Abstract
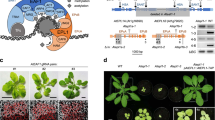
Chloroplast biogenesis is crucial in plant development, as it is essential for the transition to autotrophic growth. This process is light-induced and relies on the orchestrated transcription of nuclear and plastid genes, enabling the effective assembly and regulation of the photosynthetic machinery. Here we reveal a new regulation level for this process by showing the involvement of chromatin remodelling in the nuclear control of plastid gene expression for proper chloroplast biogenesis and function. The two Arabidopsis homologues of yeast EPL1 protein, components of the NuA4 histone acetyltransferase complex, are essential for plastid transcription and correct chloroplast development and performance. We show that EPL1 proteins are light-regulated and necessary for concerted expression of nuclear genes encoding most components of chloroplast transcriptional machinery, directly mediating H4K5ac deposition at these loci and promoting the expression of plastid genes required for chloroplast biogenesis. These data unveil a NuA4-mediated mechanism regulating chloroplast biogenesis that links the transcription of nuclear and plastid genomes during chloroplast development.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The materials generated in this work are available upon request to the corresponding authors. Sequence data related to this article can be found in the Arabidopsis information portal (https://www.araport.org/) under the accession numbers EPL1A (At1g16690), EPL1B (At1g79020), HAM1 (At5g64610), ING2 (At1g54390) and EAF6 (At4g14385). The complete genome-wide data from this publication are already deposited in the Gene Expression Omnibus database (www.ncbi.nlm.nih.gov/geo/) under accession number GSE180614. Source data are provided with this paper.
References
Arsovski, A. A., Galstyan, A., Guseman, J. M. & Nemhauser, J. L. Photomorphogenesis. Arabidopsis Book 10, e0147 (2012).
Hernandez-Verdeja, T., Vuorijoki, L. & Strand, A. Emerging from the darkness: interplay between light and plastid signaling during chloroplast biogenesis. Physiol. Plant. 169, 397–406 (2020).
Jarvis, P. & Lopez-Juez, E. Biogenesis and homeostasis of chloroplasts and other plastids. Nat. Rev. Mol. Cell Biol. 14, 787–802 (2013).
Woodson, J. D. & Chory, J. Coordination of gene expression between organellar and nuclear genomes. Nat. Rev. Genet. 9, 383–395 (2008).
Green, B. R. Chloroplast genomes of photosynthetic eukaryotes. Plant J. 66, 34–44 (2011).
Pfannschmidt, T. et al. Plastid RNA polymerases: orchestration of enzymes with different evolutionary origins controls chloroplast biogenesis during the plant life cycle. J. Exp. Bot. 66, 6957–6973 (2015).
Liebers, M. et al. Regulatory shifts in plastid transcription play a key role in morphological conversions of plastids during plant development. Front Plant Sci. 8, 23 (2017).
Deng, X. W. & Gruissem, W. Control of plastid gene expression during development: the limited role of transcriptional regulation. Cell 49, 379–387 (1987).
Baba, K. et al. Organellar gene transcription and early seedling development are affected in the rpoT;2 mutant of Arabidopsis. Plant J. 38, 38–48 (2004).
Liere, K., Weihe, A. & Borner, T. The transcription machineries of plant mitochondria and chloroplasts: composition, function, and regulation. J. Plant Physiol. 168, 1345–1360 (2011).
Xiao, J., Jin, R. & Wagner, D. Developmental transitions: integrating environmental cues with hormonal signaling in the chromatin landscape in plants. Genome Biol. 18, 88 (2017).
Servet, C., Conde E Silva, N. & Zhou, D. X. Histone acetyltransferase AtGCN5/HAG1 is a versatile regulator of developmental and inducible gene expression in Arabidopsis. Mol. Plant 3, 670–677 (2010).
Zhang, D., Li, Y., Zhang, X., Zha, P. & Lin, R. The SWI2/SNF2 chromatin-remodeling ATPase BRAHMA regulates chlorophyll biosynthesis in Arabidopsis. Mol. Plant 10, 155–167 (2017).
Zhao, L. et al. HY5 interacts with the histone deacetylase HDA15 to repress hypocotyl cell elongation in photomorphogenesis. Plant Physiol. 180, 1450–1466 (2019).
Islam, M. T., Wang, L. C., Chen, I. J., Lo, K. L. & Lo, W. S. Arabidopsis JMJ17 promotes cotyledon greening during de-etiolation by repressing genes involved in tetrapyrrole biosynthesis in etiolated seedlings. New Phytol. 231, 1023–1039 (2021).
Boudreault, A. A. et al. Yeast enhancer of polycomb defines global Esa1-dependent acetylation of chromatin. Genes Dev. 17, 1415–1428 (2003).
Chittuluru, J. R. et al. Structure and nucleosome interaction of the yeast NuA4 and Piccolo-NuA4 histone acetyltransferase complexes. Nat. Struct. Mol. Biol. 18, 1196–1203 (2011).
