Abstract
Phytochrome A (phyA) is the plant far-red (FR) light photoreceptor and plays an essential role in regulating photomorphogenic development in FR-rich conditions, such as canopy shade. It has long been observed that phyA is a phosphoprotein in vivo; however, the protein kinases that could phosphorylate phyA remain largely unknown. Here we show that a small protein kinase family, consisting of four members named PHOTOREGULATORY PROTEIN KINASES (PPKs) (also known as MUT9-LIKE KINASES), directly phosphorylate phyA in vitro and in vivo. In addition, TANDEM ZINC-FINGER/PLUS3 (TZP), a recently characterized phyA-interacting protein required for in vivo phosphorylation of phyA, is also directly phosphorylated by PPKs. We reveal that TZP contains two intrinsically disordered regions in its amino-terminal domain that undergo liquid–liquid phase separation (LLPS) upon light exposure. The LLPS of TZP promotes colocalization and interaction between PPKs and phyA, thus facilitating PPK-mediated phosphorylation of phyA in FR light. Our study identifies PPKs as a class of protein kinases mediating the phosphorylation of phyA and demonstrates that the LLPS of TZP contributes significantly to more production of the phosphorylated phyA form in FR light.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
All data supporting the findings of this study are available in the main text or the supplementary information. The biological materials used in this study are available from J.L. on reasonable request. Source data are provided with this paper.
References
Li, J., Li, G., Wang, H. & Deng, X. W. Phytochrome signaling mechanisms. Arabidopsis Book 9, e0148 (2011).
Legris, M., Ince, Y. C. & Fankhauser, C. Molecular mechanisms underlying phytochrome-controlled morphogenesis in plants. Nat. Commun. 10, 5219 (2019).
Cheng, M. C., Kathare, P. K., Paik, I. & Huq, E. Phytochrome signaling networks. Annu. Rev. Plant Biol. 72, 217–244 (2021).
Jung, J. H. et al. Phytochromes function as thermosensors in Arabidopsis. Science 354, 886–889 (2016).
Legris, M. et al. Phytochrome B integrates light and temperature signals in Arabidopsis. Science 354, 897–900 (2016).
Chen, D. et al. Integration of light and temperature sensing by liquid–liquid phase separation of phytochrome B. Mol. Cell 82, 3015–3029 (2022).
Qi, L. et al. Integration of light and temperature signaling pathways in plants. J. Integr. Plant Biol. 64, 393–411 (2022).
Bae, G. & Choi, G. Decoding of light signals by plant phytochromes and their interacting proteins. Annu. Rev. Plant Biol. 59, 281–311 (2008).
Li, J. & Hiltbrunner, A. Is the Pr form of phytochrome biologically active in the nucleus? Mol. Plant 14, 535–537 (2021).
Sharrock, R. A. & Clack, T. Patterns of expression and normalized levels of the five Arabidopsis phytochromes. Plant Physiol. 130, 442–456 (2002).
Mancinelli, A. L. in Photomorphogenesis in Plants 2nd edn (eds Kendrick, R. E. & Kronenberg, G. H. M.) 211–269 (Kluwer Academic, 1994).
Eichenberga, K. et al. Arabidopsis phytochromes C and E have different spectral characteristics from those of phytochromes A and B. FEBS Lett. 470, 107–112 (2000).
Rausenberger, J. et al. Photoconversion and nuclear trafficking cycles determine phytochrome A’s response profile to far-red light. Cell 146, 813–825 (2011).
Possart, A., Fleck, C. & Hiltbrunner, A. Shedding (far-red) light on phytochrome mechanisms and responses in land plants. Plant Sci. 217–218, 36–46 (2014).
Quail, P. H., Briggs, W. R. & Pratt, L. H. In Vivo Phosphorylation of Phytochrome Carnegie Institution Annual Report, 342–344 (Department of Plant Biology, Carnegie Institution of Washington, 1978).
Hunt, R. E. & Pratt, L. H. Partial characterization of undegraded oat phytochrome. Biochemistry 19, 390–394 (1980).
Liu, X. et al. FERONIA coordinates plant growth and salt tolerance via the phosphorylation of phyB. Nat. Plants 9, 645–660 (2023).
