Abstract
The genetic loci implicated in familial Parkinson’s disease (PD) have limited generalizability to the Indian PD population. We tested mutations and the frequency of known mutations in the SNCA gene in a PD cohort from India. We selected 298 PD cases and 301 age-matched controls for targeted resequencing (before QC), along with 363 PD genomes of Indian ancestry and 1029 publicly available whole genomes from India as healthy controls (IndiGenomes), to determine the frequency of monogenic SNCA mutations. The raw sequence reads were analyzed using an in-house analysis pipeline, allowing the detection of small variants and structural variants using Manta. The in-depth analysis of the SNCA locus did not identify missense or structural variants, including previously identified SNCA mutations, in the Indian population. The familial forms of SNCA gene variants do not play a major role in the Indian PD population and this warrants further research in the under-represented population.
Similar content being viewed by others
Introduction
The global burden of Parkinson’s disease (PD) was estimated in 2020 to be 9.4 million and about 10% of these patients live in India1,2. However, they were underrepresented in large-scale genetic studies conducted so far3. Most genetic loci implicated in familial PD were identified in the European and East-Asian populations and many of these mutations were seldom seen in Indian PD patients, including the rather frequent G2019S variant in the LRRK2 gene4. The discovery of PD caused by SNCA mutations and the presence of alpha-synuclein within Lewy bodies, the pathological hallmark of PD, triggered great interest in the pathogenic role of the SNCA-encoded protein in PD5. The p.Ala30Pro, p.Glu46Lys, p.His50Gln, p.Gly51Asp, p.Ala53Thr, p.Ala53Glu, p.Ala53Val, and most recently the p.Ala30Gly are the major SNCA mutations discovered so far6,7,8,9,10,11,12,13,14. Duplications and triplications of the SNCA locus were also reported causing PD15,16,17. However, reports of SNCA mutations from the Asian population are exceedingly rare18. In the present study, we re-sequenced the entire SNCA locus and retrieved locus-specific genetic information from an ongoing Indian PD genome sequencing study to look for known, as well as unknown mutations and/or structural variants, in a large cohort of Indian PD patients and healthy controls.
Results
Targeted resequencing and whole genome sequencing
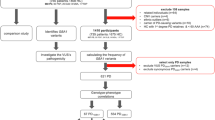
For the SNCA locus targeted resequencing cohort, the mean sequencing depth was 244x. On average, 97.85% of the 151-kbp target region was covered with at least 20× depth (Fig. 1a). For the whole-genome cohort, the mean sequencing depth was 29.6×. On average, 94.70% genome was covered with at least 20× depth (Fig. 1b). Of the 599 samples (298 cases and 301 controls), ten samples were of low quality (average target region read depth of less than 50× or more than 93% of the target region covered by less than 20×). Further, three samples that were duplicated, were excluded from further analysis. Thus, a total of 288 PD cases and 298 controls were available for the variant analysis from the targeted resequencing cohort, while 363 genomes were used for whole-genome sequencing. Taken together, in a combined total of 651 cases and 1327 controls from India, we did not identify previously described mutations and/or missense and/or structural variants of the SNCA gene in the Indian cohort.
Discussion
The Indian population because of its unique genetic makeup (due to widespread founder events) led to the accumulation of population-specific genetic variants19. Thus, the cataloging of disease-specific variants will pave the way for precision medicine in India. This is the largest study in which an in-depth analysis of the SNCA locus was performed by employing targeted resequencing as well as retrieving WGS data from our ongoing study and population-specific control genomes from India to detect unknown variants, including structural variants and assessing the prevalence of the previously identified all major SNCA mutations in the Indian population. The comprehensive analysis excluded the role of the rare forms of the previously identified major monogenic SNCA-dependent PD mutations in the Indian population (Table 1).
The first pathogenic mutation in the SNCA gene (p.Ala53Thr) was reported in 1997 in an Italian kindred and three unrelated families of Greek origin6. Later, mutations derived from a common founder were described in Finnish (p.Ala53Glu) and 3 Greek families (p.Ala30Gly)9,10. Further, duplications and triplications of the SNCA locus which correlated with disease severity, were reported to cause familial parkinsonism20,21,22. After the discovery of the p.Ala53Thr mutation, p.Ala30Pro and p.Glu46Lys mutations were identified as single families, of German and Spanish origin, respectively7,8. Another mutation, p.Thr72Met, was recently found in four members of two Turkish families23. Finally, the p.His50Gln mutation was identified in a Canadian patient of English–Welsh ancestry with PD and a positive family history of parkinsonism and dementia, and in a single English patient with sporadic, pathologically-confirmed PD11,12.
