) or hydrophilic interaction chromatography HPLC (95%–65% acetonitrile gradient in water, 60 min, for the 3 series).
Glycan microarray printing and screening
The biotinylated compounds were printed on streptavidin-coated glass slides (SuperStreptavidin Microarray Substrate Slides, ArrayIt Inc) using a Scienion sciFLEXARRAYER S3 non-contact microarray equipped with a Scienion PDC80 nozzle (Scienion Inc). Compounds were dissolved in MilliQ water at a concentration of 1 mM to prepare a stock solution. This solution was placed at −80 °C for long-term storage. A printing solution of biotinylated sialosides at a concentration of 100 µM was freshly made each time by diluting the stock solution in the printing buffer (10 mM PBS buffer, pH 7). The compounds were printed in replicates of six with a spot volume of ~400 pl at 20 °C and 50% humidity. The slides were stored at 4 °C, and blocked with TSM binding buffer (20 mM Tris·HCl, pH 7, 150 mM NaCl, 2 mM CaCl2 and 2 mM MgCl2, 0.05% Tween-20, 1% BSA) for 1 h at 4 °C prior to use. The human-Fc tagged proteins were mixed with goat anti-human IgG antibody (Alexa Fluor 647 conjugated, 109-605-008, Jackson ImmunoResearch) in a 1:1 ratio in TSM binding buffer and incubated at 4 °C for 1 h. The slides were incubated with the premixed binding solution at 4 °C for 1 h. Postincubation, the slides were washed with (1) TSM washing buffer (20 mM Tris·HCl, pH 7, 150 mM NaCl, 2 mM CaCl2 and 2 mM MgCl2 and 0.05% Tween-20), (2) TSM buffer (20 mM Tris·HCl, pH 7, 150 mM NaCl, 2 mM CaCl2 and 2 mM MgCl2) and (3) water, and were then spun dry. The slides were scanned using a GenePix 4000B microarray scanner (Molecular Devices) at the appropriate excitation wavelength. Images were analysed using GenePix Pro 7 software (version 7.2.29.2, Molecular Devices). This procedure complies with MIRAGE Glycan Array Guideline v 1.060. See Supplementary Information for the method to obtain Kd,surf.
Immunofluorescence assay of cultured cells
HEK-293T cells were grown on glass coverslips in a 24-well format and either mock transfected or transfected with pcDNA-ST8Sia1 plasmid using Fugene according to the manufacturer’s instructions. At 36 h post-transfection, cell monolayers were fixed with paraformaldehyde (3.7% in PBS, 30 min), incubated with blocking buffer (PBS, 0.05% Tween-20 and 2% BSA) for 30 min. For soluble HE, cells were treated with BCoV-LUN HE (20 μg ml–1 in blocking buffer) for 2 h at 37 °C. Cells were then incubated with S1A-mFc lectins (50 μg ml–1) for 45 min, and then for 30 min with goat-anti-mouse IgG-Alexa 594 (A32742, 1:100, Thermo Scientific) and with 4,6-diamidino-2-phenylindole (1:1,000). All the incubations were in blocking buffer. Washing was performed with PBS that contained 0.05% Tween-20. The cells were then embedded in FluorSave (Calbiochem) and examined by confocal fluorescence microscopy (Nikon STORM). Glycan biosynthesis inhibitor N-butyl-deoxygalactonojirimycin were added to culture supernatants of mock-transfected or pcDNA-ST8SIA1-transfected HEK 293 T cells at 8 h post-transfection. Incubation was continued for 72 h. Cells were then fixed and stained with S1A-mFc lectins as above.
Virolectin staining of human tissues
Human tissues were collected from anonymous individuals with their consent to secondary research uses, and stored and issued by the Erasmus MC Tissue Bank. See Supplementary Table 3 for the Tier 1 characteristics following BRISQ guidelines. Human tissue staining was performed using procedures adapted from previous reports10. Briefly, paraffin-embedded human tissue sections were dewaxed in xylene and rehydrated. Endogenous peroxidases were inactivated with peroxide (0.3% in methanol). Tissue sections were then successively incubated with S1A-mFc virolectins (50 μg ml–1) and biotinylated goat-α-mouse IgG antibodies (SAB4600004, Sigma; 1:250; lectins and antibodies all diluted in blocking buffer), with an avidin–biotin horseradish peroxidase complex (ABC-PO staining kit, Thermo Scientific) and with 3,3′-diaminobenzidine (Sigma). Blocking, incubations and washing were performed as for the immunofluorescence assays. Tissue sections were counterstained with Mayer’s haematoxylin, embedded in an Eukitt mounting medium (Fluka) and examined by standard light microscopy.
