Abstract
Mitochondrial replacement therapy (MRT) has been used to prevent maternal transmission of disease-causing mutations in mitochondrial DNA (mtDNA). However, because MRT requires nuclear transfer, it carries the risk of mtDNA carryover and hence of the reversion of mtDNA to pathogenic levels owing to selective replication and genetic drift. Here we show in HeLa cells, mouse embryos and human embryos that mtDNA heteroplasmy can be reduced by pre-labelling the mitochondrial outer membrane of a donor zygote via microinjection with an mRNA coding for a transmembrane peptide fused to an autophagy receptor, to induce the degradation of the labelled mitochondria via forced mitophagy. Forced mitophagy reduced mtDNA carryover in newly reconstructed embryos after MRT, and had negligible effects on the growth curve, reproduction, exercise capacity and other behavioural characteristics of the offspring mice. The induction of forced mitophagy to degrade undesired donor mtDNA may increase the clinical feasibility of MRT and could be extended to other nuclear transfer techniques.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$99.00 per year
only $8.25 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The main data supporting the results in this study are available within the paper and its Supplementary Information. Source data for the figures are provided with this paper. Other raw data generated during the study are available from the corresponding authors on reasonable request. Source data are provided with this paper.
References
Elliott, H. R., Samuels, D. C., Eden, J. A., Relton, C. L. & Chinnery, P. F. Pathogenic mitochondrial DNA mutations are common in the general population. Am. J. Hum. Genet. 83, 254–260 (2008).
Gorman, G. S. et al. Prevalence of nuclear and mitochondrial DNA mutations related to adult mitochondrial disease. Ann. Neurol. 77, 753–759 (2015).
Lightowlers, R. N., Taylor, R. W. & Turnbull, D. M. Mutations causing mitochondrial disease: what is new and what challenges remain? Science 349, 1494–1499 (2015).
DeLuca, S. Z. & O’Farrell, P. H. Barriers to male transmission of mitochondrial DNA in sperm development. Dev. Cell 22, 660–668 (2012).
Luo, S. M., Schatten, H. & Sun, Q. Y. Sperm mitochondria in reproduction: good or bad and where do they go? J. Genet. Genomics 40, 549–556 (2013).
Wallace, D. C. Mitochondrial genetic medicine. Nat. Genet. 50, 1642–1649 (2018).
Ou, X. H. & Sun, Q. Y. Mitochondrial replacement techniques or therapies (MRTs) to improve embryo development and to prevent mitochondrial disease transmission. J. Genet. Genomics 44, 371–374 (2017).
Stewart, J. B. & Chinnery, P. F. The dynamics of mitochondrial DNA heteroplasmy: implications for human health and disease. Nat. Rev. Genet. 16, 530–542 (2015).
Bacman, S. R. et al. MitoTALEN reduces mutant mtDNA load and restores tRNA(Ala) levels in a mouse model of heteroplasmic mtDNA mutation. Nat. Med. 24, 1696–1700 (2018).
Gammage, P. A. et al. Genome editing in mitochondria corrects a pathogenic mtDNA mutation in vivo. Nat. Med. 24, 1691–1695 (2018).
Mok, B. Y. et al. A bacterial cytidine deaminase toxin enables CRISPR-free mitochondrial base editing. Nature https://www.nature.com/articles/s41586-020-2477-4 (2020).
Greenfield, A. et al. Assisted reproductive technologies to prevent human mitochondrial disease transmission. Nat. Biotechnol. 35, 1059–1068 (2017).
Craven, L. et al. Pronuclear transfer in human embryos to prevent transmission of mitochondrial DNA disease. Nature 465, 82–85 (2010).
Hyslop, L. A. et al. Towards clinical application of pronuclear transfer to prevent mitochondrial DNA disease. Nature 534, 383–386 (2016).
Kang, E. et al. Mitochondrial replacement in human oocytes carrying pathogenic mitochondrial DNA mutations. Nature 540, 270–275 (2016).