There are scarce reports of SNCA mutations from Asian populations. A novel autosomal dominant inherited p.Met5Thr mutation was found in a recent Chinese study of 155 PD patients18. Though single nucleotide polymorphisms in the SNCA gene are known to increase the risk of sporadic PD, none of the initially identified pathogenic substitutions were found to be involved24. A novel variant, p.Val15Asp, and another unclear variant, p.Met127Ile, in the SNCA gene were found in a Chinese study of 191 sporadic PD and 200 controls25. Further, two likely pathogenic mutations, p.Ala53Val, and p.Pro117Ser were found among 3 Chinese patients in a study of 433 sporadic PD cases and 543 age-matched controls15. Two novel substitutions, p.Ala18Thr, and p.Ala29Ser, were each found in a single patient with sporadic late-onset PD in a Polish study of 629 PD patients26.
Three previous studies from India failed to detect known mutations in the SNCA gene in their PD cases. These include a study with 140 PD patients and 201 normal controls that were tested for the p.Ala53Thr, p.Ala30Pro, p.Glu46Lys mutations27, and another for the p.Gly88Cys or p.Gly209Ala mutations in 169 patients, respectively28. A third study which re-sequenced 6 exons of the SNCA gene in 100 PD patients and ethnically-matched controls, also revealed no mutations in the gene in this population29.
Based on these reports from both south and north-Indian PD cases which tested known mutations, and the resequencing and WGS data from the entire SNCA locus in the South Indian and pan-Indian PD cases respectively in the current study, it is evident that known SNCA mutations have no major role in PD in the Indian PD population. In contrast to the previously published studies from India, we re-sequenced the complete locus to identify any potential structural variants in our cohort that could have been missed in the previously published studies. The present study screened a larger cohort to assess the role of SNCA-monogenic mutations in PD patients in the Indian population. Currently, a pan-Indian study to decipher the role of common genetic variability of PD, including the role of SNCA variants, is underway in the Indian population30.
The current information on the genetic susceptibility to PD worldwide relies mostly on data from European, North- American, and East- Asian populations. To our knowledge, this is the largest study analyzing the entire coding region of the SNCA gene in a cohort of Indian patients with PD which revealed an important finding that the rare variants in the SNCA gene responsible for monogenic-PD in other populations cannot be implicated in the Indian PD population.
Methods
Targeted resequencing cohort
A total of 298 cases and 301 age-matched healthy controls of Indian ancestry were included in the study. All PD cases were diagnosed at the Movement Disorder clinic of a tertiary-care university hospital (Sree Chitra Tirunal Institute for Medical Sciences and Technology-SCTIMST) in Kerala, South India. All patients were diagnosed by a movement disorder specialist using the United Kingdom Parkinson’s Disease Brain bank criteria31. Data related to both sporadic and familial PD (n = 25) were compiled for research purposes in the data bank of the Movement Disorder clinic. The mean age at onset was 49.9 years (range, 24–80) Ethnically- matched healthy controls, unrelated to patients, were also regularly recruited to build a comprehensive control group for the study. Before inclusion in the study, the controls were examined for any neurological disorders and queried for any family history of neurodegenerative disorders.
Whole-genome sequencing cohort
From our ongoing whole genome sequencing study, a total of 363 samples were used for analysis. The mean age of onset was 52.15 ± 10.35, and the male-to-female ratio was 2.1:1. All cases were diagnosed by Movement disorder specialists using the same criteria. The institutional ethics committees of all centers approved the study. All participants signed informed consent.
Targeted resequencing of SNCA
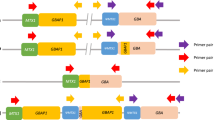
5 ml of blood was collected from each volunteer using venipuncture, and genomic DNA was extracted using the salting out method for targeted resequencing as well as for whole-genome analysis32. Resequencing was performed at the Core facility of Applied Transcriptomics and Genomics at the Institute of Medical Genetics and Applied Genomics, University Hospital of Tübingen, (Tübingen, Germany). A total of 298 cases and 301 controls were selected for resequencing. The SNCA locus resequencing was performed by using several long-range PCRs to amplify the 151 kb SNCA locus. The PCR amplicons were turned into a sequencing library using the “Nextera XT DNA Library Preparation” kit (Illumina, San Diego, CA, USA). Sequencing of the libraries was performed on the NextSeq500 sequencer (Illumina, San Diego, CA, USA) using 75 bp paired-end sequencing. Generated sequences were processed using the open-source pipeline megSAP (https://github.com/imgag/megSAP/tree/2022_08) based on the GRCh38 reference genome.