Bieluszewski, T. et al. NuA4 and H2A.Z control environmental responses and autotrophic growth in Arabidopsis. Nat. Commun. 13, 277 (2022).
Zhou, J. X. et al. The Arabidopsis NuA4 histone acetyltransferase complex is required for chlorophyll biosynthesis and photosynthesis. J. Integr. Plant Biol. 64, 901–914 (2022).
Hricova, A., Quesada, V. & Micol, J. L. The SCABRA3 nuclear gene encodes the plastid RpoTp RNA polymerase, which is required for chloroplast biogenesis and mesophyll cell proliferation in Arabidopsis. Plant Physiol. 141, 942–956 (2006).
Chen, M. et al. Arabidopsis HEMERA/pTAC12 initiates photomorphogenesis by phytochromes. Cell 141, 1230–1240 (2010).
Gao, Z. P. et al. A functional component of the transcriptionally active chromosome complex, Arabidopsis pTAC14, interacts with pTAC12/HEMERA and regulates plastid gene expression. Plant Physiol. 157, 1733–1745 (2011).
Espinosa-Cores, L. et al. Insights into the function of the NuA4 complex in plants. Front Plant Sci. 11, 125 (2020).
Bieluszewski, T. et al. AtEAF1 is a potential platform protein for Arabidopsis NuA4 acetyltransferase complex. BMC Plant Biol. 15, 75 (2015).
Tan, L. M. et al. The PEAT protein complexes are required for histone deacetylation and heterochromatin silencing. EMBO J. 37, e98770 (2018).
Galarneau, L. et al. Multiple links between the NuA4 histone acetyltransferase complex and epigenetic control of transcription. Mol. Cell 5, 927–937 (2000).
Pogson, B. J., Ganguly, D. & Albrecht-Borth, V. Insights into chloroplast biogenesis and development. Biochim. Biophys. Acta 1847, 1017–1024 (2015).
Yu, Q. B. et al. Construction of a chloroplast protein interaction network and functional mining of photosynthetic proteins in Arabidopsis thaliana. Cell Res. 18, 1007–1019 (2008).
Lian, H. et al. Photoexcited CRYPTOCHROME 1 interacts directly with G-protein beta subunit AGB1 to regulate the DNA-binding activity of HY5 and photomorphogenesis in arabidopsis. Mol. Plant 11, 1248–1263 (2018).
Waters, M. T. et al. GLK transcription factors coordinate expression of the photosynthetic apparatus in Arabidopsis. Plant Cell 21, 1109–1128 (2009).
Borner, T., Aleynikova, A. Y., Zubo, Y. O. & Kusnetsov, V. V. Chloroplast RNA polymerases: role in chloroplast biogenesis. Biochim. Biophys. Acta 1847, 761–769 (2015).
Yagi, Y. & Shiina, T. Evolutionary aspects of plastid proteins involved in transcription: the transcription of a tiny genome is mediated by a complicated machinery. Transcription 3, 290–294 (2012).
Yagi, Y. & Shiina, T. Recent advances in the study of chloroplast gene expression and its evolution. Front Plant Sci. 5, 61 (2014).
Steunou, A. L. et al. Combined action of histone reader modules regulates NuA4 local acetyltransferase function but not its recruitment on the genome. Mol. Cell. Biol. 36, 2768–2781 (2016).
Millar, C. B., Xu, F., Zhang, K. & Grunstein, M. Acetylation of H2AZ Lys 14 is associated with genome-wide gene activity in yeast. Genes Dev. 20, 711–722 (2006).
Valdes-Mora, F. et al. Acetylation of H2A.Z is a key epigenetic modification associated with gene deregulation and epigenetic remodeling in cancer. Genome Res. 22, 307–321 (2012).
Earley, K. W., Shook, M. S., Brower-Toland, B., Hicks, L. & Pikaard, C. S. In vitro specificities of Arabidopsis co-activator histone acetyltransferases: implications for histone hyperacetylation in gene activation. Plant J. 52, 615–626 (2007).
Qiu, Y. et al. HEMERA couples the proteolysis and transcriptional activity of PHYTOCHROME INTERACTING FACTORs in Arabidopsis photomorphogenesis. Plant Cell 27, 1409–1427 (2015).
Waters, M. T. & Langdale, J. A. The making of a chloroplast. EMBO J. 28, 2861–2873 (2009).
Benhamed, M., Bertrand, C., Servet, C. & Zhou, D. X. Arabidopsis GCN5, HD1, and TAF1/HAF2 interact to regulate histone acetylation required for light-responsive gene expression. Plant Cell 18, 2893–2903 (2006).
Pfalz, J., Liere, K., Kandlbinder, A., Dietz, K. J. & Oelmuller, R. pTAC2, -6, and -12 are components of the transcriptionally active plastid chromosome that are required for plastid gene expression. Plant Cell 18, 176–197 (2006).
Pfalz, J. & Pfannschmidt, T. Essential nucleoid proteins in early chloroplast development. Trends Plant Sci. 18, 186–194 (2013).