Paull, D. et al. Nuclear genome transfer in human oocytes eliminates mitochondrial DNA variants. Nature 493, 632–637 (2013).
Tachibana, M. et al. Towards germline gene therapy of inherited mitochondrial diseases. Nature 493, 627–631 (2013).
Tachibana, M. et al. Mitochondrial gene replacement in primate offspring and embryonic stem cells. Nature 461, 367–372 (2009).
Wang, T. et al. Polar body genome transfer for preventing the transmission of inherited mitochondrial diseases. Cell 157, 1591–1604 (2014).
Wu, K. et al. Polar bodies are efficient donors for reconstruction of human embryos for potential mitochondrial replacement therapy. Cell Res. 27, 1069–1072 (2017).
Wang, Z. et al. Mitochondrial replacement in macaque monkey offspring by first polar body transfer. Cell Res. 31, 233–236 (2021).
Hudson, G., Takeda, Y. & Herbert, M. Reversion after replacement of mitochondrial DNA. Nature 574, E8–E11 (2019).
Yamada, M. et al. Genetic drift can compromise mitochondrial replacement by nuclear transfer in human oocytes. Cell Stem Cell 18, 749–754 (2016).
Craven, L., Tang, M. X., Gorman, G. S., De Sutter, P. & Heindryckx, B. Novel reproductive technologies to prevent mitochondrial disease. Hum. Reprod. Update 23, 501–519 (2017).
Youle, R. J. & Narendra, D. P. Mechanisms of mitophagy. Nat. Rev. Mol. Cell Biol. 12, 9–14 (2011).
Kerr, J. S. et al. Mitophagy and Alzheimer’s disease: cellular and molecular mechanisms. Trends Neurosci. 40, 151–166 (2017).
Sandoval, H. et al. Essential role for Nix in autophagic maturation of erythroid cells. Nature 454, 232–235 (2008).
Birgisdottir, A. B., Lamark, T. & Johansen, T. The LIR motif - crucial for selective autophagy. J. Cell Sci. 126, 3237–3247 (2013).
Wu, K. et al. Mitochondrial replacement by pre-pronuclear transfer in human embryos. Cell Res. 27, 834–837 (2017).
Tsukamoto, S. et al. Functional analysis of lysosomes during mouse preimplantation embryo development. J. Reprod. Dev. 59, 33–39 (2013).
Luo, S. M. et al. Unique insights into maternal mitochondrial inheritance in mice. Proc. Natl Acad. Sci. USA 110, 13038–13043 (2013).
Bavister, B. D. & Squirrell, J. M. Mitochondrial distribution and function in oocytes and early embryos. Hum. Reprod. 15, 189–198 (2000).
Wai, T. et al. The role of mitochondrial DNA copy number in mammalian fertility. Biol. Reprod. 83, 52–62 (2010).
Cao, L. et al. The mitochondrial bottleneck occurs without reduction of mtDNA content in female mouse germ cells. Nat. Genet. 39, 386–390 (2007).
Cree, L. M. et al. A reduction of mitochondrial DNA molecules during embryogenesis explains the rapid segregation of genotypes. Nat. Genet. 40, 249–254 (2008).
Khrapko, K. Two ways to make an mtDNA bottleneck. Nat. Genet. 40, 134–135 (2008).
Lieber, T., Jeedigunta, S. P., Palozzi, J. M., Lehmann, R. & Hurd, T. R. Mitochondrial fragmentation drives selective removal of deleterious mtDNA in the germline. Nature 570, 380–384 (2019).
Wai, T., Teoli, D. & Shoubridge, E. A. The mitochondrial DNA genetic bottleneck results from replication of a subpopulation of genomes. Nat. Genet. 40, 1484–1488 (2008).
Ma, H. et al. Functional human oocytes generated by transfer of polar body genomes. Cell Stem Cell 20, 112–119 (2017).