Zhao, Y. et al. Sensory circuitry controls cytosolic calcium mediated phytochrome B phototransduction. Cell 186, 1230–1243 (2023).
Yeh, K. C. & Lagarias, J. C. Eukaryotic phytochromes: light-regulated serine/threonine protein kinases with histidine kinase ancestry. Proc. Natl Acad. Sci. USA 95, 13976–13981 (1998).
Shin, A. Y. et al. Evidence that phytochrome functions as a protein kinase in plant light signalling. Nat. Commun. 7, 11545 (2016).
Hoang, Q. T. N. et al. Protein kinase activity of phytochrome A positively correlates with photoresponses in Arabidopsis. Front. Plant Sci. 12, 706316 (2021).
Wong, Y. S., McMichael, R. W. & Lagarias, J. C. Properties of a polycation stimulated protein kinase associated with purified Avena phytochrome. Plant Physiol. 91, 709–718 (1989).
Ahmad, M., Jarillo, J. A., Smirnova, O. & Cashmore, A. R. The CRY1 blue light photoreceptor of Arabidopsis interacts with phytochrome A in vitro. Mol. Cell 1, 939–948 (1998).
Fankhauser, C. et al. PKS1, a substrate phosphorylated by phytochrome that modulates light signaling in Arabidopsis. Science 284, 1539–1541 (1999).
Colon-Carmona, A., Chen, D. L., Yeh, K. C. & Abel, S. Aux/IAA proteins are phosphorylated by phytochrome in vitro. Plant Physiol. 124, 1728–1738 (2000).
Shen, Y. et al. Phytochrome A mediates rapid red light-induced phosphorylation of Arabidopsis FAR-RED ELONGATED HYPOCOTYL1 in a low fluence response. Plant Cell 21, 494–506 (2009).
Quail, P. H. The phytochromes: a biochemical mechanism of signaling in sight? BioEssays 19, 571–579 (1997).
Quail, P. H. Phytochromes. Curr. Biol. 20, 504–507 (2010).
Ni, W. et al. PPKs mediate direct signal transfer from phytochrome photoreceptors to transcription factor PIF3. Nat. Commun. 8, 15236 (2017).
Saijo, Y. et al. Arabidopsis COP1/SPA1 complex and FHY1/FHY3 associate with distinct phosphorylated forms of phytochrome A in balancing light signaling. Mol. Cell 31, 607–613 (2008).
Zhang, S. et al. TANDEM ZINC-FINGER/PLUS3 is a key component of phytochrome A signaling. Plant Cell 30, 835–852 (2018).
Zhou, Y. et al. Hinge region of Arabidopsis phyA plays an important role in regulating phyA function. Proc. Natl Acad. Sci. USA 115, 11864–11873 (2018).
Wang, Z. et al. Osmotic stress induces phosphorylation of histone H3 at threonine 3 in pericentromeric regions of Arabidopsis thaliana. Proc. Natl Acad. Sci. USA 112, 8487–8492 (2015).
Huang, H. et al. Identification of evening complex associated proteins in Arabidopsis by affinity purification and mass spectrometry. Mol. Cell Proteom. 15, 201–217 (2016).
Liu, Q. et al. Molecular basis for blue light-dependent phosphorylation of Arabidopsis cryptochrome 2. Nat. Commun. 8, 15234 (2017).
Liu, S. et al. Differential photoregulation of the nuclear and cytoplasmic CRY1 in Arabidopsis. N. Phytol. 234, 1332–1346 (2022).
Shaw, P. J. & Brown, J. W. S. Plant nuclear bodies. Curr. Opin. Plant Biol. 7, 614–620 (2004).
Spector, D. L. SnapShot: cellular bodies. Cell 127, 1071 (2006).
Van Buskirk, E. K., Decker, P. V. & Chen, M. Photobodies in light signaling. Plant Physiol. 158, 52–60 (2012).
Pardi, S. A. & Nusinow, D. A. Out of the dark and into the light: a new view of phytochrome photobodies. Front. Plant Sci. 12, 732947 (2021).
Han, Y. J. et al. Functional characterization of phytochrome autophosphorylation in plant light signaling. Plant Cell Physiol. 51, 596–609 (2010).