Whole-genome sequencing
For DNA sequencing, 350 ng of genomic DNA was fragmented to ~450 bp pairs using the DNA PCR-Free Prep, Tagmentation (Illumina). The resulting libraries typically present a concentration of 1.5–3 ng/µl and are sequenced as paired-end 150 bp reads on an Illumina NovaSeq6000 (Illumina) with a sequencing depth of approximately 120 Gb. Generated sequences were processed using the open-source pipeline megSAP (https://github.com/imgag/megSAP/tree/2022_08) based on the GRCh38 reference genome. SNCA locus spanning 151 kb region was selected for screening PD genomes.
IndiGenomes
A publicly available database was used to search for putative variants in the control genomes ascertained from different regions of India. In brief, a total of 1029 self-declared healthy individuals underwent whole-genome sequencing to develop a comprehensive compendium of genetic variants in the Indian population. For details, please see ref. 19.
Genomic analysis
For targeted resequencing and SNCA locus data from whole genomes; the megSAP pipeline was used (https://github.com/imgag/megSAP). In brief, megSAP performs quality control, read mapping, variant detection, as well as comprehensive annotation of variants. Detailed information about tools used by megSAP and tool versions can be found in the megSAP documentation. For the main analysis steps the following tools were used: BWA-mem2 for read mapping (https://github.com/bwa-mem2/bwa-mem2), freebayes for small variant calling (https://github.com/freebayes/freebayes), and Manta for structural variant calling (https://github.com/Illumina/manta). For previously described mutations, we directly search for known mutations in our cohort. To find unknown PD variants, we filtered the detected variants using two main criteria: (1) The variant must be a protein-altering splice region, and (2) the variant should have a maximum allele frequency of 0.01% in gnomAD, including subpopulations.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The data used in this study is under restriction by the ethical committees, and thus cannot be made publicly available. However, the researcher can send a request to the corresponding author for data access. Based on the feasibility of the project, the corresponding author will provide guidelines to the researchers regarding data access to our cloud portal to perform and conduct their study for which any investigator will request our data.
Code availability
The publicly available analytical tools were used for the analysis. megSAP pipeline (https://github.com/imgag/megSAP), BWA-mem2 (https://github.com/bwa-mem2/bwa-mem2), freebayes (https://github.com/freebayes/freebayes), Manta (https://github.com/Illumina/manta).
References
GBD 2016 Parkinson’s Disease Collaborators. Global, regional, and national burden of Parkinson’s disease, 1990–2016: a systematic analysis for the Global Burden of Disease Study 2016. Lancet Neurol.17, 939–953 https://doi.org/10.1016/S1474-4422(18)30295-3 (2018).
Maserejian, N., Vinikoor-Imler, L., Dilley, A. Estimation of the 2020 global population of Parkinson’s disease (PD). Mov. Disord. 35, S79–S80 https://www.mdabstracts.org/abstract/estimation-of-the-2020-global-population-of-parkinsons-disease-pd/ (2020).
International HapMap 3 Consortium, et al. Integrating common and rare genetic variation in diverse human populations. Nature. 467,52–58 https://doi.org/10.1038/nature09298 (2010).
Kishore, A. et al. Understanding the role of genetic variability in LRRK2 in Indian population. Mov. Disord. 34, 496–505 (2019).
Henderson, M. X., Trojanowski, J. Q. & Lee, V. M. Y. α-Synuclein pathology in Parkinson’s disease and related α-synucleinopathies. Neurosci. Lett. 709, 134316 (2019).
Polymeropoulos, M. H. et al. Mutation in the alpha-synuclein gene identified in families with Parkinson’s disease. Science 276, 2045–2047 (1997).
Zarranz, J. J. et al. The new mutation, E46K, of alpha-synuclein causes Parkinson and Lewy body dementia. Ann. Neurol. 55, 164–173 (2004).
Krüger, R. et al. Ala30Pro mutation in the gene encoding alpha-synuclein in Parkinson’s disease. Nat. Genet. 18, 106–108 (1998).