VSV pseudotyping and infectivity assay
The production of OC43/HKU1 S-pseudotyped G-protein-deficient VSV particles and the infectivity assays were as described previously61. In brief, C-terminal truncated and flag-tagged OC43 and HKU1 S (wild type and W90A mutant) were cloned in the pCAGGS expression vector. A plasmid mixture of pCAGGS–OC43/HKU1–flag and pCD5–BCoV HE-Fc (molar ratios 8:1) were complexed with PEI and transfected to HEK 293T cells at 70% confluency. At 48 h post-transfection, cells were transduced with VSV-G pseudotyped VSV∆G/Fluc with an multiplicity of infection of 1. Cell-free supernatants were then harvested at 24 h after transduction, filtered through 0.45 µm membranes and the virus particles were further purified and concentrated by sucrose cushion ultracentrifugation at 29,000 r.p.m. for 2.5 h. Pelleted virions were resuspended in PBS and stored at −80 °C before usage. For the infection assay, HEK-293T or HEK-293T∆CASD1 cells were seeded in a 96-well cluster format, and were transfected or mock transfected with pcDNA-ST8SIA1 plasmid using Fugene. At 36 h post-transfection, infection was performed with equal amounts of S-pseudotyped VSVs as calculated from the VSV-N content. After 18 h, the infected cells were lysed using passive lysis buffer (Promega). Firefly luciferase expression was measured via a homemade firefly luciferase assay system as described54. Infection experiments were conducted in independent triplicates, each of which with three technical replicates.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The data supporting the findings of this study are included in this article and the Supplementary Information. Source data are provided with this paper.
Code availability
The script for Microsoft Excel Macro for batch processing glycan microarray data is uploaded to https://github.com/enthalpyliu/carbohydrate-microarray-processing.
References
Varki, A. Nothing in glycobiology makes sense, except in the light of evolution. Cell 126, 841–845 (2006).
Le Pendu, J., Nyström, K. & Ruvoën-Clouet, N. Host–pathogen co-evolution and glycan interactions. Curr. Opin. Virol. 7, 88–94 (2014).
Thompson, A. J., De Vries, R. P. & Paulson, J. C. Virus recognition of glycan receptors. Curr. Opin. Virol. 34, 117–129 (2019).
Long, J. S., Mistry, B., Haslam, S. M. & Barclay, W. S. Host and viral determinants of influenza A virus species specificity. Nat. Rev. Microbiol. 17, 67–81 (2018).
Stencel-Baerenwald, J. E., Reiss, K., Reiter, D. M., Stehle, T. & Dermody, T. S. The sweet spot: defining virus–sialic acid interactions. Nat. Rev. Microbiol. 12, 739–749 (2014).
Wasik, B. R., Barnard, K. N. & Parrish, C. R. Effects of sialic acid modifications on virus binding and infection. Trends Microbiol. 24, 991–1001 (2016).
Morse, S. S. et al. Prediction and prevention of the next pandemic zoonosis. Lancet 380, 1956–1965 (2012).
Pepin, K. et al. Identifying genetic markers of adaptation for surveillance of viral host jumps. Nat. Rev. Microbiol. 8, 802–813 (2010).
Schauer, R. & Kamerling, J. P. Exploration of the sialic acid world. Adv. Carbohydr. Chem. Biochem. 75, 1–213 (2018).
Langereis, M. A. et al. Complexity and diversity of the mammalian sialome revealed by nidovirus virolectins. Cell Rep. 11, 1966–1978 (2015).
Wasik, B. R. et al. Distribution of O-acetylated sialic acids among target host tissues for influenza virus. mSphere 2, e00379-16 (2017).
Chen, X. & Varki, A. Advances in the biology and chemistry of sialic acids. ACS Chem. Biol. 5, 163–176 (2010).
Haverkamp, J., Schauer, R., Wember, M., Kamerling, J. P. & Vliegenthart, J. F. Synthesis of 9-O-acetyl and 4,9-di-O-acetyl derivatives of the methyl ester of N-acetyl-β-d-neuraminic acid methylglycoside: their use as models in periodate oxidation studies. Hoppe Seyler Z. Physiol. Chem. 356, 1575–1583 (1975).