Trupkin, S. A., Debrieux, D., Hiltbrunner, A., Fankhauser, C. & Casal, J. J. The serine-rich N-terminal region of Arabidopsis phytochrome A is required for protein stability. Plant Mol. Biol. 63, 669–678 (2007).
Kim, J. I. et al. Phytochrome phosphorylation modulates light signaling by influencing the protein–protein interaction. Plant Cell 16, 2629–2640 (2004).
Jordan, E. T., Cherry, J. R., Walker, J. M. & Vierstra, R. D. The amino-terminus of phytochrome A contains two distinct functional domains. Plant J. 9, 243–257 (1996).
Jordan, E. T., Marita, J. M., Clough, R. C. & Vierstra, R. D. Characterization of regions within the N-terminal 6-kilodalton domain of phytochrome A that modulate its biological activity. Plant Physiol. 115, 693–704 (1997).
Medzihradszky, M. et al. Phosphorylation of phytochrome B inhibits light-induced signaling via accelerated dark reversion in Arabidopsis. Plant Cell 25, 535–544 (2013).
Nito, K., Wong, C. C., Yates, J. R. 3rd & Chory, J. Tyrosine phosphorylation regulates the activity of phytochrome photoreceptors. Cell Rep. 3, 1970–1979 (2013).
Viczian, A. et al. Differential phosphorylation of the N-terminal extension regulates phytochrome B signaling. N. Phytol. 225, 1635–1650 (2020).
Su, Y. S. & Lagarias, J. C. Light-independent phytochrome signaling mediated by dominant GAF domain tyrosine mutants of Arabidopsis phytochromes in transgenic plants. Plant Cell 19, 2124–2139 (2007).
Kaiserli, E. et al. Integration of light and photoperiodic signaling in transcriptional nuclear foci. Dev. Cell 35, 311–321 (2015).
Linding, R. et al. Protein disorder prediction: implications for structural proteomics. Structure 11, 1453–1459 (2003).
Kato, M. et al. Cell-free formation of RNA granules: low complexity sequence domains form dynamic fibers within hydrogels. Cell 149, 753–767 (2012).
Molliex, A. et al. Phase separation by low complexity domains promotes stress granule assembly and drives pathological fibrillization. Cell 163, 123–133 (2015).
Patel, A. et al. A liquid-to-solid phase transition of the ALS protein FUS accelerated by disease mutation. Cell 162, 1066–1077 (2015).
Riback, J. A. et al. Stress-triggered phase separation is an adaptive, evolutionarily tuned response. Cell 168, 1028–1040 (2017).
Fang, X. et al. Arabidopsis FLL2 promotes liquid–liquid phase separation of polyadenylation complexes. Nature 569, 265–269 (2019).
Xu, X. et al. Phase separation in plants: new insights into cellular compartmentalization. J. Integr. Plant Biol. 63, 1835–1855 (2021).
Kroschwald, S., Maharana, S. & Simon, A. Hexanediol: a chemical probe to investigate the material properties of membrane-less compartments. Matters 3, e201702000010 (2017).
Xie, D. et al. Phase separation of SERRATE drives dicing body assembly and promotes miRNA processing in Arabidopsis. Nat. Cell Biol. 23, 32–39 (2020).
Wang, J. et al. A molecular grammar governing the driving forces for phase separation of prion-like RNA binding proteins. Cell 174, 688–699.e16 (2018).
Hsieh, H. L. et al. FIN219, an auxin-regulated gene, defines a link between phytochrome A and the downstream regulator COP1 in light control of Arabidopsis development. Genes Dev. 14, 1958–1970 (2000).
Osterlund, M. T., Hardtke, C. S., Wei, N. & Deng, X. W. Targeted destabilization of HY5 during light-regulated development of Arabidopsis. Nature 405, 462–466 (2000).
Li, C. et al. Mutual upregulation of HY5 and TZP in mediating phytochrome A signaling. Plant Cell 34, 633–654 (2022).
Gao, L. et al. Blue light-induced phosphorylation of Arabidopsis cryptochrome 1 is essential for its photosensitivity. J. Integr. Plant Biol. 64, 1724–1738 (2022).
Yang, C. et al. Phytochrome A negatively regulates the shade avoidance response by increasing auxin/indole acidic acid protein stability. Dev. Cell 44, 29–41 (2018).