Pasanen, P. et al. SNCA mutation p.Ala53Glu is derived from a common founder in the Finnish population. Neurobiol. Aging 50, 168.e5–168.e8 (2017).
Liu, H. et al. A novel SNCA A30G mutation causes familial Parkinson’s disease. Mov. Disord. 36, 1624–1633 (2021).
Appel-Cresswell, S. et al. Alpha-synuclein p.H50Q, a novel pathogenic mutation for Parkinson’s disease. Mov. Disord. 28, 811–813 (2013).
Proukakis, C. et al. A novel α-synuclein missense mutation in Parkinson disease. Neurology 80, 1062–1064 (2013).
Kiely, A. P. et al. Distinct clinical and neuropathological features of G51D SNCA mutation cases compared with SNCA duplication and H50Q mutation. Mol. Neurodegener. 10, 41 (2015).
Lesage, S. et al. G51D α-synuclein mutation causes a novel parkinsonian-pyramidal syndrome. Ann. Neurol. 73, 459–471 (2013).
Guo, Y. et al. Genetic analysis and literature review of SNCA variants in Parkinson’s disease. Front. Aging Neurosci. 13, 648151 (2021).
Ibáñez, P. et al. Causal relation between alpha-synuclein gene duplication and familial Parkinson’s disease. Lancet 364, 1169–1171 (2004).
Singleton, A. B. et al. alpha-Synuclein locus triplication causes Parkinson’s disease. Science 302, 841 (2003).
Hua, P. et al. Genetic analysis of patients with early-onset Parkinson’s disease in Eastern China. Front. Aging Neurosci. 14, 849462 (2022).
Jain, A. et al. IndiGenomes: a comprehensive resource of genetic variants from over 1000 Indian genomes. Nucleic Acids Res. 49, D1225–D1232 (2021).
Mata, I. F. et al. SNCA variant associated with Parkinson disease and plasma alpha-synuclein level. Arch. Neurol. 67, 1350–1356 (2010).
Miyake, Y. et al. SNCA polymorphisms, smoking, and sporadic Parkinson’s disease in Japanese. Parkinsonism Relat. Disord. 18, 557–561 (2012).
Piper, D. A., Sastre, D., Schüle, B. Advancing stem cell models of alpha-synuclein gene regulation in neurodegenerative disease. Front. Neurosci. 12, 353462 https://www.frontiersin.org/articles/10.3389/fnins.2018.00199 (2018).
Fevga, C. et al. A new alpha-synuclein missense variant (Thr72Met) in two Turkish families with Parkinson’s disease. Parkinsonism Relat. Disord. 89, 63–72 (2021).
Farrer, M. J. Genetics of Parkinson disease: paradigm shifts and future prospects. Nat. Rev. Genet. 7, 306–318 (2006).
Zheng, R. et al. Analysis of rare variants of autosomal-dominant genes in a Chinese population with sporadic Parkinson’s disease. Mol. Genet Genom. Med. 8, e1449 (2020).
Hoffman-Zacharska, D. et al. Novel A18T and pA29S substitutions in α-synuclein may be associated with sporadic Parkinson’s disease. Parkinsonism Relat. Disord. 19, 1057–1060 (2013).
Vishwanathan Padmaja, M., Jayaraman, M., Srinivasan, A. V., Srikumari Srisailapathy, C. R. & Ramesh, A. The SNCA (A53T, A30P, E46K) and LRRK2 (G2019S) mutations are rare cause of Parkinson’s disease in South Indian patients. Parkinsonism Relat. Disord. 18, 801–802 (2012).
Nagar, S. et al. Mutations in the alpha-synuclein gene in Parkinson’s disease among Indians. Acta Neurol. Scand. 103, 120–122 (2001).
Kadakol, G. S., Kulkarni, S. S., Wali, G. M. & Gai, P. B. Molecular analysis of α-synuclein gene in Parkinson’s disease in North Karnataka, India. Neurol. India 62, 149 (2014).
Rajan, R. et al. Genetic architecture of Parkinson’s disease in the Indian population: harnessing genetic diversity to address critical gaps in Parkinson’s disease research. Front. Neurol. 11, 524 (2020).
Clarke, C. E., et al. UK Parkinson’s Disease Society Brain Bank Diagnostic Criteria. (NIHR Journals Library, 2016). https://www.ncbi.nlm.nih.gov/books/NBK379754/
Miller, S. A., Dykes, D. D. & Polesky, H. F. A simple salting out procedure for extracting DNA from human nucleated cells. Nucleic Acids Res. 16, 1215 (1988).