Anazawa, K., Furuhata, K. & Ogura, H. Synthesis of 7-O-acetyl-N-acetylneuraminic acid-derivative. Chem. Pharm. Bull. 36, 4976–4979 (1988).
Furuhata, K. & Ogura, H. Studies on sialic acids. Syntheses of partially O-acetylated 4-methylcoumarin-7-yl 5-acetamido-3,5-dideoxy-α-d-glycero-d-galacto-2-nonulopyranosidonic acids. Chem. Pharm. Bull. 37, 2037–2040 (1989).
Roth, A. & Faillard, H. Synthesis of fluorescent 7,8,9-tri-O-acetyl-N-acetyl- and 4-O-acetyl-N-acetylneuraminic acid α-thioketosides. Liebigs Ann. Chem. 1993, 485–489 (1993).
Clarke, P. A., Mistry, N. & Thomas, G. H. Synthesis of the complete series of mono acetates of N-acetyl-d-neuraminic acid. Org. Biomol. Chem. 10, 529–535 (2012).
Park, S. S. & Gervay-Hague, J. Synthesis of partially O-acetylated N-acetylneuraminic acid using regioselective silyl exchange technology. Org. Lett. 16, 5044–5047 (2014).
Yu, H. et al. Highly efficient chemoenzymatic synthesis of naturally occurring and non-natural α-2,6-linked sialosides: a P. Damsela α-2,6-sialyltransferase with extremely flexible donor–substrate specificity. Angew. Chem. Int. Ed. 45, 3938–3944 (2006).
Sugiarto, G. et al. A sialyltransferase mutant with decreased donor hydrolysis and reduced sialidase activities for directly sialylating Lewis X. ACS Chem. Biol. 7, 1232–1240 (2012).
Yu, H. et al. Chemoenzymatic synthesis of GD3 oligosaccharides and other disialyl glycans containing natural and non-natural sialic acids. J. Am. Chem. Soc. 131, 18467–18477 (2009).
Yu, H. et al. Effective one-pot multienzyme (OPME) synthesis of monotreme milk oligosaccharides and other sialosides containing 4-O-acetyl sialic acid. Org. Biomol. Chem. 14, 8586–8597 (2016).
Takayama, S., Livingston, P. O. & Wong, C.-H. Synthesis of the melanoma-associated ganglioside 9-O-acetyl GD3 through regioselective enzymatic acetylation of GD3 using subtilisin. Tetrahedron Lett. 37, 9271–9274 (1996).
Guberman, M. & Seeberger, P. H. Automated glycan assembly: a perspective. J. Am. Chem. Soc. 141, 5581–5592 (2019).
Regl, G. et al. The hemagglutinin–esterase of mouse hepatitis virus strain s is a sialate-4-O-acetylesterase. J. Virol. 73, 4721–4727 (1999).
Kamerling, J. P. et al. Migration of O-acetyl groups in N,O-acetylneuraminic acids. Eur. J. Biochem. 162, 601–607 (1987).
Chuang, H. Y. et al. Synthesis and vaccine evaluation of the tumor-associated carbohydrate antigen RM2 from prostate cancer. J. Am. Chem. Soc. 135, 11140–11150 (2013).
Komura, N. et al. Constrained sialic acid donors enable selective synthesis of α-glycosides. Science 364, 677–680 (2019).
Tanaka, H., Nishiura, Y. & Takahashi, T. An efficient convergent synthesis of GP1c ganglioside epitope. J. Am. Chem. Soc. 130, 17244–17245 (2008).
Su, S. et al. Epidemiology, genetic recombination, and pathogenesis of coronaviruses. Trends Microbiol. 24, 490–502 (2016).
Lau, S. K. et al. Discovery of a novel coronavirus, China Rattus coronavirus HKU24, from Norway rats supports the murine origin of betacoronavirus 1 and has implications for the ancestor of betacoronavirus lineage A. J. Virol. 89, 3076–3092 (2015).
Hause, B. M. et al. Isolation of a novel swine influenza virus from Oklahoma in 2011 which is distantly related to human influenza C viruses. PLoS Pathog. 9, e1003176 (2013).
Williams, R. K., Jiang, G. S. & Holmes, K. V. Receptor for mouse hepatitis virus is a member of the carcinoembryonic antigen family of glycoproteins. Proc. Natl Acad. Sci. USA 88, 5533–5536 (1991).