Bredenoord, A. L. & Appleby, J. B. Mitochondrial replacement techniques: remaining ethical challenges. Cell Stem Cell 21, 301–304 (2017).
Zhang, J. et al. Live birth derived from oocyte spindle transfer to prevent mitochondrial disease. Reprod. Biomed. Online 34, 361–368 (2017).
Reichmann, J. et al. Dual-spindle formation in zygotes keeps parental genomes apart in early mammalian embryos. Science 361, 189–193 (2018).
Xu, X., Li, L., Zhang, C. & Meng, L. Observation of two separate bipolar spindles in the human zygote. J. Assist. Reprod. Genet. 36, 601–602 (2019).
Tamashiro, K. L. et al. Phenotype of cloned mice: development, behavior, and physiology. Exp. Biol. Med. 228, 1193–1200 (2003).
Rodriguiz, R. M. & Wetsel, W. C. in Animal Models of Cognitive Impairment (eds. Levin, E. D. & Buccafusco, J. J.) Ch. 12 (CRC Press/Taylor & Francis, 2006).
Brown, R. E. Behavioural phenotyping of transgenic mice. Can. J. Exp. Psychol. 61, 328–344 (2007).
Barritt, J., Willadsen, S., Brenner, C. & Cohen, J. Cytoplasmic transfer in assisted reproduction. Hum. Reprod. Update 7, 428–435 (2001).
Miao, Y. L., Kikuchi, K., Sun, Q. Y. & Schatten, H. Oocyte aging: cellular and molecular changes, developmental potential and reversal possibility. Hum. Reprod. Update 15, 573–585 (2009).
Planchon, T. A. et al. Rapid three-dimensional isotropic imaging of living cells using Bessel beam plane illumination. Nat. Methods 8, 417–423 (2011).
Waterfall, C. M., Eisenthal, R. & Cobb, B. D. Kinetic characterization of primer mismatches in allele-specific PCR: a quantitative assessment. Biochem. Biophys. Res. Commun. 299, 715–722 (2002).
Acknowledgements
We thank those who donated gametes for this study, and R. Chen (South China Normal University, Guangzhou, China) and H. Schatten (University of Missouri-Columbia) for editing and proofreading the manuscript. This study was supported by the National Key Research and Development Program of China (2018YFC1004800), the National Natural Science Foundation of China (82071714, 81971357) and the Key-Area Research and Development Program of Guangdong Province (2019B030335001).
Author information
Authors and Affiliations
Contributions
S.-M.L. and X.-Y.F. conceived the idea. S.-M.L., X.-H.O., Q.-Y.S., X.-Y.F., L.G., L.-N.C., S.Y., C.H., L.Z., J.-Y.M., S.L., T.J., M.-X.J., W.S., Z.-J.G., Z.-B.W. and M.C. designed experiments and interpreted the results. X.-Y.F., S.-M.L., L.G., J.W., X.-H.S., F.W., C.-F.Z. and X.-H.W. carried out the experiments. S.-M.L., S.Y., X.-Y.F., C.H. and Q.-Y.S. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Biomedical Engineering thanks the anonymous reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 CISD1-RFP-SQSTM1 induces forced mitophagy and reduces the mitochondrial number.