Han, X. et al. Origin and evolution of core components responsible for monitoring light environment changes during plant terrestrialization. Mol. Plant 12, 847–862 (2019).
Mathews, S. Phytochrome evolution in green and nongreen plants. J. Hered. 96, 197–204 (2005).
Chen, M. et al. Arabidopsis HEMERA/pTAC12 initiates photomorphogenesis by phytochromes. Cell 141, 1230–1240 (2010).
Wang, X. et al. A photoregulatory mechanism of the circadian clock in Arabidopsis. Nat. Plants 7, 1397–1408 (2021).
Jiang, B. et al. Light-induced LLPS of the CRY2/SPA1/FIO1 complex regulating mRNA methylation and chlorophyll homeostasis in Arabidopsis. Nat. Plants 9, 2042–2058 (2023).
Reed, J. W., Nagatani, A., Elich, T. D., Fagan, M. & Chory, J. Phytochrome A and phytochrome B have overlapping but distinct functions in Arabidopsis development. Plant Physiol. 104, 1139–1149 (1994).
Oyama, T., Shimura, Y. & Okada, K. The Arabidopsis HY5 gene encodes a bZIP protein that regulates stimulus-induced development of root and hypocotyl. Genes Dev. 11, 2983–2995 (1997).
Li, H., Qin, X., Song, P., Han, R. & Li, J. A LexA-based yeast two-hybrid system for studying light-switchable interactions of phytochromes with their interacting partners. aBIOTECH 2, 105–116 (2021).
Yan, Y. et al. MYB30 is a key negative regulator of Arabidopsis photomorphogenic development that promotes PIF4 and PIF5 protein accumulation in the light. Plant Cell 32, 2196–2215 (2020).
Dong, X. et al. The cold response regulator CBF1 promotes Arabidopsis hypocotyl growth at ambient temperatures. EMBO J. 39, e103630 (2020).
Wang, X. et al. PUB25 and PUB26 promote plant freezing tolerance by degrading the cold signaling negative regulator MYB15. Dev. Cell 51, 222–235 (2019).
Ma, L. et al. Phytochromes enhance SOS2-mediated PIF1 and PIF3 phosphorylation and degradation to promote Arabidopsis salt tolerance. Plant Cell 35, 2997–3020 (2023).
Song, P. et al. 14-3-3 proteins regulate photomorphogenesis by facilitating light-induced degradation of PIF3. N. Phytol. 237, 140–159 (2023).
Qi, L. et al. PHYTOCHROME-INTERACTING FACTORS interact with the ABA receptors PYL8 and PYL9 to orchestrate ABA signaling in darkness. Mol. Plant 13, 414–430 (2020).
Chen, H. et al. Firefly luciferase complementation imaging assay for protein–protein interactions in plants. Plant Physiol. 146, 368–376 (2008).
Waadt, R. et al. Multicolor bimolecular fluorescence complementation reveals simultaneous formation of alternative CBL/CIPK complexes in planta. Plant J. 56, 505–516 (2008).
Liu, Z. et al. Plasma membrane CRPK1-mediated phosphorylation of 14-3-3 proteins induces their nuclear import to fine-tune CBF signaling during cold response. Mol. Cell 66, 117–128 (2017).
Clough, S. J. & Bent, A. F. Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 16, 735–743 (1998).
Navadgi-Patil, V. M. & Burgers, P. M. Yeast DNA replication protein Dpb11 activates the Mec1/ATR checkpoint kinase. J. Biol. Chem. 283, 35853–35859 (2008).
Ding, Y. et al. EGR2 phosphatase regulates OST1 kinase activity and freezing tolerance in Arabidopsis. EMBO J. 38, e99819 (2019).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Peng, J. et al. COP1 positively regulates ABA signaling during Arabidopsis seedling growth in darkness by mediating ABA-induced ABI5 accumulation. Plant Cell 34, 2286–2308 (2022).
Qiu, Y. et al. Mechanism of early light signaling by the carboxy-terminal output module of Arabidopsis phytochrome B. Nat. Commun. 8, 1905 (2017).