Linden, S. K., Sutton, P., Karlsson, N. G., Korolik, V. & McGuckin, M. A. Mucins in the mucosal barrier to infection. Mucosal Immunol. 1, 183–197 (2008).
Lang, Y. et al. Coronavirus hemagglutinin–esterase and spike proteins coevolve for functional balance and optimal virion avidity. Proc. Natl Acad. Sci. USA 117, 25759–25770 (2020).
Li, F. Structure, function, and evolution of coronavirus spike proteins. Annu. Rev. Virol. 3, 237–261 (2016).
Schauer, R., Srinivasan, G. V., Wipfler, D., Kniep, B. & Schwartz-Albiez, R. O-acetylated sialic acids and their role in immune defense. Adv. Exp. Med. Biol. 705, 525–548 (2011).
Park, S. S. Post-glycosylation modification of sialic acid and its role in virus pathogenesis. Vaccines 7, 171 (2019).
Bakkers, M. J. et al. Betacoronavirus adaptation to humans involved progressive loss of hemagglutinin–esterase lectin activity. Cell Host Microbe 21, 356–366 (2017).
Barnard, K. N. et al. Modified sialic acids on mucus and erythrocytes inhibit influenza A virus hemagglutinin and neuraminidase functions. J. Virol. 94, e01567-19 (2020).
Mühlenhoff, M., Rollenhagen, M., Werneburg, S., Gerardy-Schahn, R. & Hildebrandt, H. Polysialic acid: versatile modification of NCAM, SynCAM-1 and neuropilin-2. Neurochem. Res. 38, 1134–1143 (2013).
Volkers, G. et al. Structure of human ST8SiaIII sialyltransferase provides insight into cell-surface polysialylation. Nat. Struct. Mol. Biol. 22, 627–635 (2015).
Kiermaier, E. et al. Polysialylation controls dendritic cell trafficking by regulating chemokine recognition. Science 351, 186–190 (2016).
Elkashef, S. M. et al. Polysialic acid sustains cancer cell survival and migratory capacity in a hypoxic environment. Sci. Rep. 6, 33026 (2016).
Bobowski, M., Harduin-Lepers, A. & Delannoy, P. in Handbook of Glycosyltransferases and Related Genes (eds Taniguchi, N. et al.) 767–780 (Springer, 2014).
Tsuji, S. & Takashima, S. in Handbook of Glycosyltransferases and Related Genes (eds Taniguchi, N. et al.) 813–821 (Springer, 2014).
Takashima, S. & Tsuji, S. in Handbook of Glycosyltransferases and Related Genes (eds Taniguchi, N. et al.) 823–832 (Springer, 2014).
Jia, N. et al. The human lung glycome reveals novel glycan ligands for influenza A virus. Sci. Rep. 10, 5320 (2020).
Regina Todeschini, A. & Hakomori, S. Functional role of glycosphingolipids and gangliosides in control of cell adhesion, motility, and growth, through glycosynaptic microdomains. Biochim. Biophys. Acta 1780, 421–433 (2008).
Ewers, H. & Helenius, A. Lipid-mediated endocytosis. CSH Perspect. Biol. 3, a004721 (2011).
Herrler, G. & Klenk, H.-D. The surface receptor is a major determinant of the cell tropism of influenza C virus. Virology 159, 102–108 (1987).
Baumann, A. M. et al. 9-O-acetylation of sialic acids is catalysed by CASD1 via a covalent acetyl-enzyme intermediate. Nat. Commun. 6, 7673 (2015).
Pyrc, K. et al. Culturing the unculturable: human coronavirus HKU1 infects, replicates, and produces progeny virions in human ciliated airway epithelial cell cultures. J. Virol. 84, 11255–11263 (2010).
Hulswit, R. J. G. et al. Human coronaviruses OC43 and HKU1 bind to 9-O-acetylated sialic acids via a conserved receptor-binding site in spike protein domain A. Proc. Natl Acad. Sci. USA 116, 2681–2690 (2019).
Andersson, U., Butters, T. D., Dwek, R. A. & Platt, F. M. N-butyldeoxygalactonojirimycin: a more selective inhibitor of glycosphingolipid biosynthesis than N-butyldeoxynojirimycin, in vitro and in vivo. Biochem. Pharmacol. 59, 821–829 (2000).