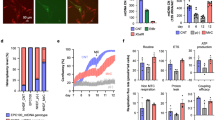
a. HeLa cells were stably transfected with GFP-LC3 (green) and stained with MitoTracker Deep Red FM (grey) to label mitochondria. CISD1-RFP-SQSTM1 (red) was transiently transfected into the HeLa cells. Images were obtained 60 hours after CISD1-RFP-SQSTM1 transfection. The insets indicate that CISD1-RFP-SQSTM1-labeled mitochondria co-localizes with GFP-LC3. b. CISD1-RFP-SQSTM1-labelled mitochondria fuse with lysosomes. HeLa cells were stably transfected with pmEmerald-Mito (grey) to label mitochondria, transiently transfected with CISD1-RFP-SQSTM1 (red), and stained with LysoTracker Blue DND-22 (green) to label lysosomes. Images were obtained 48 hours after CISD1-RFP-SQSTM1 transfection. The insets indicate that CISD1-RFP-SQSTM1-labeled mitochondria fuse with lysosomes. c. The CISD1-RFP-SQSTM1-expressed cells have fewer mitochondria than the unexpressed cells. The stably pmEmerald-Mito (green)-transfected HeLa cells were transiently transfected with CISD1-RFP-SQSTM1 (red) and stained with Hoechst 33342. Images were obtained24 and 48 hours after CISD1-RFP-SQSTM1 transfection. White arrowheads indicate that the CISD1-RFP-SQSTM1-expression cells decrease the pmEmerald-Mito fluorescence signals. In a–c, scale bars, 30 μm, and three independently repeated experiments with similar results. d. The fluorescence analysis of mitochondria in the CISD1-RFP-SQSTM1-expressed and control cells at different time points. Data were analyzed by two-tailed unpaired t-test and expressed as average ± SEM, and each point represents one cell from 3 biological replicates. For the 24 hours group, n = 106, the control value is 945 ± 84.97, and the CISD1-RFP-SQSTM1 value is 950.8 ± 90.19. For the 48 hours group, n = 67, the control value is 815.9 ± 51.32, and the CISD1-RFP-SQSTM1 value is 455.7 ± 39.86. e. The mtDNA number decreased after CISD1-RFP-SQSTM1 expression. Forty-eight hours after CISD1-RFP-SQSTM1 transfection, a single cell with or without CISD1-RFP-SQSTM1 expression was picked up, and the mtDNA quantity was analyzed by qPCR. Data were analyzed by two-tailed unpaired t-test and expressed as average ± SEM. n = 27 for control, and n = 30 for CISD1-RFP-Binding from 3 biological replicates. The value of control is 2357 ± 104.3, and the value of CISD1-RFP-Binding is 1274 ± 79.14.
Extended Data Fig. 2 CISD1-Binding expresses immediately and localizes to mitochondria after its mRNA microinjection in mouse oocytes and Zygotes.
The mRNA encoding CISD1-Binding was microinjected into mouse GV oocytes, MII oocytes, and early pronuclear embryos. Images were obtained 1.5 hours after microinjection. Red indicates CISD1-Binding proteins and Green indicates MitoTracker Deep Red FM. Pearson’s correlation coefficient assessed co-localization between the CISD1-Binding and Mitotracker. Data were analyzed from three independently repeated experiments, and each point represents one cell. n = 10 for GV oocytes; n = 11 for MII oocytes and n = 12 for zygotes. The mean ± SEM values are 0.899 ± 0.01286 (GV), 0.9491 ± 0.00667 (MII) and 0.9467 ± 0.0076 (Zygote). Bar = 30 μm.
Extended Data Fig. 3
CISD1-Binding recruits GFP-LC3 at different mouse-embryo stages. Mouse zygotes were microinjected with the CISD1-Binding (red) and GFP-LC3 (green) mRNA and cultured to 2-cell, 4-cell and 8-cell stages. The insets indicate that CISD1-Binding recruits GFP-LC3. Bar = 30 μm. Also, see Fig. 3a and c. Four independently repeated biological experiments with similar results.
Extended Data Fig. 4 CISD1-Binding-labelled mitochondria begin to fuse with lysosomes after 4-cell stages.
Mouse embryos were microinjected with the CISD1-Binding mRNA at the zygote stage and stained with Blue DND-22 to label lysosomes at 1-cell, 2-cell and 4-cell stages before imaging. The insets indicate that CISD1-Binding-labeled mitochondria fuse with lysosomes (orange only). Bar = 30 μm. Also, see Fig. 3b and e. Three independently repeated biological experiments with similar results.