Martin, K. et al. Transient expression in Nicotiana benthamiana fluorescent marker lines provides enhanced definition of protein localization, movement and interactions in planta. Plant J. 59, 150–162 (2009).
Acknowledgements
We thank Q. Wang for the ppk123 and ppk124 mutants and the ACT2p::GFP–PPK1 rdr6-11 transgenic line. This work was supported by grants from the National Natural Science Foundation of China (no. 32225006 to J.L. and no. 32200245 to L.Q.) and the Beijing Natural Science Foundation (no. 5232011 to J.L.).
Author information
Authors and Affiliations
Contributions
J.L., Z.F. and M.W. designed the research. Z.F., M.W., Yan Liu, C.L., S.Z., J.D., J.C., L.Q., Yanru Liu, H.L., J.W. and Yannan Liu performed the research. J.L., Z.F., M.W., F.T., B.Z., X.F., W.Q., Y.G. and X.W.D. discussed and interpreted the data. J.L., Z.F., M.W. and W.T. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Plants thanks Enamul Huq, Jeong-Il Kim and Cornelia Klose for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 PPKs interact with TZP in vitro and in vivo.
a, Schematic diagram of prey proteins (TZP, TZP-N, TZP-N1, TZP-C, TZP-C1 and TZP-C2 fused with AD domains). ZF, zinc finger. b, Yeast two-hybrid assays showing that PPKs interact with the C-terminal domain of TZP. c–f, Pull-down assays showing that both the zinc-finger and PLUS3 domains are required for interactions between TZP and PPK1 (c), PPK2 (d), PPK3 (e) and PPK4 (f) in vitro. g, BiFC assays showing that TZP and PPK1 interact in the nuclei of N. benthamiana plants expressing red fluorescent protein (RFP)-H2B fusion proteins89. An unrelated protein (GUS) fused with the C-terminal domain of YFP77 was used as the negative control. Super:CFP was used as the transformation control. Scale bar = 20 μm (the whole image) and 10 μm (the enlarged nuclei). h,i, LCI assays showing that TZP interacts with all four PPKs in N. benthamiana leaf cells. In (h), scale bars = 1 cm. In (i), the values are mean ± s.d. (n = 3 independent assays). P values are from two-tailed Student’s t-test. j, Co-IP assays showing the associations of TZP with phyA and PPK1 in vivo. TZP-GFP, phyA-FLAG and MYC-PPK1 were coexpressed in Col protoplasts, then total proteins were extracted and incubated with anti-GFP beads (AlpaLife). The total and precipitated proteins were subjected to immunoblotting with antibodies against TZP, phyA, MYC, HSP and GFP, respectively. The asterisk and arrowhead represent the phosphorylated and unphosphorylated phyA forms, respectively.
Extended Data Fig. 2 PPK2 phosphorylates apo- and holo-phyA in vitro.
In vitro kinase assays showing that PPK2-catalyzed phosphorylation of holo- and apo-phyA with different concentrations of phyA (a,c) or for different times (b,d). For each panel, CBB staining (top), and autoradiographs (bottom) are shown, with zinc fluorescence (Zinc) shown in the middle of a and b. PhyA-GST proteins were expressed in and purified from Saccharomyces cerevisiae BJ2168, and phycocyanobilin (PCB) extracted from Spirulina was added to the supernatant at a final concentration of 10 µM in the dark to allow the assembly of holo-phyA proteins in vitro. The holo-phyA proteins were exposed to 15 min of FR light alone, or 15 min of FR light immediately followed by 15 min of R light to allow holo-phyA to form Pr or Pfr forms in vitro, respectively.
Extended Data Fig. 3 PPKs phosphorylate TZP in vitro and in vivo.
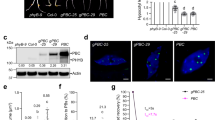
a, Identical amounts of Super:TZP-GFP and 35 S:MYC-PPK1 plasmids were transiently cotransfected into Col protoplasts. After transfection, the protoplasts were incubated in the dark for 16 h, and then treated with FR light (80 μmol m−2 s−1) for 2 h. The total proteins were extracted and treated with active (+) or inactive (boiled) or without (−) λ-PPase for 30 min, and then analyzed by immunoblots using anti-TZP, anti-MYC or anti-HSP antibodies, respectively. b, In vitro kinase assays showing that PPKs directly phosphorylate the full-length TZP proteins. CBB staining (top) and autoradiographs (bottom) are shown. c, Immunoblot assays showing the patterns of TZP proteins in Col and ppk mutant seedlings grown under FR light. Numbers below the immunoblots indicate the relative band intensities of TZP proteins normalized to those of HSP, respectively. The ratio was set to 100 for the first TZP band.