Coronaviridae Study Group of the International Committee on Taxonomy of Viruses. The species severe acute respiratory syndrome-related coronavirus: classifying 2019-nCoV and naming it SARS-CoV-2. Nat. Microbiol. 5, 536–544 (2020).
Parameswaran, R. et al. O-acetylated N-acetylneuraminic acid as a novel target for therapy in human pre-B acute lymphoblastic leukemia. J. Exp. Med. 210, 805–819 (2013).
Robinson, L. S., Lewis, W. G. & Lewis, A. L. The sialate O-acetylesterase EstA from gut Bacteroidetes species enables sialidase-mediated cross-species foraging of 9-O-acetylated sialoglycans. J. Biol. Chem. 292, 11861–11872 (2017).
Nguyen, T. et al. The role of 9-O-acetylated glycan receptor moieties in the typhoid toxin binding and intoxication. PLoS Pathog. 16, e1008336 (2020).
Liu, Y. et al. The minimum information required for a glycomics experiment (MIRAGE) project: improving the standards for reporting glycan microarray-based data. Glycobiology 27, 280–284 (2017).
Tortorici, M. A. et al. Structural basis for human coronavirus attachment to sialic acid receptors. Nat. Struct. Mol. Biol. 26, 481–489 (2019).
Acknowledgements
We thank D. van Riel and L. M. E. Leijten (Erasmus MC) for providing the formalin-fixed paraffin-embedded human tissue sections, R. Wubbolts (Utrecht University) for offering guidance on confocal fluorescence microscopy and H. Egberink and S. Zhao (Utrecht University) for providing plasmids of the ECoV spike protein. This work was supported by TOP-PUNT Grant 718.015.003 of the Netherlands Organization for Scientific Research (G.-J.B.), ECHO Grant 711.011.006 of the Council for Chemical Sciences of the Netherlands Organization for Scientific Research (R.J.d.G.) and China Scholarship Council 2014-03250042 (Y.L.).
Author information
Authors and Affiliations
Contributions
G.-J.B., R.J.d.G., Z.L. and Y.L. conceived and designed the experiments. G.-J.B. and R.J.d.G. supervised the study. Z.L. and M.I.B. performed the chemical and enzymatic syntheses. Y.L. produced the enzymes and viral glycoproteins used in the microarray analysis. L.L. performed the glycan microarray screening and wrote the script for batch-processing microarray data. G.-J.B., Z.L. and L.L. analysed the glycan microarray data. Y.L. and A.I.S. performed the immunofluorescence assay, histochemical tissue staining and infectivity assay. G.-J.B., R.J.d.G., Z.L. and Y.L. co-wrote the paper. All authors discussed the results and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Chemistry thanks Chi-Huey Wong, and the other anonymous reviewer(s), for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–14, Tables 1–3, materials and methods, detailed synthesis procedures, additional references and NMR spectra.
Supplementary Data 1
Unprocessed and background-subtracted glycan microarray signal intensities of each individual replicate spot for Supplementary Fig. 12.
Supplementary Data 2
Data for binding curve of HKU1 S1A protein (Supplementary Fig. 13).
Source data
Source Data Fig. 4
Unprocessed and background-subtracted glycan microarray signal intensities of each individual replicate spot for Fig. 4.
Source Data Fig. 5
Relative light unit (RLU) values of pseudovirus infection assay of each technical replicates in each independent experiment for Fig. 5b.
Rights and permissions
About this article
Cite this article
Li, Z., Lang, Y., Liu, L. et al. Synthetic O-acetylated sialosides facilitate functional receptor identification for human respiratory viruses. Nat. Chem. 13, 496–503 (2021). https://doi.org/10.1038/s41557-021-00655-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41557-021-00655-9
This article is cited by
-
Integrated chemoenzymatic synthesis of a comprehensive sulfated ganglioside glycan library to decipher functional sulfoglycomics and sialoglycomics
Nature Chemistry (2024)
-
Chemoenzymatic synthesis of human natural killer-1-containing glycans and application as serum antibodies probes
Nature Synthesis (2023)
-
Sialic acid O-acetylation patterns and glycosidic linkage type determination by ion mobility-mass spectrometry
Nature Communications (2023)
-
TMPRSS2 is a functional receptor for human coronavirus HKU1
Nature (2023)
-
Multivalent 9-O-Acetylated-sialic acid glycoclusters as potent inhibitors for SARS-CoV-2 infection
Nature Communications (2022)