Extended Data Fig. 5 CISD1-RFP-Binding(mutation)-labelled mitochondria do not recruit LC3 proteins and fuse with lysosomes in mouse embryos.
a. Mouse zygotes were microinjected with the mRNA encoding CISD1-RFP-Binding(mutation) and GFP-LC3. The insets indicate that the CISD1-RFP-Binding(mutation)-labelled mitochondria do not recruit GFP-LC3 in mouse embryos. b. Mouse zygotes were microinjected with the mRNA encoding CISD1-RFP-Binding(mutation) and stained with Blue DND-22 to label lysosomes before imaging. The insets indicate no fusion between the CISD1-RFP-Binding(mutation)-labelled mitochondria and lysosomes in mouse embryos. In a and b, scale bars, 30 μm, and experiments with similar results were biologically repeated twice.
Extended Data Fig. 6 The capacity of early mouse embryos to degrade CISD1-Binding-labelled mitochondria.
a. Mouse embryos were injected with or without (control) CISD1-Binding mRNA at the zygote stage, cultured to different stages, and then the whole embryos were subjected to absolute quantification of the mtDNA. Data were analyzed by two-tailed unpaired t-test and expressed as average ± SEM. Each point represents one embryo from three biological replicates. For Zygote, n = 14, the values are 2.44 ± 0.1618 (control) and 2.485 ± 0.1586 (CISD1-Binding). For 2-Cell, n = 13, the values are 2.543 ± 0.1016 (control) and 2.642 ± 0.1891 (CISD1-Binding). For 4-Cell, n = 12 (control) and n = 11 (CISD1-Binding), the values are 2.529 ± 0.07611 (control) and 2.405 ± 0.1418 (CISD1-Binding). For 8-Cell, n = 11, the values are 3.109 ± 0.1873 (control) and 2.703 ± 0.1292 (CISD1-Binding). For Morula, n = 12, the values are 3.457 ± 0.2498 (control) and 1.466 ± 0.2157 (CISD1-Binding). For Blastocyst group, n = 12, the values are 2.659 ± 0.1224 (control) and 2.143 ± 0.1093 (CISD1-Binding). b. Morula embryos expressed with or without CISD1-Bindingprotein (red) were stained with Sybgreen I (green) for observing mtDNA. The insets indicate that the embryo with CISD1-Binding protein expression has less mtDNA than in the control embryo. Bar = 40 μm. Three biologically repeated experiments with similar results.
Supplementary information
Supplementary Information
Supplementary figures and tables, and informed-consent templates.
Source data
Source Data for Fig. 2
Source data and statistics.
Source Data for Fig. 3
Source data and statistics.
Source Data for Fig. 4
Source data and statistics.
Source Data for Fig. 5
Source data and statistics.
Source Data for Fig. 6
Unprocessed gels.
Source Data for ED Fig. 1
Source data and statistics.
Source Data for ED Fig. 2
Source data and statistics.
Source Data for ED Fig. 6
Source data and statistics.
Rights and permissions
About this article
Cite this article
Fan, XY., Guo, L., Chen, LN. et al. Reduction of mtDNA heteroplasmy in mitochondrial replacement therapy by inducing forced mitophagy. Nat. Biomed. Eng 6, 339–350 (2022). https://doi.org/10.1038/s41551-022-00881-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41551-022-00881-7
This article is cited by
-
Spatiotemporal Distribution and Function of Mitochondria in Oocytes
Reproductive Sciences (2024)
-
Propionic acid induces alterations in mitochondrial morphology and dynamics in SH-SY5Y cells
Scientific Reports (2023)
-
Polyamine metabolite spermidine rejuvenates oocyte quality by enhancing mitophagy during female reproductive aging
Nature Aging (2023)
-
Operation “mitochondrial wipeout” — clearing recipient mitochondria DNA during the cytoplasmic replacement therapy
Journal of Assisted Reproduction and Genetics (2022)