Extended Data Fig. 4 Yeast two-hybrid assays showing that the T418D and C323A mutations abolished phyA interaction with TZP.
a, In vivo plate assays showing that TZP interacts with phyA(WT) and phyA(D422R), but not with phyA(T418D) in yeast cells. b, Liquid culture assays showing the interactions between TZP and phyA(WT), phyA(D422R), phyA(T418D), phyA(Y242H) and phyA(C323A) in yeast cells. Yeast cultures incubated in darkness (D) were treated with 5 min of R light alone, or 5 min of R light immediately followed by 5 min of FR light, and then incubated in D for additional 3 h. β-Galactosidase activities were measured using ONPG as the substrate (the values are mean ± s.d., n = 4 independent assays). Different letters represent significant differences indicated by one-way ANOVA with Bonferroni’s multiple comparisons test (P < 0.05). The exact P values are provided in Supplementary Table 2.
Extended Data Fig. 5 TZP promotes PPK1-mediated phyA phosphorylation in FR light.
Immunoblots showing that TZP promotion of PPK1-mediated phyA phosphorylation in Col protoplasts became more evident after longer FR light exposure (a,c,e) and with higher intensities of FR light (b,d,f). Representative pictures are shown in (a) and (b), the percentages of phosphorylated phyA are shown in (c) and (d), and the total amounts of phyA are shown in (e) and (f). In (c-f), the values are mean ± s.d. (n = 3 independent assays). P values are from two-tailed Student’s t-test. In (e) and (f), the total amounts of phyA were not significantly changed under different times of FR light or under different intensities of FR light treatments.
Extended Data Fig. 6 FRAP of TZP NBs in N. benthamiana leaf cells and in vitro.
a, FRAP assays of the whole TZP-GFP NBs in N. benthamiana leaf cells. Time 0 indicates the time of the photobleaching pulse. Scale bar = 4 μm. b, Plot showing the time course of the recovery after photobleaching the whole TZP nuclear condensates. Data are representative of ten independent experiments. Data are presented as mean ± SD (n = 10 independent experiments). c, FRAP assays of the half of TZP nuclear condensates in N. benthamiana leaf cells. The dashed lines demarcate the area of bleaching. Time 0 indicates the time of the photobleaching pulse. Scale bar = 5 μm. d, Plot showing the time course of the recovery after photobleaching the half of TZP nuclear condensates. Data are presented as the mean ± SD (n = 10 independent experiments). e, FRAP assays of the whole TZP-IDR-GFP droplets. Time 0 s indicates the time of the photobleaching pulse. Scale bar = 3 μm. f, Plot showing the time course of the recovery after photobleaching the whole TZP-IDR-GFP droplets. Data are representative of ten independent experiments. Data are presented as the mean ± SD (n = 10 independent experiments). g, FRAP assays of the half of TZP-IDR-GFP droplets. The dashed lines demarcate the area of bleaching. Time 0 s indicates the time of the photobleaching pulse. Scale bar = 5 μm. h, Plot showing the time course of the recovery after photobleaching the half of TZP-IDR-GFP droplets. Data are representative of seven independent experiments. Data are presented as the mean ± SD (n = 7 independent experiments). i, FRAP assays of FUS-IDR-NLS-TZPC-GFP NBs in N. benthamiana leaf cells. Time 0 indicates the time of the photobleaching pulse. Scale bar = 4 μm. j, Plot showing the time course of the recovery after photobleaching FUS-IDR-NLS-TZPC-GFP nuclear condensates. Data are representative of ten independent experiments. Data are presented as mean ± SD (n = 10 independent experiments).