Tambasco, N. et al. A53T in a parkinsonian family: a clinical update of the SNCA phenotypes. J. Neural Transm. 123, 1301–1307 (2016).
Acknowledgements
This study was in part funded by the Michel J Fox Foundation, USA, and the DFG (DFG SH599/16-1). NGS sequencing was performed with the support of the DFG-funded NGS Competence Center Tübingen (INST 37/1049–1). We thank all participants in the study. M.S. is supported by funding from Michael J Fox Foundation (ID;11879, MJFF 009411), GP2 (MJFF: 023430), and DFG (SH-599/16-1). T.G. obtained grants from Novartis Pharma, the Federal Ministry of Education and Research (BMBF), the Helmholtz Association, the European Commission, the German Research Foundation, the Michael J. Fox Foundation.
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
Concept and design: Kishore, Sharma. Acquisition, analysis, or interpretation of data: Kishore, Puthanveedu, Urulangodi, Krishnan, Rajan, Kamble, Debnath, Pal, Yadav, Sturm, Hakkaart, Sharma. Drafting of the manuscript: Kishore, Sharma. Critical revision of the manuscript for important intellectual content: Kishore, Sturm, Pillai, Hakkaart, Puthanveedu, Urulangodi, Krishnan, Ashok Kumar-Sreelatha, Rajan, Kamble, Debnath, Pal, Yadav, Sarma, Casadei, Gasser, Bauer, Riess, Sharma. Statistical analysis: Sturm, Hakkaart. Supervision: Sharma.
Corresponding author
Ethics declarations
Competing interests
Financial disclosures of all authors (for the preceding 12 months). A.K. discloses nothing to report. M.S. discloses nothing to report. K.S.P. discloses nothing to report. C.H. discloses nothing to report. D.K.P. discloses nothing to report. M.U. discloses nothing to report. S.K. discloses nothing to report. A.A.S. discloses nothing to report. R.R. discloses nothing to report. P.K.P. has received grants (funded to his Institute) for research projects as Principal or Co-Investigator from the Indian Council of Medical Research (ICMR), Department of Biotechnology (DBT), Department of Science & Technology (DST)-Science and Engineering Research Board, DST-Cognitive Science Research Initiative, Wellcome Trust UK-India Alliance DBT, PACE scheme of BIRAC, and Michael J. Fox Foundation, and funding (to his Institute) as a Co-Investigator for a clinical trial from ERYDEL SpA, Italy Europe. Also received honoraria as Faculty/Speaker/Author from the International Parkinson and Movement Disorder Society, and Movement Disorder Societies of Korea, Taiwan, and Bangladesh. R.Y. discloses nothing to report. G.S. discloses nothing to report. N.C. discloses nothing to report. T.G. Stock Ownership in medically related fields—NONE—Intellectual Property Rights—patent: KASPP (LRRK2) gene, its production, and its use for the detection and treatment of neurodegenerative disorders. Patent Number: EP1802749 (A2). Consultancies—Royalties for consulting activities from Cefalon Pharma and Merck-Serono—Expert Testimony—NONE. Advisory Boards—Editorial board member of Parkinsonism and Related Disorders, Journal of Parkinson’s Disease, Neurogenetics; Scientific advisory board, Joint Programming in Neurodegenerative Diseases; Advisory board, UKDP—Employment—Center for Neurology and Hertie-Institute for Clinical Brain Research, University of Tübingen and German Center of Neurodegenerative Diseases (DZNE), Tübingen. Partnerships—NONE—Contracts—NONE. Honoraria—Speaker’s honoraria from Novartis, Merck-Serono, Schwarz Pharma, Boehringer Ingelheim and Teva Pharma—Royalties—NONE. Grants—Novartis Pharma, the Federal Ministry of Education and Research (BMBF), the Helmholtz Association, the European Commission, the German Research Foundation, and the Michael J. Fox Foundation. Other—NONE. P.B. discloses nothing to report. O.R. discloses nothing to report. M.S. discloses nothing to report.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kishore, A., Sturm, M., Soman Pillai, K. et al. Resequencing the complete SNCA locus in Indian patients with Parkinson’s disease. npj Parkinsons Dis. 10, 85 (2024). https://doi.org/10.1038/s41531-024-00676-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41531-024-00676-4