Danilova, M. N. et al. Differential impact of heat stress on the expression of chloroplast-encoded genes. Plant Physiol. Biochem. 129, 90–100 (2018).
Jin, X., Zhu, J. & Zeiger, E. The hypocotyl chloroplast plays a role in phototropic bending of Arabidopsis seedlings: developmental and genetic evidence. J. Exp. Bot. 52, 91–97 (2001).
Zhao, C. et al. Molecular cloning and characterization of OsCHR4, a rice chromatin-remodeling factor required for early chloroplast development in adaxial mesophyll. Planta 236, 1165–1176 (2012).
Chua, Y. L., Brown, A. P. & Gray, J. C. Targeted histone acetylation and altered nuclease accessibility over short regions of the pea plastocyanin gene. Plant Cell 13, 599–612 (2001).
Guo, L., Zhou, J., Elling, A. A., Charron, J. B. & Deng, X. W. Histone modifications and expression of light-regulated genes in Arabidopsis are cooperatively influenced by changing light conditions. Plant Physiol. 147, 2070–2083 (2008).
Charron, J. B., He, H., Elling, A. A. & Deng, X. W. Dynamic landscapes of four histone modifications during deetiolation in Arabidopsis. Plant Cell 21, 3732–3748 (2009).
Yang, C. et al. AtINO80 represses photomorphogenesis by modulating nucleosome density and H2A.Z incorporation in light-related genes. Proc. Natl Acad. Sci. USA 117, 33679–33688 (2020).
Mao, Z. et al. Arabidopsis cryptochrome 1 controls photomorphogenesis through regulation of H2A.Z deposition. Plant Cell 33, 1961–1979 (2021).
Li, R. et al. SOAP2: an improved ultrafast tool for short read alignment. Bioinformatics 25, 1966–1967 (2009).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Czechowski, T., Stitt, M., Altmann, T., Udvardi, M. K. & Scheible, W. R. Genome-wide identification and testing of superior reference genes for transcript normalization in Arabidopsis. Plant Physiol. 139, 5–17 (2005).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25, 402–408 (2001).
Komar, D. N., Mouriz, A., Jarillo, J. A. & Pineiro, M. Chromatin immunoprecipitation assay for the identification of Arabidopsis protein-DNA interactions in vivo. J. Vis. Exp. 2016, e53422 (2016).
Ni, F. et al. Turnip yellow mosaic virus P69 interacts with and suppresses GLK transcription factors to cause pale-green symptoms in Arabidopsis. Mol. Plant 10, 764–766 (2017).
Chen, C. et al. Cytosolic acetyl-CoA promotes histone acetylation predominantly at H3K27 in Arabidopsis. Nat. Plants 3, 814–824 (2017).
Acknowledgements
We thank T. Hernández-Verdeja for the critical reading of the manuscript and also V. M. Quesada (Univ. Miguel Hernandez, Elche, Spain) and M. Cheng (Duke University, Durham, NC, USA) for kindly providing sca3-2 (ref. 20) and hmr-22 (ref. 38) mutants, respectively. This work was funded by the Spanish Ministry of Economy, Industry, and Competitiveness (grant Nos. BIO2016-77559-R and PID2019-104899GB-I00 to M.P. and J.A.J. from MCIN/AEI/10.13039/501100011033 and FEDER; FPI fellowship No. BES-2017-07992 from MCIN/AEI/10.13039/501100011033 and FSE to L.B-M.) and INIA (FPI fellowship No. FPI-SGIT-2016-08 to L.E.-C.). We acknowledge the ‘Severo Ochoa Program for Centres of Excellence in R&D’ from the Agencia Estatal de Investigación of Spain (grant No. SEV-2016-0672 (2017-2021) from MCIN/AEI/10.13039/501100011033) for supporting the scientific services used in this work.
Author information
Authors and Affiliations
Contributions
M.P. and J.A.J. conceived this project; L.E.-C. analysed protein–protein interactions between NuA4-C subunits. J.B.-G. and L.B.-M. performed the rest of the experiments. J.B.-G., M.P. and J.A.J. designed all the experiments, analysed the data and wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Plants thanks Enrique Lopez-Juez and the other, anonymous, reviewer(s) for their contribution to the peer review of this work
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–8, description of Supplementary tables and unmodified gel image of Supplementary Fig. 2.
Supplementary Tables
Combined workbook containing three supplementary tables.
Source data
Source Data Fig. 1
Unprocessed western blots.
Source Data Fig. 3
Statistical source data for Fig. 3.
Source Data Fig. 3
Unprocessed western blots.
Source Data Fig. 4
Statistical source data for Fig. 4.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Barrero-Gil, J., Bouza-Morcillo, L., Espinosa-Cores, L. et al. H4 acetylation by the NuA4 complex is required for plastid transcription and chloroplast biogenesis. Nat. Plants 8, 1052–1063 (2022). https://doi.org/10.1038/s41477-022-01229-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-022-01229-4