Extended Data Fig. 7 TZP NBs are reversible by light-to-dark transition in Arabidopsis.
a,b, Partition coefficients and numbers of TZP-GFP NBs per nucleus in response to FR light (a) or dark treatment (b) shown in Fig. 4g. Data are presented as means ± SD (n = 10 nuclei in [a] and n = 8 nuclei in [b]). c, Time-lapse images showing the behaviors of TZP NBs in one nucleus of TZPp:TZP-GFP tzp-2 seedlings in response to white light or dark treatment. White slash indicates the treatment of white light. Data are representative of ten independent experiments. Scale bar = 15 μm.
Extended Data Fig. 8 TZP phase separates into droplets in vitro.
a, Coomassie staining of MBP-His-TZP-IDR-GFP protein samples before and after TEV cleavage to remove the MBP tag. The TEV protease-treated solution was then flowed through Superdex 200 increase 10/300 column (SD200) (GE healthcare), and then the fraction containing the main peak of TZP-IDR-GFP was collected for in vitro phase separation assays shown in (b) and (c). b, In vitro phase separation assay of TZP-IDR-GFP proteins (10 μM) in the presence of 100 mM NaCl and PEG8000 (n = 3 independent assays, scale bar = 10 μm). c, Images showing formation of droplets at different concentrations of TZP-IDR-GFP and NaCl in the presence of PEG8000. Data are representative of three independent experiments. Scale bar = 5 μm.
Extended Data Fig. 9 Representative images for the subcellular localizations of FUS-IDR-NLS-TZPC-GFP, phyA-CFP and RFP-PPK1.
a,b, Confocal microscopy assays showing the subcellular localizations of GFP-, CFP- and RFP-tagged proteins coexpressed in Col protoplasts (a) and N. benthamiana leaf cells (b). The representative confocal images for each channel were subsequently merged to show colocalization. The arrow heads indicate the locations of nuclear bodies. Scale bars = 10 μm. c,d, Fluorescence profiles of FUS-IDR-NLS-TZPC-GFP/phyA-CFP/RFP-PPK1, phyA-CFP/RFP-PPK1, FUS-IDR-NLS-TZPC-GFP/phyA-CFP, FUS-IDR-NLS-TZPC-GFP/RFP-PPK1 and FUS-IDR-NLS-TZPC-GFP in Col protoplasts (c) and N. benthamiana leaf cells (d) over the white lines shown in (a) and (b), respectively. The arrow heads indicate the locations of nuclear bodies.
Extended Data Fig. 10 Evolutionary analyses of phytochromes, PPKs and TZP.
a, Evolution of the core light signaling components of Archaeplastida. Cryptochrome (CRY) was the first type of photoreceptor to originate in red algae66. PPKs first originated in the chlorophytes, and PPK1/4 and PPK2/3 separated from PPKs in lycophytes and ferns, respectively. Both phytochrome and TZP originated in the charophytes, while Arabidopsis phyA and phyB separated after the divergence of seed plants66. Figure created with BioRender.com. b, Alignment of the PLUS3 domain in multiple species from charophytes to angiosperms. The sequences were aligned using ClustalX and modified using Adobe Illustrator.
Supplementary information
Supplementary Information
Supplementary Figs. 1–12, Methods and uncropped scans of blots and gels for the supplementary figures.
Supplementary Table 1
Primers.
Supplementary Table 2
Exact P values.
Supplementary Data 1
Statistical source data for the supplementary figures.
Source data
Source Data Fig. 1
Unprocessed western blots.
Source Data Fig. 2
Unprocessed western blots.
Source Data Fig. 3
Unprocessed western blots.
Source Data Fig. 6
Unprocessed western blots.
Source Data Extended Data Fig. 1
Unprocessed western blots.
Source Data Extended Data Fig. 3
Unprocessed western blots.
Source Data Extended Data Fig. 5
Unprocessed western blots.
Source Data Figs. 1–6 and Extended Data Figs. 1, 4–7 and 9
Summary of all statistical source data.
Source Data Figs. 1 and 3–5 and Extended Data Figs. 1 and 6–9
Summary of all confocal microscopy pictures.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Feng, Z., Wang, M., Liu, Y. et al. Liquid–liquid phase separation of TZP promotes PPK-mediated phosphorylation of the phytochrome A photoreceptor. Nat. Plants 10, 798–814 (2024). https://doi.org/10.1038/s41477-024-01679-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-024-01679-y